Abstract
Glycine tomentella Hayata is a species of Glycine Willd. subgenus Glycine, and in China it is distributed along the southeast coast. In this study, 11 natural G. tomentella populations were collected and their genetic diversity levels and population structures were analyzed using 25 simple sequence repeat (SSR) markers. The number of alleles per locus averaged 7.16 and ranged from 2 to 17. The expected heterozygosity (He) per locus averaged 0.60, varying from 0.19 to 0.86. The G. tomentella populations on these Chinese islands showed a greater average genetic variation (60.96%) among populations and gene differentiation index (Gst= 0.607), and a lower average within-population genetic variation (33.47%) and gene flow (Nm= 0.162). In this study, these G. tomentella island populations were characterized by a relatively greater average multilocus outcrossing rate of 5.74%, which may the result of heterogeneity owing to the perennation of G. tomentella. A spatial autocorrelation analysis revealed that populations within a radius of approximately 30.45 km had positive and significant genetic relationships. The Neighbor-Joining (NJ) and STRUCTURE analyses strongly showed a pattern of ‘island differentiation’ for the populations on southeast islands of China and also suggested that some genetic interconnection occurred along the southeast coast of China. The F-statistics suggested that geographically different G. tomentella populations had specific population structures. We propose that when collecting this species as a genetic resource, every G. tomentella population should be sampled.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
The genus Glycine Willd. is widely distributed in Australia, the western Pacific Islands, and the surrounding islands, including Taiwan (Chung and Singh 2008). The genus Glycine contains two subgenera, termed subgenus Soja, which contains two annual species [the cultivated soybean Glycine max (L.) Merr. and its progenitor wild soybean Glycine soja Sieb. et Zucc.], and subgenus Glycine, which contains 26 perennial species. Species Glycine tomentella Hayata is one of these perennial species and has the complex karyotype containing 2n = 38, 40, 78, or 80 (Singh et al. 1987). To date, G. tomentella has been identified as having several genomes. The 2n = 38 G. tomentella contains the EE genome (D1 and D2 isozyme types); 2n = 40 G. tomentella contains the DD (D3 isozyme type), D2D2 (D5A) and H2H2 (D5B) genomes; 2n = 78 G. tomentella contains the AE (T5), D3E (T1) and EH2 (T6) genomes; and 2n = 80 G. tomentella contains the DA6 (T2), DD2 (T3) and DH2 (T4) genomes (Doyle and Brown 1985; Doyle et al. 1986; Brown et al. 2002; Chung and Singh 2008). A T7 isozyme type was found in an Indonesian tetraploid G. tomentella (2n = 80) (Kollopara et al. 1994). In phylogenetic studies of G. tomentella, Tindale (1986) classified the original G. tomentella (2n = 40, D6 isozyme type) as a species of Glycine arenaria Tindale (genome H), while Pfeil et al. (2006) recognized the original G. tomentella (2n = 40, D4 isozyme type) as a new species, Glycine syndetika B. E. Pfeil et Craven (genome A6). Moreover, one of the T2-isozyme typed G. tomentella (2n = 80, D1A, or DA6) was regarded as the species G. dolichocarpa Tateishi and Ohashi (2n = 80, D1A) (Tateishi and Ohashi 1992; Chung and Singh 2008).
Glycine tomentella is also recorded in the flora of China (Hayata 1920); however, no relevant taxonomic studies on genome classifications of these species have been performed. Two chromosome-based observational studies of G. tomentella from several places in Fujian Province, China showed that the number of chromosomes was 80 (Bau et al. 1993; Gao et al. 2002), but the genome and isozyme types of the Chinese 80-chromosome G. tomentella remain unclear.
As a wild relative of cultivated soybean, G. tomentella can enhance the genetic basis of soybean breeding and, therefore, is an important germplasm for soybean innovation (Singh et al. 1990, 1993, 1998, 2010; Singh and Nelson 2015). G. tomentella is also used as a Chinese herbal medicine for treating rheumatism and bone pain (Zhang et al. 2011). However, there have been limited studies regarding the genetic diversity of the subgenus Glycine’s perennial species. Currently, only one small-scale study of the ISSR marker-based genetic diversity in 16 G. tomentella individuals collected from three small areas in China has been performed (Chen et al. 2013). Therefore, there is a lack of geographical and genetic information on the Chinese G. tomentella complex species.
