Abstract
Purpose
The gold standard for colorectal cancer metastases detection in the peritoneum is histological evaluation of a removed tissue sample. For feedback during interventions, real-time in vivo imaging with confocal laser microscopy has been proposed for differentiation of benign and malignant tissue by manual expert evaluation. Automatic image classification could improve the surgical workflow further by providing immediate feedback.
Methods
We analyze the feasibility of classifying tissue from confocal laser microscopy in the colon and peritoneum. For this purpose, we adopt both classical and state-of-the-art convolutional neural networks to directly learn from the images. As the available dataset is small, we investigate several transfer learning strategies including partial freezing variants and full fine-tuning. We address the distinction of different tissue types, as well as benign and malignant tissue.
Results
We present a thorough analysis of transfer learning strategies for colorectal cancer with confocal laser microscopy. In the peritoneum, metastases are classified with an AUC of 97.1, and in the colon the primarius is classified with an AUC of 73.1. In general, transfer learning substantially improves performance over training from scratch. We find that the optimal transfer learning strategy differs for models and classification tasks.
Conclusions
We demonstrate that convolutional neural networks and transfer learning can be used to identify cancer tissue with confocal laser microscopy. We show that there is no generally optimal transfer learning strategy and model as well as task-specific engineering is required. Given the high performance for the peritoneum, even with a small dataset, application for intraoperative decision support could be feasible.
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
Introduction
Colorectal cancer is very common, and it is often associated with metastatic spread [1]. In particular, peritoneal carcinomatosis (PC) can arise in later stages of development which often shortens patient survival times substantially [2, 3]. Thus, early and reliable detection of metastases is crucial. Diagnosis with typical external imaging techniques such as computed tomography (CT) and magnetic resonance imaging (MRI) is difficult for PC as a very high resolution is required. For example, preoperative CT has been shown to be ineffective to detect individual peritoneal tumor deposits and the interobserver variability among experts was significant [4]. Also, integrated PET/CT did not provide sufficient information for accurate assessment [5]. For MRI, studies have shown improvement over assessment with CT only [6, 7], but overall, its resolution is still a limitation [8]. Therefore, exploratory laparoscopy is generally employed to investigate the presence of PC [9].
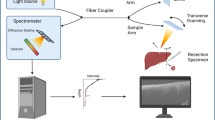
Recently, a new intraoperative device using confocal laser microscopy (CLM) has been introduced which provides submicrometer image resolution [10]. In the study, ten rats received colon carcinoma cell implants in the colon and peritoneum. After a growth period, laparotomy with in vivo CLM was performed. CLM images of healthy and malignant colon tissue, as well as healthy and malignant peritoneum, were acquired. It was shown that experts are able to distinguish different tissue types as well as healthy and malignant tissue from CLM. This raises the question whether image processing techniques can be used to automatically classify different tissue types. This could enable faster and improved intraoperative decision support with CLM.
Recently, automatic tissue characterization has been successfully addressed using deep learning methods such as convolutional neural networks (CNNs) for semantic segmentation and classification [11, 12]. For example, skin cancer classification at dermatologist-level performance was achieved [13]. However, the datasets for this and related studies are large, and commonly, datasets for medical learning tasks are small [14]. This can be problematic as insufficient data for optimal training might lead to overfitting and limited generalization. This is particularly important for deep learning models which can be prone to overfitting due to their large number of trainable parameters. To overcome this issue, transfer learning methods have been proposed where a deep learning model is first pretrained on a different, large dataset [15]. Then, information from the source domain can be transferred to the (medical) target domain using strategies such as “off-the-shelf” features, partial layer freezing, or full fine-tuning [16]. While this has been successfully applied for medical learning tasks [17], there is no single solution for all problems and the optimal transfer learning strategy is highly dependent on the imaging modality and dataset size [18].
Automatic analysis of CLM images has been proposed for different tissue types such as human skin [19], the cornea [20], or the oral cavity [21]. Recently, deep learning methods have been applied to CLM and similar modalities. For example, CNNs have been used for oral squamous cell carcinoma classification [21] and motion correction with CLM [22]. Similarly, skin images from CLM have been used with CNN-based classification [23]. For the gastrointestinal tract, CNNs have been used to distinguish three classes of Barrett’s esophagus [24]. Also, brain tumor classification with CNNs and CLM has shown promising results [25]. For example, a CNN has been used to differentiate CLM images with and without diagnostic value for a physician during surgery [26]. Also, weakly supervised localization has been used to derive local information in CLM images from image-level labels only [27].
