Abstract
Fungal endophytic species inhabiting the leaves of eucalypts are capable of utilising leaf sugars and can influence both plant growth and health. Endophytic fungal symbionts can use simple soluble sugars in leaves as their main carbon source. This study set out to determine the diversity and distribution of the endophytic filamentous fungal leaf community in the hybrid Eucalyptus urograndis due to its economic importance. The fungal leaf community was characterised using denaturing gradient electrophoresis (DGGE) and correlated with levels of leaf nutrients and sugars throughout plant development. Sequencing of DGGE bands revealed the presence of Basidiomycota and Ascomycota phyla. Fourteen species and three genera of filamentous fungi were identified, and the population structure was affected by the plant developmental stage. Levels of K, Cu, N and Mn influenced communities from the clonal garden, whereas leaves in the field had higher glucose, fructose and sucrose. Many fungi were found to be specific to a certain development stages: Diplomitoporus crustulinus, Podosphaera tridactyla and Aspergillus restrictus to the clonal garden stage; Chaetomella acutiseta and Ascotricha chartarum to the shading stage; Erratomyces patelii and Saxomyces sp. to the shading output stage; Lepidostroma sp. and Saxomyces sp. to the dispatch stage; and Mycosphaerella populicola to the field stage. Teratosphaeria toledana and Teratosphaeria acidotherma were found at more than one developmental stage. Cladosporium sp. and Rhodosporidium fluviale colonized and persisted in plants at the dispatch and field stages. This is the first report of P. tridactyla, A. restrictus, E. patelii, Saxomyces and Lepidostroma sp. as endophytes in eucalipt.
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
Introduction
Aerial parts of plants, especially the leaves, host a great diversity of endophytic and epiphytic microorganisms. Endophytes are organisms that live inside plant tissues without causing disease symptoms (Arnold and Lutzoni 2007) and, in an association with trees, may reduce damage to the leaves and reduce losses caused by pathogens (Arnold and Herre 2003). Endophytic fungi are also important components of biodiversity in several species of plants (Arnold et al. 2000). Eucalyptus spp. (Myrtaceae) are widely grown in commercial plantations around the world (FAO 2016) because of their rapid growth, adaptability and valuable wood (Brooker 2000). The diversity and abundance of filamentous fungi in eucalyptus are being elucidated using culture- or cultivation-independent methods with greater fidelity to reveal the complexity of these communities and their interactions with the plant (Kemler et al. 2013) and demonstrate that the endophytic fungal microbiota is more diverse than previously reported.
Fungal community diversity can be studied by PCR and denaturing gradient electrophoresis (DGGE) targeting the 18S rRNA gene or the internal transcribed spacer region (ITS). Specifically, fragments generated by amplification of 18S rRNA are considered practical markers in molecular approaches (Vainio and Hantula 2000; Duong et al. 2006), and comparisons generated by phylogenetic profiles based on the number of bands suggest that this region is more discriminatory than the ITS region (Evans et al. 2014).
Endophytic filamentous fungi are abundant in asymptomatic leaves of several tropical trees (Arnold 2005). These microorganisms establish different interactions with plants affecting the associated microbial community diversity and the adaptability and evolution of the host (Omacini et al. 2001; Brundrett 2006). Furthermore, these microorganisms have received significant interest due to their ability to produce compounds of industrial interest (Lumyong et al. 2002; Robl et al. 2013) including enzymes such as cellulase, xylanase, laccase, pectin lyase and hemicellulase (Robl et al. 2013).
Brazil has the most area with eucalyptus plantations in the world (FAO 2015). Although many studies have targeted the diversity of fungal communities associated with eucalyptus (Cheewangkoon et al. 2009; Sánchez Márquez et al. 2011), the dynamics of endophytic microbial communities during the development of nursery plants and in the field are unknown. Understanding the impact of seedling production practices in nurseries on the structure of endophytic microbial communities is important and can contribute to producing healthier and resistant seedlings. Thus, the aim of this study was to evaluate the diversity of endophytic filamentous fungi associated with eucalyptus leaves at different stages of development of the plant in commercial nurseries and in field, and then relate the communities to the nutrients and sugar concentrations of leaves.
Materials and methods
Plant material and leaf sampling
Leaf samples from selected seedlings of Eucalyptus grandis × Eucalyptus urophyla (eucalyptus “urograndis”) were collected at the following stages of plant development and cultivation: clonal garden, shaded, recently out from shade, dispatch and field (18-month-old plants). Plants at the clonal garden, shaded, recently out from shade and dispatch stages were selected for sampling by randomly demarcating four blocks in the nursery with each block a repetition and divided into four equal parts to locate each sampling point for a more representative sampling procedure. These points were then randomly (by lot) selected and according to the drawing, the leaves were sampled from different plants and mixed into composite samples. The leaves from the field stage were selected from four 18-month-old plants, each plant representing a repetition. The trees were located in a sub-area of 162 m2 (about 16 trees), with a spacing of 3.33 m. This area was divided into four equal parts, and a tree chosen in each plot to sample the upper, middle and lower region of the canopy. For a more representative sample from the whole canopy, leaves were collected from proximal, median and distal parts of the stem for each region, then mixed to form composite samples. Samples were stored in sterile plastic bags and immediately transferred to Styrofoam coolers containing ice, transported to the Laboratory of Microbial Ecology and stored at − 20 °C until processing for the diversity analyses. The sampling procedure and surface disinfestation of leaves were described by Miguel et al. (2016).