Presently, subgenus Glycine plants growing in China are being threatened from coastal exploration, nuclear power plant construction, and tourism, with the number and area of existing surviving populations declining. To effectively protect genetic resources of G. tomentella species in China, the genetic diversity and structure of their natural populations are required. Here, the objective was to investigate the genetic diversity and structure of the natural G. tomentella species’ populations on the southeast islands of China to establish a genetic conservation strategy.
Materials and methods
Material sampling
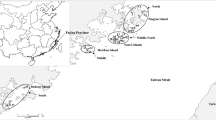
Leaves were sampled from on-the-spot field plants as the experimental materials. The leaves were stored with silica gel in zip-lock plastic bags for later analyses. In total, 291 G. tomentella individuals from 11 populations were collected on five coastal islands, Nanao, Dongshan, Jingan, Meizhou and Pingtan, along the southeast coast of China (Table 1 and Fig. 1). The geographical coordinates of each population were recorded (Table 1). The sampling distance between individuals within populations was more than 5 m, and 14–30 plants were sampled from each population, depending on population size.
DNA extraction and SSR genotyping
Total genomic DNA was extracted using a modified CTAB method (Narzary et al. 2015). Zou et al. (2004) shows that the use of soybean SSR makers is a rapid and reliable method to detect G. tomentella chromosomes. In the species having less genome sequence information, we selected usable soybean SSRs identified from 930 markers (Cregan et al. 1999; Song et al. 2010), and 25 soybean SSR markers with polymorphisms and high levels of stability were used to assess the genetic diversity of G. tomentella. The rest of 905 SSR markers were unavailable (nil-amplified, single amplification, smeared, or uncountable multi-bands). PCR amplification was carried out on a TaKaRa TP650 PCR thermal cycler (TaKaRa Bio, Japan) with a 10-μL total volume of reaction mixture, containing 2 μL of template DNA (30 ng/μL), 5 μl of 2 × Taq PCR StarMix (Genstar Biosolusiond Co., Ltd., Beijing, CN), 0.5 μL of each primer (10 mM), and 2 μL of ddH2O. PCR amplification reactions were performed with an initial denaturation at 95 °C for 5 min, 34 cycles of 30 s denaturation at 94 °C, 30 s annealing at 54 °C (the annealing temperature and cycle number varied according to different SSR primers used), and 30 s extension at 72 °C. This was followed by 10 min at 72 °C for the final extension, and samples were stored at 4 °C. The PCR products were electrophoresed on an 8% non-denaturing polyacrylamide gel and visualized by silver nitrate staining. Because most of the markers exhibited more than one band, we chose the brightest bands, clearly legible and polymorphic, and recorded them as alleles.
Data analysis
The software POPGENE1.32 (Yeh et al. 1999) was used to calculate genetic diversity parameters, including the number of alleles (Na), number of effective alleles (Ne), expected heterozygosity (He), observed heterozygosity (Ho), percentage of polymorphic loci (P), mean number of alleles per locus (A), polymorphism information content (PIC), genetic differentiation index (Gst), and gene flow (Nm), for each locus. The number of genotypes (haplotypes) was estimated using Power Marker (Liu and Muse 2005). The software FSTAT, version 2.9.3 (Goudet 2001) was used to estimate the fixing index (Fis) for each locus and population. The outcrossing rate (t) was calculated from t = (1 − Fis)/(1 + Fis) (Weir and Cockerham 1984). The F-statistics evaluating the genetic differentiation among populations and the molecular variance (AMOVA), and evaluating the among- and within-population genetic variation amounts, were calculated using ARLEQUIN, version 3.5 (Excoffier and Lischer 2010). The relationship dendrogram was constructed using the Neighbor-Joining (NJ) method based on the allele frequencies and genetic distance (Nei et al. 1983), with a bootstrap number of 1000 and a 0.95 confidence interval (CI), using PowerMarker, version 3.25 (Liu and Muse 2005). The online software IBDWS, version 3.23 (Jensen et al. 2005) (http://ibdws.sdsu.edu/~ibdws/aboutibdws.html) was used to detect the correlation between genetic and geographic distances among populations based on the Mantel test. STRUCTURE, version 2.1 (Pritchard et al. 2000) was run for the population structure analysis using a model without prior population information. Values of 1–11 K (the number of assumed clusters) were used to infer the number of clusters. Each run was conducted 20 times with independent simulations for each K value under the conditions of a 100,000 burn-in period and 100,000 MCMC (Markov Chain Monte Carlo method) replications. Furthermore, the optimal K value was inferred by the online program Structure Harvester (Earl and vonHoldt 2012) (http://taylor0.biology.ucla.edu/structureHarvester). Based on the optimal K values, the program CLUMPP, version 1.1.2 (Jakobsson and Rosenberg 2007) was used to find the optimal alignments of 20 independent runs produced by STRUCTURE. The spatial autocorrelation of the overall geographical populations was analyzed using software GenAlEx, version 6.502 (Peakall and Smouse 2012). The autocorrelogram was set using variable distance classes that spanned the full range of geographic distances among the populations. An assignment test was applied to infer the possible foreign individuals in a population using Geneclass 2 (Piry et al. 2004).
