Abstract
Silene aucheriana Boiss. belongs to the Sect. Auriculatae (Boiss.) Schischk. in the Caryophyllaceae family, widely distributed in Iran. In this study, 75 random samples from 15 populations of different geographical and climate conditions in Iran were collected. Inter Simple Sequence Repeat (ISSR) technique has been used to analyse the genetic diversity in Iranian Silene aucheriana populations. Analysis of molecular variance (AMOVA) demonstrated 87% of the genetic variation exists among populations and 13% within the populations, which is in agreement with the high genetic differentiation among population diversity (Gst = 0.8), and a low level of gene flow (Nm = 0.07). Two distinct gene pools in the main distribution areas have been identified by the STRUCTURE and the K-Means analyses. Our results suggest that genetic divergence is present amongst populations that is most likely caused by their climatic conditions.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Silene L. is the largest genus in the Caryophyllaceae family with more than 800 species found around the world (Bittrich 1993; Greuter 1995). Silene subgeneric classification has been organized in different ways according to its great diversity and number of species (Greuter 1995; Oxelman and Lidén 1995; Oxelman et al. 2001). Initially Chowdhuri (1957) establishes 44 sections, then Melzheimer (1988) recognized 21 sections but more recently, based on molecular evidence the genus has been classified into three subgenera and 34 sections (Desfeux and Lejeune 1996; Oxelman et al. 1997, 2001; Eggens et al. 2007; Popp and Oxelman 2007; Petri and Oxelman 2011; Rautenberg et al. 2012; Naciri et al. 2017). This genus is perennial, stems erect up to 40 cm, the inflorescence dichasial panicles, stem leaves linear-lanceolate, calyx ovate to cylindrical, teeth triangular, and capsule ovate (Melzheimer 1988). In Iran, section Auriculatae (Boiss.) Schischk. is one of the largest sections with 57 species, of which 27 are endemic (Jafari et al. 2019). Recent molecular analyses of the Auriculatae section show that Silene aucheriana Boiss. belongs to this section (Jafari et al. 2019; Safaeishakib et al. 2020). This species is found in Anatolia, Iraq, Persia, and Turcomania (Melzheimer 1988) and distributed in the west, northwest, the Alborz slope, northeast, east, and central part of Iran (Chowdhuri 1957; Rohrbach 1868; Melzheimer 1988; Greuter 1995; Gholipour and Amini Rad 2017).
Inter-Simple Sequence Repeats (ISSR) has been reported as a useful tool for investigating genetic variation and population structure (Zietkiewicz et al. 1994; Wang et al. 2008; Baruah et al. 2013; Coutinho et al. 2014; Ferreira et al. 2015; Hilooğlu et al. 2016; Safaei et al. 2016). Moreover, it has been used effectively to estimate the genetic differentiation and phylogenetic relationships among four subspecies of S. scabriflora from the Iberian Peninsula using ISSR and cpSSR markers (Ferreira et al. 2015). This molecular method has also been used for the mating system to develop S. vulgaris cultivars of high quality with 14 accessions based on morphological and biochemical traits (Egea-Gilabert et al. 2013). Genetic diversity may appear spatially structured at different scales, such as among populations and neighboring individuals (Escudero et al. 2003). Analysis of spatial genetic structure is a valuable method for inferring evolutionary forces like random drift and selective pressures (De Kort et al. 2013). Populations of S. aucheriana are widely distributed in different regions of Iran with different ecological conditions and therefore, it is expected to show genetic differences. Regarding the ability of ISSR markers to detect minor genetic differences (Godwin et al. 1997) we used it to illustrate genetic structure and variation among 15 populations from different regions of Iran. The main goals of this paper were to (a) estimate the genetic diversity, (b) to investigate the inter- and intra-population genetic structure of Iranian S. aucheriana, and (c) assess the geographical distribution and similarity in climate conditions for studied populations.
Materials and methods
Plant material and sampling
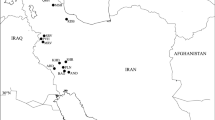
Plant descriptions from Flora Iranica (Melzheimer 1988) and Flora of Turkey (Coode and Cullen 1967) were used to identify the collected populations. The climatic conditions were used to select the population of interest for this study. For the analysis, 15 populations were divided into two groups according to their elevation (Table 1, Fig. 1). Populations with an elevation cut-off limit higher than 2800 m (mountainous area and dry cold) in one group and those with an elevation cut-off limit lower than 1800 m (semi-arid, and sub-humid) placed in another group. These cut-off limits were based on the mean elevations of localities. The assessment of these climate differences and classification was based on the Köppen-Geiger (KG) climate classification scheme (Peel et al. 2007), www.WorldClim.org, (Fick and Hijmans 2017), www.weatherunderground.com, and (Hijmans et al. 2005) definitions. These populations of S. aucheriana were selected from different climate conditions and different geographic locations. Such information would be useful not only for breeding but also for effective genetic diversity and conservation. Therefore, further characterization and identification of genetic diversities of the existing populations under different climate conditions would be beneficial for genetic diversity among and within the populations (El-Kassaby and Ritland 1996). Five specimens from each population were collected randomly at about 200 m intervals of each other, for a total of 75 specimens. Fresh leaves were collected randomly from plant specimens and dried in silica gel. Voucher specimens were deposited at the herbarium of Islamic Azad University, Tehran, Iran (IAUH). Meteorological data on localities including average temperature and rainfall were obtained from www.climate-data.org (Table 1).