Extraction of sugars and nutrients
Glucose, fructose and sucrose were measured according to Lisec et al. (2006) with modifications. About 50–60 mg of leaf were macerated and transferred to polypropylene tube with 1000 µL of methanol (100%). This mixture was heated at 70 °C with stirring at 950 rpm for 60 min, followed by centrifugation at 14,000×g for 10 min. Approximately 600 µL of the supernatant solution were transferred into tubes containing 500 µL of chloroform and 800 µL of distilled water. After centrifugation at 12,000×g for 15 min, the polar phase was collected (1000 µL), and sugars were quantified by coupled enzyme assays measuring the increase in optical density at 340 nm (Multiscan FC, Thermo Scientific, Waltham, MA, USA). Enzymes used were glucose-6-phosphatase (0.2 mg/mL), hexokinase (0.2 mg/mL), phosphoglucose isomerase (0.2 mg/mL) and invertase (1 mg/mL). Successively, these enzymes were added to the leaf extract and the concentrations of glucose, fructose and sucrose expressed in mmol kg−1 leaf dry mass. Sucrose concentration was estimated as half the hexose concentration. The data were submitted to analysis of variance at 5% probability and the averages by Tukey’s test at 5% probability using Minitab version 15 (Minitab 2006) (Minitab Inc., State College, PA, USA).
Nutrient content (Ca, Mg, K, P, K, Zn, Fe, Cu and Mn) in the leaves was analyzed by the Soil, Vegetal Tissue, and Fertilizer Analysis Laboratory in the Department of Soils located at the Universidade Federal de Viçosa. Leaves were macerated and mineralized with a 4:1 v/v nitric-perchloric mixture. N levels were then quantified in concentrated sulfuric acid by the Kjeldahl method (Bremner and Mulvaney 1982), and other nutrients were quantified by atomic absorption spectrometry. The average concentrations of the macro- and micronutrients were expressed as g kg−1 and mg kg−1 leaf dry mass, respectively.
Analysis of fungal diversity
Fungal diversity was studied using genomic DNA extracted from leaves as described by Miguel et al. (2016) as template for a nested PCR-DGGE. Primers NS1 (May et al. 2001) and EF3 (Oros-Sichler et al. 2006) targeting the region V1–V9 of the 18S rRNA gene from fungi were used in the first round of PCR. Amplicons generated in the first PCR were used as templates for a second round of PCR using primers FF390 and FR1GC (Vainio and Hantula 2000) to amplify the V7–V8 region of the rRNA gene.
Amplicons were separated by denaturing gradient electrophoresis (DGGE) (DCode System, Bio-Rad Inc., California). A mixture 16S rRNA amplified from respective pure cultures of Nocardioides thermolilacinus, Bacillus cereus, Streptomyces setonii, Clavibacter michiganensis, Pectobacterium carotovorum, Pseudomonas putida, Pseudomonas syringae, Xanthomonas vesicatoria and Ralstonia solanacearum was used as external markers to facilitate the normalization of the bands of the gels using the Bionumerics software version 7.1 (Applied Maths, Kortrijk, Belgium).
PCR products were applied in an 8% w/v polyacrylamide gel (Sigma) in denaturing gradient gel and in Tris-acetate-EDTA (TAE) 1× buffer (40 mmol L−1 Tris-hydrochloric acid, pH 8.20 mmol L−1 adjusted with acetic acid, 1 mmol L−1 EDTA, pH 8), 0.09% v/v TEMED (N, N, N, N-tetramethylene) and 0.07% w/v ammonium persulfate. The denaturant gradient was optimized to 35–55% (filamentous fungi) urea/formamide (100% denaturant contains 7 mol L−1 urea and 40% v/v formamide). The electrophoresis was performed in TAE 1× buffer at 60 V for 20 h at constant temperature of 60 °C. DNA fragments in the gel were stained for 20 min in 1× TAE buffer containing the dye SYBR Gold 1X (Invitrogen, Carlsbad, California, USA), and the images obtained by UV light using Molecular Imaging system LPIX Chemi (Loccus Biotechnology, São Paulo, SP, Brazil).
