Abstract
Studies on microbial populations in near-ground aerosols during fog–haze days could help enhance our knowledge concerning the relationship between fog–haze and human health. In this study, microbial populations in aerosols near-ground during a fog–haze event in Beijing were analyzed using clone library methods. Results showed that the bacterial diversity in aerosols during fog–haze days was lower than that after fog–haze days. Proteobacteria alone and Proteobacteria with Firmicutes were respectively detected in 1.5 and 20 m aerosols during fog–haze days. In addition to Proteobacteria and Firmicutes, Bacteroidetes alone and Acidobacteria with Verrucomicrobia were respectively found in aerosols at 20 and 1.5 m after fog–haze days. The fungal species observed during fog–haze days were completely different from those detected after fog–haze days. Ascomycota and Basidiomycota were respectively detected during and after fog–haze days. The distribution of microbial diversity in aerosols exhibited meteorological and site-associated variations. The same potential pathogenic microorganisms were detected at different heights during fog–haze days. This study on the characteristics of microbial population in aerosols could provide a comprehensive understanding of the factors causing the harmful effects of particles on humans during fog–haze days.
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
Introduction
Fog–haze events exhibit an episodic tendency in China (Sun et al., 2006; Liu et al., 2008; Zhang et al., 2012; Li et al., 2013a; Zhao et al., 2013; Tao et al., 2012, 2013; Xu et al., 2016; He et al., 2016; Yin et al., 2017). Aerosol particles during fog–haze days can affect human health and cause several diseases, particularly respiratory diseases (Salma et al., 2002; Spindler et al., 2010; Pope et al., 2011; Atkinson et al., 2012; Nirmalkar and Deb, 2016). As such, the properties of aerosols and their microbial composition have been extensively investigated.
Microbial concentration and size distribution in aerosols vary under different weather conditions. Hu et al. (2015) demonstrated that cultured bacterial and fungal concentrations in aerosols are lower on haze days than on non-haze days. Conversely, Gao et al. (2015) observed that cultured bacterial and fungal concentrations in aerosols are higher on haze days than on non-haze days. The total microbial concentration in aerosols on foggy and hazy days is also higher than that on non-hazy days (Dong et al., 2016). Cultivation methods initially revealed that Gram-positive bacteria were the dominant ones in aerosols (Di Giorgio et al., 1996; Shaffer and Lighthart, 1997). With the development of molecular biological techniques based on DNA analysis, Proteobacteria, Firmicutes, and Actinobacteria are often observed in aerosols (Ranjard et al., 2000; Radosevich et al., 2002; Wilson et al., 2002; Maron et al., 2005). In addition to bacterial populations, fungal populations in aerosols have also been explored (Medrela-Kuder, 2003; Fröhlich-Nowoisky et al., 2009; Dannemiller et al., 2014).
Considering the relationship between microorganisms in aerosols and human health, researchers examined the microbial characteristics of aerosols from near-ground areas. Bowers et al. (2011) observed that bacterial communities in aerosols sampled from different land uses are related to sources but not to local meteorological conditions. They also investigated the seasonal variability of microbial diversity in aerosols and showed that microbial diversity exhibits significant seasonality patterns (Bowers et al., 2013). Microbial populations in aerosols at 2 m near-ground are characterized by a strong seasonal trend related to general climatic variables, but these populations are not significantly affected by urban development, as indicated by anthropogenic pollutants and human population levels (Woo et al., 2013). Gou et al. (2016) sampled Particulate Matter (PM) from 5 m above ground without surrounding barriers and analyzed the microbial populations in PM1 and PM10. They found that Proteobacteria, Firmicutes, and Actinobacteria are the main communities in PM while Ascomycota and Basidiomycota are the main fungal groups in PM. Cao et al. (2014) used metagenomic methods and observed that the relative abundance of respiratory microbial allergens and pathogens increases as the concentrations of PM pollution increase during a severe smog event in January in Beijing.