Topographic map of the populations distribution of S. aucheriana in Iran showing the STRUCTURE plot based on k = 2. (Map designed using Arc GIS 10.2). Brown-colored segments reflect elevation cut-off limit higher than 2800 m, lower average temperature, and less rainfall (8.66 °C, 566.34 mm), whereas yellow-colored segments show different climate conditions with an elevation cut-off limit lower than 1800 m, higher average temperature, more average rainfall, (15.57 °C, 600 mm). Local names are presented in Table 1
DNA isolation and ISSR-PCR amplifications
Total genomic DNA was extracted from silica-dried leaves (approximately 0.5 g material per sample) using CTAB activated charcoal protocol (Doyle and Dickson 1987; Doyle and Doyle 1987). The quality and integrity of extracted DNA was determined by running on 1% agarose gel electrophoresis. High-quality DNA samples were stored at -20 °C until required. A set of 16 ISSR primers were selected according to Agarwal et al. (2008) and Sharma et al. (2008) studies. These primers include UBC-801 (AT)8 T, UBC-802 (AT)8G, UBC-803(AT)8C, UBC-804 (TA)8A, UBC-805 (TA)8C, UBC-806 (TA)8G, UBC-807 (AG)8 T, UBC-808 (AG)8C, UBC-809 (AG)8G, UBC-810 (CA)8 T, UBC-812 (GA)8A, UBC-818 (CA)8G, UBC-834 (AG)8YT, UBC-848 (CA)8RG, UBC-855 (AC)8YT, UBC-861 (ACC)6 commercialized by UBC (the University of British Columbia). All of these primers were preliminarily assayed on a limited number of individuals. This screening allowed us to identify eight primers producing more variable bands: UBC-807 (AG)8 T, UBC-808 (AG)8C, UBC-810 (CA)8 T, UBC-818 (CA)8G, UBC-834 (AG)8YT, UBC-848 (CA)8RG, UBC-855 (AC)8YT, UBC-861 (ACC)6.
Polymerase Chain Reaction (PCR) were carried out in a 25 μl volume containing 10 mM Tris–HCl buffer at pH 8.0 mM KCl; 1.5 mM MgCl2; 0.2 mM of each dNTP (Bioron, Germany); 0.2 μM of each primer; 20 ng genomic DNA and three U of Taq DNA polymerase (Bioron, Germany). The PCR amplification was performed with Techne thermocycler (Sensoquest, Göttingen, Germany) using the following program profile: Pre-heating at 94 °C for 5 min, 30 cycles of denaturation for 40 s at 94 °C, annealing for 1 min at 48 °C, the extension for 1 min at 72 °C, and 7 min final extension at 72 °C (Doyle 1991). As a standard for determining the size of the amplified fragments, the 100 bp DNA marker size was used. The quality of the amplification products was initially checked on 1% agarose gel. The PCR products of NED, PET, 6-FAM, and VIC labeled products were pooled (2 pools for 8 primers). 2 μl of each compound product was added to 7.75 μl of HiDi formamide (Applied Biosystems, Foster City, CA, USA) and 0.25 ll ROX-500 internal size standard (Applied Biosystems) were then injected to a 3730xl capillary sequencer (Applied Biosystems-Hitachi). Raw ISSR data were visualized with GENE MAPPER version 4.0 (Applied Biosystems, Foster City, CA, USA). Output ISSR files were aligned with the internal size standard using GeneMarker version 1.95 (GeneMarker, SoftGenetics, State College, Pennsylvania). Each peak with a signal intensity of more than 500 RFU (Relative Fluorescence Units) was scored as the present. The presence (1)/absence (0) binary matrixes of two primer sets were combined and prepared for analysis (Hulce et al. 2011).
