Abstract
The main objective was to evaluate if different planes of maternal nutrition during late gestation and weaning age alter microRNA (miRNA) and target gene expression in offspring longissimus muscle (LM). Early (EW) and normal weaned (NW) Angus × Simmental calves (n = 30) born to cows that were grazing endophyte-infected tall fescue and red clover pastures with no supplement [low plane of nutrition (LPN)], or supplemented with 2.3 and 9.1 kg of dried distiller’s grains with solubles and soy hulls [medium and high plane of nutrition (MPN, HPN), respectively] during the last 105 ± 11 days of gestation were used. Biopsies of LM were harvested at 78 (early weaning), 187 (normal weaning) and 354 days of age. Results indicate a role of pro-adipogenic miRNA in the control of adipogenesis in LM of NW-MPN steers between 78 and 187 days of age through upregulation of (1) miR-103 which inhibits CAV1, a protein that destabilizes INSR and leads to insulin resistance; (2) miR-143 which inhibits DLK1, a protein that inhibits adipocyte differentiation; and (3) miR-21 which impairs TGFBR2-induced inhibition of adipocyte differentiation. Among the studied anti-adipogenic miRNA, cow plane of nutrition resulted in downregulation of miR-34a expression in MPN steers compared with HPN and LPN at 78 days of age. Data for miR-34a provided a potential sign of epigenetic regulation of LM in beef offspring due to the cow plane of nutrition during late gestation.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Nutritional programming of the offspring can occur via epigenetics, i.e., changes in gene expression unrelated to the DNA sequence of the animal [1]. The conformational change of the structure of histones is a key epigenetic mechanism that regulates gene expression via changes in transcriptional activation, chromatin remodeling, nucleosome loosening, chromatin condensation, and chromatin stabilization [2]. Furthermore, non-coding microRNA (miRNA), a functional RNA molecule that is not translated into a protein, also is considered an important mechanism of epigenetic regulation of gene expression [3]. A miRNA is a short ribonucleic acid molecule of ~21–23 nucleotides. MicroRNA are post-transcriptional regulators that bind to complementary sequences on target messenger RNA transcripts (mRNA), usually resulting in translational repression, target degradation and gene silencing [4].
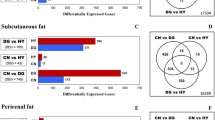
Studies of adipose tissue miRNA profiling in beef cattle revealed different patterns according to fat depot location, with 80, 66 and 63 miRNA, respectively, detected in abdominal subcutaneous, back subcutaneous and rump subcutaneous tissues [5]. Fat depot features also influence miRNA profiles, i.e., high backfat thickness vs. low backfat thickness; 42 miRNA were differentially expressed between these two groups, with miR-378 being the most differentially expressed [6].
In a study using rats, it was determined that late gestation (rapid fetal growth) is a physiological point at which maternal nutrition could elicit programming of adipose tissue in the offspring [7]. Thus, the hypothesis we sought to test was that maternal plane of nutrition during late gestation leads to greater expression of skeletal muscle metabolic target genes related to adipogenesis, lipogenesis, and adipokine synthesis in the offspring at least in part through alterations in the expression of pro-adipogenic and anti-adipogenic miRNA. The objective of this study was to assess the effect of a high, medium, and low plane of nutrition (LPN) of the cow during the last 90 days of pregnancy on selected pro-adipogenic and anti-adipogenic miRNA and their associated target genes in LM samples of offspring that were early weaned or normal weaned followed by a common high-grain diet during the finishing period [8].
Materials and Methods
Treatments and Skeletal Muscle Biopsy
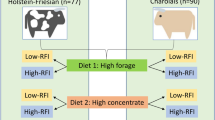
A portion of the animals used in the present study was used for longissimus muscle transcriptomics [8] under a protocol approved by the Institutional Animal Care and Use Committee of the University of Illinois. Briefly, a subset of 30 steers from Angus × Simmental beef cows from the University of Illinois Dixon Springs Agriculture Center (Simpson, IL, USA) was selected from a group of animals (n = 326) utilized in a parallel study [9]. Main effects evaluated were maternal plane of nutrition during late gestation and postnatal management of the offspring (Fig. 1). Three months prior to the projected parturition date, cows were assigned to treatments (low, medium or high plane of nutrition) in a split-plot design. Dietary treatments were formulated to provide 70 % (nutrient restriction), 100 % (maintenance), or 130 % (nutrient excess) of NRC requirements for the stage of gestation for crude protein and energy with feeding level adjusted to provide the desired plane of nutrition. LPN was achieved by grazing endophyte-infected tall fescue/red clover pastures during July, August, and September with no supplementation of dried distiller’s grains. Medium and high plane of nutrition cow diets (MPN, HPN) were achieved by grazing endophyte-infected tall fescue/red clover pastures supplemented with 2.3 and 9.1 kg per animal per day of dried distiller’s grains with solubles and soy hulls (70–30 %), respectively.
Composition of the diet [dry matter (DM) basis] fed to early weaned (EW) steers upon arrival to the feedlot and prior to normal weaning (NW) is presented in Table 1. Cow supplementation was initiated at 103 ± 11 days prepartum while on pasture and it was halted at the midpoint of parturition (2 ± 11 days postpartum). Cows were consolidated into three pastures, each containing 1 LPN, 1 MPN, and 1 HPN cow group. Following consolidation, cows were fed 2.14 kg of DDGS + Co-Product Balancer (Purina Animal Nutrition®, Saint Paul, MN: 700 g/ton Monensin; 25 % crude protein; 2 % crude fat; 8 % crude fiber; 13 % Ca; 3.5 % NaCl; 0.1 % K; 24,000 IU/lb. Vitamin A) per cow each day [9].
The mentioned subset of 30 Angus × Simmental steer calves were randomly assigned to early or normal weaning (EW or NW) treatments within each gestational treatment. This selection allowed for ten animals for each prenatal treatment and five animals of each of the interactions of gestational × postnatal treatments. At 78 ± 2 days postpartum, EW offspring were weaned, weighed, hip height measured, transported to the University of Illinois Beef and Sheep Field Laboratory, and adapted to a high-grain diet until they had ad libitum consumption. At 187 ± 2 days postpartum, NW offspring were weaned and transported to the University of Illinois Beef and Sheep Field Laboratory. All offspring were co-mingled among treatments. LM biopsies were taken from the first lumbar vertebra region and the subsequent biopsy sites were identified alongside the back on the left side of the spine. Samples were immersed in liquid N right after collection. LM biopsies were sampled from a subset of five animals per gestational × postnatal treatment at ~78 days of age, ~187 days, and during the last week prior to harvest (~354 days). After normal weaning, all offspring were placed on a common, corn grain-based finishing diet that is typical of industry management. All the steers in the study were harvested at a commercial packing plant when they reached the selected end point target back fat thickness of 1.1 cm. Reported final body weight (BW) was calculated from hot carcass weight using a 62 % dressing percentage.
Selection of steer progeny for biopsy was performed based on 2 criteria: first, offspring for biopsy were selected based on their dam’s performance. Only offspring from cows whose BW and BW change during late gestation was within ½ of a standard deviation on either side of the mean were considered for biopsy. Selecting based on cow BW and BW change ensured that only calves from cows that were representative of their treatment were utilized. The final selection of steers for biopsy was based on steer BW. Only steers whose BW was within ½ of a standard deviation on either side of the mean were utilized. This selection strategy minimized the effects of variation in dam’s milk production, which was not significantly different between treatments [10]. The biopsy date for the last biopsy was chosen based on ultrasound measurements taken one or two days prior to the biopsy date. Biopsies were performed in the animals when they reached the end-point for back fat thickness of 1.1 cm. In terms of days prior to slaughter, the approach we used leads to variability among animals for the last biopsy. Longissimus muscle transcriptomics was only performed in EW and NW steers in the HPN and LPN groups [8].