So far, deep learning-based classification of colorectal cancer from CLM images has not been addressed. Also, while several approaches have used CLM and CNNs for other problems [28], there is no analysis of transfer learning properties for colorectal cancer with CLM. Therefore, we study deep learning-based colon cancer classification from CLM images with a variety of transfer learning methods from the ImageNet dataset. We consider training from scratch, partial layer freezing, “off-the-shelf” features and full fine-tuning to investigate how transferable ImageNet features are to CLM. We perform this study with the classic models VGG-16 [29] and Inception-V3 [30] as well as the state-of-the-art architectures Densenet [31] and squeeze-and-excitation networks [32] to analyze the consistency of transfer strategies across architectures. We consider the classes healthy colon (HC), malignant colon (MC), healthy peritoneum (HP) and malignant peritoneum (MP). Based on these classes, we address three binary classification tasks with CLM. First, we consider the differentiation of organs (HP vs. HC). Then, we study the detection of malignant tissue in two types of organs (HP vs. MP and HC vs. MC). This allows us to study variations across different classification tasks for CLM. A preliminary version of this paper was presented at the BVM Workshop 2019 [33]. We substantially revised the paper, extended the review of the literature and performed more experiments with additional transfer strategies and more architectures. This paper is structured as follows: First, we describe our models and transfer learning strategies and the dataset we use in “Methods” section. Then, we report our results in “Results” section and discuss them in “Discussion” section. Last, we conclude in “Conclusion” section.
Methods
Model architectures and training strategies
First, we consider the classic model VGG-16 [29] with the addition of batch normalization which enables faster training of the architecture by reducing the internal covariate shift [34]. The model itself is simple as it consists of several stacked convolutional layers without further augmentation. In between blocks of two to three convolutional layers with kernel sizes of \(3\times 3\) and \(1\times 1\), max pooling reduces the spatial dimensions. Subsequent convolutions double the number of feature maps. A building block of the architecture is shown in Fig. 1 (top left). Due to its simple structure, the architecture can serve as a baseline.
Second, we employ Inception-V3 [30]. The model consists of multiple Inception blocks which follow two core design principles. First, the blocks have a multi-path structure, i.e., the input feature maps are processed in parallel by different convolution and pooling operations. At the block’s output, the feature maps from all paths are concatenated. Second, the convolutional paths perform a reduction operation that downsizes the feature map dimension with \(1 \times 1\) kernels. Then, computationally more expensive \(3\times 3\) convolutions process the lower-dimensional representations. The output feature map size is increased if the spatial dimensions are reduced inside the block which avoids representational limitations.
The idea of reduction and expansion has also found its way into the Resnet architecture [35] which is a core component of the next two models. Resnets learn a residual instead of a full feature transformation by using skip connections. In detail, a Resnet block (ResBlock) computes
where \(x_{(l)}\) is the block output, \(x_{(l-1)}\) is the block input, a is a ReLU activation [36] and \(\mathcal {F}\) represents two convolutional layers with parameters \(\theta _{(l)}\). The skip connection enables better gradient propagation for improved training.
Third, we consider Densenet121 [31], a state-of-the-art architecture which strives for more efficiency by introducing extensive feature reuse. In particular, within one DenseBlock, features computed in previous layers are also fed into the subsequent layers. To keep the feature map sizes moderate, compression blocks reduce the feature maps between DenseBlocks. The DenseBlock is shown in Fig. 1 (bottom left).
Fourth, we adopt the architecture SE-Resnext50 [32]. At its core, the model uses Resnext blocks [37] which are an extension of Resnet. Here, the single convolutional path \(\mathcal {F}\) is split into multiple paths with individual layers which increases representational power. The key addition in SE-Resnext50 is the use of squeeze-and-excitation (SE) modules which recalibrate the feature maps learned by Resnext blocks. These modules have shown improved performance with only a minimal increase in the number of parameters. The concept is shown in Fig. 1 (bottom right).