Data analysis
The achieved ISSR bands were coded as binary datamatrix and used for genetic diversity analysis. Genetic diversity parameters including the average number of alleles per locus (Na), the number of effective alleles (Ne) (Kimura and Crow 1964), Shannon’s information index (I) (Lewontin 1972), average observed heterozygosity (Ho), average expected heterozygosity (He), number of bands (NB), the percentage of polymorphic loci (P%), unbiased heterozygosity (uHe) (Weising et al. 2005; Freeland et al. 2011) were calculated using GenAlEx 6.4 (Peakall and Smouse 2006). Additionally, the Nei’s genetic distance matrix (Nei 1973) were analysed using the POPGEN version 1.32 (Yeh et al. 1997). Moreover, population genetic differentiation of the studied genotypes was performed in GenoDive version 3 (Meirmans and Tienderen 2004). The correlation between the populations' genetic and geographic distance was estimated by the Mantel test (Mantel 1967) as implemented in PAST version 3.18 (Podani 2000; Hammer 2012) performing 5,000 permutations. Analysis of molecular variance (AMOVA) test (1000 permutations) (Excoffier et al. 1992) were performed using of GenAlex version 6.4 (Peakall and Smouse 2006). Furthermore, the genetic differentiation was determined using PhiPT, a measure that permits intra-individual variation to be suppressed and is therefore ideal for comparing codominant and binary data with 999 permutations (Assoumane et al. 2012). The Bayesian clustering approach implemented in STRUCTURE version 2.3.4 (Falush et al. 2007) was used to infer groups or subpopulations in the dataset. The analysis was performed with an admixture linkage model with correlated allele frequencies. To evaluate the discontinuous group (K) numbers, the K number was set between 1 and 15 with 10 replicates, and the length of the burn-in period was set to 100,000 times (Evanno et al. 2005; Zhong et al. 2019). The online STRUCTURE HARVESTER tool (Earl and vonHoldt 2012) was used to apply the Evanno method (Evanno et al. 2005) to detect the value of the optimal K that best fit the data. In K-Means clustering, two summary statistics, pseudo-F, and Bayesian Information Criterion (BIC), provide the best fit for k (Meirmans 2012). A Neighbor -joining (NJ) phylogenetic tree was constructed based on the Nei’s (1973) genetic distances for determining the genetic relationship among populations and principal coordinate analysis (PCoA) determined the relative genetic distance between individuals by using PAST version 3.18 (Hamer et al. 2001).
Results
Genetic diversity among the populations
The size of the amplified fragments ranged from 100 to 1700 bp. The UBC-807 primer amplified the largest number of loci (19) and the UBC-848 primer the smallest (11). The total number of loci amplified by the eight ISSR primers was 138, with an average of 15 loci per primer. For evaluating the genetic diversity of S. aucheriana populations a high number of reproducible and reliable bands from primers were achieved. The Tarom (Pop 13) population exhibited the highest number of private and common bands. While in the populations from Emamzadeh Hashem (Pop 1), Zangoleh (Pop 6), and Dizin (Pop 14) the value of the private bands became zero. The highest values of He (0.060), uHe (0.066), and Ho (0.041) were generated in the Klishom population (Pop 9). Whereas the lowest values of He (0.011), uHe (0.012), Ho (0.009), I (0.015), and Na (0.020) were achieved in Gorsfid (Pop 2) population. Furthermore, Silvana (Pop 7), and Klishom (Pop 9) had the highest values of (I = 0.088, Na = 1.112) respectively. The other genetic diversity parameters (NB = 57, Ne = 0.500) showed the highest value in the Tarom (Pop 13) population and the lowest values of (NB = 23, Ne = 0.196) were gained in the Dona population (Pop 11). Additionally, analysis of the present data displayed that the percentage of polymorphic loci (P%) ranged from (2.17%) in the Gorsfid population (Pop 2) to (16.67%) in the Silvana population (Pop 7) with a mean of (8.74%) (Table 2). The Genetic diversity within populations (Hs) and total expected heterozygosity (Ht) in the populations were (0.03, 0.27) respectively. A high level of genetic differentiation between populations (Gst = 0.87) with low and restricted gene flow (Nm = 0.07) confirmed divergence and differentiation have occurred among populations (Table 3).
Populational genetic structure
The Mantel test results reflected a highly significant correlation between genetic and geographic distances (r = 0.2356, p = 0.0002) after 5,000 replications. According to the pairwise population matrix of Nei’s genetic identity and the genetic distance determined among the studied populations show that the highest genetic distance (0.583) occurred between Gorsfid (Pop 2), and Tarom (Pop 13) populations while the shortest genetic distance and close affinity (0.165) happened between Tochal (Pop12), and Zangoleh (Pop 6) populations (Fig. 2). These analyses were consistent with the result of AMOVA which estimate that 87% of the genetic variation exists among populations and 13% within the populations. PhiPT values were statistically significant and were assessed by AMOVA demonstrating more contribution by interpopulation variation (PhiPT value = 0.868, p = 0.001) (Table 4). STRUCTURE utilizes a model-based clustering analysis that groups individuals into genetic clusters (K) therefore, populations were divided into two groups. Method of Evanno et al. (2005) was resulted in finding K = 2 (Fig. 3A). The STRUCTURE plot revealed the Tarom (Pop 13), Klishom (Pop 9), Heydareh (Pop 3), Dona (Pop 11), Silvana (Pop 7), Qoshchi (Pop 5), Gorsfid (Pop 2), Golestan Kooh (Pop 4) populations were grouped together while the other studied populations such as Zangoleh (Pop 6), Emamzadeh Hashem (Pop 1), Rineh (Pop 10), Gaduk (Pop 8), Hezar Masjed (Pop 15), Dizen (Pop 14), and Tochal (Pop12) were placed in the same group. However, it can be acknowledged that there is a little allelic combination in the Klishom population (Pop 9) (Fig. 3B). By considering the environmental patterns it can be noticed that Tochal (Pop12), Zangoleh (Pop 6), Emamzadeh Hashem (Pop 1), Rineh (Pop 10), Hezar Masjed (Pop 15), Gaduk (Pop 8), Dizin (Pop 14) populations are placed together and distributed in elevation cut-off limit higher than 2800 m with less average rainfall, and lower average temperature (566.34 mm, and 8.66 °C) respectively. In contrast, Tarom (Pop 13), Klishom (Pop 9), Heydareh (Pop 3), Dona (Pop 11), Silvana (Pop 7), Qoshchi (Pop 5), Gorsfid (Pop 2), Golestan Kooh (Pop 4) populations separated based on the similarity in climates with more average rainfall, higher average temperature (600 mm, 15.57 °C), and elevation cut-off limit lower than 1800 m (Fig. 1).