Blood Sampling and Analyses
These details were reported previously [8]. Briefly, blood was collected from the jugular vein at 78, 187 and 296 days of age prior to the morning feeding to isolate serum for insulin (Bovine Insulin ELISA kit, Cat No. 10-1201-01, Mercodia AB, Uppsala, Sweden), glucose (Cayman Glucose Colorimetric Assay Kit, Diagnostics Laboratory, College of Veterinary Medicine, University of Illinois, Urbana, USA) and adiponectin. The latter was determined using a liquid RIA (Millipore, LA, USA) following a protocol previously described [11]. The sensitivity of the adiponectin assay was 1.6 ng/mL. The intra-assay coefficient of variation for low (9.9 ng/mL) and high (55.2 ng/mL) concentrations were 10.2 and 13.4 %, respectively. Biomarker data from steers in the HPN and LPN in both the EW and NW groups were reported previously [8].
Feed Intake and Growth
Individual intakes were obtained utilizing the Grow-Safe® feed monitoring system (GrowSafe Systems Ltd, Canada) after normal weaning. Animals were weighed at birth, early weaning, midpoint between early and normal weaning (109 days), normal weaning, and every 2 weeks during the finishing phase. Ultrasonic measurements were taken for intramuscular fat deposition score and 12th rib fat thickness using ultrasound 1 day prior to early weaning and normal weaning, and during late finishing phase to sort the cattle into harvest groups with a target harvest backfat thickness of 1.1 cm.
Total and MicroRNA Analysis
LM biopsied samples represent a combination of mainly myocytes and adipocytes. In terms of microRNA and mRNA extraction, because the amount of tissue harvested using our biopsy procedure is no more than 500 mg it would be difficult (if not impossible) to separate both cell types without running into problems of sample degradation. By selecting microRNA and target genes with higher expression in adipocytes, the risk of confounding during interpretation is reduced. Total and miRNA were extracted using a mir-Vana™ miRNA isolation kit (Ambion, USA). Two hundred mg of LM tissue was homogenized in 10 vol of mirVana Lysis/Binding Solution using a Tissue-Tearor™ (BioSpec Products, Inc.) homogenizer at maximum speed. Then 1/10 vol of miRNA homogenate additive was added, mixed thoroughly via vortex for 30 s and incubated on ice for 10 min. An equal vol of Acid-Phenol:Chloroform:IAA (125:24:1; Ambion) was then added to each aliquot. The resulting solutions were vortexed for 1 min and centrifuged for 5 min at 10,000×g at room temperature. The aqueous phase was carefully removed and then mixed thoroughly with 1.25 vol of 100 % ethanol and passed through a mirVana Filter Cartridge in sequential 700 µL aliquots. The Filter Cartridge was washed according to the manufacturer’s protocol, and RNA was eluted in nuclease-free water at 95 °C. Purity of RNA was assessed by ratio of optical density OD260 mm (oligonucleotide absorbance)/OD280 mm (protein absorbance) using a NanoDrop ND-1000 (NanoDrop Technologies, Rockland, DE).
Quantitative RT-PCR of miRNA expression was performed using the NCode™ miRNA First-Strand Synthesis and quantitative RT-PCR (qPCR) kits (Invitrogen, USA), according to the manufacturer’s protocol. For cDNA synthesis, 500 ng of purified RNA sample were added to MIX1, consisting of 5.0 µL 5 × miRNA Reaction Buffer, 2.5 µL 25 mM MnCl2, 1 µL diluted ATP, 0.5 µL Poly A Polymerase and 11 µL of DNase/RNase free water was then added. The mixture was incubated at 37 °C for 15 min. 4 µL polyadenylated RNA (from the above) were added to MIX2, consisting of 1 µL of Annealing Buffer and 3 µL of Universal RT Primer. The mixture was incubated at 65 °C for 5 min, and then placed on ice for 1 min. A third mix (MIX3) was added to the above mixture, consisting of 10 µL of 2 × First-Strand Reaction Mix and 2 µL of SuperScript™ III RT/RNaseOUT™ Enzyme Mix. The mixture was incubated at 85 °C for 5 min to stop the reaction.
The cDNA was then diluted 1:10 with DNase/RNase-free water and from this solution a pool of cDNA was made for standard curve preparation using a 1:5 dilution. A combination of 1 µL of diluted cDNA with 9 µL of the mix of 5 µL of PerfecCTa SYBR Green Super Mix (Cat#95054-02K), 0.2 µL each of 10 µM sequence-specific forward primer and Universal qPCR Primer, and 3.6 µL of DNase/RNase free water was added to each well of a MicroAmp Optical 384-Well Reaction Plate (Applied Biosystems). The reactions were performed in an ABI Prism 7900 HT SDS instrument (Applied Biosystems) under the following thermocycler conditions: 95 °C for 5 min (Stage 1), and 40 cycles of 95 °C for 10 s and 60 °C for 30 s (Stage 2). Stage 3 consisted of 95 °C for 15 s, 60 °C for 15 s and 95 °C for 15 s that allows seeing the dissociation curve of each reaction of the qPCR plate. MiRNA forward primers sequences for Bos Taurus (Additional Table 3) were obtained from the microRNA database (http://www.mirbase.org) for the selected miRNA (Table 1). For better interpretation of the miRNA results, transcriptome results from a microarray analysis [8] using the same LM samples of the present study were used to evaluate other significant miRNA targets that were not considered at the beginning of this study (Additional Figure 3).
Statistical Analysis
Quantitative PCR data were analyzed using the MIXED procedure of SAS (SAS 9.1 Institute, Cary, NC, USA). Before statistical analysis, qPCR data were normalized using the geometric mean of miR-16b, miR-let-7a and miR-181a as internal control genes for miRNA and the geometric mean of UXT, MTG1 and RPS15A as internal control genes for the chosen miRNA targets. Fixed effects in the statistical model for each variable analyzed (i.e., miRNA and target genes) included plane of nutrition [diet (D)], weaning age (W), time on experiment (T), D × W, D × T, W × T, and W × D × T interactions. Gene expression data analysis included a repeated-measures statement with an autoregressive covariate structure. Animal performance, carcass quality parameters, blood biomarkers and ultrasound data also were analyzed using the MIXED procedure of SAS with diet as the fixed effect in the statistical model. The random effect in all models was steer.
The statistical model used was: Y ijklm = μ + C i + T j + B k +S l + (T × D)ij + (D × W)jk + (T × W)ik + (D × T × W)ijk + ε ijklm; where Y ijklm is the background-adjusted normalized gene expression and blood metabolites data; μ is the overall mean; T i is the fixed effect of time (3 levels); D j is the fixed effect of diet (3 levels); W k is the fixed effect of wean (2 levels); S l is the random effect of steer nested within weaning time; T × D, D × W, T × W are the interactions of time by diet, diet by wean and time by wean, respectively; D × T × W is the interaction or third order for the main effects; and ε ijklm is the random error (0, σ 2e ) associated with Y ijklm. Statistical significant differences were declared at P < 0.05.
Results
Animal Performance
Performance data for the entire group from which the animals for this study were selected are reported elsewhere [10]. Cow initial BW did not differ among supplementation level. Cow body condition score among the three supplement levels were within 0.2 units. Cows fed the HPN had a 28 kg and 15 kg greater precalving BW compared with LPN and MPN cows [10]. Ultrasound measurements performed at EW did not reveal differences between treatments (P > 0.05) in marbling (intramuscular fat) and back fat thickness (Additional Table 2). The EW steers had greater body weight as compared with NW steers during the finishing period (Additional Table 2). There were no significant differences (P > 0.05) in BW between treatments at early weaning, i.e., all possible combinations between weaning age and cow plane of nutrition (Additional Table 2). The EW steers had greater hip height (P = 0.05) compared with NW steers (Additional Table 2). After normal weaning and between 0 and 29 days on feedlot (P = 0.03) average daily gain (ADG) was greater in HPN steers compared with other treatments, and between 29 and 57 days on feedlot (P = 0.05) MPN steers had greater ADG compared with other treatments. An overall weaning age effect was detected at the beginning (0–29 days) of the finishing phase for ADG because of higher values for EW steers (P = 0.02), and at the end (84–113 days) of the finishing phase because of higher values for NW steers (P = 0.04) (Additional Table 2). Dry matter intake (DMI) during the finishing phase was lower (diet P = 0.04) in LPN as compared with MPN and HPN steers (Additional Table 2). Lastly, none of the carcass quality parameters had significant differences between treatments (Additional Table 2).
