Abstract
In the clinical research, three-dimensional/volumetric anatomical structure of the human body is very significant for diagnosis, computer-aided surgery, surgical planning, patient follow-up, and biomechanical applications. Medical imaging procedures including MRI (Magnetic Resonance Imaging), CT (Computed Tomography), and CBCT (Cone-beam computed tomography) have certain drawbacks such as radiation exposure, availability, and cost. As a result, 3D reconstruction from 2D X-ray images is an alternative way of achieving 3D models with significantly low radiation exposure to the patient. The purpose of this study is to provide a comprehensive view of 3D image reconstruction methods using X-ray images, and their applicability in the various anatomical sections of the human body. This study provides a critical analysis of the computational methods, requirements and steps for 3D reconstruction. This work includes a comparative critical analysis of the state-of-the-art approaches including the feature selection along with their benefits and drawbacks. This review motivates the researchers to work for 3D reconstruction using X-ray images as only a limited work is available in the area. It may provide a solution for many experts who are looking for techniques to reconstruct 3D models from X-ray images for clinical purposes.
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
1 Introduction
A human body is a real three-dimensional anatomical structure. Invasive methods are not always recommendable for the disease diagnosis and treatment planning of a patient. Therefore, several imaging modalities are used to visualize the internal anatomy of the human body such as X-ray, CT (Computed Tomography), CBCT (Cone-beam Computed Tomography), Ultra-sound imaging, MRI (Magnetic Resonance Imaging), PET (Positron Emission Tomography), etc. These modalities provide 2D images as well as 3D images based on the type of modality. Every imaging modality has its specific use and applicability for clinical use. X-ray imaging is a popular imaging modality and is widely used to visualize the anatomical structures inside the body. X-ray imaging modality is the most available and low-cost modality compared to the other modalities. Therefore, the use of the X-ray image is very high and acceptable.
However, X-ray images have many drawbacks [1, 2]. The X-ray image provides the projected 2D view of the real 3D anatomical structure which may not be much useful for the radiologist in a few severe cases. The anatomical geometry is overlapped in X-ray images which may create confusion to understand the real anatomical structure. The measurements are obtained projected measurements from 3 to 2D which loses the real measurements. The image calibration is required to read the X-ray images.
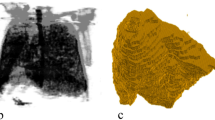
To overcome the drawbacks of the X-ray images [3], 3D images like CT and CBCT are used for clinical practices [4]. But there are certain drawbacks with 3D images like CT and CBCT. These imaging modalities are very less available, costly, and expose the patient to higher radiation compared to the X-ray images [5]. A conventional way of visualization of the anatomical volume through the CT scan is shown in Fig. 1.
To solve the problems of 3D imaging modalities, diagnosis and treatment planning can depend on 2D X-ray images in non-severe cases where radiologists and surgeons can understand the 3D anatomy by looking at the X-ray images. But in serious cases, diagnosis and treatment planning becomes difficult using X-ray images only. Therefore, many methods were involved that can construct the 3D image using X-ray images. These methods are overcoming the drawbacks of 2D as well as 3D imaging modalities and are now available at an early stage. Such methods are widely required for the possible anatomical regions.
Three-dimensional image construction for an anatomical region is used for the patients in terms of medical diagnosis, treatment planning, and follow-up [6,7,8,9,10]. There are several diseases associated with different anatomical regions [11] of the human body where image-based assessment is important (for example., Knee bone fracture, Femur bone fracture, arthritis, obstructive sleep apnea syndrome, sinusitis, lesions detections in the chest, inflammatory diseases, bone fractures, etc.). Diagnostic images enable the radiologist to view the anatomical structure of the region to figure out causes of illness, injury and to confirm a disease [12] with proper follow-up [13].
Thus, 3D construction from X-ray images is the most favorable solution for visualizing the 3D anatomy of the patient for clinical analysis. It is cost-effective, widely available and exposes a patient to less radiation. The number of radiographs taken should be less in number viz., one or two per patient as it is a primary concern of 3D reconstruction from 2D X-ray images.
In this paper, a comprehensive review of the many available approaches or techniques for 3D image reconstruction from X-ray images is presented in further sections. The requirements, concepts and classifications of 3D reconstruction methods from X-ray images are discussed. Then their applications, benefits, and drawbacks are assessed. Finally, the accuracy of various methods is mentioned. Hence, this review can help researchers in this field to find the best technique as per their work and needs. It also makes recommendations or gives ideas to potential researchers on this subject for their future work.
2 Literature Review
3D imaging modalities such as CT, CBCT has certain drawbacks for a variety of reasons, as discussed above. As a result, several studies have been conducted to introduce a computational method that avoids these constraints yet provides precise 3D information. Various techniques, methodology, extracted features and reconstruction of an image are listed in Table 1 as available in the literature.
Initially, a comprehensive and comparative review on 3D medical imaging was presented by Stytz et al. [16] in 1991. Humbert et al. [17] presented an application of parametric models for spine reconstruction from biplane X-rays. The authors performed the two-level reconstruction. The first level estimates a fast 3D reconstruction of the bone. The second level was applied for obtaining fine adjustments to the model for precisely accurate 3d reconstruction. In the first level, the length of the spinal curve, the depth of the L1 and L5 endplates were chosen as the predictors for describing eight other parametric predictors of each vertebra. These eight specific measurements then infer the 19 important anatomical 3D points for each parametric vertebra. Approximately 2000 points cloud model were generated and then projected onto the 2D plane to view the 3D reconstructed model of the spine. In the second level, fine adjustment was performed by correcting the anatomical features using the control points of the vertebral body, which results in a parametric model to self-improve.
Chaibi et al. [18] used a parametric model to present a rapid 3D reconstruction method of femur bone from biplanar X-rays utilizing parametric and statistical models. The author obtained a simplified personalized parametric model (SPPM) using geometric features such as cylinders, 3D points, spheres, etc. (for a femur; the femoral neck-shaft angle (FNSA), the femoral head and posterior condyles representing spheres) and a full 3D morpho-realistic personalized parametric model (MPPM) of the lower limb obtained by correcting the SPPM. The Kriging methodology was used for global deformation and local adaptation. The as-rigid-as-possible deformation method was used for local adaptation which is based on the moving least-squares approach. Then 3D image reconstruction method was applied in two steps; a fast 3D lower limb reconstruction and a full 3D lower limb reconstruction. This was the first 3D modeling approach for the whole lower limb using biplanar X-rays with FT (femoral torsion) and TT (Angle between Tibial Plates Axis and the Bimalleolar Axis) calculations.
Cresson et al. [19] tried to overcome the drawback of overlapping regions to infer the information of location and shape of the hidden portions of the spine. The algorithm operates using two iterating processes. The first process uses a sophisticated 2D/3D registration procedure. Boundary edges were retrieved from radiographs to create a customized model of a vertebra. The second process refines the reconstruction of all quasi sections of the spine by registering estimates of anatomical features using a statistical model. The suggested technique can be a dependable alternative when compared with state-of-the-art technologies.
Convolution Neural Network (CNN) is an effective solution in a variety of applications viz; image processing [20, 21], image segmentation [22, 23], pattern recognition [24, 25] and computer vision [20]. The author Kasten et al. [26] addressed the issues of an absence of heuristic knowledge and dimensional expansion with conventional differential layers. It employed a dimensional augmentation technique where each pair of matching epipolar lines was back-projected into a two-channeled epipolar plane by bi-planar X-rays. A 3D structure was produced by combining an augmentation technique with a deep learning architecture. This method to generate 3D representations of the various bones maintains the geometric limits of the two views. To achieve a more robust, accurate and efficient result, domain adaptation is used.
Dixit et al. [27] used a machine learning approach for the construction of 3D models from 2D X-ray images. Features extracted are the color, size and depth of the femur bone. Depth information was used to create a mesh point cloud. The image was then converted into STL (Stereolithography) representation. A 3D model was created using CNN.
From X-ray projection images obtained in an upright position, the author Akkoul et al. [28] created a 3D customized model of the femur. The strategy was based on a two-stage pseudo-stereo matching process. The coordinates of a 3D contour are determined in the first stage using two sequential projections. The contours of the proximal femur are derived from the 2D X-ray images. The city-block, the Euclidian 2D spatial distances and the Chessboard were compared to match the points of two contours. Then, point pairs estimated are used to build a collection of three-dimensional points. The surface model was reconstructed using a meshing approach from the cloud of points based on Poisson's equation, resulting in a closed 3D surface. To compensate for the absence of information and increase the quality of the rebuilt 3D surface, a 3D reference surface of the proximal femur was employed in the second stage to inject extracted points locally into the reconstructed surface. Thus, in this technique of merging numerous 3D points created from a reference model obtained from a CT scan, the use of an X-ray stereo model in conjunction with a shape constraint improves accuracy and reduces error.