Population genetic structure of S. aucheriana. (A) Population genetic structure of the estimated delta K value with respect to K, according to the calculation method by Earl and vonHoldt (2012), graph showing the best value of K = 2. (B) Population structure estimated by STRUCTURE (k = 2)
Populations grouping based on genetic data
Further analysis with PCoA to ensure the correct grouping of populations was done along the first two axes. These first two axes (69.45% and 19.76%) explained 89.21% of the total variance (Fig. 4A). According to NJ phylogenetic tree and PCoA plot based on ISSR molecular data Hezar Masjed (Pop 15), Rineh (Pop 10), Emamzadeh Hashem (Pop 1), Tochal (Pop12), Zangoleh (Pop 6), and Dizin (Pop 14) populations were placed together in a closer distance. Also, Gorsfid (Pop 2), Golestan Kooh (Pop 4), Qoshchi (Pop 5), Dona (Pop 11), Tarom (Pop 13), Silvana (Pop 7), Heydareh (Pop 3) populations were placed in the different clusters (Fig. 4A, B). Klishom population (Pop 9) samples were placed near other populations which appears to show a slight gene flow in both populations which support by the PCoA and NJ approaches. The genetic cluster comprises all of the populations from different regions without geographical proximity.
A) Principal coordinates analysis (PCoA) using the Nei’s genetic distance computed from ISSR data. For an explanation of numbers, see Table 1. (B) Neighbor -joining (NJ) dendrogram based on Nei’s genetic distance showing relationships among the S. aucheriana populations Climate conditions were labeled by colors that reflect groupings derived from STRUCTURE analysis
Discussion
Populations of S. aucheriana have been reported from various regions of Iran in the present study. Thus, the investigation of genetic diversity should provide basic information about their current status in this country. According to the results, the value of gene flow was extremely low (Nm = 0.07). In the absence of a high gene flow dispersion of genetic drift mounting, decreasing the share of within-population gene diversity favors among-population genetic diversity intensifying the differentiation (Setsuko et al. 2007). Among the investigated populations Tarom (Pop 13) population (belonging to lower elevation) showed the highest genetic diversity in terms of the private allele while Emamzadeh Hashem (Pop 1), Zangoleh (Pop 6), and Dizin (Pop 14) populations (belonging to higher elevation) exhibited the least diversity, with no private alleles indicating a founder effect followed by genetic drift (Nei et al. 1975; Slatkin and Takahata 1985). The present study confirms molecular markers are the major technique to detect the variability and differences within and among populations at the DNA level (Freeland 2020). The Mantel test offered clear geographic inclination in the distribution of genetic variation. Therefore, displayed the isolation of populations by geographic distances in shaping the present genetic structure of S. aucheriana populations. This geographical isolation played a major role in the high genetic differentiation among populations (Slatkin 1987). One hypothesis is that there is a geographic trend of higher elevation populations (brown) that are closer to the Caspian Sea (NE) against the yellow populations that have a more southerly (SW) trend, except for the Gorsfid (Pop 2) population that breaks this trend and could indicate a recent introduction or a relict population of the yellow group. As Ferreira et al. (2015) concluded two subspecies of Silene scabriflora Brot. were closer to the sea; the Mediterranean and Atlantic coasts, and influenced by very specific and characteristic environments which can partially explain their divergence compared to the other subspecies. Similarly, As Brothers et al. (2016) have reported the population divergence in Silene latifolia Poir. was observed in two types of climate regions, which indicated differences in their morphology and phenology. Among-population differentiation in phenotypic traits and allelic variation is expected to occur as a result of isolation, drift, founder effects, and local selection (Jolivet and Bernasconi 2007). Consequently, based on the results conducted on European populations of S. latifolia, investigating molecular and genetic divergence is a pre-requisite for researches of local adaptation in response to selection under variable environmental conditions (Richards et al. 2003; Jolivet and Bernasconi 2007). Furthermore, new genotypes should be handled with care because of local adaptation to environmental and ecogeographic conditions (Vander Mijnsbrugge et al. 2010). According to the study by Karrenberg et al. (2018) based on environmental data containing 19 bioclimatic variables conducted on populations of Silene dioica (L.) Clairv. and S. latifolia, they found that ecological divergence has been the driver of speciation. Principally, climate conditions and some related parameters in a short time frame show the genetic composition of plants (Allard 1988; Perez de la Vega et al. 1994). It can also be expressed that breeding systems represent an important effect on the amount and distribution of genetic variation among and within populations (Charlesworth 2003). The importance assigned to marker characterization (particularly, to the influence of the environment on the type of character) has caused a diminution in the importance of morphological descriptors for genetic diversity (Egea-Gilabert et al. 2013). In addition, the levels of the breeding system (inbreeding or outbreeding) and pollination modes can seriously affect the genetic structure of the populations (Hamrick and Godt 1996). Subsequently, ecogeographic variables associated with genetic variation would help the study of future breeding programs (Granberg et al. 2008; Hansen 2003; Taylor et al. 1999). The importance of this issue is undeniable that genetic variation and population structure are strongly correlated with population size, systems of mating, natural selection, life form, ecological tolerance, dispersal of seeds, and also the pattern of gene flow (Colautti et al. 2010). Environmental factors have also been shown to influence flower size, and other reproductive traits (Worley et al. 2000; Delph et al. 2004). By providing more evidence for this research, it can be concluded that the divergence of populations has been revealed under the influence of two different types of climate. In our study, the clustering of each population was consistent with their climatic conditions, indicating genetic divergence along with their geographic distribution.