Blood Biomarkers
Serum glucose concentration had a significant time effect (P < 0.05) with lower values at 296 days of age. Insulin concentration was affected by weaning age, time and weaning × time interaction (P < 0.05) with a noticeable increase in concentration between 187 and 296 days of age for all treatments (Fig. 2). Adiponectin concentration had a significant diet × time interaction (P < 0.05). The HPN steers had markedly lower adiponectin concentration as compared with MPN and LPN steers. The general pattern of serum adiponectin concentration was an increase between 78 and 187 days of age followed by a decrease in concentration after 187 days of age for all treatments. It was noteworthy that in EW-HPN the concentration increased between 187 and 296 days of age (Fig. 2).
Glucose, adiponectin and insulin serum concentration in Angus × Simmental steers born to cows that received a low (LPN), medium (MPN) or a high (HPN) plane of nutrition during the late gestation period. Weaning times are early weaning (EW) and normal weaning (NW). asterisk Diet, double asterisk Weaning, triple asterisk Time, hash diet × time and triple hash time × weaning. Significant differences were declared at P < 0.05
Pro-adipogenic MicroRNA
MiR-103
MiR-103 had a time × weaning interaction (P = 0.04) and a time × diet interaction (P = 0.07). MiR-103 expression increased between 78 and 187 days of age and decreased between 187 and 354 days of age in NW steers (Table 5).
The adipogenic nuclear receptor PPARG, a target gene for miR-103 (Table 2), had a significant diet × weaning × time interaction, a time × diet interaction and a weaning × time interaction (Tables 3, 5). PPARG expression increased to the greatest extent in EW-HPN steers between 78 and 187 days of age. Other treatments also had an up-regulation of PPARG but to a lesser extent. In contrast, NW-HPN steers had a decrease in PPARG expression during this time. Between 187 and 354 days of age, PPARG expression decreased significantly for EW-HPN and EW-MPN steers but remained relatively constant for other treatments (Tables 3, 5).
MiR-143
MiR-143 had a similar miRNA relative abundance compared with miR-103 (Additional Figure 1). MiR-143 had a significant weaning × time (P < 0.01), and a diet × weaning interaction (P < 0.01) (Tables 4, 5). MiR-143 had a slight increase in its expression between 78 and 187 days of age in NW-MPN and NW-LPN steers, and between 187 and 354 days of age it decreased while in other treatments an opposite pattern of expression was detected (Tables 4, 5). NW-HPN steers had an unchanged expression pattern of miR-143 throughout the study. After normal weaning time (187 days of age), the period when NW steers began to receive the high-grain diet, steers had a decrease in miR-143 expression, especially NW-MPN steers.
The expression of PPARG, CEBPA and FABP4 (all targets of miR-143; Table 2) had a significant diet × weaning × time interaction, a time × weaning interaction and time × diet interaction (Tables 3, 5). These three miRNA targets had a similar pattern of expression with an upregulation in EW steers between 78 and 187 days of age (Table 5).
MiR-21-5p
MiR-21-5p (5p denotes from the 5′ arm) was the pro-adipogenic miRNA with lowest relative miRNA abundance (Additional Figure 1). MiR-21-5p had a significant diet × time interaction (P = 0.05). The difference in miR-21-5p expression for NW-HPN steers at 78 days of age as compared with other treatments (Tables 4, 5) was not statistically significant (Additional Figure 2).
FABP4 was the target gene selected for miR-21-5p (Table 2). FABP4 had a significant time × weaning × diet interaction and it had similar expression pattern for NW-HPN when compared with miR-21-5p expression (Tables 3, 4, 5).
MiR-378
MiR-378 had a significant time × diet × weaning interaction (P = 0.05) and it also was the pro-adipogenic miRNA with highest miRNA relative abundance (Additional Figure 1). Expression of miR-378 differed in NW-MPN steers as compared with other treatments (Tables 3, 4, 5).
PPARG was the target gene selected for miR-378 (Table 2). MiR-378 had higher relative miRNA abundance as compared with PPARG (Additional Figure 1). MiR-378 expression was greater for NW steers as compared with EW steers (Table 4) but variable throughout time and plane of nutrition (Tables 3, 5).
Anti-adipogenic MicroRNA
MiR-27a/b
MiR-27a/b was affected by cow plane of nutrition and weaning time (Table 4). MiR-27a/b had a significant time × diet × weaning interaction (P < 0.01), a time × weaning interaction (P < 0.01), a diet × weaning interaction (P = 0.01) and it was the anti-adipogenic miRNA with highest relative miRNA abundance (Tables 4, 5 and Additional Figure 1). MiR-27a/b expression was lower for EW-HPN steers as compared with other treatments (Table 4). At 187 days of age, miR-27a/b expression was greater for NW steers (Table 5).
C/EBPA and PPARG were the target genes measured for miR-27a/b (Table 2). They had similar expression pattern with a significant time × weaning × diet interaction (P < 0.01), a weaning × time interaction (P < 0.01) and a diet × time interaction (P < 0.01). PPARG and C/EBPA had higher expression level at 187 days of age in EW-HPN steers (Tables 3, 4, 5).
MiR-130a
MiR-130a had a significant time × diet × weaning interaction (P = 0.03 (Tables 3, 4, 5). MiR-130a relative miRNA abundance was negligible compared with other miRNA and miRNA targets analyzed (Additional Figure 1). Steers born from cows that received MPN had an increase in expression of miR-130a between 78 and 187 days of age, while other treatments had undetectable changes (Tables 3, 4, 5). In contrast, between 187 and 354 days of age, MPN and HPN steers had a significant decrease in miR-130a expression, while in LPN steers it remained constant (Table 3).
PPARG was the target gene selected for miR-130a (Table 2). PPARG was significantly upregulated (P < 0.05) between 78 and 187 days of age, while miR-130a did not change during that time (Tables 3, 5). NW-LPN steers had lower (P < 0.05) miR-130a expression (Table 4) while PPARG had a slight increase (P < 0.05) between 187 and 354 days of age (Table 5).
MiR-34a
When all biopsy time points were considered in the statistical analysis, miR-34a was not affected due to time, weaning age or cow plane of nutrition (Tables 3, 4, 5). However, when only the early weaning time (78 days of age) was considered in the statistical analysis, miR-34a had a significant cow plane of nutrition effect (P < 0.01) with higher expression in HPN and LPN steers (Additional Figure 2). Furthermore, miR-34a was the second anti-adipogenic miRNA with higher relative miRNA abundance among those studied (Additional Figure 1). In general, the anti-adipogenic miRNA measured had lower miRNA relative abundance as compared to the pro-adipogenic miRNA (Additional Figure 1).
Silent information regulator 1 (SIRT1) was the target gene selected for miR-34a (Table 2). SIRT1 expression decreased between 78 and 187 days of age for NW steers and between 187 and 354 days of age for EW steers (Table 5).
MiR-369-5p
MiR-369-5p relative abundance was negligible as compared with other miRNA and its targets (Additional Figure 1). MiR-369-5p was only affected by weaning age (P = 0.06), with an important numerical increase in its expression level for NW-MPN steers compared with other treatments (Table 4).
ADIPOQ and FABP4 were the target genes selected for miR-369 (Table 2). ADIPOQ expression was higher between 78 and 187 days of age especially in EW steers (Table 5). ADIPOQ had a significant time × diet × weaning interaction (P < 0.01), a weaning × time interaction (P < 0.01) and a diet × time interaction (P < 0.01) (Tables 3, 4, 5). ADIPOQ expression reached its higher level at 187 days of age for MPN and HPN steers but LPN steers had higher level of ADIPOQ expression later on at 354 days of age (Table 3).