After excision of DGGE bands, amplicons were eluted from the gel in 30 μL of Milli-Q sterile water kept overnight at 4 °C. Seven microliters from the eluate of each band was used as a template for PCR using oligonucleotides initiators FF390 and FR1. Amplicons were sequenced by Macrogen, Inc. Korea, and their sequences were compared with those in the GenBank database (NCBI) using the BLASTn algorithm (http://www.ncbi.nlm.nih.gov/BLAST) for nucleotides (Altschul et al. 1990). Sequences generated in this study are available in the GenBank database under the GenBank ID numbers KX063837 to KX063850.
DGGE band profiles were analyzed using Bionumerics version 7.1. The variable richness from filamentous fungi was estimated by the program based on the binary matrix, in which the presence of the band corresponding to each operational taxonomic unit (OTU) was coded one (1), and the absence, zero (0). The structure of the community was evaluated based on the Dice similarity coefficient (DSC) and average clustering method between clusters (UPGMA). Richness and diversity were analyzed using PAST software (Hammer et al. 2001), the diversity was estimated from Shannon-Winner index, and the graphics were generated in the SigmaPlot software (Systat Software, Inc. 2008).
Phylogenetic analysis
Sequences in the database sharing more than 97% identity with those generated in this study were imported with Mega 6.0 and aligned using ClustalW. The alignments were manually adjusted, and a phylogenetic analysis was performed using the neighbor-joining method (Saitou and Nei 1987). The phylogenetic distance was computed using the p-distance method, and the robustness of the resulting trees and the statistical significance levels of the interior nodes were obtained by bootstrap analysis with 1000 replicates. Bootstrap values greater than 70% are shown, since nodes supported by values below this number are considered of poor resolution (Schneider 2007).
Statistical analyses
To verify any correlation between sugar (glucose, fructose and sucrose) content and the richness and diversity indexes at 5% probability (Systat Software, Inc. 2008), the Pearson correlation test was performed. The data were subjected to analysis of variance at 5% probability, and the averages were compared by Tukey’s test at 5% probability in Minitab software version 15.
Correlations among the occurrence of DGGE bands from endophytic fungi and the environmental data were determined using principal component analysis (PCA) with the Canoco software (version 4.5, Biometris, Wageningen, Netherlands). Distinct bands observed in the DGGE gels were considered as unique OTUs, and their relative intensities were considered as the frequency of occurrence. Nutrient (K, Cu, N, Mn, Fe, P, Mg, Ca and Zn) and sugar (glucose, fructose and sucrose) levels were considered as environmental variables.
Results
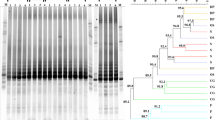
The PCR-DGGE analysis allowed the identification of distinct band profiles among samples representing changes in the endophytic fungal community influenced by the developmental stages of eucalyptus (Fig. 1). The analysis of diversity considered presence/absence and intensity of individual bands, allowing calculation of diversity indexes.
DGGE electrophoretic profile of the endophytic fungal community during the development of eucalyptus plants. a Gel from the DGGE with four replications for each stage (1–4). The letter F followed by the numerals indicate the location of the excised bands. CG clonal garden, S shading, OS recently out from shading, DP plants for dispatch, F plants in the field, M molecular weight markers. b Dendrogram constructed from the DGGE amplicons in A
Hierarchical clustering analysis by UPGMA allowed identification of more related samples based on band profile (Fig. 1b). Different clusters of samples sharing at least 25% similarity were identified (Fig. 1), with those more related sharing up to 64.7% similarity.
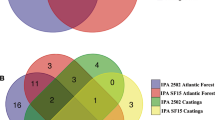
Leaves of plants from the garden clonal stage and from those newly removed from shading clustered together, sharing 43.9% similarity, whereas samples from the shading stage and the dispatch stage formed at least one clade with 43.9 and 64.2% similarity (Fig. 1). Although most OTUs (Fig. 2) were common among stages of development, each sample had some unique OTUs, confirming differences in fungal community (Fig. 2). In spite of being small, these changes indicate that developmental stages of eucalyptus harbor different communities of endophytic fungi. The diversity and richness of endophytic fungi, however, were not influenced by the phases of eucalyptus development (Fig. 3).
Shannon diversity index (a) and richness (b) of endophytic fungi at different stages of eucalyptus development. CG clonal garden, S plants in shade, OS plants newly out from shade, DP plants for dispatch, F plants in the field. Patterns with the same letter do not differ significantly by Tukey’s test at P < 0.05
Glucose and fructose contents were variable in the seedling stage and field stage samples, while sucrose levels were more similar. Highest levels of glucose and fructose were found in leaves of plants in the dispatch and the field stages, while sucrose was found only in the field (Fig. 4). Higher levels of N and P were found in plants from the garden stage and the shaded stage, while the levels of K in those at recently out from the shade were minors. Significant differences (Tukey’s test; p < 0.05) among levels of Ca and Mg were detected only in plants at the field stage. When micronutrients are considered, Cu was highest in plants from the clonal garden, Mn was highest in those recently out from the shade, and iron (Fe) was highest at the dispatch stage (Table 1).