The statistical shape model (SSM) is a mathematical model incorporating information on the shape as well as its variance. Lamecker et al. [40] used a 3D Statistical shape Model to reconstruct the 3D shape from a few digital X-ray images. The author extracted the silhouettes from the projection using statistical shape training model thickness and the simulated X-rays. The approach optimizes a similarity measure analyzing the difference between the projections of the shape model and X-ray images by measuring the distance between the object silhouettes in the projections.
Zhu et al. [32] constructed 3D geometric surface models of the human knee joint. This work developed an enhanced SSM approach for predicting the 3D joint surface model that only utilizes 2D images of the joint. A total of 40 human knee distal femur models were used to create the SSM. A series validation and parametric analysis indicated that the SSM requires more than 25 distal femur models; each distal femur should be specified using at least 3000 nodes in space; and the 3D surface shape prediction should be based on two 2D fluoroscopic images obtained in 450 directions.
Zheng et al. [34] used a 2D/3D reconstruction technique based on a single image to create a scaled, patient-specific 3D surface model of the pelvis from a single standard AP X-ray radiograph. This single-image 2D/3D reconstruction method uses a hybrid 2D/3D deformable registration strategy that combines landmark-to-ray registration with SSM-based 2D/3D reconstruction.
Ehlke et al. [31] attempted to increase the reliability of the reconstruction process by considering as much information as possible about the anatomy of interest. Deformation of a volumetric tetrahedral mesh with density information generated by digitally reconstructed radiographic (DRR) deformations were projected to represent possible candidates for patient-specific shapes. Compare the X-ray attenuation of clinical X-rays with the pixel intensity of virtual X-rays to find the best candidate. This method transfers computations from the CPU to the GPU, providing interactive frame rates.
Gamage et al. [37] created a 3D reconstruction of the bone models using salient anatomical edges and contours computed from orthogonal radiographs. The method employs an iterative non-rigid 2D point matching methodology as well as thin-plate spline-based deformation. Noise, outliers, distortion and occlusions don't affect the non-rigid registration system. This method was unique in that it encompasses not only the exterior contours but also several significant inner edges to ensure adaptability to various bone anatomies and increase customization accuracy.
Gunay et al. [38] provided a cost and time-effective computational technique for generating a 3D bone structure from numerous X-ray images. When projected onto a two-dimensional (2D) plane, this technology scales and deforms a pre-set 3D template bone structure that is clinically normal and scaled to an average size until the distorted shape gives an image equivalent to an input X-ray picture. Sequential quadratic programming (SQP) was used to achieve multi-dimensional optimization by minimizing the error between the input X-ray image and the image projected from the deformed template shape.
Mahfouz et al. [39] provided a methodology for reconstructing lumbar vertebrae from orthogonal views to obtain precise contour projections. Reconstruction was performed by deforming the 3D bone model through a robust 3D-2D registration technique constrained through extracted 2D morphometric measurements utilizing biplanar X-ray images and statistical atlas of bone based on Principal Component Analysis (PCA).
Karade et al. [30] used Laplacian mesh deformation and self-organizing maps for an effective and precise 3D femur bone model reconstruction by reforming a 3D template mesh model to fit bone shape.
By gathering feature information, performing alignment, and SSM fitting using CNN, Kim et al. [29] have developed a superior technique for 3D reconstruction of leg bones using only front X-ray images. The bounding boxes are recognized by CNN. In situations where boundary outlines are difficult to extract, feature ellipses and feature points are used to recognize boundaries, which unifies feature information detection and saves time and money over manually defining feature information. It's also more exact and consistent than the manual assignment.
Koh et al. [33] used three CT images in combination with two X-ray images and the free form deformation method for 3D reconstruction of patient-specific femurs. It takes very little time to complete the bone segmentation of three CT images, the proposed reconstruction approach can be viewed as being similarly time and cost-effective as the reconstruction with X-ray images.
Mitton et al. [41] proposed an accurate 3D personalized model of the pelvis from biplanar X-rays. The method makes use of both stereo & non- stereo corresponding landmarks, anatomical atlas (prior knowledge) and contour identification. The landmarks identified are used to deform the fast computed initial solution. These retro projected landmarks on the radiographs are then best matched with the X-rays contours. Finally, utilizing the corresponding 2D/3D contours as constraints, an elastic 3D surface deformation mechanism based on the kriging method is used. The fact that several anatomical characteristics of the pelvis are not visible on the two radiographic views from which the reconstruction is performed is a limitation of the approach.
Three-dimensional reconstruction proved to be very effective and useful for preoperative planning and is becoming more important with time. Despite the number of techniques mentioned above in Table 1 in various literature, the choice of the method and technology clearly depends upon the requirement and the region reconstructed.
3 Imaging Requirements of 3D Reconstruction
Requirements of 3D reconstruction from X-ray images should be addressed while constructing 3D models, which are as follows:
-
i.
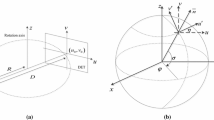
Acquisition of X-ray image(s): 3D reconstruction from 2D X-ray images can be performed using single X-ray images [40, 43, 44], two X-ray images [28, 30, 33, 37, 38] or more X-ray images for several anatomical regions such as femur, tibia, fibula, pelvis and spine. With an increase in the number of X-ray images, the information provided for the reconstruction will be increased. Figure 2 represents a conventional way of X-ray image acquisition. A portion of the X-ray beam emerging from the X-ray tube pass through the person’s body where they are absorbed by the internal structures and the remaining are transmitted to a detector. It is then use for further recording or processing by a computer.
-
ii.
Image Enhancement: X-ray images' are low in intensity, poor in contrast [47]. The quality of the X-ray image can be improved by applying image enhancement [48] to improve the assessment. Many approaches for improving the quality of X-ray images have been proposed such as histogram equalization (HE) [49], adaptive histogram equalization (AHE) [50], wavelet transform coefficients (WT) [51], and the unsharp masking method (USM) [52, 53]. Huang et al. [54] employed an adaptive median filter and a bilateral filter to suppress mixed noise, which includes both Gaussian and impulsive noise while keeping the image structures (edges). After that, gray-level morphology and contrast limiting histogram equalization (CLAHE) were used to increase the image contrast. The CLAHE approach enhances fine details, texture, and local contrast in images. Adaptive contrast enhancement (ACE) [55] is another well-known local enhancement method that uses Contrast gains (CG) to change the high-frequency components of images.
-
iii.
Immobilization during image acquisition: Immobilization means fixing a body part to reduce or eliminate the motion of a patient during the image acquisition process [5]. During image acquisition, it is necessary to sustain the rigid relationship between the original anatomy and the acquisition device. Zheng et al. [56] developed their new immobilization device including all the anatomical structures.
4 Steps of 3D Reconstruction from 2D X-ray Images
There are several types of approaches available for 3D reconstruction using X-ray images such as based on a single generic model [17, 41, 57,58,59], based on statistical shape and deformation models [60,61,62,63,64,65]. The former approaches deform a generic model to build a patient-specific 3D model, but the SSM-based methods build an SSM to produce the statistically probable models and decrease the number of parameters to optimize. Hybrid approaches [66] combine the SSM-based methods with generic model-based methods. Steps for doing 2D-3D reconstruction are as follows and are also shown in Fig. 3:
-
i.
2D X-rays Image acquisition: Input images for 3D reconstruction derived from X-ray images can be obtained by acquiring single X-ray images [40, 43, 44], two X-ray images [28, 30, 33, 37, 38] and more number of X-ray images for femur, tibia, fibula, pelvis and spine regions. The amount of information will increase in 3D reconstruction if the number of X-ray images increases.
-
ii.
Calibration: Due to the non-uniform magnification of the X-ray images, calibration is necessary for obtaining linear measurements. Calibration can be performed for a radiograph using the scale as available over the radiograph. The scale present on the top right corner of the craniofacial lateral image helps to calibrate the lateral image easily. These linear measurements are important for radiologists, for patients’ diagnosis and treatment planning. The measurement result can differ if the calibration is not performed correctly [5].
-
iii.
Contour Extraction (Segmentation): To extract the edges or contours from the input images, semi-automatic or manual [40, 58] solutions are available. Different methods have been offered by various authors [28, 34, 65, 72, 73] for precise contours extraction such as Thresholding based, Region-based segmentation, watershed segmentation, etc. A Canny edge detector can be applied to detect the edges. The active Contour method (ACM) provides smooth and closed contours which are suitable for medical images.
-
iv.
2D/2D correspondences: The process of identifying correspondences between all points or extracted contours of two or more X-ray images of the same scene is known as image correspondences. The point pair matching or correspondences could be developed by extracting the points of interest along with some descriptors and using some (dis)similarity measure over the descriptors [30]. Pruning the correspondences by finding the global minimum. Another method could be finding the iterative distance (Euclidean distance, city block distance, etc.) between each point to every point in the other image [28].
-
v.