Conclusions
Present data provide information on the genetic diversity and population structure of S. aucheriana from different regions showing significant levels of genetic variation among populations. These results indicate that all individuals of populations shared two genetic pools as revealed by STRUCTURE. The geographic and genetic distance showed a significant correlation. Our results indicated two types of biogeographical groups have been formed (in terms of elevation) and what has been seen in relation to the climate data. In this regard, seven populations with an elevation cut-off limit higher than 2800 m were placed in one group, and the other eight populations with an elevation cut-off limit lower than 1800 m were placed together.
Data availability
The data are available from the corresponding author upon reasonable contact.
References
Agarwal M, Shrivastava N, Padh H (2008) Advances in molecular marker techniques and their applications in plant sciences. Plant Cell Rep 27:617–631. https://doi.org/10.1007/s00299-008-0507-z
Allard RW (1988) Genetic changes associated with the evolution of adaptedness in cultivated plants and their wild progenitors. J Hered 79:225–238. https://doi.org/10.1093/oxfordjour.jhered.3
Assoumane A, Zoubeirou AM, Rodier-Goud M, Favreau B, Bezancon G, Verhaegen D (2012) Highlighting the occurrence of tetraploidy in Acacia senegal (L.) Willd. and genetic variation patterns in its natural range revealed by DNA microsatellite markers. Tree Genet Genomes 9:93–106. https://doi.org/10.1007/s11295-012-0537-0
Baruah C, Laskar MA, Sharma DK (2013) Inter-simple sequence repeat (ISSR) polymorphismbased analysis of diversity in the freshwater turtle genus Pangshura. Afr J Biotechnol 12(3):238–248. https://doi.org/10.5897/AJB12.1273
Bittrich V (1993) Caryophyllaceae Juss. In: Kubitzki, K., Bittrich, V. Rohwer, J.G. (Eds.) The families and genera of vascular plants, vol. 2. Springer, Berlin, pp. 206–236. https://doi.org/10.1007/978-3-662-02899-5_21
Brothers A, Weingartner L, Delph L (2016) Genetically based population divergence of Silene latifolia from two climate regions. Evol Ecol Res 17:637–650
Charlesworth D (2003) Effects of inbreeding on the genetic diversity of populations. Philos 358:1051–1070. https://doi.org/10.1098/rstb.2003.1296
Chowdhuri PK (1957) Studies in the genus Silene. Notes from the Royal Botanical Garden Edinburgh 22:221–278
Colautti RI, Eckert CG, Barrett SCH (2010) Evolutionary constraints on adaptive evolution during range expansion in an invasive plant. Proc R Soc B 277:1799–1806. https://doi.org/10.1098/rspb.2009.2231
Coode M, Cullen J (1967) Silene L. Flora of Turkey and the East Aegean Islands Edinburgh Univ Press, Edinburgh, pp 179–242
Coutinho J, Carvalho A, Lima-Brito J (2014) Genetic diversity assessment and estimation of phylogenetic relationships among 26 Fagaceae species using ISSRs. Biochem Syst Ecol 54, 247– 256. https://doi.org/10.1016/J.BSE.2014.02.012
De Kort H, Vandepitte K, Honnay O (2013) A meta-analysis of the effects of plant traits and geographical scale on the magnitude of adaptive differentiation as measured by the difference between QST and FST. Evol Ecol 27:1081–1097. https://doi.org/10.1007/s10682-012-9624-9
Delph LD, Gehring JL, Frey FM, Arntz AM, Levri M (2004) Genetic constraints on floral evolution in a sexually dimorphic plant revealed by artificial selection. Evolution 58:1936–1946. https://doi.org/10.1111/j.0014-3820.2004.tb00481.x
Desfeux C, Lejeune B (1996) Systematics of Euro mediterranean Silene (Caryophyllaceae) evidence from a phylogenetic analysis using ITS sequences. Sciences De La Vie 319(4):351–358
Doyle JJ, Dickson EE (1987) Preservation of plant samples for DNA restriction endonuclease analysis. Taxon 36:715–722. https://doi.org/10.2307/1221122
Doyle JJ, Doyle JL (1987) A rapid DNA isolation procedure for small quantities of fresh leaf tissue. Bull Phytochem 19:11–15 Doyle JJ, Dickson EE (1987) Preservation of plant samples for DNA restriction endonuclease analysis. Taxon 36:715– 722. https://doi.org/10.2307/1221122