Box plots showing sugar concentrations in eucalyptus leaves (mmol kg−1 dry mass leaves) at different stages of development. a Glucose, b fructose and c sucrose. CG clonal garden, S plants in shade, OS plants newly out from shade, DP plants for dispatch, F plants in the field. Patterns with the same letter do not differ significantly by Tukey’s test at P < 0.05
Correlation analysis between levels of carbohydrates and the Shannon diversity index detected a moderate negative correlation for samples clonal garden, shading, newly removed from shading and dispatch (− 0.8 < r ≤ − 0,5) (p > 0.05). Multivariate analysis demonstrated 73.8% of the variation in species distribution in samples was explained by environmental factors, with principal component 1 and 2 explaining 48.6% of the variation in the community of fungi at different stages of eucalyptus development (Fig. 5). Fungal communities in samples from the clonal garden were affected by levels of K, Cu, N and Mn from leaves, whereas glucose, fructose and sucrose contents influenced those communities in samples in field (Fig. 5).
The OTU identification was carried out by sequencing 18S rRNA after DGGE-band excision and amplification. When sequences were compared with those available in the NCBI databases, all OTUs were classified within Ascomycota or Basidiomycota, among three genera and four species of filamentous fungi (Fig. 6).
Interaction between endophytic fungi and eucalyptus leaves was dependent on the stage of plant development, since most species were exclusively found in one stage, except OTUs classified as Cladosporium sp. and T. acidotherma, which were present in more than one stage (Table 2). The observed difference can be the result of distinct environmental conditions at each stage of development, such as sugar concentration, bacterial competition and others. The fewest endophytic fungi identified by 18S rRNA sequencing were from the shaded stage, and they all were specific to that development stage (Table 2).
The OTU classifications by 18S rRNA sequencing was confirmed by phylogeny reconstruction of sequences in which sequences formed distinct clades corresponding to Ascomycota and Basidiomycota sequences (Fig. 6). Sequences obtained in this study grouped together with other sequences with moderate to strong bootstrap values (Schneider 2007) support of the analysis. OTU B07 clustered with Podosphaera genus, OTU B12 clustered with Ascotricha, and OTU B25 clustered with Mycosphaerella lateralis. These clusters with high bootstraps demonstrate that these UTOs belong to those genera or species, corroborating the comparison using BLASTn. UTO B10 clustered with the two fungi from the Diplomitoporus genus, the OTU B17 grouped with Erratomyces patelli and Conidiosporomyces ayresii species and OTU B21 clustered with Cortinarius (Fig. 6).
Discussion
Investigation of the diversity of the endophytic fungal leaf community using DGGE revealed the presence of OTUs with distinct intensities that reflect differences in the abundance of those populations. The DGGE technique has been shown to be suitable for evaluating microbial diversity in various environments (Bresolin et al. 2010; Kittelmann et al. 2012; Oliveira et al. 2013), including eucalyptus leaves (Miguel et al. 2016). This analysis proved to be more robust for analyzing complex microbial communities if combined with nested PCR (Oliveira et al. 2013), providing good resolution of less-abundant populations. The DGGE analysis using UPGMA provides fingerprinting patterns that can be obtained quickly (Fromin et al. 2002) and results in dendrograms that graphically show the similarities between samples (Laplante and Derome 2011).
In this study, nested-PCR-DGGE was a suitable technique to detect the community structures of endophytic filamentous fungi at various stages of eucalyptus development during cultivation, allowing the calculation of diversity and richness indexes, which ranged from 1.5 to 3.5, values commonly found in literature (Gazis and Chaverri 2010). Although differences were observed between DGGE band profiles and sequenced OTUs, no significant differences in diversity and richness were detected. However, the distribution of endophytic fungi among the different stages of eucalyptus development was distinct and can be related to the age and environmental conditions of the plant host (Jia et al. 2016). Nutrient content in plants is determined by many factors: nutrient requirement and availability, efficiency of nutrient absorption and utilization, and nutrient mobility (Marschener 1998). Endophytes can modify the kinetics of nutrient absorption by colonized plants (Rho and Kim 2017). The inoculation of aerial parts of Lolium with the endophytic fungus Neotyphodiun led to decreased levels of P, Ca, S and B, evidence that even colonization in aerial parts of a plant can affect nutrient absorption in roots (Soto-Barajas et al. 2016).
Leaves of eucalyptus harbored diverse communities of endophytic fungi in all stages of development studied. Fungi from the Myrtaceae family are considered to be an important source of fungal biodiversity, typically those associated with leaves of Corymbia, Syzygium spp. (Cheewangkoon et al. 2009) and Eucalyptus (Cheewangkoon et al. 2009; Miguel et al. 2017). The presence of endophytic fungi in leaves expands our understanding of endophytic colonization in eucalyptus (Miguel et al. 2017).