MiR-448
MiR-448 had a significant time × weaning interaction (P < 0.01) and a significant time × diet interaction (P < 0.01) (Tables 3, 4, 5). It was the miRNA with lowest relative miRNA abundance compared with both other miRNA and miRNA targets analyzed in this study (Additional Figure 1). EW-MPN steers were the only ones with a differential pattern of miR-448 expression (Table 4). EW-MPN steers had an increase in miR-448 expression between 78 and 187 days of age, followed by downregulation between 187 and 354 days of age. All other treatments had a relatively constant expression level. There were no target genes measured for miR-448 in this study (Table 2).
Discussion
Animal Performance
Overall, steer feedlot performance in terms of ADG was positively influenced by the plane of nutrition of the dam, and coincided with results from a previous study of similar design [12]. However, unlike another study where cow precalving body condition score was manipulated through diet [13], the MPN offspring had better performance during the first-half and lower performance during the second-half of the growing phase. More specifically, between 78 and 187 days of age, access to a high-grain diet was advantageous to EW-MPN steers which had a higher growth performance compared with NW-MPN steers. The latter were grazing and nursing their dams during the same time-frame. This difference in nutrition during the growing phase resulted in more efficient growth in EW-MPN steers; in contrast, NW-MPN steers remained longer in the feedlot due to a lower efficiency of growth during the growing phase, i.e., grazing pastures and nursing “delayed” their target back fat thickness end point during the finishing phase.
Blood Biomarkers
The decrease in glucose concentration between 187 and 296 days of age might reflect a greater rate of glucose uptake from the bloodstream. Higher plasma insulin levels between 187 and 296 days of age are common in animals fed diets that result in greater production of propionate (i.e., high-starch diets). Similar to a previous study [14], serum glucose and insulin concentrations were not affected by weaning time.
Adiponectin is exclusively secreted from white adipose tissue [15]. Levels of this adipokine are usually inversely correlated with body fat percentage and positively correlated with body size in humans [16]. Alterations in plasma adiponectin levels are inversely correlated with biomarkers of adipose tissue mobilization (i.e., non-esterified fatty acids), some of which might be caused by a local inflammatory response [17]. The lower plasma adiponectin levels measured in HPN steers could be related to the presence of an inflammatory response produced via macrophage infiltration in visceral or subcutaneous adipose tissue [18, 19]. Adiponectin also increases tissue sensitivity to insulin in non-ruminants, hence, greater adiponectin concentrations may facilitate a biological response to lower concentrations of insulin [20]. The lower serum adiponectin concentrations in HPN steers could be taken as an indication of reduced insulin sensitivity [21].
Pro-adipogenic MicroRNA
MiR-103
In a mouse study with two different obese models (diet-induced obesity and leptin-deficient ob/ob), the treatment of animals with a miR-103 antagonist (i.e., TNF-α) confirmed the role of miR-103 in inducing adipogenesis because triacylglycerol accumulation increased during early stages of adipogenesis. In contrast, in vitro, in both obesity models TNF-α treatment of differentiated adipocytes led to miR-103 downregulation [22]. In another study utilizing guinea pig offspring with intrauterine growth restriction, miR-103 was greater in low-birth weight male offspring with greater visceral adiposity [23].
MiR-103 is an intronic miRNA that resides in the sense orientation (5′–3′) in intron 5 of pantothenate kinase family member genes (PANK), hence, miR-103 plays a role in the rate-limiting step of CoA generation [24]. Phosphorylation of pantothenate (Vitamin B5) [25] is governed by PANK, and constitutes an important step in the overall metabolism of pyruvate, lactate, glucose and the citric acid cycle, or its incorporation into the acyl-carrier-protein needed for fatty acids synthesis [26]. In a previous study, miR-103 was co-regulated with PANK during 3T3-L1 adipogenesis [22].
Additional data supporting a pro-adipogenic role for miR-103 (after ectopic expression) during adipocyte development in 3T3-L1 cells include an increase in the rate of triacylglycerol accumulation in adipocytes and also upregulation of the expression of important transcription factors such as PPARG, key cell cycle regulators such as G0/G1 switch 2 (G0S2), and molecules associated with lipid metabolism (FABP4), glucose homeostasis (SLC2A4) and endocrine functions (ADIPOQ) of adipocytes [22]. The upregulation between 78 and 187 days of age of SLC2A4 coupled with downregulation of G0S2 (Additional Figure 3) argues against an effect of miR-103 on bovine longissimus cell cycle through G0S2. Rather, it suggests that miR-103 helps regulate glucose homeostasis through SLC2A4 in NW-MPN steers.
In previous studies, miR-103 also was associated with insulin and glucose levels through inhibition of caveolin 1 (CAV1) which influences lipid raft signaling, through destabilization of the insulin receptor (INSR), leading to increased insulin resistance in the adipose tissue [27, 28]. CAV1 inhibition supports this mechanism for NW-MPN steers between 78 and 187 days of age (Additional Figure 3). Taken together, data suggest that miR-103 and PANK1 could activate adipogenesis in LM of NW-MPN steers but, alternatively, they could have increased insulin insensitivity in the adipose tissue of NW-MPN steers through downregulation of INSR due to CAV1 inhibition.
MiR-143
In preadipocytes, miR-143 was upregulated on day 4 of culture and increased further on day 8 after induction of differentiation, indicating that miRNA-143 normally promotes intramuscular preadipocyte differentiation; it was also reported that miR-143 is highly-expressed in bovine intramuscular fat [29]. Furthermore, in cells transfected with an miR-143 inhibitor, C/EBPA and FABP4 expression was downregulated [30]. The transfection of the fibroblast-like preadipocytes with miRNA-143 antisense inhibitor induced a significant suppression of differentiation, indicated by a decrease in storage of lipid droplets [30]. In contrast, cell proliferation was increased with the transfection of a miRNA-143 inhibitor. Taken together, the above results and those of the present study confirm the importance of miR-143 in bovine intramuscular fat development.
Greater miRNA-143 expression in NW steers compared with EW steers before weaning (187 days of age) suggests that in those steers epigenetic control of adipogenesis via miR-143 was more pronounced. In human adipocytes, free fatty acids downregulate miR-143 [31]. At late terminal differentiation stages of adipogenesis, miR-143 leads to downregulation of delta-like 1 (DLK1), an inhibitor of adipocyte differentiation through ERK1/2 activation [32]. Furthermore, hyperinsulinemia significantly suppressed DLK1 [33]. Before 187 days of age, the fact that NW steers were nursing their dam, consuming pasture, and receiving some grain meant that insulin secretion was less pronounced compared with EW steers fed the high-grain diet (Fig. 2). The expression of DLK1 was downregulated between 78 and 187 days of age (Additional Figure 3), hence, it could be possible that miR-143 activation in NW steers and the higher serum insulin concentration in EW steers were mechanisms that ensured downregulation of DLK1.
In contrast to what was expected, miR-143 target genes (PPARG, C/EBPA and FABP4) were downregulated when miR-143 was upregulated suggesting that miR-143 did not influence these targets genes. Taken together, the main mechanism of action of miR-143 for promoting adipogenesis in LM appears to be through blocking adipogenic repressors (DLK1) more than promoting pro-adipogenic gene activation.
MiR-21-5p
MiR-21 is upregulated in white adipose tissue (WAT) of obese humans as compared with lean controls [34]. MiR-21 also controls adipogenesis by modulating one of its targets, transforming growth factor beta receptor 2 (TGFBR2) [35]. In human adipose tissue-derived mesenchymal stem cells (hASC) induced to overexpress miR-21, the TGF-β-induced inhibition of adipocyte differentiation was significantly dampened. In our study, TGFBR2, TGF-β2 and TGF-β3 were affected by time (Additional Figure 3) with activation of the receptor between 187 and 354 days and inhibition of TGF-β3 before 187 days and TGF-β2 between 187 and 354 days. Judging by their expression pattern, TGFBR2 seems to be more susceptible to miR-21 control because its expression did not change when miR-21 was upregulated (i.e., NW-MPN steers).