Registration: Image registration is required in image analysis applications that include several images of a scene. It aligns images of a single scene captured from different angles together to get all the necessary information without replication. A rigid registration can be used to register images that are related by rotation, translation, or scaling. In the registration of medical images, a 2D/3D correspondence is built between a 3D model (SSM or Mesh model) and 2D X-ray images to obtain the best match. The features taken from 2D X-ray images and those recovered from 3D models led to the 2D/3D registration. In the Non-rigid image registration category, the images to be registered include geometric variations. SOM (Self-organizing method) is one of the non-rigid registration methodology that Ferrarini et al. [74] used successfully in their GAME approach.
-
vi.
Shape deformation: The last step is to deform the template in such a manner that when its silhouette vertices are projected on a 2D plane, its projection contours resemble the X-ray image contours. There are different types of techniques [75] available for deformation such as Kriging algorithm, Thin-plate splines deformation, Laplacian and Modified Laplacian [76], Torsional spring method [77, 78], etc.
5 Computational Approaches for 3D reconstruction from 2D X-rays
Models, anatomical knowledge, architectures and applications were used to classify 3D reconstruction approaches as shown in Fig. 4. The categorization of 3D image reconstruction methods are as follows:
5.1 Based on the Information Utilised for Reconstruction
Intensity-Based Methods: Intensity-based approaches evaluate the significance of pixel intensities in 2D radiographs including internal intensity distributions between anatomical boundaries. To capture the heterogeneity of the region of interest, volumetric density information is used in the representation of the 3D anatomical structures. 3D/2D registrations based on intensity rely entirely on data contained in voxels and pixels of 3D and 2D images, respectively. The coinciding points in this method are regarded as corresponding points and the similarity measure is determined via pixel-wise comparison instead of the distance. Mutual information, cross-correlation and a sum of square differences are the most commonly used similarity measures for DRR-based 3D/2D registrations. One such work of Yao et al. [79] has developed a deformable density atlas for bone anatomy and use it in a range of applications, the steps for generating atlas are as follows:
-
Construct Tetrahedral Mesh Models from extracted contours as it is simple, easy to deform and density information is also stored in the tetrahedron.
-
Apply an analytical density function to each tetrahedron instead of keeping the density value of each pixel in the model. As a result, computing the measurements such as integral, differentiating, interpolating and deforming is simple.
-
Once Tetrahedral Mesh Model is built, apply the deformation technique to it
5.2 Based on the Type of Model Used
Statistical shape modelling: 3D Model built by statistically analyzing a set of shapes is known as a SSM [32, 60, 63, 80, 81]. The mathematical model contains information on the shape and its changes. The SSM contains data on the shape geometry and variances. A Statistical shape model-based methods are more considered as automated reconstruction algorithms. These approaches need a large shape learning database. A Statistical shape modelling system learns from shape variations of an object or region. The statistical surface model approach was presented by Cootes et al. [82] and produced a deformable shape template. The approach integrates a priori knowledge about the geometrical shape and its morphological variability. Cootes et al. [82] introduced the SSM idea as point distribution models (PDMs), which are deformable representations that convey the mean shape as well as shape variations. PDMs are created by performing a PCA on the locations of landmarks that correlate to different shapes. PCA is used to determine the variations in the positions of the subject's paired points using a covariance matrix to create an SSM.
A covariance matrix (Cov) is constructed using the sample set (Si) as in Eq. (1):
where S is the population, \(\overline{S }\) is a population mean, p is the number of subjects in the sample set starting from I, (x, y, z) is the nodes of the sample set in three views and n is the node number of each model. Then the eigenvalues and the eigenvector of the (Cov) are computed as shown in Eq. (2).
Using the eigenvalues and eigenvectors, the PCA for subject models captures geometric features of the models. The first principal component represents the highest variation in all models based on direction and location. A new SSM surface \({S}^{\prime}\) can be obtained via eigen analysis in Eq. (3).
where M is a matrix, \(\overline{S }\) is a population mean, \({P}_{j}\) is an eigenvector, \({a}_{j}\) is a weight vector used to produce a new shape model (SSM), and \({M}_{i}\) is a transformation matrix used to shift the \({i}^{th}\) model in the SSM from its local coordinate system to the global coordinate system.
Articulated anatomical Models: Articulated statistical shape models (ASSMs) were first recommended by Heap et al. [83] as an extended version of the statistical shape model. It includes a structure of the joint and an analytical model of a joint to describe the degree of freedom of joint motion. It allows the model to determine both the shape variation of individual joint components as well as their usual range of motion separately. ASSMs are frequently used in the representation of bones from X-rays. ASSMs convey relative poses of bones with extra characteristics to represent diverse postures. An ASSM of the hip joint provided by Kainmueller et al. [84] used a ball-and-socket joint to link the relative transformations between the pelvis and the femur.
5.3 Based on the Method of Model Deformation
Free form deformation: Deformation techniques apply an adaptable deformation to a template model without taking into account any statistical data. A geometric approach called free-form deformation (FFD) [85] is used for basic deformations of rigid objects. It is built on the concept of enclosing a template inside a cube (lattice or hull) and altering it when the cube deforms. The control lattice is designed to be perpendicular to the model's principal ray. This idea is called hyper-patches, which are analogues of parametric curves like Bezier curves [86, 87], B-splines [88, 89], or NURBs [90, 91]. The template shape deforms until it produces an image similar to the X-ray image when projected onto the 2D plane. 3D template model within the lattice can be modified by changing the control volume parameters until its projection boundary fits the region contours retrieved from the X-ray images. The FFD-based technique demonstrated by Koh et al. [33] used volumetric data in the form of sparse CT data.
Hierarchical Deformation: The control lattice in hierarchical deformation [38] is hierarchically segmented in the regions. The distortion is applied to each region separately until the template projection resembles the X-ray image contours. Gunay et al. [38] demonstrated a technique of Hierarchical FFD. The bounding box is repeatedly split into smaller boxes, each of which was individually adjusted to get a form that is closest to the desired shape. Carl Shimer [92] describes the FFD block as a cubic structure with a hyper patch on each face. The vectors (S, T, and U) were used to represent the three sides. The FFD block was defined as an array of \(\left(3l +1\right)\times \left(3m +1\right)\times \left(3n +1\right)\) hyper patches. This is essentially a stack of \(1\times m\times n\) hyper.
-
To complete, create the lattice space which is a local parametric coordinate system within the FFD block. The lattice space is defined as follows in Eq. (4):
where \(X\left(s,t,u\right)\) is the local coordinate system's origin, while S, T, and U are the edges of the FFD block. Any point inside the lattice \(0<s<1, 0<t<1 and 0<u<1\) is valid. The following equation Eq. (5) is used to define the control points on the lattice:
where \({P}_{ijk}\) is a control points, (S, T and U) are the edges of the FFD block and (l, m and n) are the integer component.
-
After determining the points, the hyper patch deforms by moving the control points and \(l \times s, m \times t, n\times u\) are save as \(is, it, and iu\) to determine the location of the hyper patch within the FFD block.
-
Local coordinate of the system is determined by Eq. (6):
-
Calculating position of deformed lattice by putting back the \(\left(u,v,w\right)\) values into the hyper patch formula in Eq. (7):
where B(u) is a blending function, \({P}_{ijk}\) are three-dimensional control points,
The FFD transformations will apply to all neighboring regions that tend to deform the object. These approaches appear to be good alternatives, but they are not adequately useful in areas where the noise level is high and the useful data is difficult to distinguish. The FFD approach is constrained by the fact that it ignores the topology while deforming it.
5.4 Based on the Architecture Used
Convolutional Neural Network: CNN has lately demonstrated its effectiveness in a variety of applications, including image segmentation and classification. CNN for 3D reconstruction used by Kasten et al. [26] developed a dimensional expansion method that back-projects each pair of related epipolar lines into a two-channeled epipolar plane using bi-planar X-rays. This representation was combined with a deep learning architecture that generates 3D representations of a bone. The procedure followed by the Kasten et al. [26] was as follows:
-
For training purposes: Each pair of X-ray images was coupled with aligned ground truth. X-ray images were created by drawing DRRs from annotated CT scans.
-
For controlling the loss function of each training sample; a spatial 3D Distance Weight Map (DWM) with a size equal to the ground truth volume and its value on voxel i was determined by Eq. (8):
where d is a distance transform that describes the distance between each voxel and each bone surface, and γ = 8, σ = 10 were constants for all the training samples.
-
Then, DWM was used for weighting the voxel-wised cross-entropy loss.
-
An unsupervised reconstruction loss was specified to align the network prediction of the bone’s probability map with the input X-ray images.
-
The overall Loss function is Eq. (9):
CNN as a feature Detection: CNN could be used for automatic feature analysis [27]. The approach by Kim et al. [29] extends automation by doing automatic feature analysis of femur bone using CNN. The structure's position and shape can be simply determined in the X-ray image as feature elements. The femur's feature information compensates two proximal feature points, five distal feature points, and one ellipse, which is the distinctive shape of the femoral head. For feature information detection, two CNN-based modules were used: YOLO (You only look once) v3 [93] and Faster R-CNN [94] with ROI (Region of Interest) alignment. The image was fed into these two CNNs, which subsequently gives a bounding box as an output.