Doyle J (1991) DNA protocols for plants. In: Hewitt, G., Johnston, A.W.B. & Young, J.P.W. (eds.) Molecular techniques in taxonomy. Springer Heidelberg, Berlin, pp. 283–293. https://doi.org/10.1007/978-3-642-83962-7_18
Earl DA, VonHoldt BM (2012) STRUCTURE HARVESTER: a website and program for visualizing STRUCTURE output and implementing the Evanno method. Conserv Genet Res 4:359–361
Eggens F, Popp M, Nepokroeff M, Wagner W, Oxelman B (2007) The origin and number of introductions of the Hawaiian endemic Silene species (Caryophyllaceae). Am J Bot 94:210–218. https://doi.org/10.3732/ajb.94.2.210
Egea-Gilabert C, Ninirola E, Conesa M, Fernandez J (2013) Agronomical use as baby leaf salad of Silene vulgaris based on morphological biochemical and molecular traits. Sci Hort 152:35–43. https://doi.org/10.1016/j.scienta.2013.01
El-Kassaby YA, Ritland K (1996) Genetic variation in low elevation Douglas-fir of British Columbia and its relevance to gene conservation. Biodivers Conserv 5:779–794. https://doi.org/10.1007/BF00051786
Escudero A, Iriondo JM, Torres ME (2003) Spatial analysis of genetic diversity as a tool for plant conservation. Biol Conserv 113:351–365
Evanno G, Regnaut S, Goudet J (2005) Detecting the number of clusters of individuals using the software STRUCTURE: a Simulation study. Mol Ecol 14(8):2611–2620. https://doi.org/10.1111/j.1365
Excoffier L, Smouse PE, Quattro JM (1992) Analysis of molecular variance inferred from metric distances among DNA haplotypes: application to human mitochondrial DNA restriction data. Genetics 131:479–491
Ferreira V, Castro I, Rocha J, Crespí A, Pinto-Carnide O, Amich F, Carnide V (2015) Chloroplast and nuclear DNA studies in Iberian Peninsula endemic Silene scabriflora Subspecies using cpSSR and ISSR markers Genetic diversity and phylogenetic relationships. Biochem Syst Ecol 61:312–318. https://doi.org/10.1016/j.bse.2015.06.029
Fick SE, Hijmans RJ (2017) WorldClim 2: New 1-km spatial resolution climate surfaces for global land areas: New climate surfaces for global land areas. Int J Climatol 37(12):4302–4315. https://doi.org/10.1002/joc.5086
Falush D, Stephens M, Pritchard JK (2007) Inference of population structure using multilocus genotype data: dominant markers and null alleles. Mol Ecol Notes 7(4):574–578. https://doi.org/10.1111/j.1471-8286.2007.01758.x
Freeland JR, Petersen SD, Kirk H (2011) Molecular ecology, 2nd edn. Wiley, Chichester
Freeland JR (2020) Molecular ecology, 3rdedn. Wiley-Blackwell, London, p 363
Gholipour A, Amini Rad M (2017) Silene capitellata and S. retinervis (Caryophyllaceae) new records for Iran. Iran J Bot 23(2): 91–97. https://doi.org/10.22092/ijb.2017.1157.1184
Godwin ID, Aitken EAB, Smith LW (1997) Application of intersimple sequence repeat (ISSR) markers to plant genetics. Electrophoresis 18(9):1524–1528. https:// doi. org/10. 1002/elps. 1150180906
Granberg A, Carlsson-Graner U, Arnqvist P,Giles BE (2008) Variation in breeding system traits within and among Populations of Microbotryum violaceum on Silene dioica. Int J PlantSci 169: 293–303.https://doi.org/10.1086/523964
Greuter W (1995) Silene (Caryophyllaceae) in Greece, A subgeneric and sectional Classification. Taxon 44:543–558. https://doi.org/10.2307/1223499
Hamer Y, Harper D, Ryan P (2001) PAST: Paleontological statistics software package for ducation and data analysis. Palaeontol Electron 4:9
Hansen T (2003) Quantitative genetics of sexual allocation and inbreeding depression in hermaphroditic plants with special reference to Silene nutans (Caryophyllaceae). PhD Thesis, University of Copenhagen
Hamrick J, Godt M (1996) Effects of life history traits on genetic diversity in plant species. Philosophical Transactions of the Royal Society of London. Proc Biol Sci 351:1291–1298. http://doi.org/https://doi.org/10.1098/rstb.1996.0112
Hammer Ø (2012) PAST: paleontological statistics version 2.17, reference manual. atural History Museum, University of Oslo, Oslo
Hijmans RJ, Cameron SE, Parra JL, Jones PG, Jarvis A (2005) Very high resolution nterpolated climate surfaces for global land areas. Int J Climatol 25:1965–1978. https://doi.org/10.1002/joc.1276
