Abstract
Purpose
Long-term excessive application of nitrogen fertilizer under diverse land uses has caused serious environmental problems on the North China Plain. Current studies focus on denitrifiers and bacterial communities in topsoil under diverse land-use types; however, few studies have studied denitrifiers and microbial communities in subsoils.
Materials and methods
The variations in soil bacterial communities and denitrifiers with soil profiles (0–300 cm) under crop, apple orchard, and vegetable fields were investigated through high-throughput sequencing and quantitative PCR technologies.
Results and discussion
NO3−-N largely accumulated in the deeper soil layers (100–300 cm) in the apple orchard, resulting in a higher risk of NO3−-N leaching. The soil bacterial community structure at the 0–100 cm soil depth had a marketable difference from that at 100–300 cm under these three land-use types, and the C:N ratio was the main driving factor for their vertical distribution. The bacterial α-diversity decreased with soil depth; the crop field had the highest α-diversity across all horizons except 100–200 cm, and total carbon was the most important factor driving α-diversity. In addition, the absolute abundance of the nirK, nirS, and nosZ genes decreased with soil depth and varied with land-use type, which was deeply affected by multiple soil properties, such as soil organic matter and total nitrogen.
Conclusion
Our findings highlighted that potentially important and unique functions remain to be revealed in subsoils, which may provide new insights into mitigating nitrate leaching in various land-use types.
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
1 Introduction
Nitrogen (N) fertilizers are commonly applied to improve crop yields in agricultural activities (Lv et al. 2022; Wang et al. 2019). However, with the development of agricultural intensification, repeated and excessive N application in agricultural production has resulted in serious N loss, soil acidification, and groundwater nitrate pollution, which are potentially harmful to human health (Wheeler et al. 2015). In addition, various land-use types have differences in fertilization, irrigation, and cropping practices, thus causing various nitrate N (NO3−-N) accumulation patterns in soils (Ju et al. 2006). The annual average N input in the North China Plain was reported to be approximately 1358 kg N ha−1 in a greenhouse vegetable field, 2.5 and 2.1 times higher than that in a grainfield and apple orchard, respectively (Ju et al. 2006). The long-term surplus application of N fertilizer in wheat‒maize fields, orchards, and greenhouse vegetables has accelerated N accumulation in the 0–90 cm, 0–100 cm, and 0–120 cm soil profiles, which is approximately 184, 594, and 881 kg N ha−1, respectively (Fan et al. 2014; Gao et al. 2021; Wang et al. 2014). The accumulated N could be more quickly and easily leached into deep soil, even vadose soil, after heavy rainfall or flood irrigation (Gao et al. 2021).
Soil contains the most abundant species of microorganisms on Earth, and it is one of the most important habitats for microorganisms, providing essential nutrients for microbial colonization and growth (Lian et al. 2022). Recent researches have discovered the response of soil microbial communities to fertilization, especially using high amounts of N (Boot et al. 2016; Fierer et al. 2012; Wang et al. 2018). A previous study revealed that chemical fertilization (NPK) significantly reduced bacterial richness in the 0–10 cm soil layer (Liu et al. 2020b), and N amendment significantly reduced the relative abundance of Acidobacteria and enhanced the relative abundance of Proteobacteria (Zhou et al. 2017). Acidobacteria and Proteobacteria belong to oligotrophic and copiotrophic groups, respectively (Fierer et al. 2012). Apart from N inputs, soil types and anthropogenic long-term production activities such as irrigation and tillage also affect soil bacterial communities (Muhammad et al. 2022; Zhang et al. 2022). Early reports indicated that land-use types could induce significant variations in the soil microbial community structure, alpha- or beta-diversity, and dominant taxa (Lauber et al. 2008; Romdhane et al. 2022). Keystone species are highly related taxa in the microbial community structure and play a uniquely pivotal role in the microbial community (Xun et al. 2021). The removal of keystone species may lead to dramatic changes in the function and composition of the microbial community (Banerjee et al. 2018). Notably, soil microbes have crucial roles in the element biogeochemical cycle around the earth, and variations in soil microbial communities will inevitably affect soil ecological functions, such as carbon and N cycling (Bowles et al. 2014; Lian et al. 2022). Within the N cycle, denitrification, catalyzed by microorganisms, is a process of reducing NO3−-N to gaseous nitrogen, and the functional genes for denitrification (nirK, nirS, and nosZ) usually act as indicators of the denitrifying ability (Li et al. 2019; Wang et al. 2021). There is research demonstrating that long-term N inputs can increase the relative abundances of denitrification genes, thus stimulating the denitrification capacity due to the abundance of the substrate nitrate for denitrifiers (Wang et al. 2018). Previous research has revealed that organic fertilizer amendments improve the efficiency and activity of soil denitrifiers in orchards and reduce N pollution, suggesting that denitrifiers can help alleviate soil nitrate leaching (Kramer et al. 2006). A subsequent study in another microcosmic culture experiment has showed similar results (Chen et al. 2018).