Building blocks of the models we use. The building blocks from CNN architectures as indicated in Fig. 2. We employ VGG-16 (a), Inception-V3 (b), Densenet121 (c) and SE-Resnext50 (d). F denotes the number of feature maps in each block. The Conv blocks also contain ReLU activations and batch normalization for VGG-16 (a). SE-Resnext50 is shown in simplified form without its bottleneck in the SE module. FC-\(\sigma \) is a fully connected layer with sigmoid activation. C. is an abbreviation for convolutional layers. Note that Inception-V3 employs multiple block variants and we show one example
Different transfer learning scenarios we investigate. Model Block refers to one of the blocks shown in Fig. 1. Green indicates that blocks are retrained. Red indicates that blocks are frozen with their weights having been trained on ImageNet
Due to the small dataset size, we study several transfer learning strategies where the above-mentioned models are trained on ImageNet. We cut off the last layer of all models and replace it with a fully connected layer with two outputs for binary classification. We apply a softmax layer on top, and the final classification output is the class with the highest probability. We train a separate model for each of our binary classification tasks.
As a baseline, we consider training from scratch, i.e., all weights are randomly initialized. Then, we use several different transfer learning strategies illustrated in Fig. 2. The first transfer approach follows the “off-the-shelf” features idea. Here, only the new classifier is retrained on features extracted by the pretrained CNN. We also consider two partial freezing methods, where an initial part of the network remains frozen and the part closer to the classifier is retrained. We chose the freezing points block-wise, i.e., we do not cut into building blocks. Last, we consider full fine-tuning where all weights in the network are retrained with a small learning rate. The different strategies represent different abstractions of feature transfer between ImageNet and CLM images.
To further improve generalization, we employ online data augmentation with random image flipping and random changes in brightness and contrast. Furthermore, we use random cropping with crops of size \(224\times 224\) (\(299\times 299\) for Inception-V3) taken from the full images of size \(384\times 384\). We use the Adam algorithm for optimization. We adapt learning rates and the number of training epochs for the different transfer scenarios. We use a cross-entropy loss function with additional weighting to account for the slight class imbalance. In detail, we multiply the loss of a training example by \(N/n_i\) where N is the total number of training examples in the current fold and \(n_i\) is the number of examples belonging to class i in the current fold. In this way, underrepresented classes receive a higher weighting in the loss function. During evaluation, we use mutli-crop evaluation with \(N_c = 36\) evenly spread crops over the images. This ensures that all image regions are covered with large overlaps between crops. The final predictions are averaged over the \(N_c\) crops. We implement our models in PyTorch.
Dataset and experiments
The dataset was collected in a previous study conducted at the University Hospital Schleswig-Holstein in Lübeck where expert assessment of CLM images in the colon area was evaluated [10]. A custom intraoperative device with integrated CLM (Karl Storz GmbH & Co KG, Tuttlingen, Germany) was built. The image resolution was \(384\times 384\) pixels which covers a field of view of \({300}\,{\upmu \hbox {m}} \times {300}\,{\upmu \hbox {m}}\). In the study, ten rats received colon adenocarcinoma cell implantation in the colon and peritoneum with a growth time of seven days. Then, laparotomy was conducted and images of healthy colon tissue, malignant colon tissue, healthy peritoneum tissue and malignant peritoneum tissue were obtained. Example CLM images for each tissue type are shown in Fig. 3. After removal of low-quality images, 1577 images remained with 533 belonging to class HC, 309 belonging to class MC, 343 belonging to class HP and 392 belonging to class MP. Note that some subjects are missing classes such that, on average, six subjects per class remain. Ground-truth annotation of all images was obtained by tissue removal of the scanned areas and subsequent histological evaluation.
Due to the small dataset size, we chose a cross-validation scheme where images from one subject are left for evaluation and training is performed on the remaining ones. Thus, all reported results are the mean value of, on average, six training scenarios with six different folds. Based on the four classes, we address three binary classification problems. First, we consider HC versus HP, i.e., we investigate the feasibility of distinguishing the different organs in CLM. Then, we consider the differentiation of healthy and malignant tissue with the two binary classification problems HP versus MP and HC versus MC. We report the accuracy, sensitivity, specificity, F1-score and AUC. We use the AUC as the main metric as it is threshold independent.