Results
Performance of soybean SSR markers
In this study, 25 pairs of SSR markers could be amplified with polymorphisms, and they produced 179 bands (alleles) in the 291 G. tomentella individuals. The mean Na was 7.16 per loci, ranging from 2 (Satt153, Satt504 and TAB 16) to 17 (sat_142). The number of genotypes (haplotype) was 11.32 per locus, ranging from 3 (Satt153, Satt504 and TAB 16) to 25 (Satt563). The mean He and PIC per locus showed similar trends. The mean He value was 0.595 per locus, with a range of 0.186 (Satt421) to 0.862 (Satt339 and Satt563), and the mean PIC value was 0.562 per locus, with a range of 0.174 (Satt421) to 0.846 (Satt339 and Satt563) (Table 2). The fixation index (Fis) for the degree of allelic fixation was 0.842 per loci, ranging from 0.649 (Sat_421) to 1.00 (Sat_142) among loci.
Population genetic variation
In the 11 populations, the mean P was 58.6% for populations, ranging from 24% (pop. 7) to 100% (pop. 4). These populations averaged 13.82 haplotypes, ranging from 3 (pop. 7) to 30 (pop. 3). The mean A for populations was 2.18, ranging from 1.24 (pop. 7) to 4.28 (pop. 3). The He per population was 0.232, ranging from 0.020 (pop. 7) to 0.545 (pop. 3). The average PIC was 0.203 for the populations, ranging from 0.02 (pop. 7) to 0.50 (pop. 3). Among these populations, pops-3, 4, and 9 had the greatest levels of genetic variation, as characterized by their greater genetic parameters (P = 88.0–100.0, Hap = 19–30, A = 3.400–4.280, He = 0.408–0.545, Ho = 0.080–0.156 and PIC = 0.369–0.497, Table 3). Additionally, they usually occupied greater land areas (Table 1).
Relatively greater outcrossing rate
Outcrossing can cause genetic recombination and heterogeneity in species and populations. The outcrossing rate is different among plant species or categories. This is related to the species’ nature and is also affected by geographical distance, weather, and pollinators. Single locus (ts) and multiloci (tm) outcrossing rates were estimated for populations (Table 3). The ts showed a mean of 8.2% per locus, ranging from 0 (sat_142) to 18.6% (sat_421). The tm exhibited a mean of 5.74% per population, ranging from 0 (pops. 1, 7) to 16.7% (pop. 3). This value seemed to be greater for self-pollinating plants. The mean Fis across loci in populations was 0.897. Three populations, pops. 2, 3, and 4, had lower Fis values (0.772–0.787) and two populations, pops. 1 and 7, were fixed (Fis= 1.00) in the loci. Only two populations, pops. 1 and 7, had no occurrence of outcrossing, suggesting that there was genetic recombination within most populations.
Population genetic differentiation
Population genetic differentiation in G. tomentella species on the southeast islands of China was estimated. It was characterized by an average Gst of 0.607 per locus among populations, ranging from 0.393 (sat_262) to 0.755 (sat_111) (Table 2). The number of loci with a Gst value over 0.5 was 21, implying that an overwhelming majority of the loci had greater among-population variation levels.
The AMOVA analysis showed that 60.96% of genetic variation in the G. tomentella species on the southeast islands occurred among populations (d.f. = 10; P < 0.0001), while 33.47% of genetic variation existed within populations (d.f. = 280; P < 0.0001) (Table 4). The result was in accordance with the Gst value, indicating a greater among-population genetic variation than within-population genetic variation in this G. tomentella species.
F-statistics for population differentiation showed significant differences between populations (Table 5), suggesting that all of the populations had their own genetic structures, even those at shorter distances (197 m between pops. 8 and 9; 408 m, pops. 10 and 11; 436 m, pops. 6 and 7; and 602 m, pops. 5 and 6; Table 5).