The microbial properties of aerosols from near-ground are of different states under various weather conditions. However, microbial populations in aerosols from human breathing height under special weather conditions, such as fog–haze days, have been rarely examined. The crucial connection between urban atmosphere pollution and human health remains poorly understood because of insufficient intensive investigations on microorganisms. In our study, microbial populations in aerosols from near-ground (1.5 and 20 m) were investigated. Aerosols were sampled during and after fog–haze days from January 29 to February 2, 2013. A cloning library was prepared to analyze the bacterial and fungal diversities and contents in aerosols from different heights under various weather conditions. The characteristics of the microorganisms in aerosols during and after fog–haze days were compared. Our results could provide valuable data to improve knowledge regarding aerosol properties in different pollution episodes in Beijing.
Materials and methods
Sample collection
All aerosol samples were consecutively collected at 1.5 m (breathing height) and 20 m (height of no surrounding obstacles) above ground during and after fog-haze days in northern China (116.34° E, 40.01° N) (Fig. 1, Fig. S1 and Table 1). An impingement airborne microorganism sampler (SKC BioSampler, SKC Inc. USA) was used to collect aerosols (Han et al., 2012).The collection vessel of this sampler was filled with 20 ml of sterile phosphate buffered saline as the collection medium. Air samples were transferred into the collection medium by drawing the gas at a flow rate of 12.5 l/min. During fog-haze days, the samples were collected eight hours each days and Continuous three days. After fog-haze days, the samples were collected twelve hours each days and Continuous two days. Several meteorological parameters were observed. Temperature and relative humidity were recorded by using a dewpoint thermohygrometer (WD-35612, OAKTON, Germany). Wind speed and irradiance were measured with a portable anemometer (HD2303, Delta OHM, Italy) and irradiance meter (HD2302.0, Delta OHM, Italy), respectively.
Clone library
The total DNA of aerosols was isolated using a DNA autoplate (TanBead, Taiwan) by Automatic Platform for Magnetic System-16 (TanBead, Taiwan). DNA bands were observed through 1.0% agarose gel electrophoresis.
All of the samples were assayed in duplicate. The universal primers F16S-27/R16S-1492 and ITS1/ITS4 were used to amplify the bacterial 16S rRNA and fungal 18S rRNA segment, respectively (Madsen et al., 2015). PCR amplification was performed using the Gene AmpR PCR System (9700, AB, USA) at a final volume of 50 μl. The reaction mixture contained 2.0 μl of both primers, 2.0 μl of DNA template, 4.0 μl of each dNTPs, 5.0 μl of 10 × PCR buffer (TaKaRa, China), 0.25 μl of Taq DNA polymerase (TaKaRa, China) and 36.75 μl DNA-free water. Temperature cycling conditions of bacterial PCR were 94 °C for 5 min, followed by 25 cycles of 94 °C for 30 s, 56 °C for 30 s, and 72 °C for 90 s. A final extension at 72 °C for 10 min was used. Fungal PCR included 95 °C for 5 min, followed by 36 cycles of 95 °C for 30 s, 55 °C for 30 s, and 72 °C for 1 min, followed by 72 °C for 10 min. The PCR products were purified, ligated to the vector, and transformed into competent cells. The positive clones were submitted for sequencing by using an ABI 3730DXL DNA sequencer (AB, USA).
All of the sequences from the clone libraries were trimmed to exclude vector sequences and then checked for chimeras with Bellerophon via Greengenes (http://greengenes.lbl.gov/). The aligned sequences (DNAMAN V6) were grouped into universal operational taxonomic units (OTUs) at a threshold of 97% minimum similarity. Individual sequences were also grouped into OTUs to identify microbial distribution in aerosols from different sampling sites. Representative sequence of each OTU in the libraries was aligned to the NCBI database by the BLAST website. Phylogenetic affiliations of all OTUs were identified by constructing a phylogenetic tree including representative sequences of each OTU and related sequences from the previous NCBI database using the neighbor-joining algorithm by MEGA version 6.1.
Coverage and diversity of clone libraries from different samples
The coverage of each clone library was calculated according to the following equation (Good, 1953):
- n :
-
is the number of unique clones
- N :
-
is the total number of sequences examined
Whether the coverage of a clone library was large enough to represent the diversity of an original community was determined. The diversity of each sample was estimated using the Shannon Wiener index (Ding et al., 2008).