Genetic Algorithm (GA): The GA method is popular in the field of Artificial Intelligence because of its speedy search capability and resilience technique [95]. The algorithm operates in the form of a population, which evolves several viable solutions over a while (generations) until it finds the most suitable individual (solution) [96]. Mahfouz et al. [39] employ a genetic algorithm in conjunction with their 3D-2D score metric as an optimization technique to optimize the population, form and alignment of X-ray images for the reconstruction of lumbar and femur bone. The procedure used by Mahfouz et al. was as follows:
-
Automatic morphometric and surgical axis measurements were performed on the bone for the registration process to assess the shape and size of the bone.
-
An atlas bone model was aligned with the X-ray images that provide the initial position.
-
The initial population of bone models was created by combining the initial position with the average model.
-
Then, a genetic algorithm was used in conjunction with a pose score metric as a selection function along with crossover and mutation. It will improve the population's structure and alignment with the X-ray images.
-
Once the genetic algorithm has converged to a rigid alignment, the second optimization stage has reached.
-
The best member of the population was used as a genetic dopant.
-
After the alignment and registration; affine transformations (rotation, translation and scaling) are applied for deformation.
5.5 Hybrid Methods:
This method was derived through the combination of different approaches. Although these methods have some properties with the preceding methods, their qualities distinguish them as a separate class known as hybrid methods. This class's methods can have a variety of attributes depending on how they're combined and used. The Kadoury et al. [97] work is an example of a hybrid approach. He presented a method for biplanar spine reconstruction that integrated statistical and image-based methodologies. Their hybrid 3D reconstruction method merged statistical knowledge with information derived from images.
5.6 Based on the Anatomical Knowledge Used for Reconstruction
Point-Based Approaches: Point-based approaches work by recognizing and matching points on radiographs. Point-based methods are divided into SCP (Stereo-Corresponding Points) and NSCP (Non-Stereo Corresponding Points).
Stereo-Corresponding Point-Based Techniques: SCP technique is implemented by identifying stereo corresponding points (SCPs) in radiographs.
-
(a)
The first step is to find the points in two X-ray images that correlate to each other.
-
(b)
Then, rebuilt in 3D using methods such as Discrete Linear Transform (DLT).
The DLT approach can be used to find linear mappings between any two data sets if there are a certain amount of corresponding data points between them. The number of SCPs used determines the quality of the results. The more SCPs used, the better the results [98] but finding more SCP results is a slow process. Pearcy et al. [99] used SCP and generate 3D data using the DLT algorithm. Bony formations with no visible edges are not suited for SCP-based approaches. Aubin et al. [98] use this method for vertebra reconstruction utilizing more points (21 SCPs and 6 SCPs for each vertebra in each radiograph for comparison). The results were better with 21 SCPs but time-consuming. This approach is constrained by a number of matching anatomical landmarks on radiographs.
Non-Stereo Corresponding Point-Based Techniques: To overcome the limitation of the SCP-based approach, the NSCP based method (only points visible on a single radiograph which are having no correspondence with another image) is introduced as an improvement. This method is based on the idea that the NSCP is the part of the line connecting the X-ray source and the point projection in one view. Steps could be as follows:
-
(a)
The first step in the NSCP method is to calculate the initial solution. The anatomical area of the general model is specified.
-
(b)
The 2D contours are then manually identified on the radiograph. The original 3D model is used to generate 2D contours from each radiograph. The target surface is then projected on the 2D plane.
-
(c)
The 2D relationship between two given points is based on the distance between points and contour derivation, so 2D and 3D contours match. Then the initial response is optimized in the next step.
-
(d)
Finally, the optimal solution is transformed by applying the Kriging algorithm to it. The depth finding model used in the kriging algorithm as described by Keaomanee et al. [100] is depicted in Eq. (10) and Eq. (11), respectively:
where \({c}_{0}\) called nugget and \({c}_{1}\) called sill and range a are kriging parameters determined empirically and \(\delta \) is a semi-variance used to describe the degree of spatial dependency between two points (correspondence between 3D model point and 2D X-ray point).
-
(e)
Finally, the reconstructed shape is obtained by repeating the last step until the distance between the two given points is greater than the given accuracy value.
The NSCP-based approach for 3D spine reconstruction has been proposed by Mitton et al. [101]. Calibration was performed according to the detected spots. The Shape was then reconstructed in 3D (using the DLT method).
Parametric Based: Parametric modeling adjusts the shape of model geometry by changing the dimensional values. Instead of employing a whole set of points as in SSM, parametric models examine anatomical descriptive parameters (DPs) taken from the surface of interest [102]. Depending on the structure considered, the parametric model consists of several geometric primitives such as lines, spheres, points and circles. For example, parametric spine models consist of points and axes. Humbert et al. [17] suggested a parametric technique based on transverse and longitudinal inferences for the 3D reconstruction of lumbar and thoracic radiographs [103]. The procedure is stated as follows:
-
The parameters used are: length (curvature of the spine), depth, the width of the curve passing through the center of the vertebral body, the location along the spine curve of each vertebral endplate.
-
These are all then used to build a parametric model of the spine.
-
The parametric model was then back-projected on the X-ray plane to visualize the 3D geometry of the required structure.
Contour-Based Methods: Point-based methods cannot be employed for structures with continuous shapes such as the knee joint due to their time-consuming and non-reproducible nature. Instead of using points, a deformable generic model with matching contours is used. The non-stereo corresponding contour (NSCC) method's principle is to link identifiable 2D contours from radiographs to 3D contour. The steps are as follows:
-
Calculate the initial point to initiate contours segmentation manually or automatically.
-
Manually identify 2D outlines on radiographs of the anatomical area of interest from a generic model.
-
On the corresponding radiograph, 3D contours of the model surface are projected. Then, for each radiograph, 2D contours should be extracted from the 3D initial solution object.
-
After that, an association between these two set points is conducted. This 2D association is built on point-to-point distances and contours. This allows for the creation of a correlation between the 2D and 3D contours.
-
The initial solution is then optimized in the next stage.
-
The final step is to deform the optimum solution using the Kriging technique as described by Laporte et al. [57].
-
The reconstructed object is achieved by iterating as long as the distance between two specified points is greater than a predetermined accuracy value.
For 2D/3D reconstruction, non-statistical geometric parameters using the contour-based method have been proposed by Zeng et al. [104] and Karade et al. [30]. Karade et al. proposed a novel template reconstruction algorithm that preserves the local properties of the template shape during transformation. Karade et al. [30] used Laplacian surface deformation (LSD) to reconstruct 3D structure from two X-ray images (taken in Mediolateral (ML) and Anteroposterior (AP) directions). Compared to other deformation algorithms such as FFD or TPS (Splines in Sheet Metal), LSD is easier to build and takes less computation time. The input parameters used in the algorithm were:
-
Simulated contours of femur X-ray images derived from perspective ML (Medio Lateral) and AP (Anterior–Posterior) views.
-
Projections of the 3D model (derived from CT data of the femur).
-
The 3D model of the bone in the form of a 3D mesh with triangular elements from a clinically normal person. The model was a 3D point cloud and its projection was also a 2D image plane point cloud.
As an output, the template model will be rebuilt into a form that fits the input contours of the 2D image. There were three steps for reconfiguring the template model. The first stage involves aligning the template with the input contours. The second stage was to determine the 2D–3D correlation between input contour points (in both the ML and AP planes) and silhouette vertices. In the third step, SOM non-rigid registration was used to find the 2D-2D correspondences between projection contours of the model and the input contours to finally have the 2D-3D correspondences. The steps involved in the process as mentioned by the Karade et al. [30] are as follows:
“Let ‘K’ be the total number of input contours with \({p}_{k}^{c}\) be the \({k}^{th}\) input contour (k = 1, 2… K).
Calculate the distance between template projection contours and input contours as Eq. (12):
where M = total number of template projection points with (m = 1, 2… M;) and \({p}_{m}^{c}\) = mth template projection contour point and \({p}_{k}^{c}\)= \({k}^{th}\) input contour point.
The best match between template projection contours and input contours was considered as \({p}_{winner}^{p}\) (“nearest to the input contour”) was selected as Eq. (13):
Each template projection contour point was updated as Eqs. (14), (15), (16), (17) and (18):
where, \({t}_{frac} =current iteration number /total number of iterations\) and
and,
SOM output was the adapted template projection contour points (pp’) onto the input contour. The 2D–2D correspondence gives the required 2D–3D correspondence because the template projection contour points are directly associated with the silhouette vertices (projection)”. The template mesh was deformed using LSD as per the estimated 2D–3D correspondences. The template was deformed such that the projections of the template's silhouette on both image planes (ML and AP) have the same shape as the corresponding input contours. Laplacian surface deformation as a template deformation approach retains mesh topology and shape properties. As an output, the deformed mesh with its silhouette vertices updated to its desired final positions, so that its projection matches the shape of the input contours.