Population structure and spatial distribution
The structure analysis showed that there were two sharp peaks at k = 2 and k = 5 (Fig. 2). When k = 2 (Fig. 3a, b), group-1 included pops. 1, 2, 8, 10, 11 and a majority of pops. 3 and 4, and a minority of pop. 9 and group-2 included pops. 5, 6, 7 and a minority of pops. 3 and 4, and a majority of pop. 9. The results by STRUCTURE analysis could be understood as having two geographical groups, i.e. northeastern group (pops. 8, 10 and 11) and southwestern group (pops. 5, 6 and 7) (Fig. 1). The southwestern pops. 1 and 2 were only genetically consanguineous to the northeastern group; pops. 3 and 4 on Dongshan Island and pop. 9 on Meizhou Island were consanguineously mixed populations, being assigned to the northeastern and southwestern groups, respectively, for their individual plants.
Genetic structure analysis under a model without prior population information for the 11 natural G. tomentella populations on the islands off the southeastern coast of China. When k = 2, there were two genetic structural groups (a, b); when k = 5, all plants were assigned to five genetic structural groups just as island populations, group-1 (pops. 10, 11), group-2 (pops. 1, 2), group-3 (pops. 5, 6, 7), group-4 (pops. 3, 4) and group-5 (pops. 8, 9) (c, d)
When k = 5 (Fig. 3c, d), all 291 G. tomentella plant samples were assigned to five genetic structural groups just as island populations (Fig. 1), group-1 (pops. 10 and 11, Pingtain Island), group-2 (pops. 1 and 2, Nanao Island), group-3 (pops. 5, 6 and 7, Jingan Island), group-4 (pops. 3 and 4, Dongshan Island) and group-5 (pops. 8 and 9, Meizhou Island), which genetically demonstrated ‘island differentiation’. However, most populations exhibited geographical genetic exchange; pops. 5 and 9 were genetically more complicated (Fig. 3d), and included genetic composition from the southwest and northeast.
The NJ clusters based on Nei et al.’s (1983) genetic distance revealed that the 11 populations were first clustered as the sampling islands geographically (Fig. 4), suggesting that there was a genetically stronger island differentiation effect. A closer genetic kinship could be observed between geographically far northeastern and southwestern regions (Nanao, Meizhou and Pingtan Islands) (Fig. 4b), suggesting that there was a genetic connection between the northeastern and southwestern G. tomentella.
Neighbor-Joining clustering of the 11 G. tomentella populations based on allele frequencies and Nei et al.’s (1983) distance with 1000 bootstrap replicates (0.95 confidence interval). The island populations of Chinese G. tomentella had a genetically stronger island differentiation pattern (a). Genetic kinship appeared to exist between the northeastern and southwestern regions (Nanao, Meizhou, and Pingtan Islands) (b)
A Mantel test was performed to determine whether genetic distances were related to the geographic distances among G. tomentella populations. A significant correlation (r = 0.295*, P < 0.001) existed between the two, and the regression analysis (y) also showed that the correlation decreased as the geographical distances increased between populations (Fig. 5). The spatial autocorrelation analysis showed a positive correlation within 40.59 km, and a significant positive correlation within 30.45 km (Fig. 6).
Spatial autocorrelogram for G. tomentella’s geographical distribution on the southeast islands of China. A positive correlation occurred within a radius of 40.59 km, and a significantly positive correlation occurred within a radius of 30.45 km. The r indicates correlation coefficient; U and L indicate upper and lower 95% confidence intervals of the null hypothesis, respectively
Discussion
Distribution of perennial G. tomentella species in China
Four subgenus Glycine species have been reported to exist in Taiwan, i.e. G. tabacina (Labill.) Benth. (2n = 80, BB1, BB2 B1B2), 80-chromosome typed G. tomentella Hayata (2n = 80, DA6; DH2), G. pescadrensis Hayata (2n = 80, AB1), and G. dolichocarpa Tateishi et Ohashi (2n = 80, D1A) (Pfeil et al. 2006; Tateishi and Ohashi 1992; Chung and Singh 2008). Thus far, only one 80-chromosome typed G. tomentella has been reported to grow on the southeast islands of China (Bau et al. 1993; Gao et al. 2002), while the 38-chromosome, 40-chromosome and 78-chromosome G. tomentella have not been found. Our partial cytological analyses showed that the G. tomentella plants from the southeast islands of China were also 80-chromosomes, but that their genome types remain unknown. Geographically, the southeast islands of China were close to Taiwan. Since genome-DA6, and DH2 typed G. tomentella (2n = 80) exists in Taiwan, the G. tomentella that appears on the southeast islands of China could also belong to the DA6, or/and DH2 genomes (Chung and Singh 2008), this remains to be confirmed in our future research. However, based on morphological comparison, particularly pods, the G. tomentella on the islands of southeastern China greatly differs from the G. dolichocarpa in Taiwan, (Pfeil et al. 2006; Tateishi and Ohashi 1992; Chung and Singh 2008).