Results
Coverage of clone libraries
A total of 400 sequences for bacteria and 200 sequences for fungi were obtained. After alignment was completed, 7, 2, 22, and 15 OTUs were identified in HYP-FZB20, HYP-FZB1.5, HYP-CB20, and HYP-CB1.5 clone libraries, respectively, and 7, 13, 1, and 4 OTUs were determined in the 50 screened fungal clones from HYP-FZF20, HYP-FZF1.5, HYP-CF20, and HYP-CF1.5 clone libraries, respectively. Coverage analysis indicated that the aerosol libraries represented approximately 83.00%–99.00% of the total number of the examined clones. Thus, a dependable inventory of bacterial 16S rRNA and fungal 18S rRNA gene sequences present in aerosols is provided.
Microbial diversity in aerosols during and after fog–haze days
Shannon–Wiener indexes (Fig. S2a) of bacterial species were 1.32, 0.23, 2.47, and 2.23 for HYP-FZB20, HYP-FZB1.5, HYP-CB20, and HYP-CB1.5, respectively. The species richness of bacteria in aerosols collected from clear days (after fog–haze days in our study) was higher than that in aerosols sampled from fog–haze days. The bacterial diversity in 20 m aerosols was also richer than in 1.5 m aerosols during and after fog–haze days. The change in the bacterial diversity in 1.5 m to 20 m aerosols during fog–haze days was more apparent than that after fog–haze days. Shannon–Wiener indexes (Fig. S2b) of the fungal species were 1.83, 2.15, 0, and 0.42 for HYP-FZF20, HYP-FZF1.5, HYP-CF20, and HYP-CF1.5, respectively. The species richness of the fungal diversity in aerosols during fog–haze days at 1.5 m height was the highest among the four fungal clone libraries. The fungal diversity in aerosols was different from bacterial diversity. The fungal diversity in aerosols during fog–haze days was richer than that after fog–haze days, and the fungal diversity in aerosols at 1.5 m height was richer than that at 20 m height.
Microbial population structure in aerosols during and after fog–haze days
Sequences were assigned to bacterial and fungal affiliations according to their position in the phylogenetic tree (Figs. S3 and S4 and Tables S1 and S2).
During fog–haze days, most of the bacteria in 20 m aerosols were assigned to uncultured Burkholderia (51.92%) and uncultured bacterium (28.85%); the minor groups were Enhydrobacter, Achromobacter, Anaerovorax, and Sporosarcina (Fig. S3a and Table S1). Under the same weather conditions and in the same sampling sites, six fungal genera, namely, Stemphylium (25.93%), Penicillium (22.22%), Curvularia (18.52%), Chalara (14.81%), Aureobasidium (14.81%), and Aspergillus (3.71%),were detected (Fig. S4a and Table S2). During fog–haze days, only two bacterial genera were observed at human breathing height (1.5 m). As shown in Fig. S3b and Table S1, Burkholderia was the most represented bacterial group (94.00%), and the other OTUs were related to uncultured Achromobacter (6.00%). The dominant fungal groups at 1.5 m were Chalara (35.48%), Penicillium (19.37%), Curvularia (12.90%), Aspergillus (9.68%), and Stemphylium (9.68%). The other OTUs were related to uncultured Talaromyces, Cadophora, Geomyces, orAureobasidium (Fig. S4b and Table S2).
After fog–haze days, the represented bacterial groups in HYP-CB20 were Achromobacter (47.17%), Stenotrophomonas (16.98%), uncultured bacterium (9.45%), and Sporosarcina (5.66%). The other minor OTUs were Idiomarina, Nevskia, Brevundimonas, Tsukamurella, Carnobacterium, Chryseomicrobium, Bacillus, Halomonas, Sphingomonas, Microvirga, and Legionellaceae (Fig. S3c and Table S1). The fungal group in HYP-CF20 was Cryptococcus (100%). The bacterial groups in HYP-CB1.5 were uncultured bacterium (37.74%), Pseudomonas (22.64%), Psychrobacter (11.32%), and Acinetobacter (9.43%). The other minor OTUs were Clostridium, Acidobacteria, Rhodocyclus, Hyphomiceobiaceae, Carnobacterium, and Verrucomicrobiales (Fig. S3d and Table S1). The represented fungal group in HYP-CF1.5 was uncultured Cryptococcus (96.77%). The other minor OTU was Clavulina.