This approach can be applied to the problem of cavities that are not clearly seen when viewed perpendicular to the imaging direction. This approach by Karade et al. [30] may be used in any area of the skeleton.
6 Evaluation of 3D Reconstruction Methods of the Literature Surveyed
In terms of the aforementioned features, each recommended approach in this review paper has some benefits and drawbacks that should be evaluated before implementation. Not one method can be chosen as the best for all applications as every method has its pros and cons which make them useful as per the situation and need. Table 2 describes various 3D reconstruction methods for different structures along with their computation time, accuracy and validation technique used by authors as mentioned in the literature.
7 Challenges In 2d/3d Reconstruction Techniques:
Even though the reconstruction of a 3D model from 2D X-rays images has received a lot of attention in recent years. Yet, the suggested methods commonly suffer from one or more of the following practical challenges:
-
i.
Relevant Dataset: Relevant dataset related to the region of interest is not easily available. To evolve the 3D reconstruction method, one/multiple X-ray images along with CT/CBCT is required from the same patient. Multiple X-ray images and the CT/CBCT from the same patient are generally not available in clinics due to the nature of images i.e., radiation exposure to the patient and overlapping area through multiple images. To acquire such type of clinical data for perspective studies is not ethical. Therefore, this is a major challenge to work in this area.
-
ii.
Feature extraction: Extraction of contours from radiographs is a complex procedure typically performed manually [105] or semi-automatically [35, 40]. Manual segmentation of the anatomy is time-consuming, takes more effort and requires expertise. As the operator has to manually outline the contours of the anatomy on the radiograph. Semi-automatic segmentation requires an expert to mark the landmarks using the software.
-
iii.
Large Dataset: When using deep learning algorithms for 3D reconstruction, a bigger dataset of X-ray images for training is usually required which is not easily available [106, 107]. Mostly 3D/2D registration approach is based on DRRs which are simulated X-ray projection images because of the difficulty of getting a real dataset.
8 Discussion
Different authors have employed a variety of strategies for 3D reconstruction of the anatomy of interest utilizing one, two or more X-ray images. This review paper includes some of them that are often used for 3D reconstruction. This paper also includes the steps used for the 3D reconstruction and challenges encountered during reconstruction. Although researchers continue to develop new approaches, the core requirements for 3D reconstruction remain the same for all the techniques. 3D imaging modalities such as CT and CBCT rely solely on X-ray images. 3D image reconstruction from X-ray images can be used to overcome the drawbacks associated with 3D images like CT and CBCT such as cost and exposing the patient to higher radiation compared to the X-ray images. Some reconstruction approaches use single X-ray images only, while others use X-ray images along with silhouette-based models, mathematical models and deformation model techniques. Silhouette-based [108] reconstruction approaches are easier to use and are more reliable, that is why many researchers choose to utilize this approach in their work. 3D template deformation method uses shape information as a template model, which is then transformed using the contour information collected from the calibrated X-ray images. FFD has also emerged as an option for template deformation for developing 3D structure reconstruction applications. However, the FFD with few control points is not effective in preserving the fine features of the shape. Hierarchical FFD (for accurate shape matching) is a more complex and time-consuming approach compared to FFD, but it does not ignore the topology of the anatomy. Feature-based methods are mostly used compared to intensity-based methods; the features used are contours, points and parameters. The point-based procedures are dependent on the operator's expertise. On multi-view radiographs, it's difficult to precisely detect and match points. As a result, point-based approaches do not guarantee repeatability. Furthermore, due to the limited number of corresponding anatomical landmarks identifiable on radiographs, SCP based methods have limited accuracy when used. Point-based approaches cannot be employed for structures with continuous form because of the lack of anatomical landmark points. The time taken for 3D reconstruction using this procedure is roughly 2–4 h due to the necessary identification of points. Thus, these approaches are time-consuming. This would be a significant drawback that should not be overlooked, particularly in therapeutic applications and in severe cases. For a quick and reliable 3D reconstruction, parametric techniques are ideal. They increased the algorithms' resilience and convergence. This method results are improved by repeatability. The intensity-based method [60] is used to optimize the similarity between real radiographs and DRR. A volume-based statistical shape and intensity model (SSIM) developed by Ehlke et al. [31] was used to provide additional information on the volumetric bone density of the 3D model. Besides, deep learning methodologies are also nowadays gaining popularity among researchers. Deep learning algorithms learn by constructing a more abstract representation of data, the model extracts feature automatically and produces greater accuracy outcomes [106]. On a more precise level, the techniques are separated into deep learning, silhouette-based, single X-ray based, feature-based, intensity-based procedures and model deformation-based. Then there are hybrid approaches. If hybrid strategies are applied consistently, they can enhance the outcomes [109]. Concerning the above characteristics, each approach recommended by the author in the literature has some advantages and disadvantages to consider before implementation. It is not possible to choose the best one for all applications. Table 3 shows all these 3D reconstruction methods, each with its own set of benefits and limitations. It should be noted that some of the approaches discussed utilized the EOS system for image acquisition rather than typical radiography equipment. EOS (Electro-Optical System) device [110,111,112] can be an alternative between 2D radiographs and 3D CT scans at lower radiations but it is not readily available to all clinicians. Its cost is also dependent upon the anatomy of interest. This has an impact on the accuracy and speed of reconstruction approaches. If EOS systems are used, more time may be saved. Photogrammetry for building 3D models from radiography images has also been suggested as a potential alternative technique due to its accuracy and low irradiating dose. Various photogrammetry-based approaches for 3D image reconstruction from X-ray images have been proposed and evaluated in [113,114,115]. DRR images generated from the perspective projection of a three-dimensional image onto a two-dimensional plane can be used to create a relevant dataset of the anatomy of interest, but it does not guarantee the method's applicability in real scenarios. Less radiation dosage and low computation cost with little user supervision are important factors for the acceptability of 3D reconstruction approaches. 2D/3D reconstruction methods have the potential to take a place of CT or CBCT imaging in a non-severe case, if they didn't have access to CT or MRI equipment. The choice of this reasonably inexpensive and conveniently accessible method will be advantageous. Furthermore, a patient's implied risk of cancer can be decreased by reducing their exposure to ionizing radiation (compared to CT & CBCT). Reconstruction of 3D patient-specific models from X-ray images is, therefore, an essential and powerful approach in medical imaging and is worthy of further investigation.
9 Conclusion
This review critically analyses the various available approaches for the reconstruction of 3D anatomical structures from X-ray images. The pros and cons of each method were analyzed along with the steps of reconstruction. The techniques of 3D reconstruction can be used for any anatomical region of the human body provided the applicability and complexity of the anatomical region.
Even with the implementation of various methods, there is still scope to solve this challenging problem and improve the required precision in the measurements. CT and CBCT images are commonly used for obtaining a volumetric view of the anatomical structure. However, due to their difficulty in accessibility, high cost and high radiation exposure, robust methods are still required for the construction of 3D models from X-ray images to use in clinical analysis. Further research and analysis tools are required more specifically for each of the human anatomical region.