Endophytic fungal–plant interactions have been studied in other plant families such as the Malvaceae, Cecropiaceae, Lauraceae, Fabaceae, Lecythidaceae and Celastraceae (Cannon and Simmons 2002). Currently, studies focusing on the diversity of fungal communities associated with the surface of eucalyptus leaves have used culture-dependent approaches (Lupo et al. 2001; Cheewangkoon et al. 2009; Kharwar et al. 2010), which can underestimate microbial diversity and abundance.
Analyses by 18S rRNA sequencing resulted in the identification of OTUs derived from a single plant, suggesting that more endophytic fungi in eucalyptus leaves can be identified if more plants are sampled. Simple soluble sugars such as glucose and fructose may function as the main carbon source from symbiotic fungi of plants (Bago et al. 2000; Chambergo et al. 2002). Moreover, metabolic pathways in microorganisms in association with plant can contribute to sucrose absorption and breakdown, which might represent an evolutionary adaptation conferring a competitive advantage to organisms involved in the interaction. Nutritional strategies for sucrose degradation are diverse in fungi (Doidy et al. 2012). Endophytic species, in general, feature different nutritional strategies to degrade sucrose (Doidy et al. 2012).
Lower concentrations of antimicrobial defensive chemicals in plants during the dispatch and field stages can explain the predominance of some endophytic fungi species in these samples (Arnold and Herre 2003). Humidity also affects fungal growth and consequently microbial diversity (Rayner and Boddy 1986). The leaves from the garden clonal, shading and newly out from shading were irrigated daily, according to the humidity conditions in the region, while plants in the dispatch and field stages depended on rainfall and could explain the differences in both prevalence and frequency of fungi at the various development phases of eucalyptus.
Most fungi identified in association with eucalyptus were classified within Ascomycota, including T. acidotherma and Cladosporium sp., species found in more than one development stage. New Teratosphaeria species were detected in leaves of eucalyptus with symptoms of Mycosphaerella leaf disease in Australia (Andjic et al. 2010), but in this study T. acidotherma and T. toledana were found in healthy leaves, suggesting that certain genera of fungi can be either endophytic or pathogenic in a certain plant (Saikkonen et al. 2004). In addition, differences between the endophytic and pathogenic stages may be due to abiotic factors or plant age (Saikkonen et al. 2004; Porras-Alfaro and Bayman 2011).
T. toledana and T. acidotherma were found in more than one seedling production stage, suggesting that they can adapt to different nutritional and environmental conditions. The presence of Cladosporium sp. and R. fluviale in leaves in more than one development stage could be related to their ability to establish an endophytic interaction with the host (Hardoim et al. 2008) to positively influence plant physiology.
The PCA demonstrated that the influence of eucalyptus leaf development stages on the structure of the fungal community is more related to leaf nutrient content, rather than sugar content. Plants in field conditions are more likely to encounter higher levels of stress and thus accumulate sugars, as shown in plants that tolerate drought (Hoekstra and Buitink 2001), and may have influenced the diversity of endophytic fungi in this stage of development. The fact that water, oxygen, nutrients, and pH are controlled during nursery cultivation of eucalyptus in the absence of toxic chemicals (Guerrini and Trigueiro 2004) and that the commercial substrate in the nursery is richer in nutrients than in the field may explain the differing clusters of endophytic fungi at these stages.
Fungi can benefit plants by increasing tolerance to biotic and abiotic factors, by improving growth and survival in adverse conditions, by competing with pathogens (Hubbard et al. 2014) and by producing wide range of growth regulators, e.g., gibberellins, abscisic acid and auxin (You et al. 2012). Many species of filamentous fungi are specific to some developmental stages of eucalyptus. D. crustulinus, P. tridactyla restrictus and Aspergillus are specific to the clonal garden stage, C. acutiseta and A. chartarum to shading, E. patelii and Saxomyces sp. in plants newly removed from shading, Lepidostroma sp. and M. lateralis in the dispatch and M. populicola in the field stage.
Phylogenetic analysis of 18S rDNA sequences confirmed the formation of distinct clades within Basidiomycota and Ascomycota with high bootstraps values. The majority of the clusters exhibhadited moderate to strong bootstrap values (Schneider 2007), thus supporting the phylogenetic analysis.
Among the Ascomycota, the best-resolved clusters (with high bootstrap values) were from OTU B07 with Podosphaera, B12 with Ascotricha and B25 with M. lateralis, demonstrating that these OTUs belong to those genera or species and confirming the comparison between sequences using BLASTn. Among the Basidiomycota, the best-resolved clusters (with high bootstrap values) were from UTO B10, with two fungi belonging to Diplomitoporus, the B17 with E. patelli and C. ayresii, and B21 in Cortinarius.