Pathogenic property in aerosols during and after fog–haze days
Among these identified bacterial and fungal species, some microorganisms are potential pathogens (Table S3). During fog–haze days, Burkholderia fungorum (Peeters et al., 2016) and Achromobacter sp. (Coward et al., 2016) were also detected at 20 and 1.5 m height. The respective relative percentages of Burkholderia fungorum were 1.92% and 94% at 20 and 1.5 m height, and the respective relative percentages of Achromobacter sp. were 5.77% and 6% at 20 and 1.5 m height. After fog–haze days, several species, including 47.17% Achromobacter sp., 1.89% Sphingomonas aurantiaca (Babic et al., 2015), 1.89% Nevskia sp. (Low et al., 2016), 1.89% Idiomarina sp. (Cosseau et al., 2016), 3.78% Legionella bacterium (Alum and Isaacs, 2016), and 1.89% Tsukamurella tyrosinosolvens (Romano et al., 2011), were detected at 20 m height. Two potential pathogens, namely, 3.77% Acinetobacter lwoffii (Baldeo et al., 2015) and 11.32% Psychrobacter sp. (Benhamed et al., 2014) were detected at 1.5 m height. For fungi, 14.81% and 3.23% Aureobasidium sp. (Oliveira et al., 2013) were respectively found at 20 and 1.5 m height during fog–haze days, and 100% and 90.32% Cryptococcus albidus (Liu et al., 2014) were respectively observed at 20 and 1.5 m height after fog–haze days.
Discussion
Studies on microbial populations in aerosols have indicated that bacteria affiliated with Proteobacteria, Firmicutes, Actinobacteria, and Bacteroidetes and fungi associated with Ascomycota and Basidiomycota are detected in near-ground aerosols (Bowers et al., 2013). However, the precise characteristics of microorganisms in aerosols on human breathing height have yet to be identified under unique weather conditions, such as fog–haze days. In our work, aerosols were sampled from human breathing height (1.5 m) and at a height at which no surrounding barriers were observed (20 m) during and after fog-haze days. At the phylum level, the microbial populations were different between during and after fog-haze days.
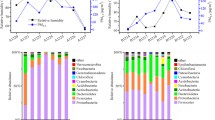
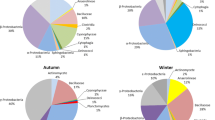
In Fig. 2, only Proteobacteria and Firmicutes were detected in aerosols during fog–haze days. Five bacterial phyla, namely, Proteobacteria, Firmicutes, Actinobacteria, Bacteroidetes, and Verrucomicrobia, were detected in aerosols after fog–haze days. As the dominant bacterial phylum, Proteobacteria also varied under different weather conditions with deepening classification (Fig. 4). During fog–haze days, only Betaproteobacteria were detected. Gammaproteobacteria, Alphaproteobacteria, and Betaproteobacteria were found in aerosols after fog–haze days. The difference in fungal groups was more evident. All of the fungi detected during fog–haze days were affiliated with Ascomycota, and all of the fungi found after fog–haze days were associated with Basidiomycota (Fig. 3).
Proportion of bacterial population in aerosols during and after fog-haze days (HYP-FZB1.5: bacteria in 1.5 m aerosols during fog-haze days; HYP-FZB20: bacteria in 20 m aerosols during fog-haze days; HYP-CB1.5: bacteria in 1.5 m aerosols after fog-haze days; HYP-CB20: bacteria in 20 m aerosols after fog-haze days)
Our study found that microbial populations in aerosols demonstrated site-related characteristics under different weather conditions. In Fig. 4, the bacteria at 1.5 m height were also found at 20 m height during fog–haze days, but this phenomenon did not appear after fog–haze days. During fog–haze days, Burkholderiales was observed at 1.5 m height. Burkholderiales, Rhodocyclales, and Pseudomonadales were also found at 20 m height. After fog–haze days, three bacterial orders, namely, Rhodocyclales, Pseudomonadales, and Rhizobiales, were detected in 1.5 m aerosols, and seven bacterial orders, namely, Burkholderiales, Oceanospirillales, Alteromonadales, Legionellales, Xanthomonadales, Caulobacterales, and Sphingomonadales, were observed in 20 m aerosols.