Results
First, we compare the different transfer learning scenarios described in “Methods” section across all architectures for each classification scenario, see Fig. 4. In general, the AUC is very high for the differentiation of different healthy tissue types and healthy and malignant peritoneum tissue. The AUC for classifying malignant colon tissue is substantially lower. Also, the standard deviation is higher for this task. Training from scratch performs worst for all architectures and classification scenarios.
AUC values of all applied architectures for the different classification problems. We evaluate the following training types: (1) retrain classifier, (2) partial freeze 1, (3) partial freeze 2, (4) full fine-tuning, (5) training from scratch. For each value, the standard deviation over multiple folds is represented by an error bar
Regarding the transfer learning scenarios, training from scratch performs worst for all classification scenarios. For two of the three scenarios, only retraining the classifier shows substantially lower performance than other transfer scenarios. There are no clear trends between the partial freezing and fine-tuning scenarios.
Second, we go into more details for the classification task HP versus MP. Figure 5 shows the ROC curves for all models with all transfer learning scenarios for the classification task. Operating points with a good trade-off in the upper left corner vary for each model. For VGG-16, retraining the classifier only stands out. For Densenet121, partial freezing performs well. For Inception-V3 and SE-Resnext50, partial freezing and fine-tuning perform similar.
Third, an overview of the best performing transfer strategies is shown in Table 1. Comparing individual results for each architecture, no model clearly outperforms the others consistently. In general, Densenet121 performs slightly better across the tasks. The optimal transfer strategy differs across models and classification tasks. For HC versus HP and for Densenet121 in general, the partial freezing method performs best.
Last, we provide training times for all architectures and training scenarios, see Fig. 6. In general, freezing more weights during training reduces the overall training time. Furthermore, training time loosely scales with the number of trainable parameters as VGG-16 contains the most parameters and shows the longest training times, followed by SE-Resnext50.
Discussion
We study deep transfer learning methods for CLM images for three binary classification problems. Automatic decision support with CLM during interventions could improve workflow with immediate feedback on the tissue properties. For this purpose, we investigate the use of CNNs with four different architectures and five training scenarios.
The three classification tasks As a baseline, differentiating healthy colon and peritoneum tissue works well with an AUC over 0.90 for partial freezing across all models, see Fig. 4. This indicates that discriminative features for different organs can be learned from CLM images. Similarly, for classification of metastases in the peritoneum the AUC is around 0.90 for all transfer learning scenarios. However, classifying healthy and malignant colon tissue performs substantially worse with an AUC of \(\approx 0.70\) for partial freezing and fine-tuning. The task appears to be more difficult which is also reflected in a slightly higher standard deviation. This indicates higher uncertainty of model predictions. This could be caused by the heterogeneous appearance of colon tissue in different parts of the colon which complicates the learning task in conjunction with the small dataset size. Furthermore, during development, colon carcinoma cells transform from a healthy stage to adenoma and then carcinoma. At earlier stages, healthy and malignant cells can still have similar appearance which complicates the learning task.
Transfer learning scenarios Figure 4 also provides an overview of the transfer strategies across all models. Clearly, transfer learning substantially outperforms training from scratch across all classification tasks which supports the effectiveness of transfer learning for medical image classification problems [38]. The results indicate that meaningful feature transfer from the natural image domain to CLM images is possible, although the images have a vastly different appearance. However, comparing transfer strategies, only retraining the classifier performs worse than other scenarios in two out of three classification tasks. This agrees with results of a previous study on transfer learning with CLM images in neurosurgery [28]. Here, the authors found that full fine-tuning outperforms retraining of the classifier only. However, in our case, retraining the classifier only also shows a high performance for the task HP versus MP. This could be caused by fragile co-adaptation of weights [39] which leads to large performance differences between the different classification tasks. For some tasks (e.g., HP vs. MP), recovery and reuse of potentially coadapted weights might be feasible while reuse is impaired for other tasks (e.g., HC vs. MC). The partial freezing and fine-tuning strategies appear to be more consistent across tasks; however, the optimal strategy still differs. Overall, our results indicate that the transferability of features not only depends on the imaging modality but also the classification task. This adds to previous insights on transfer learning in the medical domain where the optimal transfer strategy was found to be modality and dataset size dependent [18]. Comparing the partial freezing and fine-tuning strategies, performance is very close and there is no optimal strategy for each of the tasks. However, training times are also an aspect to consider for the different transfer learning strategies. As shown in Fig. 6, freezing more parameters inside the architecture leads to reduced training times. Thus, partial freezing can be generally seen as advantageous as it often achieves similar performance as full fine-tuning while requiring less training time. For application, this insight could be useful when adopting and retraining models for cancer classification in other organs or when newer architectures are introduced.