In a study using hASC, STAT3 was detected as the most likely target gene for miR-21. MiR-21 reduced proliferation of hASC by impairing total and phospho-STAT3 expression which inhibits self-renewal of stem cells [36]. In contrast, it has been reported that miR-21 increases proliferation in cancer cells through inhibition of programmed cell death protein 4 (PDCD4) and phosphatase and tensin homolog (PTEN) [37]. By means of its interaction with STAT3, in the present study, miR-21 inhibition between 78 and 187 days of age for NW-HPN and EW-HPN might have generated an increase in cell proliferation during the clonal stage of adipogenic expansion; between 187 and 354 days of age, when the rate of fat accumulation was likely greater in the steers (especially EW), miR-21 could have interacted with PDCD4 leading to an anti-apoptotic effect. As such, the adipocyte differentiation process would have continued undisrupted.
Subedi et al. [38] suggested that the miR-21/PDCD4 axis may be a novel target mediating adiponectin-induced biological responses because miR-21 increased adiponectin mRNA and protein expression levels [39]. In that context, the fact that EW-HPN and NW-HPN steers had a lower level of serum adiponectin and a simultaneous decrease in miR-21 expression suggests the existence of a similar effect.
MiR-378
In beef cattle, miR-378 is located on intron 1 of peroxisome proliferator-activated receptor gamma coactivator 1 beta (PGC-1β) [40, 41] and is strongly and positively associated with back fat thickness [42]. Mitogen-activated protein kinase 1 (MAPK1) is a target of miR-378 [43], and it can mediate phosphorylation of PPARG leading to a reduction in transcriptional activity [42]. These facts indicate that miR-378 may induce both PPARG and PGC-1β to allow both to function synergistically in the regulation of lipid metabolism. Importantly, a recent study identified a role of miR-378 as a regulator of bovine preadipocyte differentiation [44].
In general, we observed that when PPARG was activated, miR-378 was inhibited, hence, casting doubt on a positive effect of miR-378 on PPARG activation. MiR-378 activation leads to inhibition of ADIPOQ [45], but only in the case of NW-MPN steers, PPARG, ADIPOQ and miR-378 had a similar expression profile (Tables 3, 4, 5). The data suggest that miR-378 activation in NW-MPN steers before 187 days of age might have been influenced by a higher proportion of preadipocytes in the LM or by a higher intracellular concentration of free fatty acids in LM which were previously reported to activate both miR-378 and adipocyte differentiation [46].
MiR-378 is considered a novel target for controlling adipose tissue inflammation because secretion of adipokines and cytokines (i.e., IL-6, TNF-α) due to onset of obesity upregulates miR-378 expression through SREBP and C/EBP binding sites [47]. In the microarray analysis of LM from these animals [8], data revealed a clear innate immune response in EW steers between 78 and 187 days of age, hence, it is unlikely that the marked upregulation of miR-378 through 187 days of age in NW-MPN steers was related to IMF inflammation. Furthermore, SREBF1 upregulation was not significant (P = 0.08) (Additional Figure 3) while C/EBPA was upregulated when miR-378 was downregulated (Tables 3, 4, 5). Thus, the data could suggest that secretion of cytokines within the LM might control miR-378 expression more likely through its C/EBP binding sites.
Overall, at 187 days NW-MPN steers had the highest level of activation of all pro-adipogenic miRNA analyzed. Because the NW-MPN steers remained in the feedlot for a longer period of time to achieve the desired threshold of backfat for slaughter (Additional Table 2), it is possible that in these animals adipogenic corepressors in LM hampered the activity of the pro-adipogenic microRNA. Hence, the marked upregulation of these miRNA.
Anti-adipogenic MicroRNA
MiR-27 a/b
MiR-27 inhibits adipogenesis partly by repressing the early adipogenic transcription factor cAMP response element-binding protein (CREB) [48]. Tumor necrosis factor alpha treatment upregulates miR-27 through the NF-κB pathway [48]. The utilization of an anti-miR-27 reduces the TNF-α-induced inhibition of adipogenesis. The levels of miR-27 expression decrease in mature adipocytes of obese mice when compared with lean mice [48]. In the present study, the downregulation of miR-27 a/b when TNF was activated (Tables 3, 4, 5 and Additional Figure 3) suggests that control of adipogenesis through miR-27 after 187 days of age could have been impaired by the cytokine, especially in NW-MPN and NW-LPN steers. This hypothesis is supported not by the degree of change in miR-27a/b expression but because miR-27 a/b was highly abundant as compared to other anti-adipogenic miRNA. As such, it can be surmised that a slight upregulation of miR-27a/b would be enough to produce the inhibitory effect on adipogenesis.
The negative effect of miR-27 on adipogenesis is counterbalanced by activation of Wnt/β-catenin signaling [49] leading to myocyte differentiation [50]. In a previous study, marked upregulation of miR-27b partly explained muscle hypertrophy in Piedmontese cattle due to its linkage with myostatin (MSTN) [51]. MiR-27a over expression resulted in reduced MSTN expression, skeletal muscle hypertrophy and an increase in the number of activated satellite cells [52]. In the present study, the upregulation of miR-27a/b could have affected MSTN expression but MSTN downregulation was not significant (P = 0.41) between 78 and 187 days of age (Additional Figure 3). Taken together, miR-27a/b expression had certain response to treatments in NW-MPN steers between 187 and 354 days of age but, in general, there was no consistency in miR-27a/b response to treatments.
MiR-130a
A previous study provided evidence of miR-130 as a potential negative regulator of adipogenesis by inhibiting PPARG [53]. In another study with adipocytes [54], pro-inflammatory responses mediated by TNF-α treatment resulted in adipocyte dysfunction by down-regulating PPARG expression. Inhibition of miR-130a and miR-130b using specific miRNA inhibitors significantly restored PPARG expression upon TNF-α stimulation. These results indicate that miR-130 induced by the TNF-α-mediated inflammatory response could downregulate PPARG expression. Taken together, miR-130a action in MPN steers could have impaired normal PPARG activation between 78 and 187 days of age, and subsequently between 187 and 354 days of age. Therefore, through TNF stimulation, miR-130a could have maintained PPARG expression at a constant level (i.e., impairing its activation) in steers that received less dietary starch in utero and postpartum (Tables 3, 4, 5 and Additional Figure 3).
MiR-34a
The different pattern of expression of miR-34a in response to maternal plane of nutrition at 78 days of age provides the closest evidence for epigenetic regulation of LM in offspring steers. In human subcutaneous adipose tissue, miR-34a was positively upregulated during adipogenesis, and correlated with body mass index [55]. MiR-34a inhibits SIRT1 expression through a miR-34a-binding site within the 3′UTR of SIRT1 [56]. TP53 induces expression of miR-34a which suppresses SIRT1 [57]. SIRT1 also regulates TP53-dependent apoptosis through deacetylation and stabilization of TP53. Furthermore, TP53 and miR-34 function in parallel to activate p21 to induce cell cycle arrest [58]. Between 78 and 354 days of age, miR-34a expression was not affected by treatment (Tables 3, 4, 5), but it was significantly different at 78 days of age (Additional Figure 2). In contrast, SIRT1 remained unchanged in EW steers while TP53 was upregulated at this time (Additional Figure 3). These data suggest that miR-34a has a role on the activation of cell cycle arrest in LM by suppressing SIRT1 activation, which leads to TP53 activation between 78 and 187 days of age.
MiR-369-5p
The relative miRNA abundance of miR-369-5p was the second lowest in the present study, thus, its relevance to intramuscular adipogenesis appears doubtful. It was previously determined that miR-369-5p directly targets FABP4 in human mesenchymal stem cells [59]. MiR-369-5p binds directly to the 3′UTR of FABP4 to decrease gene expression and thereby inhibit adipogenesis. This miRNA inhibits the adipogenic factors ADIPOQ and FABP4 and causes an overall reduction in lipid droplet formation [59]. In addition to its low miRNA abundance, the fact that expression of C/EBPA and PPARG did not correlate with that of miR-369-5p is further support that this miRNA is not relevant for intramuscular fat deposition in steers.