For fungi, the species detected at 20 m height were nearly detected at 1.5 m height during and after fog–haze days (Fig. 5). The classes and genera were also different at various heights during and after days. After fog–haze days, Tremellomycetes were detected at 20 m height, whereas Agaricomycetes and Tremellomycetes were found at 1.5 m height. Class taxonomic levels of Dothideomycetes, Leotiomycetes, and Eurotiomycetes did not differ during fog–haze days at 1.5 and 20 m height. Conversely, these fungi varied in the genus taxonomic level at different heights. The following genera were detected at 20 m height: Stemphylium, Aureobasidium, and Curvularia with Dothideomycetes; Chalara with Leotiomycetes; and Penicillium and Aspergillus with Eurotiomycetes. In addition to these genera, Cadophora and Geomyces with Leotiomycetes and Talaromyces with Eurotiomycetes were detected at 1.5 m height.
Potential pathogenic bacteria, such as Burkholderia fungorum and Achromobacter sp., were also detected in aerosols at 20 and 1.5 m height during fog–haze days, and the percentages of these two species increased from 1.92% to 94% and from 5.77% to 6% at 20 and 1.5 m height, respectively. As fog–haze disappears, completely different pathogenic bacteria were observed in 20 and 1.5 m aerosols. At 20 m, six potential pathogenic bacteria, namely, 47.17% Achromobacter sp., 1.89% S. aurantiaca, 1.89% Nevskia sp., 1.89% Idiomarina sp., 1.89% Legionella-like amoebal pathogen, and 1.89% Tsukamurella tyrosinosolvens and two potential pathogenic bacteria, namely, 11.32% Psychrobacter sp. and 3.77% Acinetobacter lwoffii, were observed. For potential pathogenic fungi, 14.81% and 3.23% Aureobasidium sp. were respectively detected in aerosols at 20 and 1.5 m height during fog–haze days, and 100% and 90.32% Cryptococcus sp. were respectively found in aerosols at 20 and 1.5 m height after fog–haze days. These results suggested that the distribution of microbial populations in aerosols presented at different heights near-ground during fog-haze days compared with that after fog-haze days.
The difference in microbial distribution in near-ground aerosols under various weather conditions can be determined on the basis of meteorological conditions. In our work, particulate matter concentration, irradiance, wind speed, temperature, and relative humidity were recorded and monitored. In Table 2, these weather conditions mainly differed in terms of irradiance and wind speed during and after fog–haze days. During fog–haze days, irradiance and wind speed were 2.54–5.60 and 0.02–0.07 m/s, respectively. After fog–haze days, irradiance and wind speed increased to 65.16–438.50 W/m2 and 1.10–7.83 m/s. A previous study indicated that the formation of fog–haze is closely related to meteorological conditions (Quan et al., 2011), particularly wind speed. Wind speed is a key factor that affects aerosol concentration during fog–haze days. Low wind speeds likely affect the diffusion and transport of aerosols Li et al., 2013b) and therefore accounted for the similarity of microbial populations in aerosols between 20 and 1.5 m during fog–haze days in our study. After fog–haze days, microbial populations, especially bacterial population, differed between 20 and 1.5 m because of particle diffusion as wind speed increased (1.10–7.83 m/s).
As an important microbial carrier, aerosols possibly affect human health (Tie et al., 2009; Xu et al., 2013). In our study, potential pathogenic bacteria and fungi detected at 20 m could be detected at 1.5 m during fog–haze days. The relative contents of potential pathogenic bacteria, such as Burkholderia fungorum and Achromobacter sp., at 1.5 m were higher than those at 20 m. However, this phenomenon was not observed after fog–haze days. This result implied that these potential pathogenic microorganisms posed greater harm to humans during fog–haze days than after fog–haze days.