References
Gupta A, Kharbanda OP, Sardana V, Balachandran R, Sardana HK (2015) A knowledge-based algorithm for automatic detection of cephalometric landmarks on cbct images. Int J Comput Assist Radiol Surg 10:1737–1752. https://doi.org/10.1007/s11548-015-1173-6
Gupta A, Kharbanda OP, Sardana V, Balachandran R, Sardana HK (2016) Accuracy of 3d cephalometric measurements based on an automatic knowledge-based landmark detection algorithm. Int J Comput Assist Radiol Surg 11:1297–1309. https://doi.org/10.1007/s11548-015-1334-7
Henderson R (1995) The potential and limitations of neutrons, electrons and X-rays for atomic resolution microscopy of unstained biological molecules. Q Rev Biophys 28:171–193. https://doi.org/10.1017/S003358350000305X
Gupta A, Kharbanda O, Balachandran R, Sardana V, Kalra S, Chaurasia S, Sardana H (2017) Precision of manual landmark identification between as-received and oriented volume-rendered cone-beam computed tomography images. Am J Orthod Dentofac Orthop 151:118–131. https://doi.org/10.1016/j.ajodo.2016.06.027
Gupta A (2020) Challenges for computer aided diagnostics using X-ray and tomographic reconstruction images in craniofacial applications. Int J Comput Vis Robot 10:11. https://doi.org/10.1504/IJCVR.2020.10029170
Ehlke M. 3d reconstruction of anatomical structures from 2d X-ray images. Doctoral Thesis, Technische Universität Berlin
Tomazevic D, Likar B, Pernus F (2006) 3-d/2-d registration by integrating 2-d information in 3-d. IEEE Trans Med Imaging 25:17–27. https://doi.org/10.1109/TMI.2005.859715
Neelapu BC, Kharbanda OP, Sardana V, Gupta A, Vasamsetti S, Balachandran R, Rana SS, Sardana HK (2017) A pilot study for segmentation of pharyngeal and sino-nasal airway subregions by automatic contour initialization. Int J Comput Assist Radiol Surg 12:1877–1893. https://doi.org/10.1007/s11548-017-1650-1
Neelapu BC, Kharbanda OP, Sardana V, Gupta A, Vasamsetti S, Balachandran R, Sardana HK (2018) Automatic localization of three-dimensional cephalometric landmarks on cbct images by extracting symmetry features of the skull. Dentomaxillofac Radiol 47:1–12. https://doi.org/10.1259/dmfr.20170054
Neelapu BC, Kharbanda OP, Sardana HK, Gupta A, Vasamsetti S, Balachandran R, Rana SS, Sardana V (2017) The reliability of different methods of manual volumetric segmentation of pharyngeal and sinonasal subregions. Oral Surg Oral Med Oral Pathol Oral Radiol 124:577–587. https://doi.org/10.1016/j.oooo.2017.08.020
Tu JY, Inthavong K, Ahmadi G (2013) Computational fluid and particle dynamics in the human respiratory system. Springer. https://doi.org/10.1007/978-94-007-4488-2
(2017) What are the top 5 benefits of advanced medical imaging? https://www.trivitron.com/blog/what-are-the-top-5-benefits-of-advanced-medical-imaging/. Accessed 2 Mar 2017
What is diagnostic imaging? https://www.healthimages.com/what-is-diagnostic-imaging/
Li L, Wu W, Yan G, Liu L, Liu H, Li G, Li J, Liu D (2016) Analogue simulation of pharyngeal airflow response to twin block treatment in growing patients with class ii1 and mandibular retrognathia. Sci Rep 6:26012. https://doi.org/10.1038/srep26012
Huang R, Li X, Rong Q (2013) Control mechanism for the upper airway collapse in patients with obstructive sleep apnea syndrome: a finite element study. Sci China Life Sci. https://doi.org/10.1007/s11427-013-4448-6
Stytz MR, Frieder G, Frieder OJACS (1991) Three-dimensional medical imaging: algorithms and computer systems. ACM Comput Surv 23:421–499
Humbert L, De Guise JA, Aubert B, Godbout B, Skalli W (2009) 3d reconstruction of the spine from biplanar X-rays using parametric models based on transversal and longitudinal inferences. Med Eng Phys 31:681–687. https://doi.org/10.1016/j.medengphy.2009.01.003
Chaibi Y, Cresson T, Benjamin A, Hausselle J, Neyret P, Hauger O, de Guise J, Skalli W (2012) Fast 3d reconstruction of the lower limb using a parametric model and statistical inferences and clinical measurements calculation from biplanar X-rays. Comput Methods Biomech Biomed Eng 15:457–466. https://doi.org/10.1080/10255842.2010.540758
Cresson T, Chav R, Branchaud D, Humbert L, Godbout B, Aubert B, Skalli W, De Guise JA (2009) Coupling 2d/3d registration method and statistical model to perform 3d reconstruction from partial X-rays images data. Annu Int Conf IEEE Eng Med Biol Soc 2009:1008–1011. https://doi.org/10.1109/IEMBS.2009.5333869
Gupta A (2019) Current research opportunities of image processing and computer vision. Comput Sci 20:387–410. https://doi.org/10.7494/csci.2019.20.4.3163
Trivedi M, Gupta A (2022) A lightweight deep learning architecture for the automatic detection of pneumonia using chest X-ray images. Multimed Tools Appl 81:5515–5536. https://doi.org/10.1007/s11042-021-11807-x
Pandey M, Gupta A (2021) A systematic review of the automatic kidney segmentation methods in abdominal images. Biocybern Biomed Eng. https://doi.org/10.1016/j.bbe.2021.10.006
Ashok M, Gupta A (2021) A systematic review of the techniques for the automatic segmentation of organs-at-risk in thoracic computed tomography images. Arch Comput Methods Eng 28:3245–3267. https://doi.org/10.1007/s11831-020-09497-z
Maken P, Gupta A (2021) A method for automatic classification of gender based on text- independent handwriting. Multimed Tools Appl 80:24573–24602. https://doi.org/10.1007/s11042-021-10837-9
Maken P, Gupta A, Gupta MK (2019) A study on various techniques involved in gender prediction system: a comprehensive review. Cybern Inf Technol 19:51–73. https://doi.org/10.2478/cait-2019-0015
Kasten Y, Doktofsky D, Kovler I (2020) End-to-end convolutional neural network for 3d reconstruction of knee bones from bi-planar X-ray images. Machine learning for medical image reconstruction. Springer, Cham, pp 123–133
Dixit S, Pai VG, Rodrigues VC, Agnani K, Priyan SRV (2019) 3d reconstruction of 2d X-ray images. In: 2019 4th International Conference on Computational Systems and Information Technology for Sustainable Solution (CSITSS). pp 1–5
Akkoul S, Hafiane A, Rozenbaum O, Lespessailles E, Jennane R (2017) 3d reconstruction of the proximal femur shape from few pairs of X-ray radiographs. Signal Process 59:65–72. https://doi.org/10.1016/j.image.2017.03.014
Kim H, Lee K, Lee D, Baek N (2019) 3d reconstruction of leg bones from X-ray images using cnn-based feature analysis. In: 2019 International Conference on Information and Communication Technology Convergence (ICTC). pp 669–672
Karade V, Ravi B (2015) 3d femur model reconstruction from biplane X-ray images: a novel method based on Laplacian surface deformation. Int J Comput Assist Radiol Surg 10:473–485. https://doi.org/10.1007/s11548-014-1097-6
Ehlke M, Ramm H, Lamecker H, Hege HC, Zachow S (2013) Fast generation of virtual X-ray images for reconstruction of 3d anatomy. IEEE Trans Visual Comput Graphics 19:2673–2682. https://doi.org/10.1109/tvcg.2013.159
Zhu Z, Li G (2011) Construction of 3d human distal femoral surface models using a 3d statistical deformable model. J Biomech 44:2362–2368. https://doi.org/10.1016/j.jbiomech.2011.07.006
Koh K, Kim YH, Kim K, Park WM (2011) Reconstruction of patient-specific femurs using X-ray and sparse ct images. Comput Biol Med 41:421–426. https://doi.org/10.1016/j.compbiomed.2011.03.016
Zheng G (2010) Statistical shape model-based reconstruction of a scaled, patient-specific surface model of the pelvis from a single standard ap X-ray radiograph. Med Phys 37:1424–1439. https://doi.org/10.1118/1.3327453
Zheng G (2009) Statistical deformable model-based reconstruction of a patient-specific surface model from single standard X-ray radiograph. Springer, Berlin, pp 672–679
Zheng G (2009) Statistically deformable 2d/3d registration for accurate determination of post-operative cup orientation from single standard X-ray radiograph. Springer, Berlin, pp 820–827
Gamage P, Xie SQ, Delmas P, Xu P (2009) 3d reconstruction of patient specific bone models from 2d radiographs for image guided orthopedic surgery. In: 2009 digital image computing: techniques and applications. pp 212–216
Gunay M, Shim MB, Shimada K (2007) Cost- and time-effective three-dimensional bone-shape reconstruction from X-ray images. Int J Med Robot. https://doi.org/10.1002/rcs.162
Mahfouz M, Badawi A, Abdel Fatah E, Kuhn M, Merkl B (2006) Reconstruction of 3d patient-specific bone models from biplanar X-ray images utilizing morphometric measurements. In: Proceedings of the 2006 international conference on image processing, computer vision, & pattern recognition. Las Vegas, Nevada, USA, pp 26–29