MiR-448
MiR-448-mediated repression of Krüppel-like factor 5 (KLF5) at its 3′ UTR contributes to inhibition of adipocyte differentiation [60], which was confirmed by the lower expression of adipogenic genes and intracellular triacylglycerol [61]. KLF5 is activated by C/EBPB and C/EBPD and, in concert with these C/EBP [62], contributes to PPARG induction which regulates adipocyte differentiation [60]. The expression of C/EBPA and PPARG decreased in cells transfected with miR-448. In contrast, C/EBPB and C/EBPD were not affected [61]. From the microarray data it can be surmised that KLF5 upregulation between 187 and 354 days of age in EW-MPN steers (Additional Figure 3) was partly due to the downregulation of miR-448. It is noteworthy that miR-448 relative expression was quite low as compared with other miRNA, but its expression in EW-MPN between 78 and 187 days increased substantially. Clearly, additional research needs to be performed to better understand the functional cause of miR-448 activation in LM of EW-MPN steers.
In conclusion, miR-34a was the sole miRNA studied in offspring LM that had a significant change in expression at 78 days of age due to cow plane of nutrition. Among the miRNA studied, this fact makes miR-34a the one with a potential sign of epigenetic regulation of LM of beef offspring due to cow plane of nutrition during late gestation. Early weaning age was expected to potentiate the dam supplementation effect but our results did not reveal a clear and concise response to make this hypothesis true. Furthermore, there was a lack of difference in carcass quality traits. A possible reason for this could be that the level of activation of the microRNA studied was not enough to produce the expected change in the steer carcass conformation. Nevertheless, consistent activation of the pro-adipogenic microRNA in general denoted that epigenetic control of LM metabolism was positively affected by selected pro-adipogenic miRNA between 78 and 187 days of age, especially in animals that had a delay in reaching target back fat thickness (NW-MPN) during the finishing phase.
Abbreviations
- ADIPOQ:
-
Adiponectin
- ADG:
-
Average daily gain
- CAV1:
-
Caveolin 1
- C/EBPA:
-
CCAAT/enhancer binding protein (C/EBP), alpha
- C/EBPB:
-
CCAAT/enhancer binding protein (C/EBP), beta
- C/EBPD:
-
CCAAT/enhancer binding protein (C/EBP), delta
- CREB:
-
cAMP response element binding protein
- DLK1:
-
Delta-like 1 homolog
- DM:
-
Dry matter
- DMI:
-
Dry matter intake
- ERK1/2:
-
Mitogen-activated protein kinase 3
- EW:
-
Early wean(ed)
- FABP4:
-
Fatty acid binding protein 4
- G0S2:
-
G0/G1 switch 2
- hASC:
-
Human adipose tissue-derived mesenchymal stem cells
- HPN:
-
High plane of nutrition
- IL-6:
-
Interleukin 6
- INSR:
-
Insulin receptor
- KLF5:
-
Kruppel-like factor 5
- LM:
-
Longissimus muscle
- LPN:
-
Low plane of nutrition
- MAPK1:
-
Mitogen-activated protein kinase 1
- miRNA:
-
MicroRNA
- miR-16b:
-
MicroRNA 16a
- miR-let-7a:
-
MicroRNA let7a
- miR-181a:
-
MicroRNA 181a
- miR-103:
-
MicroRNA 103
- miR-143:
-
MicroRNA 143
- miR-21-5p:
-
MicroRNA 21-5p
- miR-378:
-
MicroRNA 378
- miR-27a/b:
-
MicroRNA 27a/b
- miR-130a:
-
MicroRNA 130a
- miR-34a:
-
MicroRNA 34a
- miR-369-5p:
-
MicroRNA 369-5p
- miR-448:
-
MicroRNA 448
- MPN:
-
Medium plane of nutrition
- MSTN:
-
Myostatin
- MTG1:
-
Mitochondrial ribosome-associated GTPase 1
- NF-κB:
-
Nuclear factor of kappa light polypeptide gene enhancer in B-cells
- NW:
-
Normal wean(ed)
- PANK:
-
Pantothenate kinase
- PDCD4:
-
Programmed cell death 4
- PGC-1β:
-
Peroxisome proliferator-activated receptor gamma coactivator 1 beta
- PPARG:
-
Peroxisome proliferator-activated receptor gamma
- PTEN:
-
Phosphatase and tensin homolog
- qPCR:
-
Quantitative RT-PCR
- RPS15A:
-
Ribosomal protein S15a
- SIRT1:
-
Sirtuin 1
- SLC2A4:
-
Solute carrier family 2 (facilitated glucose transporter), member 4
- SREBP:
-
Sterol responsive element binding protein
- STAT3:
-
Signal transducer and activator of transcription 3
- TGFBR2:
-
Transforming growth factor, beta receptor II
- TNF:
-
Tumor necrosis alpha
- TNF-α:
-
Tumor necrosis alpha
- TP53:
-
Tumor protein p53
- UTR:
-
Untranslated region
- UXT:
-
Ubiquitously-expressed
References
Wilkins JF (2005) Genomic imprinting and methylation: epigenetic canalization and conflict. Trends Genet 21:356–365
Munshi A, Shafi G, Aliya N, Jyothy A (2009) Histone modifications dictate specific biological readouts. J Genet Genom 36:75–88
McKay JA, Mathers JC (2011) Diet induced epigenetic changes and their implications for health. Acta Physiol (Oxf) 202:103–118
Bartel DP (2009) MicroRNAs: target recognition and regulatory functions. Cell 136:215–233
Jin W, Grant JR, Stothard P, Moore SS, Guan LL (2009) Characterization of bovine miRNAs by sequencing and bioinformatics analysis. BMC Mol Biol 10:90
Basu U, Romao JM, Guan LL (2012) Adipogenic transcriptome profiling using high throughput technologies. J Genom 1:22–28
Erhuma A, Salter AM, Sculley DV, Langley-Evans SC, Bennett AJ (2007) Prenatal exposure to a low-protein diet programs disordered regulation of lipid metabolism in the aging rat. Am J Physiol Endocrinol Metab 292:E1702–E1714
Moisa SJ, Shike DW, Shoup L, Rodriguez-Zas SL, Loor JJ (2015) Maternal plane of nutrition during late gestation and weaning age alter Angus × Simmental offspring longissimus muscle transcriptome and intramuscular fat. PLoS One 10:e0131478
Shoup LM, Wilson TB, González-Peña D, Ireland FA, Rodriguez-Zas S, Felix TL, Shike DW (2015) Beef cow prepartum supplement level and age at weaning: II. Effects of developmental programming on performance and carcass composition of steer progeny. J Anim Sci 93(10):4936–4947. doi:10.2527/jas2014-8565
Shoup LM, Kloth AC, Wilson TB, González-Peña D, Ireland FA, Rodriguez-Zas S, Felix TL, Shike DW (2015) Prepartum supplement level and age at weaning: I. Effects on pre- and postpartum beef cow performance and calf performance through weaning. J Anim Sci 93(10):4926-35. doi: 10.2527/jas2014-8564
Raddatz J (2008) Measurement of adiponectin and insulin in lactating and non-lactating Holstein cows in Animal Sciences. North Carolina State University, Raleigh, p 79. http://repository.lib.ncsu.edu/ir/handle/1840.1816/1845
Funston RN, Larson DM, Vonnahme KA (2010) Effects of maternal nutrition on conceptus growth and offspring performance: implications for beef cattle production. J Anim Sci 88:E205–E215
Bohnert DW, Stalker LA, Mills RR, Nyman A, Falck SJ, Cooke RF (2013) Late gestation supplementation of beef cows differing in body condition score: effects on cow and calf performance. J Anim Sci 91:5485–5491