Different architectures for CLM To analyze the different transfer strategies further, we consider the ROC curves of each architecture for the HP versus MP task, see Fig. 5. For this task, using “off-the-shelf” features and only retraining the classifier performed considerably better than for the other tasks. As discussed before, this indicates that transfer learning scenarios are classification task dependent. In detail, the ROC curves reveal that VGG-16 stands out in particular where retraining the classifier only performs best out of all transfer strategies. In transfer learning research, VGG-16 is still a popular general purpose feature extractor for numerous tasks [11, 40]. For the other architectures, the optimal strategy differs. For example, for Densenet121, the partial freezing methods show good operating points in the upper, left corner of the ROC curve. For Inception-V3 and SE-Resnext50, partial freezing and fine-tuning perform similar with no clearly superior method. This indicates that the choice of transfer learning method depends on the architecture. This should be expected, as the models have very different block types and each freezing type fixes a different number of parameters. The detailed results in Table 1 with additional metrics underline this insight. There is no optimal transfer learning strategy, and the best performing strategy varies for different architectures and classification tasks. Overall, we demonstrate that transfer learning has an impact on performance; however, there is no simple rule of thumb for optimal transfer learning with CLM. Our results show that examining different freezing strategies can considerably improve performance for individual models.
Conclusion
We investigate the feasibility of colon cancer classification in CLM images using CNNs and multiple transfer learning scenarios. Using in vivo images of healthy and malignant colon and peritoneum tissue obtained from ten subjects, we adopt four architectures and five transfer learning scenarios for three classification problems with CLM. Our results show that different organs as well as healthy and malignant peritoneum tissue can be classified with deep transfer learning. We show that transfer learning from ImageNet is successful with CLM, but the transferability of features is limited. We find that there is no single optimal model or transfer strategy for all CLM classification problems and that task-specific engineering is likely required for application. For future work, our results could be extended to more classification problems with CLM.
References
Torre LA, Bray F, Siegel RL, Ferlay J, Lortet-Tieulent J, Jemal A (2015) Global cancer statistics, 2012. CA Cancer J Clin 65(2):87–108
Verwaal VJ, van Ruth S, Witkamp A, Boot H, van Slooten G, Zoetmulder FA (2005) Long-term survival of peritoneal carcinomatosis of colorectal origin. Ann Surg Oncol 12(1):65–71
Franko J, Shi Q, Goldman CD, Pockaj BA, Nelson GD, Goldberg RM, Pitot HC, Grothey A, Alberts SR, Sargent DJ (2012) Treatment of colorectal peritoneal carcinomatosis with systemic chemotherapy: a pooled analysis of north central cancer treatment group phase III trials N9741 and N9841. J Clin Oncol 30(3):263
de Bree E, Koops W, Kröger R, van Ruth S, Witkamp AJ, Zoetmulder FA (2004) Peritoneal carcinomatosis from colorectal or appendiceal origin: correlation of preoperative CT with intraoperative findings and evaluation of interobserver agreement. J Surg Oncol 86(2):64–73
Dromain C, Leboulleux S, Auperin A, Goere D, Malka D, Lumbroso J, Schumberger M, Sigal R, Elias D (2008) Staging of peritoneal carcinomatosis: enhanced CT vs. PET/CT. Abdom Imaging 33(1):87–93
Low RN, Semelka RC, Worawattanakul S, Alzate GD (2000) Extrahepatic abdominal imaging in patients with malignancy: comparison of MR imaging and helical CT in 164 patients. J Magn Reson Imaging 12(2):269–277