Hilooğlu M, Poyraz I, Poyraz I, Ataşlar E, Sözen E ( 2016) Genetic relationships among some Turkish Petrorhagia (Ser.) Link (Caryophyllaceae) taxa using ISSR markers. Phytotaxa 272: 165–172. https://doi.org/10.11646/phytotaxa.272.2.8
Hulce D, Li X, Snyder-Leiby T, Johathan Liu CS (2011) GeneMarkerR Genotyping software: tools to increase the statistical power of DNA fragment analysis. J Biomol Tech 22:S35–S36
Jafari F, Mirtadzadini M, Gholipour A, Rabeler R, Oxelman B, Zarre S (2019) Notes on the genus Silene (Caryophyllaceae, Sileneae) in Iran. Phytotaxa 425 (1): 35–48. https://doi.org/10.11646/phytotaxa.425.1.3
Jolivet C, Bernasconi G (2007) Molecular and quantitative genetic differentiation in European populations of Silene latifolia (Caryophyllaceae). Ann Bot 100:119–127. https://doi.org/10.1093/aob/mcm088
Karrenberg S, Liu X, Hallander E, Favre A, Herforth-Rahmé J, Widmer A (2018) Ecological divergence plays an important role in strong but complex reproductive isolation in campions (Silene). Evol Int j Org Evol 73:245–326. https://doi.org/10.1111/evo.13652
Kimura M, Crow JF (1964) The number of alleles that can be maintained in a finite population. Genetics 49:725–738
Lewontin RC (1972) Testing the theory of natural selection. Nature 236:181–182. https://doi.org/10.1038/236181a0
Mantel N (1967) The detection of disease clustering and a generalized regression approach. Cancer Res 27:209–220
Meirmans PG, Van Tienderen PH (2004) Genotype and Genodive: two programs for the analysis of genetic diversity of asexual organisms. Mol Ecol Notes 4:792–794. https://doi.org/10.1111/j.1471-8286.2004.00770.x
Meirmans PG (2012) AMOVA-based clustering of population genetic data. J Hered 103:744–750. https://doi.org/10.1093/jhered/ess047
Melzheimer V (1988) Silene L. in: Rechinger, K. H. (ed.), Flora Iranica, 163 Graz, pp 341–508
Naciri Y, Du Pasquier P, Lundberg M, Jeanmonod D, Oxelman B (2017) A phylogenetic circumscription of Silene sect. Siphonomorpha (Caryophyllaceae) in the Mediterranean Basin. Taxon 66(1): 91–108. https://doi.org/10.12705/661.5
Nei M (1973) Analysis of gene diversity in subdivided populations. PNAS (12): 3321–3323.https://doi.org/10.1073/pnas.70.12.3321
Nei M, Maruyama T, Chakraborty R (1975) The bottleneck effect and genetic variability of populations. Evolution 29:1–10. https:// doi. org/ 10. 1111/j. 1558- 5646. 1975. tb008 07.x
Oxelman B, Lidén M (1995) Generic boundaries in the tribe Sileneae (Caryophyllaceae) as inferred from nuclear rDNA sequences. Taxon 44:525–542. https://doi.org/10.2307/1223498
Oxelman B, Lidén M, Berglund D (1997) Chloroplast rps16 intron phylogeny of the tribe Sileneae (Caryophyllaceae). Pl Syst Evol 206:39–410. http://dx.doi.org/https://doi.org/10.1007/BF00987959
Oxelman B, Lidén M, Rabeler RK, Popp M (2001) A revised generic classification of the tribe Sileneae Caryophyllaceae). Nord J Bot 20:515–518. https://doi.org/10.1111/j.1756-1051.2000.tb00760.x
Peakall R, Smouse P (2006) GenAlEx 6: genetic analysis in Excel. Population genetic software for teaching and research. Mol Ecol Notes 6(1): 288–295
Peel MC, Finlayson BL, McMahon TA (2007) Updated world map of the Köppen-Geiger climate classification. Hydrol Earth Syst Sci 11:1633–1644
Perez de la Vega M, Saenz Miera LE, Allard RW (1994) Ecogeographical distribution and differential adaptedness of multilocus allelic associations in Spanish oat Avena sativa L. Theor Appl Genet 88:56–64. https://doi.org/10.1007/BF00222394
Petri A, Oxelman B (2011) Phylogenetic relationships within Silene (Caryophyllaceae) section Physolychnis. Taxon 60:953–968. https://doi.org/10.1002/tax.604002
Podani J (2000) Introduction to the exploration of multi-variate data English translation. Backhuyes Publishers, p 407
Popp M, Oxelman B (2007) Origin and evolution of North American polyploidy Silene Caryophyllaceae). Am J Bot 94:330–349. https://doi.org/10.3732/ajb.94.3.330
Rautenberg A, Sloan D, AldenV OB (2012) Phylogenetic relationships of Silene multinervia and Silene section Conoimorpha (Caryophyllaceae). Syst Biol 1:226–237. https://doi.org/10.1600/036364412X616792