Mycosphaerella species are often reported as residents of aerial parts of plants, with some being host-specific (Maxwell et al. 2005). M. lateralis can form a pathogenic interaction with eucalyptus in Australia, causing necrotic lesions on leaves (Maxwell et al. 2000). However, in the present study, this fungus was identified in asymptomatic leaves, which can be an indication that the interaction is dependent on factors such as plant species and age, cultivation area, and recognition of host factors that determine a pathogenic or endophytic interaction (Saikkonen et al. 2004; Porras-Alfaro and Bayman 2011).
Aureobasidium pullulans is cosmopolitan and has been identified as a pathogen in various plants (Andrews et al. 1994). In the present study, it was found as an endophyte in an 18-month-old eucalyptus plant, but not at other stages of development.
A wide variety of endophytic fungi was found in eucalyptus from various stages of plant development during nursery production of seedlings. The results of our study raise the possibility that endophytic interactions in leaves can affect commercial seedling production systems. The relevance and the impact of such interactions established in plant leaves can be further assessed using different methodologies. For example, taxonomic identification of microbial communities can be deeply analyzed by sequencing rRNA or ITS fragments using high-throughput sequencing technologies. Also, metatranscriptomic studies can provide more precise information about what microbial and plant genes are up- or downregulated during the interaction. These strategies, in association with a larger sampling approach and considering the effect of environmental condition, can help further elucidate the role of associated and symbiotic microorganisms in the nutrition and adaptation of plants in the environment.
Conclusions
The endophytic fungal community in eucalyptus leaves throughout the various stages of plant development during nursery cultivation comprised 14 species and three genera of filamentous fungi. T. toledana and T. acidotherma colonized more than one phase of development, and Cladosporium sp. and R. fluviale persisted in the dispatch and field stages. Fungal communities in leaves sampled in the clonal garden were more closely related to levels of K, Cu, N and Mn, whereas those in the field samples were more closely related to the levels of glucose, fructose and sucrose.
References
Altschul SF, Gish W, Miller W, Myers EW, Lipman DJ (1990) Basic local alignment search tool. J Mol Biol 215:403–410
Andjic V, Whyte G, Hardy G, Burgess T (2010) New Teratosphaeria species occurring on eucalypts in Australia. Fungal Diver 43:27–38
Andrews JH, Harris RF, Spear RN, Lau GW, Nordheim EV (1994) Morphogenesis and adhesion of Aureobasidium pullulans. Can J Microbiol 40:6–17
Arnold AE (2005) Diversity and ecology of fungal endophytes in tropical forests. In: Deshmukh S (ed) Current trends in mycological research. Oxford & IBH Publishing Co. Pvt. Ltd., New Delhi, pp 49–68
Arnold AE, Herre EA (2003) Canopy cover and leaf age affect colonization by tropical fungal endophytes: ecological pattern and process in Theobroma cacao (Malvaceae). Mycol 95:388–398
Arnold AE, Lutzoni F (2007) Diversity and host range of foliar fungal endophytes: are tropical leaves biodiversity hotspots? Ecology 88:541–549
Arnold AE, Maynard Z, Gilbert GS, Coley PD, Kursar TA (2000) Are tropical fungal endophytes hyperdiverse? Ecol Lett 3:267–274
Bago B, Pfeffer PE, Shachar-Hill Y (2000) Carbon metabolism and transport in arbuscular mycorrhizas. Plant Physiol 124:949–958
Bremner JM, Mulvaney CS (1982) Total nitrogen. In: Page AL (ed) Methods of soil analysis. American Society of Agronomy, Madison, pp 595–624
Bresolin JD, Bustamante MMC, Krüger RH, Silva MRSS, Perez KS (2010) Structure and composition of bacterial and fungal community in soil under soybean monoculture in the Brazilian cerrado. Braz J Microbiol 41:391–403
Brooker MIH (2000) A new classification of the genus Eucalyptus L’Her (Myrtaceae). Aust Syst Bot 13:79–148
Brundrett MC (2006) Understanding the roles of multifunctional mycorrhizal and endophytic fungi. In: Schulz BJE, Boyle CJC, Sieber TN (eds) Microbial root endophytes. Springer, Berlin, pp 281–293
Cannon PF, Simmons CM (2002) Diversity and host preference of leaf endophytic fungi in the Iwokrama Forest Reserve, Guyana. Mycol 94:210–220