Iafrate F, Ciolina M, Sammartino P, Baldassari P, Rengo M, Lucchesi P, Sibio S, Accarpio F, Di Giorgio A, Laghi A (2012) Peritoneal carcinomatosis: imaging with 64-MDCT and 3T MRI with diffusion-weighted imaging. Abdom Imaging 37(4):616–627
González-Moreno S, González-Bayón L, Ortega-Pérez G, González-Hernando C (2009) Imaging of peritoneal carcinomatosis. Cancer J 15(3):184–189
Ishigami S, Uenosono Y, Arigami T, Yanagita S, Okumura H, Uchikado Y, Kita Y, Kurahara H, Kijima Y, Nakajo A, Maemura K, Natsugoe S (2014) Clinical utility of perioperative staging laparoscopy for advanced gastric cancer. World J Surg Oncol 12(1):350
Ellebrecht DB, Kuempers C, Horn M, Keck T, Kleemann M (2019) Confocal laser microscopy as novel approach for real-time and in-vivo tissue examination during minimal-invasive surgery in colon cancer. Surg Endosc 33(6):1811–1817
Litjens G, Kooi T, Bejnordi BE, Setio AAA, Ciompi F, Ghafoorian M, van der Laak JA, Van Ginneken B, Sánchez CI (2017) A survey on deep learning in medical image analysis. Med Image Anal 42:60–88
Goceri E, Goceri N (2017) Deep learning in medical image analysis: recent advances and future trends. In: International conferences computer graphics, visualization, computer vision and image processing, pp 305–311
Esteva A, Kuprel B, Novoa RA, Ko J, Swetter SM, Blau HM, Thrun S (2017) Dermatologist-level classification of skin cancer with deep neural networks. Nature 542(7639):115
Shen D, Wu G, Suk HI (2017) Deep learning in medical image analysis. Ann Rev Biomed Eng 19:221–248
Bengio Y (2012) Deep learning of representations for unsupervised and transfer learning. In: Proceedings of ICML workshop on unsupervised and transfer learning, pp 17–36
Hoo-Chang S, Roth HR, Gao M, Lu L, Xu Z, Nogues I, Yao J, Mollura D, Summers RM (2016) Deep convolutional neural networks for computer-aided detection: CNN architectures, dataset characteristics and transfer learning. IEEE Trans Med Imaging 35(5):1285
Gessert N, Lutz M, Heyder M, Latus S, Leistner DM, Abdelwahed YS, Schlaefer A (2019) Automatic plaque detection in IVOCT pullbacks using convolutional neural networks. IEEE Trans Med Imaging 38(2):426–434
Tajbakhsh N, Shin JY, Gurudu SR, Hurst RT, Kendall CB, Gotway MB, Liang J (2016) Convolutional neural networks for medical image analysis: full training or fine tuning? IEEE Trans Med Imaging 35(5):1299–1312
Rajadhyaksha M, Grossman M, Esterowitz D, Webb RH, Anderson RR (1995) In vivo confocal scanning laser microscopy of human skin: melanin provides strong contrast. J Investig Dermatol 104(6):946–952
Niederer RL, Perumal D, Sherwin T, McGhee CN (2007) Age-related differences in the normal human cornea: a laser scanning in vivo confocal microscopy study. Br J Ophthalmol 91(9):1165–1169
Aubreville M, Knipfer C, Oetter N, Jaremenko C, Rodner E, Denzler J, Bohr C, Neumann H, Stelzle F, Maier A (2017) Automatic classification of cancerous tissue in laser endomicroscopy images of the oral cavity using deep learning. Sci Rep 7(1):11979
Aubreville M, Stoeve M, Oetter N, Goncalves M, Knipfer C, Neumann H, Bohr C, Stelzle F, Maier A (2019) Deep learning-based detection of motion artifacts in probe-based confocal laser endomicroscopy images. Int J Comput Assist Radiol Surg 14(1):31–42
Wiltgen M, Bloice M (2016) Automatic interpretation of melanocytic images in confocal laser scanning microscopy. In: Microscopy and analysis. InTech
Hong J, Park By, Park H (2017) Convolutional neural network classifier for distinguishing Barrett’s esophagus and neoplasia endomicroscopy images. In: 2017 39th annual international conference of the IEEE engineering in medicine and biology society (EMBC), IEEE, pp 2892–2895