Moisa SJ, Shike DW, Faulkner DB, Meteer WT, Keisler D, Loor JJ (2014) Central role of the PPARgamma gene network in coordinating beef cattle intramuscular adipogenesis in response to weaning age and nutrition. Gene Regul Syst Bio 8:17–32
Hoffstedt J, Arvidsson E, Sjolin E, Wahlen K, Arner P (2004) Adipose tissue adiponectin production and adiponectin serum concentration in human obesity and insulin resistance. J Clin Endocrinol Metab 89:1391–1396
Smith J, Al-Amri M, Sniderman A, Cianflone K (2006) Leptin and adiponectin in relation to body fat percentage, waist to hip ratio and the apoB/apoA1 ratio in Asian Indian and Caucasian men and women. Nutr Metab (Lond) 3:18
Kabara E, Sordillo LM, Holcombe S, Contreras GA (2014) Adiponectin links adipose tissue function and monocyte inflammatory responses during bovine metabolic stress. Comp Immunol Microbiol Infect Dis 37:49–58
Ji P, Drackley JK, Khan MJ, Loor JJ (2014) Overfeeding energy upregulates peroxisome proliferator-activated receptor (PPAR)gamma-controlled adipogenic and lipolytic gene networks but does not affect proinflammatory markers in visceral and subcutaneous adipose depots of Holstein cows. J Dairy Sci 97:3431–3440
Ji P, Drackley JK, Khan MJ, Loor JJ (2014) Inflammation- and lipid metabolism-related gene network expression in visceral and subcutaneous adipose depots of Holstein cows. J Dairy Sci 97:3441–3448
Arita Y, Kihara S, Ouchi N, Takahashi M, Maeda K, Miyagawa J, Hotta K, Shimomura I, Nakamura T, Miyaoka K, Kuriyama H, Nishida M, Yamashita S, Okubo K, Matsubara K, Muraguchi M, Ohmoto Y, Funahashi T, Matsuzawa Y (2012) Paradoxical decrease of an adipose-specific protein, adiponectin, in obesity. 1999. Biochem Biophys Res Commun 425:560–564
Ohtani Y, Takahashi T, Sato K, Ardiyanti A, Song SH, Sato R, Onda K, Wada Y, Obara Y, Suzuki K, Hagino A, Roh SG, Katoh K (2012) Changes in circulating adiponectin and metabolic hormone concentrations during periparturient and lactation periods in Holstein dairy cows. Anim Sci J 83:788–795
Xie H, Lim B, Lodish HF (2009) MicroRNAs induced during adipogenesis that accelerate fat cell development are downregulated in obesity. Diabetes 58:1050–1057
Sarr O, Thompson JA, Zhao L, Lee TY, Regnault TR (2014) Low birth weight male guinea pig offspring display increased visceral adiposity in early adulthood. PLoS One 9:e98433
Polster BJ, Westaway SK, Nguyen TM, Yoon MY, Hayflick SJ (2010) Discordant expression of miR-103/7 and pantothenate kinase host genes in mouse. Mol Genet Metab 101:292–295
Wilfred BR, Wang WX, Nelson PT (2007) Energizing miRNA research: a review of the role of miRNAs in lipid metabolism, with a prediction that miR-103/107 regulates human metabolic pathways. Mol Genet Metab 91:209–217
Ragaller V, Lebzien P, Sudekum KH, Huther L, Flachowsky G (2011) Pantothenic acid in ruminant nutrition: a review. J Anim Physiol Anim Nutr (Berl) 95:6–16
Rottiers V, Naar AM (2012) MicroRNAs in metabolism and metabolic disorders. Nat Rev Mol Cell Biol 13:239–250
Trajkovski M, Hausser J, Soutschek J, Bhat B, Akin A, Zavolan M, Heim MH, Stoffel M (2011) MicroRNAs 103 and 107 regulate insulin sensitivity. Nature 474:649–653
Wang H, Xiao S, Wang M, Kim NH, Li H, Wang G (2015) In silico identification of conserved microRNAs and their targets in bovine fat tissue. Gene 559(2):119–128
Li H, Zhang Z, Zhou X, Wang Z, Wang G, Han Z (2011) Effects of microRNA-143 in the differentiation and proliferation of bovine intramuscular preadipocytes. Mol Biol Rep 38:4273–4280
Zhu L, Shi C, Ji C, Xu G, Chen L, Yang L, Fu Z, Cui X, Lu Y, Guo X (2013) FFAs and adipokine-mediated regulation of hsa-miR-143 expression in human adipocytes. Mol Biol Rep 40:5669–5675
Kim YJ, Min TS, Seo KS, Kim SH (2015) Expression of pref-1/dlk-1 is regulated by microRNA-143 in 3T3-L1 cells. Mol Biol Rep 42:617–624
Kavalkova P, Touskova V, Roubicek T, Trachta P, Urbanova M, Drapalova J, Haluzikova D, Mraz M, Novak D, Matoulek M, Lacinova Z, Haluzik M (2013) Serum preadipocyte factor-1 concentrations in females with obesity and type 2 diabetes mellitus: the influence of very low calorie diet, acute hyperinsulinemia, and fenofibrate treatment. Horm Metab Res 45:820–826
Keller P, Gburcik V, Petrovic N, Gallagher IJ, Nedergaard J, Cannon B, Timmons JA (2011) Gene-chip studies of adipogenesis-regulated microRNAs in mouse primary adipocytes and human obesity. BMC Endocr Disord 11:7
Kim YJ, Hwang SJ, Bae YC, Jung JS (2009) MiR-21 regulates adipogenic differentiation through the modulation of TGF-beta signaling in mesenchymal stem cells derived from human adipose tissue. Stem Cells 27:3093–3102
Kim YJ, Hwang SH, Cho HH, Shin KK, Bae YC, Jung JS (2012) MicroRNA 21 regulates the proliferation of human adipose tissue-derived mesenchymal stem cells and high-fat diet-induced obesity alters microRNA 21 expression in white adipose tissues. J Cell Physiol 227:183–193
Frankel LB, Christoffersen NR, Jacobsen A, Lindow M, Krogh A, Lund AH (2008) Programmed cell death 4 (PDCD4) is an important functional target of the microRNA miR-21 in breast cancer cells. J Biol Chem 283:1026–1033
Subedi A, Kim MJ, Nepal S, Lee ES, Kim JA, Sohn DH, Song K, Lee SH, Park WS, Jeong BS, Park PH (2013) Globular adiponectin modulates expression of programmed cell death 4 and miR-21 in RAW 264.7 macrophages through the MAPK/NF-kappaB pathway. FEBS Lett 587:1556–1561
Kang M, Yan LM, Zhang WY, Li YM, Tang AZ, Ou HS (2013) Role of microRNA-21 in regulating 3T3-L1 adipocyte differentiation and adiponectin expression. Mol Biol Rep 40:5027–5034
Eichner LJ, Perry MC, Dufour CR, Bertos N, Park M, St-Pierre J, Giguere V (2010) miR-378(*) mediates metabolic shift in breast cancer cells via the PGC-1beta/ERRgamma transcriptional pathway. Cell Metab 12:352–361
Romao JM, Jin W, He M, McAllister T, le Guan L (2014) MicroRNAs in bovine adipogenesis: genomic context, expression and function. BMC Genom 15:137
Jin W, Dodson MV, Moore SS, Basarab JA, Guan LL (2010) Characterization of microRNA expression in bovine adipose tissues: a potential regulatory mechanism of subcutaneous adipose tissue development. BMC Mol Biol 11:29
Ganesan J, Ramanujam D, Sassi Y, Ahles A, Jentzsch C, Werfel S, Leierseder S, Loyer X, Giacca M, Zentilin L, Thum T, Laggerbauer B, Engelhardt S (2013) MiR-378 controls cardiac hypertrophy by combined repression of mitogen-activated protein kinase pathway factors. Circulation 127:2097–2106