Rohrbach P (1868) Monographie Der Gattung Silene. Leipzig, Wilhelm Engelmann, 249 pp. [in German] https://doi.org/10.5962/bhl.title.15462
Richards CM, Emery SN, McCauley DE (2003) Genetic and demographic dynamics of small populations of Silene latifolia. Heredity 90:181–186
Safaei M, Sheidai M, Alijanpour B, Noormohammadi Z (2016) Species delimitation and genetic diversity analysis in Salvia with the use of ISSR molecular markers. Acta Bot Croat 75(1):45–52. https://doi.org/10.1515/botcro-2016-0005
Safaeishakib M, Assadi M, Mehregan I, Ghazanfar S (2020) Phylogenetic study of Silene sections Auriculatae, Spergulifoliae, Ampullatae, and Lasiocalycinae in Iran. Phytotaxa 472 (2):169–183. https://doi.org/10.11646/phytotaxa.472.2.7
Setsuko S, Ishida K, Ueno S, Tsumura Y, Tomaru N (2007) Population differentiation and gene flow within a metapopulation of athreatened tree, Magnolia stellata (Magnoliaceae). Am J Bot 94:128–136. https://doi.org/10.3732/ajb.94.1.128
Sharma K, Agrawal V, Gupta S, Kumar R, Prasad M (2008) ISSR marker-assisted selection of male and female plants in a promising dioecious crop: jojoba (Simmondsia chinensis). Plant Biotechnol Rep 2:239–243. https://doi.org/10.1007/s11816-008-0070-7
Slatkin M, Takahata N (1985) The average frequency of private alleles in a partially isolated population. Theor Popul Biol 28:314–331
Slatkin M (1987) Gene flow and the geographic structure of natural-populations.Science 236: 787–792.
Taylor D, Trimble S, McCauley D (1999) Ecological genetics of gynodioecy in Silene vulgaris: relative fitness of females and hermaphrodites during the colonization process. Evolution 53:745–751. https://doi.org/10.1111/j.1558-5646.1999.tb05368.x
Vander Mijnsbrugge K, Bischoff A, Smith B (2010) A question of origin: Where and how to collect seed for ecological restoration. Basic Appl Ecol 11:300–311. https://doi.org/10.1016/j.baae.2009.002
Wang C, Zhang H, Qian Z, Zhao G (2008) Genetic differentiation in endangered Gynostemma pentaphyllum Makino based on ISSR polymorphism and its implications for conservation. Biochem Syst Ecol 36:699–705. https://doi.org/10.1016/j.bse.2008007.004
Weising K, Nybom H, Wolff K, Kahl G (2005) DNA fingerprinting in plants. Principles, methods, and applications, 2nd edn, Boca Rayton, Fl., CRC Press, USA, p 472
Worley AC, Baker AM, Thompson JD, Barrett S (2000) Floral display in Narcissus: variation in flower size and number at the species, population, and individual levels. Int J Plant Sci 161:69–79. https://doi.org/10.1086/314225
Yeh FC, Yang R, Boyle TB, Ye ZH, Mao JX (1997) POPGENE, the user friendly shareware for population genetic analysis molecular biology and biotechnology Centre. University of Alberta, Canada, p 10
Zhong Y, Yang A, Li Z, Zhang H, Liu L, Wu Z, Li Y, Xu LT, M, Yu F, (2019) Genetic diversity and population genetic structure of Cinnamomum camphora in South China revealed by EST-SSR markers. Forests 10:1019. https://doi.org/10.3390/f10111019
Zietkiewicz E, Rafalski A, Labuda D (1994) Genome fingerprinting by simple sequence repeat (SSR)-anchored polymerase chain reaction amplification. Genomics 20:176–183. https://doi.org/10.1006/geno.1994.115
Acknowledgements
Masoumeh Safaeishakib would like to thank Dr. Abbas Gholipour to check the identification of collected populations.
Funding
This work was supported by Islamic Azad University, Tehran, Iran Science and Research Branch for providing the facilities necessary to carry out the research.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Consent for publication
Consent for publication submission is approved by all authors.
Ethical approval
The article does not contain any studies with human participants or animals performed by any of the authors.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Safaeishakib, M., Assadi, M., Ghazanfar, S.A. et al. Cryptic molecular-geographical divergence among Iranian Silene aucheriana populations inferred from ISSR and climatic data. Biologia 79, 1225–1235 (2024). https://doi.org/10.1007/s11756-023-01390-x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11756-023-01390-x