Izadyyazdanabadi M, Belykh E, Mooney MA, Eschbacher JM, Nakaji P, Yang Y, Preul MC (2018) Prospects for theranostics in neurosurgical imaging: empowering confocal laser endomicroscopy diagnostics via deep learning. Front Oncol 8:240
Izadyyazdanabadi M, Belykh E, Martirosyan N, Eschbacher J, Nakaji P, Yang Y, Preul MC (2017) Improving utility of brain tumor confocal laser endomicroscopy: objective value assessment and diagnostic frame detection with convolutional neural networks. In: Medical imaging 2017: computer-aided diagnosis, vol. 10134. International Society for Optics and Photonics, p 101342J
Izadyyazdanabadi M, Belykh E, Cavallo C, Zhao X, Gandhi S, Moreira LB, Eschbacher J, Nakaji P, Preul MC, Yang Y (2018) Weakly-supervised learning-based feature localization for confocal laser endomicroscopy glioma images. In: International conference on medical image computing and computer-assisted intervention. Springer, pp 300–308
Izadyyazdanabadi M, Belykh E, Mooney M, Martirosyan N, Eschbacher J, Nakaji P, Preul MC, Yang Y (2018) Convolutional neural networks: ensemble modeling, fine-tuning and unsupervised semantic localization for neurosurgical CLE images. J Vis Commun Image Represent 54:10–20
Simonyan K, Zisserman A (2014) Very deep convolutional networks for large-scale image recognition. arXiv preprint arXiv:1409.1556
Szegedy C, Vanhoucke V, Ioffe S, Shlens J, Wojna Z (2016) Rethinking the inception architecture for computer vision. In: CVPR, pp 2818–2826
Huang G, Liu Z, Weinberger KQ, van der Maaten L (2016) Densely connected convolutional networks. arXiv preprint arXiv:1608.06993
Hu J, Shen L, Sun G (2018) Squeeze-and-excitation networks. In: Proceedings of the IEEE conference on computer vision and pattern recognition, pp 7132–7141
Gessert N, Wittig L, Drömann D, Keck T, Schlaefer A, Ellebrecht DB (2019) Feasibility of colon cancer detection in confocal laser microscopy images using convolution neural networks. In: Bildverarbeitung für die Medizin 2019
Ioffe S, Szegedy C (2015) Batch normalization: accelerating deep network training by reducing internal covariate shift. In: ICML
He K, Zhang X, Ren S, Sun J (2016) Deep residual learning for image recognition. In: CVPR, pp 770–778
Nair V, Hinton GE (2010) Rectified linear units improve restricted Boltzmann machines. In: ICML, pp 807–814
Xie S, Girshick R, Dollár P, Tu Z, He K (2017) Aggregated residual transformations for deep neural networks. In: 2017 IEEE conference on computer vision and pattern recognition (CVPR). IEEE, pp 5987–5995
Shin HC, Roth HR, Gao M, Le Lu, Xu Z, Nogues I, Yao J, Mollura D, Summers RM (2016) Deep convolutional neural networks for computer-aided detection: CNN architectures, dataset characteristics and transfer learning. IEEE Trans Med Imaging 35(5):1285–1298
Yosinski J, Clune J, Bengio Y, Lipson H (2014) How transferable are features in deep neural networks? In: Advances in neural information processing systems, pp 3320–3328
Herath S, Harandi M, Porikli F (2017) Going deeper into action recognition: a survey. Image Vis Comput 60:4–21
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of Interest
The authors declare that they have no conflict of interest.
Ethical approval
All procedures performed in studies involving animals were in accordance with the ethical standards of the institution or practice at which the studies were conducted.
Informed consent
Informed consent was obtained from all individual participants included in the study.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Gessert, N., Bengs, M., Wittig, L. et al. Deep transfer learning methods for colon cancer classification in confocal laser microscopy images. Int J CARS 14, 1837–1845 (2019). https://doi.org/10.1007/s11548-019-02004-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11548-019-02004-1