Our field survey in 2013–2017 found that the distribution range of G. tomentella in China was contoured by about 114°44′07″–119°52′57″E, for the west–east limit, on the eastern Guangdong coast (Dayawa Bay) to the southeastern Fujian coast (Pingtan Island) and the westernmost site was located in Dayawan (22°35′18″N, 114°44′07″E). Here, we only sampled the natural populations on the islands (Fig. 1), but Pingtain and Meizhou Islands also had distributions of Glycine tabacina species. Another Chinese perennial species, G. tabacina has a small geographical range, being confined to the southeast coast of Fujian Province, whereas G. tomentella has a more extensive geographical range, owing to its stronger environmental adaptability. Chinese G. tomentella has strong drought and salt tolerance levels, and prefers sunny locations. It grows in a wide variety of environments, such as roadsides, sand dunes, grass, thickets, hillsides, rocky hills and graveyards on the islands, but it rarely grows in shady places, tall grasses, or little shrubs. Commonly, G. tomentella has stoloniferous stems, with plant heights of about 10–200 cm, procumbent or creeping along the ground. In the field, G. tomentella grows commonly prostrate along the ground, and rarely twists or climbs on accompanying plants.
Does the creeping growth habit reduce genetic diversity in Chinese G. tomentella compared with the sympatric G. tabacina (Labill.) Benth.?
In general, the natural G. tomentella populations had a relatively lower mean genetic diversity (P = 58.6%, He= 0.232 and PIC = 0.203) than the G. tabacina populations (P = 78.57%, He= 0.272 and PIC = 0.237) in this sympatric region (all the analytic results about G. tabacina will appear elsewhere). The causes of this lower genetic diversity level in G. tomentella than in G. tabacina is unknown. However, G. tomentella and G. tabacina grow in the same ecological environments in the same region, but the two species exhibited differences in genetic diversity levels. The creeping growth habit of G. tomentella beneath the companion species, which was inferior for its Nm, may influence seed dispersal and lead to the lower genetic diversity in comparison with that of the sympatric G. tabacina.
The implications of a relatively high outcrossing rate and lower N m in Chinese G. tomentella
Commonly, self-pollinating plants have a lower outcrossing rate, such as annual wild soybean (G. soja), which has an outcrossing rate of less than 4% (Kiang et al. 1992; Fujita et al. 1997; Kuroda et al. 2006; Wang and Li 2012). Compared with the sympatric perennial G. tabacina in the same island zone, G. tabacina had a greater tm of 6.75% and an Nm of 0.315. In this study, G. tomentella also had a relatively high average tm of 5.74% (Table 3). Of 11 populations, pops. 2, 3, 4, and 9 had exceptionally high tm values of 10.87–16.70%, and pop. 1 had no occurrence of outcrossing (Table 3). This study did not consider the age structure of individual plants in the populations because determining age is difficult for plants when sampling in the field. Additionally, an isogenous family, or lineage samples of assorted ages, would decrease the detected outcrossing rate. We hypothesize that the relatively high outcrossing rate did not always reflect the actual outcrossing occurring among G. tomentella populations, and it may have resulted from heterogeneity owing to the perennation that maintains historical mutations and genetic recombination in G. tomentella.
Nm is composed of the components of seed dispersal or individual migration and introgression by hybridization. The relatively lower Nm (mean 0.162 Nm, Table 3) implied that individual dispersal seemed be restricted among populations, particularly islands, as shown by the F-statistics in which all populations had significant genetic differentiation (Table 5). Additionally, the NJ-clustering showed a pattern of stronger island differentiation (Fig. 4).
We hypothesize that the stoloniferous habit of G. tomentella along the ground limits the exchanges between populations or islands through the activities of birds, animals and natural factors. This influences the Nm, resulting in a lower Nm (Table 2).