Lamecker H, Wenckebach T, Hege H-C (2006) Atlas-based 3d-shape reconstruction from X-ray images. In: Proc Int Conf of Pattern Recognition (ICPR2006). IEEE Computer Society, pp 371–374
Mitton D, Deschênes S, Laporte S, Godbout B, Bertrand S, de Guise JA, Skalli W (2006) 3d reconstruction of the pelvis from bi-planar radiography. Comput Methods Biomech Biomed Eng 9:1–5. https://doi.org/10.1080/10255840500521786
Laporte A, Mitulescu D, Mitton J, Dubousset JAdG, W. Skalli (2001) 3d personalized geometric modeling of the pelvis using stereo x rays. In: VIIIth Congr Biomech. pp 186
Zheng G (2010) Statistically deformable 2d/3d registration for estimating post-operative cup orientation from a single standard ap X-ray radiograph. Ann Biomed Eng 38:2910–2927. https://doi.org/10.1007/s10439-010-0060-0
Novosad J, Cheriet F, Petit Y, Labelle H (2004) Three-dimensional (3-d) reconstruction of the spine from a single X-ray image and prior vertebra models. IEEE Trans Biomed Eng 51:1628–1639. https://doi.org/10.1109/TBME.2004.827537
Kabaliuk N, Nejati A, Loch C, Schwass D, Cater JE, Jermy MC (2017) Strategies for segmenting the upper airway in cone-beam computed tomography (cbct) data. Open J Med Imaging 07:196–219. https://doi.org/10.4236/ojmi.2017.74019
Jena M, Mishra S, Mishra D (2018) A survey on applications of machine learning techniques for medical image segmentation. Int J Eng Technol 7:4489–4495. https://doi.org/10.14419/ijet.v7i4.19005
Wang X, Wong BS, Guan TC (2005) Image enhancement for radiography inspection. SPIE
Koonsanit K, Thongvigitmanee SS, Pongnapang N, Thajchayapong PJtBEIC (2017) Image enhancement on digital X-ray images using n-clahe. In: 10th Biomedical Engineering International Conference (BMEiCON). pp 1–4
Ahmad SAB, Taib MN, Khalid NEA, Taib H (2012) Analysis of image quality based on dentists' perception cognitive analysis and statistical measurements of intra-oral dental radiographs. In: 2012 International Conference on Biomedical Engineering (ICoBE). pp 379–384
Zeng M, Li Y, Meng Q, Yang T, Liu J (2012) Improving histogram-based image contrast enhancement using gray-level information histogram with application to X-ray images. Optik 123:511–520. https://doi.org/10.1016/j.ijleo.2011.05.017
Öktem H, Egiazarian K, Niittylahti J, Lemmetti J (2003) An approach to adaptive enhancement of diagnostic X-ray images. EURASIP J Adv Signal Process 2003:635640. https://doi.org/10.1155/S1110865703211069
Sezn MI, Teklap AM, Schaetzing R (1989) Automatic anatomically selective image enhancement in digital chest radiography. IEEE Trans Med Imaging 8:154–162. https://doi.org/10.1109/42.24863
Deng G (2011) A generalized unsharp masking algorithm. IEEE Trans Image Process 20:1249–1261. https://doi.org/10.1109/TIP.2010.2092441
Huang R-Y, Dung L-R, Chu C-F, Wu Y-Y (2016) Noise removal and contrast enhancement for X-ray images. Br J Healthcare Med Res 3:56. https://doi.org/10.14738/jbemi.31.1893
Dah-Chung C, Wen-Rong W (1998) Image contrast enhancement based on a histogram transformation of local standard deviation. IEEE Trans Med Imaging 17:518–531. https://doi.org/10.1109/42.730397
Zheng G, Schumann S, Akcoltekin A, Jaramaz B, Nolte L (2016) Patient-specific 3d reconstruction of a complete lower extremity from 2d X-rays. https://doi.org/10.1007/978-3-319-43775-0_37
Laporte S, Skalli W, de Guise JA, Lavaste F, Mitton D (2003) A biplanar reconstruction method based on 2d and 3d contours: application to the distal femur. Comput Methods Biomech Biomed Engin 6:1–6. https://doi.org/10.1080/1025584031000065956
Le Bras A, Laporte S, Bousson V, Mitton D, De Guise JA, Laredo JD, Skalli W (2004) 3d reconstruction of the proximal femur with low-dose digital stereoradiography. Comput Aid Surg 9:51–57. https://doi.org/10.3109/10929080400018122
Yu W, Zheng G (2015) 2d-3d regularized deformable b-spline registration: Application to the proximal femur. In: 2015 IEEE 12th international symposium on biomedical imaging (ISBI). pp 829–832
Baka N, Kaptein B, de Bruijne M, Walsum T, Giphart J, Niessen WJ, Lelieveldt B (2011) 2d–3d shape reconstruction of the distal femur from stereo X-ray imaging using statistical shape models. Med Image Anal 15:840–850. https://doi.org/10.1016/j.media.2011.04.001
Ahmad O, Ramamurthi K, Wilson KE, Engelke K, Prince RL, Taylor RH (2010) Volumetric dxa (vxa): a new method to extract 3d information from multiple in vivo dxa images. J Bone Mineral Res 25:2744–2751. https://doi.org/10.1002/jbmr.140
Fleute M, Lavallée S (1999) Nonrigid 3-d/2-d registration of images using statistical models. Springer, Berlin, pp 138–147
Benameur S, Mignotte M, Parent S, Labelle H, Skalli W, de Guise J (2003) 3d/2d registration and segmentation of scoliotic vertebrae using statistical models. Comput Med Imaging Graph 27:321–337. https://doi.org/10.1016/s0895-6111(03)00019-3
Sadowsky O, Chintalapani G, Taylor RH (2007) Deformable 2d–3d registration of the pelvis with a limited field of view, using shape statistics. Springer, Berlin, pp 519–526
Schumann S, Liu L, Tannast M, Bergmann M, Nolte LP, Zheng G (2013) An integrated system for 3d hip joint reconstruction from 2d X-rays: a preliminary validation study. Ann Biomed Eng 41:2077–2087. https://doi.org/10.1007/s10439-013-0822-6
Zheng G, Gollmer S, Schumann S, Dong X, Feilkas T, González Ballester MA (2009) A 2d/3d correspondence building method for reconstruction of a patient-specific 3d bone surface model using point distribution models and calibrated X-ray images. Med Image Anal 13:883–899. https://doi.org/10.1016/j.media.2008.12.003
Çallı E, Sogancioglu E, van Ginneken B, van Leeuwen KG, Murphy K (2021) Deep learning for chest X-ray analysis: a survey. Med Image Anal 72:102125. https://doi.org/10.1016/j.media.2021.102125
Munawar F, Azmat S, Iqbal T, Grönlund C, Ali H (2020) Segmentation of lungs in chest X-ray image using generative adversarial networks. IEEE Access. https://doi.org/10.1109/ACCESS.2020.3017915
Cao F, Zhao H (2021) Automatic lung segmentation algorithm on chest X-ray images based on fusion variational auto-encoder and three-terminal attention mechanism. In: JÄNTSCHI DBL (ed), p 814. https://doi.org/10.3390/sym13050814
Chen H-J, Ruan S-J, Huang S-W, Peng Y-T (2020) Lung X-ray segmentation using deep convolutional neural networks on contrast-enhanced binarized images. Mathematics 8:545. https://doi.org/10.3390/math8040545
Xie L, Siqi C, Xie L, Chen G, Zhou H (2017) Development of a computer-aided design and finite-element analysis combined method for customized nuss bar in pectus excavatum surgery. Sci Rep. https://doi.org/10.1038/s41598-017-03622-y
Ozanian TO, Phillips R (2000) Image analysis for computer-assisted internal fixation of hip fractures. Med Image Anal 4:137–159. https://doi.org/10.1016/s1361-8415(00)00010-4
Behiels G, Maes F, Vandermeulen D, Suetens P (2002) Evaluation of image features and search strategies for segmentation of bone structures in radiographs using active shape models. Med Image Anal 6:47–62. https://doi.org/10.1016/s1361-8415(01)00051-2
Ferrarini L, Olofsen H, Palm WM, van Buchem MA, Reiber JH, Admiraal-Behloul F (2007) Games: growing and adaptive meshes for fully automatic shape modeling and analysis. Med Image Anal 11:302–314. https://doi.org/10.1016/j.media.2007.03.006
Selim M, Koomullil R (2016) Mesh deformation approaches – a survey. J Phys Math. https://doi.org/10.4172/2090-0902.1000181
Burg C (2006) Analytic study of 2d and 3d grid motion using modified Laplacian. Int J Numer Meth Fluids 52:163–197. https://doi.org/10.1002/fld.1173
Farhat C, Degand C, Koobus B, Lesoinne M (1998) Torsional springs for two-dimensional dynamic unstructured fluid meshes. Comput Methods Appl Mech Eng 163:231–245. https://doi.org/10.1016/S0045-7825(98)00016-4
Degand C, Farhat C (2002) A three-dimensional torsional spring analogy method for unstructured dynamic meshes. Comput Struct 80:305–316. https://doi.org/10.1016/S0045-7949(02)00002-0
Yao J, Taylor R (2000). Tetrahedral mesh modeling of density data for anatomical atlases and intensity-based registration. https://doi.org/10.1007/978-3-540-40899-4_54
Tang TS, Ellis RE (2005) 2d/3d deformable registration using a hybrid atlas. Med Image Comput Comput Assist Intervent 8:223–230. https://doi.org/10.1007/11566489_28
Fleute M, Lavallée S (1999) Nonrigid 3-d/2-d registration of images using statistical models. In: Proceedings of the second international conference on medical image computing and computer-assisted intervention. Springer, pp 138–147