Chambergo FS, Bonaccorsi ED, Ferreira AJ, Ramos AS, Ferreira JR, Abrahao-Neto J, Farah JP, El-Dorry H (2002) Elucidation of the metabolic fate of glucose in the filamentous fungus Trichoderma reesei using expressed sequence tag (EST) analysis and cDNA microarrays. J Biol Chem 277:13983–13988
Cheewangkoon R, Groenewald JZ, Summerell BA, Hyde KD, To-Anun C, Crous PW (2009) Myrtaceae, a cache of fungal biodiversity. Persoonia 23:55–85
Doidy J, Grace E, Kühn C, Simon-Pas F, Casieri L, Wipf D (2012) Sugar transporters in plants and in their interactions with fungi. Trends Plant Sci 17:413–422
Duong LM, Jeewon R, Lumyong S, Hyde KD (2006) DGGE coupledwith ribosomal DNA gene phylogenies reveal uncharacterizedfungal phylotypes. Fungal Divers 23:121–138
Evans TN, Watson G, Rees GN, Seviour RJ (2014) Comparing activated sludge fungal community population diversity using denaturing gradient gel electrophoresis and terminal restriction fragment length polymorphism. Antonie Van Leeuwenhoek 105:559–569
FAO (2016) State of the world’s forests 2016. Forests and agriculture: land-use challenges and opportunities. Food and Agriculture Organization of the United Nations, Rome
Fromin N, Hamelin J, Tarnaw Ski S, Roesti D, Jourdain-Miserez K, Forestier N, Teyssier-Cuvelle S, Gillet F, Aragno M, Rossi P (2002) Statistical analysis of denaturing gel electrophoresis (DGE) fingerprinting patterns. Environ Microbiol 11:634–643
Gazis R, Chaverri P (2010) Diversity of fungal endophytes in leaves and stems of wild rubber trees (Hevea brasiliensis) in Peru. Fungal Ecol 3:240–254
Guerrini IA, Trigueiro RM (2004) Atributos físicos e químicos de substratos compostos por biossólidos e casca de arroz carbonizada. Rev Bras Cienc Solo 28:1069–1076
Hammer Ø, Harper DAT, Ryan PD (2001) PAST: paleontological statistics software package for education and data analysis. Palaeontol Electron 4:9
Hardoim PR, Overbeek LSV, Elsas KDV (2008) Properties of bacterial endophytes and their proposed role in plant growth. Trends Microbiol 16:463–471
Hoekstra FA, Buitink J (2001) Mechanisms of plant desiccation tolerance. Trends Plant Sci 8:431–438
Hubbard M, Germida JJ, Vujanovic V (2014) Fungal endophytes enhance wheat heat and drought tolerance in terms of grain yield and second—generation seed viability. J Appl Microbiol 116:109–122
Jia M, Chen L, Xin HL, Zheng CJ, Rahman K, Han T, Qin LP (2016) A friendly relationship between endophytic fungi and medicinal plants: a systematic review. Front Microbiol 7:1–14
Kemler M, Garnas J, Wingfield MJ, Gryzenhout M, Pillay KA, Slippers B (2013) Ion torrent PGM as tool for fungal community analysis: a case study of endophytes in Eucalyptus grandis reveals high taxonomic diversity. PLoS ONE 8:e81718
Kharwar RN, Gond SK, Kimar A, Misha A (2010) A comparative study of endophytic and epiphytic fungal association with leaf of Eucalyptus citriodora Hook, and their antimicrobial activity. World J Microb Biot 26:1941–1948
Kittelmann S, Naylor GE, Koolaard JP, Janssen PH (2012) A proposed taxonomy of anaerobic fungi (Class Neocallimastigomycetes) suitable for large-scale sequence-based community structure analysis. PLoS ONE 7:1–13
Laplante K, Derome N (2011) Parallel changes in the taxonomical structure of bacterial communities exposed to a similar environmental disturbance. Ecol Evol 4:489–501
Lisec J, Schaeur N, Kopka J, Willmitzer L, Fernie AR (2006) Gas chromatography mass spectrometry-based metabolite profiling in plants. Nat Protoc 1:387–396
Lumyong S, Lumyong P, Mckenzie EH, Hyde KD (2002) Enzymatic activity of endophytic fungi of six native seedling species from oi Suthep-Pui National Park, Thailand. Can J Microbiol 48:1109–1112
Lupo S, Tiscornia S, Bettucci L (2001) Endophytic fungi from flowers, capsules and seeds of Eucalyptus globulus. Rev Iberoam Micol 18:38–41
Marschener H (1998) Role of root growth, arbuscular mycorrhiza, and root exudates for the efficiency in nutrient acquisition. Field Crops Res 56:203–207
Maxwell A, Hardy GESTJ, Wingfield MJ, Dell B (2000) First record of Mycosphaerella lateralis on Eucalyptus in Australia. Australas Plant Pathol 29:279
Maxwell A, Jackson SL, Dell B, Hardy GESJ (2005) PCR-identification of Mycosphaerella species associated with leaf diseases of Eucalyptus. Mycol Res 109:992–1004
May LA, Smiley B, Schmidt MG (2001) Comparative denaturing gradient gel electrophoresis analysis of fungal communities associated with whole plant corn silage. Can J Microbiol 47:829–841