Population genetic variation and differentiation in Chinese G. tomentella
A typical feature of autogamous plants is that they have greater inter-population than intra-population genetic variation (Reif et al. 2003; Guo et al. 2012; He et al. 2012; Wang et al. 2014, 2017), in contrast to typical allogamous plants (Persson and Bothmer 2002; Hang et al. 2004; Costa et al. 2013; Maggioni et al. 2014; Hao et al. 2015). This study revealed that island populations of Chinese G. tomentella had the typical feature of autogamous plants in terms of genetic variation, with a high inter-populations genetic variation of 60.96% (Table 4; and 60.7% variation in Gst, Table 2). This differed from G. tabacina, which exhibited the opposite trend, like an allogamous plant, in terms of population genetic variation.
The NJ cluster and STRUCTURE assignation showed that island populations of Chinese G. tomentella had a genetically stronger island differentiation pattern based on the clustering of populations and individual samples (Figs. 3c, 4b). The closer genetic kinship between northeastern and southwestern regions (Nanao, Meizhou, and Pingtan Islands) in G. tomentella suggested that some genetic interconnection occurred along the southeast coast of China in ancient times. The spatial autocorrelation analysis showed a significant positive correlation within a radius of about 30.45 km (Fig. 5). The assignment test inferred possible individual dispersal (Table 6). The STRUCTURE analysis showed some geographical consanguinity (Fig. 3a). G. tomentella populations might have spread among the islands along the coast while the southeast islands of China and the Taiwan Strait were joined as a continuous land form during the last glacial period (approximately 70,000–11,500 years ago) (Lin 1980), and the island differentiation pattern was formed as the islands separated at the end of the glacial period, approximately 15,000 years ago (Yao et al. 2009). It is possible that some dispersal would be disseminated by birds and sea seawater or by human activities.
The F-statistics suggested that geographically different G. tomentella populations had unique structures, even distance–near neighboring populations in the same islands, such as pops. 1 and 2, and pops. 3 and 4 (Table 5). Based on the idiosyncrasies of the genetically-based geographical differentiation among populations of G. tomentella, we propose that when collecting genetic resources of this species, every G. tomentella population should be sampled.
References
Bau T, Xu B, Zhuang BC (1993) A brief report of cytology study on Chinese subgenus Glycine. J Jilin Agric Univ 15:218 (in Chinese)
Brown AHD, Doyle JL, Grace JP, Doyle JJ (2002) Molecular phylogenetic relationships within and among diploid races of Glycine tomentella (Leguminosae). Aust Syst Bot 15:37–47
Chen LL, Liu XD, Zhao HK, Yuan CP, Wang YN, Xu MZ, Wang YM (2013) Genetic diversity of G. tabacina from Meizhou Island of Fujian Province. Soybean Sci 32:286–290 (in Chinese)
Chung G, Singh RJ (2008) Broadening the genetic base of soybean: a multidisciplinary approach. Crit Rev Plant Sci 27:295–341
Costa TRD, Filho PSV, Gonçalvesvidigal MC, Galván MZ, Lacanallo GF, Silva LID, Kvitschal MV (2013) Genetic diversity and population structure of sweet cassava using simple sequence repeat (SSR) molecular markers. Afr J Biotechnol 12:1040–1048
Cregan PB, Jarvik T, Bush AL, Shoemaker RC, Lark KG, Kahler AL, Kaya N, VanToai TT, Lohnes DG, Chung J, Specht JE (1999) An integrated genetic linkage map of the soybean genome. Crop Sci 39:1464–1490
Doyle MJ, Brown AHD (1985) Numerical analysis of isozyme variation in Glycine tomentella. Biochem Syst Ecol 13:413–419
Doyle MJ, Grant JE, Brown AHD (1986) Reproductive isolation between isozyme groups of Glycine tomentella (Leguminosae), and spontaneous doubling in their hybrids. Aust J Bot 34:523–535
Earl DA, vonHoldt BM (2012) STRUCTURE HARVESTER: a website and program for visualizing STRUCTURE output and implementing the Evanno method. Conserv Genet Resour 4:359–361