Conclusions
The characteristics of microbial populations in aerosols during fog–haze outbreak periods in Beijing are examined. Although bacterial diversity in aerosols during fog–haze days is lower than that after fog–haze days, the distribution of bacterial populations in aerosols from different sampling sites varies under different weather conditions. The bacterial population in aerosols is similar between 20 and 1.5 m during fog–haze days. After fog–haze days, the bacteria in aerosols detected at 20 and 1.5 m height also differ. Only Carnobacterium sp. was also detected in 20 and 1.5 m aerosols. The fungal diversity in aerosols during fog–haze days is higher than that after fog–haze days, and the fungal population in aerosols is similar during and after fog–haze days. This phenomenon is also observed in potential pathogenic microorganisms. Two potential pathogenic bacteria, namely, Burkholderia fungorum and Achromobacter sp., are detected at 20 and 1.5 m height, and their percentages increase as height decreases during fog–haze days.
References
Alum A, Galahad Zachariah Isaacs GZ (2016) Aerobiology of the built environment: synergy between legionella and fungi. Am J Infect Control 44:S138–S143
Atkinson RW, Cohen A, Mehta S, Anderson HR (2012) Systematic review and meta-analysis of epidemiological time-series studies on outdoor air pollution and health in Asia. Air Qual Atmos Health 5:383–391
Babic MN, Zalar P, Zenko B, Schroers HJ, Dzeroski S, Gundecimerman N (2015) Candida and Fusarium species known as opportunistic human pathogens from customer-accessible parts of residential washing machines. Fungal Biol 119:95–113
Baldeo C, Isache C, Baldeo C, Bajwa A (2015) A case of disseminated intravascular coagulation secondary to Acinetobacter lwoffii and Acinetobacter baumannii bacteremia. IDCases 2:70–71
Benhamed S, Guardiola FA, Mars M, Esteban MA (2014) Pathogen bacteria adhesion to skin mucus of fishes. Vet Microbiol 171:1–12
Bowers RM, McLetchie S, Knight R, Fierer N (2011) Spatial variability in airborne bacterial communities across land-use types and their relationship to the bacterial communities of potential source environments. ISME J 5:601–612
Bowers RM, Clements N, Emerson JB, Wiedinmyer C, Hannigan MP, Fierer N (2013) Seasonal variability in bacterial and fungal diversity of the near-surface atmosphere. Environ SciTechnol 47:12097–12106
Cao C, Jiang W, Wang B, Fang J, Lang J, Tian G, Jiang J, Zhu TF (2014) Inhalable microorganisms in Beijing’s PM2.5 and PM10 pollutants during a severe smog event. Environ SciTechnol 48:1499–1507
Cosseau C, Romano-Bertrandb S, Duplan H, Lucas O, Ingrassia I, Pigasse C, Roques C, Jumas-Bilak E (2016) Proteobacteria from the human skin microbiota: species-level diversity and hypotheses. One Health 2:33–41
Coward A, Kenna DTD, Perry C, Martin K, Doumith M, Turton JF (2016) Use of nrdA gene sequence clustering to estimate the prevalence of different Achromobacter species among cystic fibrosis patients in the UK. J Cyst Fibros 15:479–485
Dannemiller KC, Lang-Yona N, Yamamoto N, Rudich Y, Peccia J (2014) Combining real-time PCR and next-generation DNA sequencing to provide quantitative comparisons of fungal aerosol populations. Atmos Environ 84:113–121
Di Giorgio C, Krempff C, Guiraud A, Binder H, Tiret P, Dumenil C (1996) Atmospheric pollution by airborne microorganisms in the city of Marseilles. Atmos Environ 30(1):155–160
Ding Y, Wu WX, Han ZY, Chen YX (2008) Correlation of reactor performance and bacterial community composition during the removal of trimethylamine in three-stage biofilters. Biochem Eng J 38(2):248–258
Dong L, Qi J, Shao C, Zhong X, Gao D, Cao W, Gao J, Bai R, Long G, Chu C (2016) Concentration and size distribution of total airborne microbes in hazy and foggy weather. Sci Total Environ 541:1011–1018