Miguel PSB, de Oliveira MNV, Delvaux JC, de Jesus GL, Chaer AC, Tótola MR, Neves JCL, Costa MD (2016) Diversity and distribution of the endophytic bacterial community at different stages of Eucalyptus growth. Antonie Van Leeuwenhoek 109(6):755–771
Miguel PSB, Delvaux JC, Oliveira MNV, Moreira BC, Freitas FS, Costa MD, Tótola MR, Neves JCL, Costa MD (2017) Diversity and distribution of the endophytic fungal community in eucalyptus leaves. Afr J Microbiol 11:92–105
Minitab I (2006) MINITAB statistical software. Version: Release. 15
Oliveira MNV, Santos TMA, Vale HMM, Delvaux JC, Cordero AP, Ferreira AB, Miguel PSB, Tótola MR, Costa MD, Moraes CA, Borges AC (2013) Endophytic microbial diversity in coffee cherries of Coffea arabica from southeastern Brazil. Can J Microbiol 59:221–230
Omacini M, Chaneton EJ, Ghersa CM, Muller CB (2001) Symbiotic fungal endophytes control insect host-parasite interaction webs. Nature 409:78–81
Oros-Sichler M, Gomes NCM, Neuber G, Smalla K (2006) A new semi-nested PCR protocol to amplify large 18S rRNA gene fragments for PCR-DGGE analysis of soil fungal communities. J Microbiol Meth 65:63–75
Porras-Alfaro A, Bayman P (2011) Hidden fungi, emergent properties: endophytes and microbiomes. Phytopathol 49:291
Rayner ADM, Boddy L (1986) Population structure and the infection biology of wood-decay fungi in living trees. Adv Plant Pathol 5:119–160
Rho H, Kim SH (2017) Endophyte effects on photosynthesis and water use of plant hosts: a meta-analysis. In: Doty S (ed) Functional importance of the plant microbiome. Springer, Cham
Robl D, Da Delabona P, Mergel CM, Rojas JD, Dos Costa P, Pimentel IC, Padilha G (2013) The capability of endophytic fungi for production of hemicellulases and related enzymes. BMC Biotechnol 13:1–12
Saikkonen K, Wali P, Helander M, Taeth SH (2004) Evolution of endophyte-plant symbioses. Trends Plant Sci 9:275–280
Saitou N, Nei M (1987) The neighbor-joining method: a new method for reconstructing phylogenetic trees. Mol Biol Evol 4:406–425
Sánchez Márquez S, Bills GF, Zabalgogeazcoa I (2011) Fungal species diversity in juvenile and adult leaves of Eucalyptus globulus from plantations affected by Mycosphaerella leaf disease. Ann Appl Biol 158:177–187
Schneider H (2007) Métodos de análise filogenética: um guia prático [Methods for phylogenetic analysis: a practical guide]. Holos Editora e Sociedade Brasileira de Genética, Ribeirão Preto
Soto-Barajas MC, Iñigo Z, Gómez-Fuertes J, González-Blanco V, Vázquez de Aldana BR (2016) Epichloë endophytes affect the nutrient and fiber content of Lolium perenne regardless of plant genotype. Plant Soil 405:265–277
Systat Software, Inc (2008) Sigma plot for windows, version 11.0
Vainio EJ, Hantula J (2000) Direct analysis of wood-inhabiting fungi using denaturing gradient gel electrophoresis of amplified ribosomal DNA. Mycol Res 104:927–936
You YH, Kang SM, Choo YS, Lee JM (2012) Fungal diversity and plant growth promotion of endophytic fungi from six halophytes in Suncheon Bay. J Microbiol Biotechnol 22:1549–1556
Acknowledgements
The authors thank the Brazilian Federal Agency for Support and Evaluation of Graduate Education (Coordenação de Aperfeiçoamento de Pessoal de Nível Superior–CAPES) and Celulose NipoBrasileira (CENIBRA) for financial support and the Minas Gerais State Research Foundation (Fundação de Amparo à Pesquisa do Estado de Minas Gerais–FAPEMIG) for the grant provided to P. S. B. Miguel. The authors also gratefully acknowledge the Laboratório de Ecologia Microbiana (LEM), of the Microbiology Department at the Universidade Federal de Viçosa, for operational support.
Author information
Authors and Affiliations
Corresponding author
Additional information
Project funding: The work was financially supported by the Evaluation of Graduate Education (Coordenação de Aperfeiçoamento de Pessoal de Nível Superior–CAPES) and Celulose NipoBrasileira (CENIBRA).
The online version is available at http://www.springerlink.com
Corresponding editor: Tao Xu.
Rights and permissions
About this article
Cite this article
Miguel, P.S.B., Miguel, F.B., Moreira, B.C. et al. Diversity of the endophytic filamentous fungal leaf community at different development stages of eucalyptus. J. For. Res. 30, 1093–1103 (2019). https://doi.org/10.1007/s11676-018-0610-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11676-018-0610-0