Excoffier L, Lischer HEL (2010) Arlequin suite ver 3.5: a new series of programs to perform population genetics analyses under Linux and Windows. Mol Ecol Resour 10:564–567
Fujita R, Ohara M, Okazaki K, Shimamoto Y (1997) The extent of natural cross-pollination in wild soybean (Glycine soja). J Hered 88:124–128
Gao X, Qian J, Ma YH, Zheng SZ (2002) Research on chromosomes of perennial wild soybeans in China. J Fudan Univ (Nat Sci) 41:717–719 (in Chinese)
Goudet J (2001) FSTAT, a program to estimate and test gene diversities and fixation indices (version 2.9.3). My Publications
Guo J, Liu YF, Wang YS, Chen JJ, Li YH, Huang HW, Qiu LJ, Wang Y (2012) Population structure of the wild soybean (Glycine soja) in China: implications from microsatellite analyses. Ann Bot 110:777–785
Hang Y, Jin Y, Lu BR (2004) Genetic diversity of the endangered species Psathyrostachys huashanica in China and its strategic conservation. J Fudan Univ (Nat Sci) 43:260–266 (in Chinese)
Hao D, Zhang Z, Cheng Y, Chen G, Lu H, Mao Y, Shi M, Huang X, Zhou G, Xue L (2015) Identification of genetic differentiation between waxy and common maize by SNP genotyping. PLoS ONE 10:e0142585
Hayata B (1920) Icones Plantarum Formosanarum nec non et Contributiones ad Floram Formosanam; or, Icones of the Plants of Formosa, and Materials for a Flora of the Island, based on a study of the collections of the botanical survey of the Government of Formosa, vol 9. Bureau of Forestry, Industries, Government of Formosa, Taihoku, p 26
He S, Wang Y, Volis S, Li D, Yi T (2012) Genetic diversity and population structure: implications for conservation of wild soybean (Glycine soja Sieb. et Zucc.) based on nuclear and chloroplast microsatellite variation. Int J Mol Sci 13:12608–12628
Jakobsson M, Rosenberg NA (2007) CLUMPP: a cluster matching and permutation program for dealing with label switching and multimodality in analysis of population structure. Bioinformatics 23:1801–1806
Jensen JL, Bohonak AJ, Kelley ST (2005) Isolation by distance, web service. BMC Genet 6:13
Kiang YT, Chiang YC, Kaizuma N (1992) Genetic diversity in natural populations of wild soybean in Iwate Prefecture, Japan. J Hered 83:325–329
Kollopara KP, Singh RJ, Hymowitz T (1994) Genomic diversity and multiple origins of tetraploid (2n = 78, 80) Glycine tomentella. Genome 37:448–459
Kuroda Y, Kaga A, Tomooka N, Vaughan DA (2006) Population genetic structure of Japanese wild soybean (Glycine soja) based on microsatellite variation. Mol Ecol 15:959–974
Lin DG (1980) Eustacy in Fujian during the Quaternary Period. Chin Sci Bull 25:1134–1136 (in Chinese)
Liu KJ, Muse SV (2005) POWERMARKER: integrated analysis environment for genetic marker data. Bioinformatics 21:2128–2129
Maggioni L, von Bothmer R, Poulsen G, Branca F, Bagger Jørgensen R (2014) Genetic diversity and population structure of leafy kale and Brassica rupestris Raf. in south Italy. Hereditas 151:145–158
Narzary D, Verma S, Mahar KS, Rana TS (2015) A rapid and effective method for isolation of genomic DNA from small amount of silica-dried leaf tissues. Nat Acad Sci Lett 38:441–444
Nei M, Tajima F, Tateno Y (1983) Accuracy of estimated phylogenetic trees from molecular data II. Gene frequency data. J Mol Evol 19:153–170
Peakall R, Smouse PE (2012) GenAlEx 6.5: genetic analysis in Excel. Population genetic software for teaching and research—an update. Bioinformatics 28:2537–2539
Persson K, Bothmer RV (2002) Genetic diversity amongst landraces of rye (Secale cereale L.) from northern Europe. Hereditas 136:29–38
Pfeil BE, Craven LA, Brown AHD, Murray BG, Doyle JJ (2006) Three new species of northern Australian Glycine (Fabaceae, Phaseolae), G. gracei, G. montis-douglas and G. syndetika. Austral Syst Bot 19:245–258
Piry S, Alapetite A, Cornuet JM, Paetkau D, Baudouin L, Estoup A (2004) GENECLASS2: a software for genetic assignment and first-generation migrant detection. J Hered 95:536–539