Cootes TF, Taylor CJ, Cooper DH, Graham J (1992) Training models of shape from sets of examples. In: Hogg D, Boyle R (eds) BMVC92. Springer, London, pp 9–18
Heap T, Hogg DC (1995) Extending the point distribution model using polar coordinates. In: CAIP
Kainmueller D, Lamecker H, Zachow S, Hege HC (2009) An articulated statistical shape model for accurate hip joint segmentation. Annu Int Conf IEEE Eng Med Biol Soc 2009:6345–6351. https://doi.org/10.1109/iembs.2009.5333269
Jack D, Pontes JK, Sridharan S, Fookes C, Shirazi S, Maire F, Eriksson A (2019) Learning free-form deformations for 3d object reconstruction. Springer International Publishing, Lecture Notes in Computer Science. https://doi.org/10.1007/978-3-030-20890-5_21
Bézier PJCAGD (1974) Mathematical and practical possibilities of unisurf. pp 127–152
Böhm W, Farin G, Kahmann J (1984) A survey of curve and surface methods in cagd. Comput Aid Geometr Des 1:1–60. https://doi.org/10.1016/0167-8396(84)90003-7
Yu W, Tannast M, Zheng G (2017) Non-rigid free-form 2d–3d registration using a b-spline-based statistical deformation model. Pattern Recogn 63:689–699. https://doi.org/10.1016/j.patcog.2016.09.036
Zheng G, Yu W (2017) Chapter 12 - statistical shape and deformation models based 2d–3d reconstruction. In: Zheng G, Li S, Székely G (eds) Statistical shape and deformation analysis. Academic Press, New York, pp 329–349
Mohd Ali M, Jaafar NN, Abdul Aziz F, Nooraizedfiza Z (2014) Review on non uniform rational b-spline (nurbs): concept and optimization. Adv Mater Res 903:338–343. https://doi.org/10.4028/www.scientific.net/AMR.903.338
Sanchez-Reyes J (1997) A simple technique for nurbs shape modification. IEEE Comput Graphics Appl 17:52–59. https://doi.org/10.1109/38.576858
Shimer C. Free form deformation and extended free form deformation. https://web.cs.wpi.edu/~matt/courses/cs563/talks/freeform/free_form.html
Redmon J, Farhadi A (2018) Yolov3: an incremental improvement. Computer Vision and Pattern Recognition. ArXiv Prepr. arXiv:1804.02767v1
He K, Gkioxari G, Dollár P, Girshick R (2017) Mask r-cnn. In: 2017 IEEE International Conference on Computer Vision (ICCV). pp 2980–2988
Siddique MT, Zakaria MN (2010) 3d reconstruction of geometry from 2d image using genetic algorithm. In: 2010 international symposium on information technology. pp 1–5
Kabolizade M, Ebadi H, Mohammadzadeh A (2012) Design and implementation of an algorithm for automatic 3d reconstruction of building models using genetic algorithm. Int J Appl Earth Obs Geoinf 19:104–114. https://doi.org/10.1016/j.jag.2012.05.006
Kadoury S, Cheriet F, Labelle H (2009) Personalized X-ray 3-d reconstruction of the scoliotic spine from hybrid statistical and image-based models. IEEE Trans Med Imaging 28:1422–1435. https://doi.org/10.1109/tmi.2009.2016756
Aubin CE, Dansereau J, Parent F, Labelle H, de Guise JA (1997) Morphometric evaluations of personalised 3d reconstructions and geometric models of the human spine. Med Biol Eng Comput 35:611–618. https://doi.org/10.1007/bf02510968
Pearcy MJ (1985) Stereo radiography of lumbar spine motion. Acta Orthop Scand Suppl 212:1–45. https://doi.org/10.3109/17453678509154154
Keaomanee Y, Heednacram A, Youngkong P (2020) Implementation of four kriging models for depth inpainting. ICT Express 6:209–213. https://doi.org/10.1016/j.icte.2020.05.004
Mitton D, Landry C, Véron S, Skalli W, Lavaste F, De Guise JA (2000) 3d reconstruction method from biplanar radiography using non-stereocorresponding points and elastic deformable meshes. Med Biol Eng Compu 38:133–139. https://doi.org/10.1007/BF02344767
Quijano S, Serrurier A, Aubert B, Laporte S, Thoreux P, Skalli W (2013) Three-dimensional reconstruction of the lower limb from biplanar calibrated radiographs. Med Eng Phys 35:1703–1712. https://doi.org/10.1016/j.medengphy.2013.07.002
Pomero V, Mitton D, Laporte S, de Guise JA, Skalli W (2004) Fast accurate stereoradiographic 3d-reconstruction of the spine using a combined geometric and statistic model. Clin Biomech 19:240–247. https://doi.org/10.1016/j.clinbiomech.2003.11.014
Zeng X, Wang C, Zhou H, Wei S, Chen X (2014) Low-dose three-dimensional reconstruction of the femur with unit free-form deformation. Med Phys 41:081911. https://doi.org/10.1118/1.4887816
Dworzak J, Lamecker H, von Berg J, Klinder T, Lorenz C, Kainmüller D, Seim H, Hege HC, Zachow S (2010) 3d reconstruction of the human rib cage from 2d projection images using a statistical shape model. Int J Comput Assist Radiol Surg 5:111–124. https://doi.org/10.1007/s11548-009-0390-2
Sharma S, Kumar V (2022) 3d face reconstruction in deep learning era: a survey. Arch Comput Methods Eng. https://doi.org/10.1007/s11831-021-09705-4
Yuniarti A, Suciati N (2019) A review of deep learning techniques for 3d reconstruction of 2d images. In: 2019 12th International Conference on Information & Communication Technology and System (ICTS). pp 327–331
Mulayim AY, Yılmaz U, Atalay MV (2003) Silhouette-based 3-d model reconstruction from multiple images. IEEE Trans Syst Man Cybern B 33:582–591. https://doi.org/10.1109/TSMCB.2003.814303
Hosseinian S, Arefi H (2015) 3d reconstruction from multi-view medical X-ray images – review and evaluation of existing methods. The International Archives of the Photogrammetry, Remote Sensing and Spatial Information Sciences XL-1/W5: 319-326. https://doi.org/10.5194/isprsarchives-XL-1-W5-319-2015
Cresson T, Branchaud D, Chav R, Godbout B, de Guise J (2010) 3d shape reconstruction of bone from two X-ray images using 2d/3d non-rigid registration based on moving least-squares deformation. Progr Biomed Opt Imaging. https://doi.org/10.1117/12.844098
McKenna C, Wade R, Faria R, Yang H, Stirk L, Gummerson N, Sculpher M, Woolacott N (2012) Eos 2d/3d X-ray imaging system: a systematic review and economic evaluation. Health Technol Assess 16:1–188. https://doi.org/10.3310/hta16140
Kalifa G, Charpak Y, Maccia C, Fery-Lemonnier E, Bloch J, Boussard JM, Attal M, Dubousset J, Adamsbaum C (1998) Evaluation of a new low-dose digital X-ray device: first dosimetric and clinical results in children. Pediatr Radiol 28:557–561. https://doi.org/10.1007/s002470050413
Gajic D, Mihic S, Dragan D, Petrovic V, Anisic Z (2019) Simulation of photogrammetry-based 3d data acquisition. Int J Simulat Model 18:59–71. https://doi.org/10.2507/IJSIMM18(1)460
Zhang X, Li L, Chen G, Lytton R (2015) A photogrammetry-based method to measure total and local volume changes of unsaturated soils during triaxial testing. Acta Geotech 10:55–82. https://doi.org/10.1007/s11440-014-0346-8
Gesslein T, Scherer D, Grubert J (2017) Bodydigitizer: an open source photogrammetry-based 3d body scanner
Hufnagel H (2011) Current methods in statistical shape analysis. A probabilistic framework for point-based shape modeling in medical image analysis. Vieweg+Teubner, Wiesbaden, pp 7–25
Frysz M, Gregory JS, Aspden RM, Paternoster L, Tobias JH (2019) Describing the application of statistical shape modelling to dxa images to quantify the shape of the proximal femur at ages 14 and 18 years in the avon longitudinal study of parents and children. Wellcome Open Res 4:24–24. https://doi.org/10.12688/wellcomeopenres.15092.2
Malekzadeh M, Gul M, Kwon I-B, Catbas N (2014) An integrated approach for structural health monitoring using an in-house built fiber optic system and non-parametric data analysis. Smart Struct Syst 14:917–942. https://doi.org/10.12989/sss.2014.14.5.917
Oswald M, Toeppe E, Nieuwenhuis C, Cremers D (2013) A survey on geometry recovery from a single image with focus on curved object reconstruction. In: Proceedings of the 2011 conference on innovations for shape analysis: models and algorithms. pp 343–378
Prasad M, Fitzgibbon AW, Zisserman A (2005) Fast and controllable 3d modelling from silhouettes. In: Eurographics
Sintini I. Statistical shape and intensity modeling of the shoulder. University of Denver
Ourselin S, Roche A, Prima S, Ayache N (2000) Block matching: A general framework to improve robustness of rigid registration of medical images. Springer, Berlin, pp 557–566
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Maken, P., Gupta, A. 2D-to-3D: A Review for Computational 3D Image Reconstruction from X-ray Images. Arch Computat Methods Eng 30, 85–114 (2023). https://doi.org/10.1007/s11831-022-09790-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11831-022-09790-z