Fröhlich-Nowoisky J, Pickersgill DA, Després VR, Pöschl U (2009) High diversity of fungi in air particulate matter. P Natl Acad Sci USA 106(31):12814–12819
Gao M, Jia RZ, Qiu TL, Han ML, Song Y, Wang XM (2015) Seasonal size distribution of airborne culturable bacteria and fungi and preliminary estimation of their deposition in human lungs during non-haze and haze days. Atmos Environ 118:203–210
Good IJ (1953) The population frequencies of species and the estimation of population parameters. Biometrika 40(3/4):237–264
Gou H, Lu J, Li S, Tong Y, Xie C, Zheng X (2016) Assessment of microbial communities in PM1 and PM10 of Urumqi during winter. Environ Pollut 214:202–210
Han Y, Li L, Liu J, Zhang M (2012) Microbial structure and chemical components of aerosols caused by rotating brushes in a wastewater treatment plant. Environ Sci Pollut Res 19:4097–4108
He Q, Zhou G, Geng F, Gao W, Yu W (2016) Spatial distribution of aerosol hygroscopicity and its effect on PM2.5 retrieval in East China. Atmos Res 170:161–167
Hu L, Zhang K, Wang H, Li N, Wang J, Yang W, Yin Z, Jiao Z, Wen Z, Li J (2015) Concentration and particle size distribution of microbiological aerosol during haze days in Beijing. Environ Sci 36:3144–3149
Li J, Wang Z, Huang H, Hu M, Meng F, Sun Y, Wang X, Wang Y, Wang Q (2013a) Assessing the effects of trans-boundary aerosol transport between various city clusters on regional haze episodes in spring over East China. Tellus B 65(1):60–73
Li J, Peng XT, Zhou HY, Li JT, Sun ZL (2013b) Molecular evidence for microorganisms participating in Fe, Mn, and S biogeochemical cycling in two low-temperature hydrothermal fields at the southwest Indian ridge. J Geophys Res-Biogeosciences 118(2):665–679
Liu Z, Liu D, Huang J, Vaughan M, Uno I, Sugimoto N, Kittaka C, Trepte C, Wang Z, Hostetler C, Winker D (2008) Airborne dust distributions over the Tibetan plateau and surrounding areas derived from the first year of CALIPSO lidar observations. Atmo Chem Phys 8(16):5045–5060
Liu Y, Man S, Wang X, Xu W, Tang J (2014) Cryptococcus Albidus encephalitis in newly diagnosed HIV-patient and literature review. Med Mycol Case Reports 3:8–10
Low A, Ng C, He J (2016) Identification of antibiotic resistant bacteria community and a GeoChip based study of resistome in urban watersheds. Water Res 106:330–338
Madsen AM, Zervas A, Tendal K, Nielsen JL (2015) Microbial diversity in bioaerosol samples causing ODTS compared to reference bioaerosol samples as measured using Illumina sequencing and MALDI-TOF. Environ Res 140:255–267
Maron P-A, Lejon DPH, Carvalho E, Bizet K, Lemanceau P, Ranjard L, Mougel C (2005) Assessing genetic structure and diversity of airborne bacterial communities by DNA fingerprinting and 16S rDNA clone library. Atmos Environ 39(20):3687–3695
Medrela-Kuder E (2003) Seasonal variations in the occurrence ofculturable airborne fungi in outdoor and indoor air in Craców. Int Biodeterior Biodegr 52(4):203–205
Nirmalkar J, Deb MK (2016) Impact of intense field burning episode on aerosol mass loading and its possible health implications in rural area of eastern central India. Air Qual Atmos Health 9:241–249
Oliveira BR, Barreto Crespo MT, San Romǎo MV, Benoliel MJ, Samson RA, Pereira VJ (2013) New insights concerning the occurrence of fungi in water sources and their potential pathogenicity. Water Res 47:6338–6347
Peeters C, Depoorter E, Praet J, Vandamme P (2016) Extensive cultivation of soil and water samples yields various pathogens in patients with cystic fibrosis but not Burkholderia multivorans. J Cyst Fibros 15:769–775
Pope CA III, Brook RD, Burnett RT, Dockery DW (2011) How is cardiovascular disease mortality risk affected by duration and intensity of fine particulate matter exposure? An integration of the epidemiologic evidence. Air Qual Atmos Health 4:5–14
Quan J, Zhang Q, He H, Liu J, Huang M, Jin H (2011) Analysis of the formation of fog and haze in North China plain (NCP). Atmos Chem Phys 11(15):8205–8214