Pritchard JK, Stephens M, Donnelly P (2000) Inference of population structure using multilocus genotype data. Genetics 155:945–959
Reif JC, Melchinger AE, Xia XC, Warburton ML, Hoisington DA, Vasal SK, Srinivasan G, Bohn M, Frisch M (2003) Genetic distance based on simple sequence repeats and heterosis in tropical maize populations. Crop Sci 43:1275–1282
Singh RJ (2010) Methods for producing fertile crosses between wild and domestic soybean species: US, US7842850
Singh RJ, Nelson RL (2015) Intersubgeneric hybridization between Glycine max and G. tomentella: production of F1, amphidiploid, BC1, BC2, BC3, and fertile soybean plants. Theor Appl Genet 128:1117–1136
Singh RJ, Kollipara KP, Hymowitz T (1987) Polyploid complexes of Glycine tabacina (Labill.) Benth. and G. tomentella Hayata revealed by cytogenetic analysis. Genome 29:490–497
Singh RJ, Kollipara KP, Hymowitz T (1990) Backcross-derived progeny from soybean and Glycine tomentella Hayata intersubgeneric hybrids. Crop Sci 30:871–874
Singh RJ, Kollipara KP, Hymowitz T (1993) Backcross (BC2–BC4)-derived fertile plants from Glycine max and G. tomentella intersubgeneric hybrids. Crop Sci 33:1002–1007
Singh RJ, Kollipara KP, Hymowitz T (1998) Monosomic alien addition lines derived from Glycine max (L.) Merr. and G. tomentella Hayata: production, characterization, and breeding behavior. Crop Sci 38:1483–1489
Song QJ, Jia GF, Zhu YL, Grant D, Nelson RT, Hwang EY, Hyten DL, Cregan PB (2010) Abundance of SSR motifs and development of candidate polymorphic SSR markers (BARCSOYSSR_1.0) in soybean. Crop Sci 50:1950–1960
Tateishi Y, Ohashi H (1992) Taxonomic studies on Glycine of Taiwan. J Jpn Bot 67:127–147
Tindale M (1986) Taxonomic notes on three Australian and Norfolk Island species of Glycine Willd. (Fabaceae: Phaseolae) including the choice of a neotype for G. clandestina Wendl. Brunonia 9:179
Wang KJ, Li XH (2012) Genetic characterization and gene flow in different geographical-distance neighbouring natural populations of wild soybean (Glycine soja Sieb. & Zucc.) and implications for protection from GM soybeans. Euphytica 186:817–830
Wang KJ, Li XH, Yan MF (2014) Microsatellite markers reveal genetic diversity of wild soybean in different habitats and implications for conservation strategies (Glycine soja) in China. Conserv Genet 15:605–618
Wang Y, Ghouri F, Shahid MQ, Naeem M, Baloch FS (2017) The genetic diversity and population structure of wild soybean evaluated by chloroplast and nuclear gene sequences. Biochem Syst Ecol 71:170–178
Weir BS, Cockerham CC (1984) Estimating F-statistics for the analysis of population structure. Evolution 38:1358–1370
Yao YT, Harff J, Meyer M, Zhan WH (2009) Reconstruction of paleocoastlines for the northwestern South China Sea since the last glacial maximum. Sci China Ser D Earth Sci 52:1127–1136
Yeh FC, Yang RC, Boyle T (1999) Popgene version 1.32: Microsoft Windows-based freeware for population genetic analysis. University of Alberta, Edmonton
Zhang R, Liu JQ, Yang M, Zhu GH (2011) The extracts from Glycine tomentella roots used to produce traditional Chinese medicine for the treatment of rheumatoid arthritis: Patent, China, CN102119954A (in Chinese)
Zou JJ, Singh RJ, Hymowitz T (2004) SSR marker and ITS cleaved amplified polymorphic sequence analysis of soybean × Glycine tomentella intersubgeneric derived lines. Theor Appl Genet 109:769–774
Acknowledgements
This work was financially supported by project “Sci and Tech Innovation Program of Chinese Academy of Agricultural Sciences” and “Crop Germplasm Resources Protection (Item No. 2130135-09) from the Ministry of Agriculture of the People’s Republic of China.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Wang, XD., Li, XH., Zhang, ZW. et al. Characterization of genetic diversity and structures in natural Glycine tomentella populations on the southeast islands of China. Genet Resour Crop Evol 66, 47–59 (2019). https://doi.org/10.1007/s10722-018-0694-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10722-018-0694-6