Liu SY, Zhang YY, Gao Y, Zhang LJ, Chen HY, Zhou Q, Chai ML, Li QY, Jiang H, Yuan B, Dai LS, Zhang JB (2015) MiR-378 Plays an Important Role in the Differentiation of Bovine Preadipocytes. Cell Physiol Biochem 36:1552–1562
Ishida M, Shimabukuro M, Yagi S, Nishimoto S, Kozuka C, Fukuda D, Soeki T, Masuzaki H, Tsutsui M, Sata M (2014) MicroRNA-378 regulates adiponectin expression in adipose tissue: a new plausible mechanism. PLoS One 9:e111537
Gerin I, Bommer GT, McCoin CS, Sousa KM, Krishnan V, MacDougald OA (2010) Roles for miRNA-378/378* in adipocyte gene expression and lipogenesis. Am J Physiol Endocrinol Metab 299:E198–E206
Jiang X, Xue M, Fu Z, Ji C, Guo X, Zhu L, Xu L, Pang L, Xu M, Qu H (2014) Insight into the effects of adipose tissue inflammation factors on miR-378 expression and the underlying mechanism. Cell Physiol Biochem 33:1778–1788
Zhu Y, Zhang X, Ding X, Wang H, Chen X, Zhao H, Jia Y, Liu S, Liu Y (2014) miR-27 inhibits adipocyte differentiation via suppressing CREB expression. Acta Biochim Biophys Sin (Shanghai) 46:590–596
Qin L, Chen Y, Niu Y, Chen W, Wang Q, Xiao S, Li A, Xie Y, Li J, Zhao X, He Z, Mo D (2010) A deep investigation into the adipogenesis mechanism: profile of microRNAs regulating adipogenesis by modulating the canonical Wnt/beta-catenin signaling pathway. BMC Genom 11:320
Zhao JX, Hu J, Zhu MJ, Du M (2011) Trenbolone enhances myogenic differentiation by enhancing beta-catenin signaling in muscle-derived stem cells of cattle. Domest Anim Endocrinol 40:222–229
Miretti S, Martignani E, Accornero P, Baratta M (2013) Functional effect of mir-27b on myostatin expression: a relationship in Piedmontese cattle with double-muscled phenotype. BMC Genom 14:194
McFarlane C, Vajjala A, Arigela H, Lokireddy S, Ge X, Bonala S, Manickam R, Kambadur R, Sharma M (2014) Negative auto-regulation of myostatin expression is mediated by Smad3 and microRNA-27. PLoS One 9:e87687
Guilherme A, Virbasius JV, Puri V, Czech MP (2008) Adipocyte dysfunctions linking obesity to insulin resistance and type 2 diabetes. Nat Rev Mol Cell Biol 9:367–377
Kim C, Lee H, Cho YM, Kwon OJ, Kim W, Lee EK (2013) TNFalpha-induced miR-130 resulted in adipocyte dysfunction during obesity-related inflammation. FEBS Lett 587:3853–3858
Ortega FJ, Moreno-Navarrete JM, Pardo G, Sabater M, Hummel M, Ferrer A, Rodriguez-Hermosa JI, Ruiz B, Ricart W, Peral B, Fernandez-Real JM (2010) MiRNA expression profile of human subcutaneous adipose and during adipocyte differentiation. PLoS One 5:e9022
Chen F, Hu SJ (2012) Effect of microRNA-34a in cell cycle, differentiation, and apoptosis: a review. J Biochem Mol Toxicol 26:79–86
Yamakuchi M, Lowenstein CJ (2009) MiR-34, SIRT1 and p53: the feedback loop. Cell Cycle 8:712–715
Yamakuchi M, Ferlito M, Lowenstein CJ (2008) miR-34a repression of SIRT1 regulates apoptosis. Proc Natl Acad Sci USA 105:13421–13426
Bork S, Horn P, Castoldi M, Hellwig I, Ho AD, Wagner W (2011) Adipogenic differentiation of human mesenchymal stromal cells is down-regulated by microRNA-369-5p and up-regulated by microRNA-371. J Cell Physiol 226:2226–2234
Oishi Y, Manabe I, Tobe K, Tsushima K, Shindo T, Fujiu K, Nishimura G, Maemura K, Yamauchi T, Kubota N, Suzuki R, Kitamura T, Akira S, Kadowaki T, Nagai R (2005) Kruppel-like transcription factor KLF5 is a key regulator of adipocyte differentiation. Cell Metab 1:27–39
Kinoshita M, Ono K, Horie T, Nagao K, Nishi H, Kuwabara Y, Takanabe-Mori R, Hasegawa K, Kita T, Kimura T (2010) Regulation of adipocyte differentiation by activation of serotonin (5-HT) receptors 5-HT2AR and 5-HT2CR and involvement of microRNA-448-mediated repression of KLF5. Mol Endocrinol 24:1978–1987
Farmer SR (2006) Transcriptional control of adipocyte formation. Cell Metab 4:263–273
Pratt SL, Burns TA, Curry E, Duckett SK (2010) Expression of microRNA during bovine adipogenesis. J Nucleic Acids Investig 1:12
Hulsmans M, De Keyzer D, Holvoet P (2011) MicroRNAs regulating oxidative stress and inflammation in relation to obesity and atherosclerosis. FASEB J 25:2515–2527
Kang M, Yan LM, Li YM, Zhang WY, Wang H, Tang AZ, Ou HS (2013) Inhibitory effect of microRNA-24 on fatty acid-binding protein expression on 3T3-L1 adipocyte differentiation. Genet Mol Res 12:5267–5277
McGregor RA, Choi MS (2011) microRNAs in the regulation of adipogenesis and obesity. Curr Mol Med 11:304–316
Romao JM, Jin W, Dodson MV, Hausman GJ, Moore SS, le Guan L (2011) MicroRNA regulation in mammalian adipogenesis. Exp Biol Med (Maywood) 236:997–1004
Lin Q, Gao Z, Alarcon RM, Ye J, Yun Z (2009) A role of miR-27 in the regulation of adipogenesis. FEBS J 276:2348–2358
Sacco J, Adeli K (2012) MicroRNAs: emerging roles in lipid and lipoprotein metabolism. Curr Opin Lipidol 23:220–225
Lee J, Kemper JK (2010) Controlling SIRT1 expression by microRNAs in health and metabolic disease. Aging (Albany NY) 2:527–534
John E, Wienecke-Baldacchino A, Liivrand M, Heinaniemi M, Carlberg C, Sinkkonen L (2012) Dataset integration identifies transcriptional regulation of microRNA genes by PPARgamma in differentiating mouse 3T3-L1 adipocytes. Nucleic Acids Res 40:4446–4460
Esau C, Kang X, Peralta E, Hanson E, Marcusson EG, Ravichandran LV, Sun Y, Koo S, Perera RJ, Jain R, Dean NM, Freier SM, Bennett CF, Lollo B, Griffey R (2004) MicroRNA-143 regulates adipocyte differentiation. J Biol Chem 279:52361–52365
Kajimoto K, Naraba H, Iwai N (2006) MicroRNA and 3T3-L1 pre-adipocyte differentiation. RNA 12:1626–1632
Acknowledgments
We gratefully acknowledge the help from J. Segers, T. B. Wilson and L. Shoup at the time of collecting tissue samples, as well as the rest of the staff at the University of Illinois Beef and Sheep Field Laboratory for animal handling and care. Supported in part by a competitive grant from the “Division of Nutritional Sciences: Vision 20/20’’ (University of Illinois, Urbana-Champaign) interdisciplinary nutrition-related research program. Project # ILLU-971-352 (National Institute of Food and Agriculture, USA).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
Author(s) disclose no potential conflicts of interest.
Electronic supplementary material
Below is the link to the electronic supplementary material.
About this article
Cite this article
Moisá, S.J., Shike, D.W., Shoup, L. et al. Maternal Plane of Nutrition During Late-Gestation and Weaning Age Alter Steer Calf Longissimus Muscle Adipogenic MicroRNA and Target Gene Expression. Lipids 51, 123–138 (2016). https://doi.org/10.1007/s11745-015-4092-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11745-015-4092-y