Radosevich JL, Wilson WJ, Shinn JH, DeSantis TZ, Andersen GL (2002) Development of a high-volume aerosol collection system for the identification of air-borne micro-organisms. Lett Appl Microbiol 34(3):162–167
Ranjard L, Poly F, Nazaret S (2000) Monitoring complex bacterial communities using culture-independent molecular techniques: application to soil environment. Res Microbiol 151(3):167–177
Romano L, Spanu T, Calista F, Zappacosta B, Mignogna S, Sali M, Fiori B, Fadda G (2011) Tsukamurella tyrosinosolvens and rhizobium radiobacter sepsis presenting with septic pulmonary emboli. Clin Microbiol Infec 17:1032–1052
Salma I, Balashazy I, Winkler-Heil R, Hofmann W, Zaray G (2002) Effect of particle mass size distribution on the deposition of aerosols in the human respiratory system. J Aerosol Sci 33:119–132
Shaffer BT, Lighthart B (1997) Survey of culturable airborne bacteria at four diverse locations in Oregon: urban, rural, forest, and coastal. Microb Ecol 34(3):167–177
Spindler G, Bruggemann E, Gnauk T, Gruner A, Muller K, Herrmann H (2010) A four-year size-segregated characterization study of particles PM10, PM2.5 and PM1 depending on air mass origin at Melpitz. Atmos Environ 44:164–173
Sun Y, Zhuang G, Tang A, Wang Y, An Z (2006) Chemical characteristics of PM2.5 and PM10 in haze-fog episodes in Beijing. Environ SciTechnol 40(10):3148–3155
Tao M, Chen L, Su L, Tao J (2012) Satellite observation of regional haze pollution over the North China plain. J Geophys Res 117(D12):12203
Tao M, Chen L, Wang Z, Tao J, Su L (2013) Satellite observation of abnormal yellow haze clouds over East China during summer agricultural burning season. Atmos Environ 79:632–640
Tie X, Wu D, Brasseur G (2009) Lung cancer mortality and exposure to atmospheric aerosol particles in Guangzhou, China. Atmos Environ 43(14):2375–2377
Wilson KH, Wilson WJ, Radosevich JL, Desantis TZ, Viswanathan VS, Kuczmarski TA, Andersen GL (2002) High-density microarray of small subunit ribosomal DNA probes. Appl Environ Microb 68(5):2535–2541
Woo AC, Brar MS, Chan Y, Lau MCY, Leung FCC, Scott JA, Vrijmoed LLP, Zawar-Reza P, Pointing SB (2013) Temporal variation in airborne microbial populations and microbially-derived allergens in a tropical urban landscape. Atmos Environ 74:291–300
Xu J, Xu H-H, Xiao H, Tong L, Snape CE, Wang C-J, He J (2016) Aerosol composition and sources during high and low pollution periods in Ningbo, China. Atmos Res 178-179:559–569
Xu Z, Yao M (2013) Monitoring of bioaerosol inhalation risks in different environments using a six-stage Andersen sampler and the PCR-DGGE method. Environ Monit Assess 185(5):3993–4003
Yin D, Zhao S, Qu J (2017) Spatial and seasonal variations of gaseous and particulate matter pollutants in 31 provincial capital cities, China. Air Qual Atmos Health 10(3):359–370
Zhang XY, Wang YQ, Niu T, Zhang XC, Gong SL, ZhangYM SJY (2012) Atmospheric aerosol compositions in China: spatial/temporal variability, chemical signature, regional haze distribution and comparisons with global aerosols. Atmos Chem Phys 12(2):779–799
Zhao XJ, Zhao PS, Xu J, Meng W, Pu WW, Dong F, He D, Shi QF (2013) Analysis of a winter regional haze event and its formation mechanism in the North China plain. Atmos Chem Phys 13(11):5685–5696
Acknowledgements
This work was supported by National Natural Science Foundation of China (NO.51308527); Young Scientists Fund of RCEES (RCEES-QN-20130006F); and China Welfare Funds for Environmental Protection (No. 201509008). We express our gratitude to John J. for revised the grammar and sentence.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Han, Y., Zhang, M., Li, L. et al. Microbial population structure in near-ground aerosols during fog-haze days in northern China. Air Qual Atmos Health 10, 1113–1121 (2017). https://doi.org/10.1007/s11869-017-0498-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11869-017-0498-4