Although studies have highlighted that soil microbial communities can be affected by land-use types, including microbial denitrifiers, most studies only focus on topsoil (Braker et al. 2015; Ding et al. 2022; Lauber et al. 2008). However, the response of soil microbes in deep soil to fertilization under different land-use types remains poorly understood. Soil properties, including pH, soil organic matter, and total carbon, differ considerably with depth (Liu et al. 2020b), which further impact the structure of the microbial community and denitrification activities (Merloti et al. 2019). A previous study found a strong correlation between vertical separation of microbial communities and TN (Liu et al. 2020b). Early reports also found that denitrification gene abundance decreased with increasing soil depth, which was coherent with the change in soil organic matter in winter wheat and summer maize fields (Liu et al. 2020b; Wang et al. 2021). However, in the North China Plain, the impact of fertilization on the vertical distribution of soil microbes and denitrifiers under other typical orchard and vegetable land-use types is less understood. In this study, based on soil samples from 0 to 300 cm under three typical land-use types, a winter wheat‒summer maize field, an apple orchard, and a vegetable field, we aimed to (1) investigate nitrate distribution with depth under the above three typical land-use types, (2) explore the variations in the bacterial communities and denitrifiers with depth under the above three typical land-use types, and (3) clarify the responses between soil properties and the structural and functional variances of bacterial communities under above three typical land-use types.
2 Materials and methods
2.1 Soil sampling
Field sites were situated in Luancheng, Shijiazhuang, Hebei Province, China (37°88′N, 114°65′E), with Cambisol soil and a warm temperate monsoon-type climate. Three typical land-use types with more than 20 years of continuous fertilization were selected: a winter wheat‒summer maize field (WM) (600 kg compound fertilizer ha−1 y−1), an apple orchard (OR) (750 kg potassium phosphate, 1244 kg diammonium hydrogen phosphate, 1244 kg urea, and 10,495 kg pig manure ha−1 y−1) and a tomato vegetable field (VE) (68,216 kg pig manure and 300 kg compound fertilizer ha−1 y−1). Soil samples (0–300 cm depth) were collected in October 2016 from each land-use type using specially designed soil columns (43 mm diameter by 120 cm long) with a Geoprobe (Geoprobe 54DT, USA). Each soil core was split into four layers: 0–30 cm, 30–100 cm, 100–200 cm, and 200–300 cm. In all, 36 samples (three land-use types × four soil layers × three replicates) were collected and delivered to the lab on ice in labeled sterile polythene bags. Each soil sample was passed through a 2-mm sieve and then separated into two parts: one part was used for soil property analysis (stored at 4°C), and the other part was used for molecular analysis (stored at − 80 °C).
2.2 Soil property analysis
The determination of soil properties relied on previous studies (Sun et al. 2018; Wang et al. 2022). Soil samples were dried at 105°C for 12 h to a constant weight to determine the soil water content. Soil pH was assessed at a soil/water (carbon dioxide-free) ratio of 1:2.5 by a pH meter (PHS–3C, Shanghai INESA). The dichromate oxidation method was used to analyze soil organic matter (SOM). An element analyzer (Vario PYRO cube, Elementar, Germany) was used to measure soil total nitrogen (TN) and total carbon (TC). Soil nitrate (NO3−–N) was extracted with 2 M potassium chloride and then determined using a spectrophotometer (UV–6100S, Shanghai METASH).
2.3 Soil DNA extraction and quantification of denitrification genes
The total microbial DNA of each soil sample was extracted with 0.5 g fresh soil according to the manufacturer’s instructions using a FastDNA Spin Kit for Soil (MP Biomedicals, USA). DNA purity and concentration were determined by a Nanodrop spectrophotometer (NanoDrop™ One, Thermo Fisher Scientific, USA). The extracted DNA was stored at − 20°C before further analysis.
Three genes (nirK, nirS, and nosZ), as indicators of the denitrification processes, were quantified using the SYBR Green approach by quantitative polymerase chain reaction (qPCR) based on a former study (Sun et al. 2015). The primers used in this study were FlaCu (5′-ATCATGGTSCTGCCGCG-3′) and R3Cu (5′-GCCTCGATCAGRTTGTGGTT-3′) for the nirK gene, cd3aF (5′-GTSAACGTSAAGGARACSGG-3′) and R3cd (5′-GASTTCGGRTGSGTCTTGA-3′) for the nirS gene, and nosZF (5′-CGYTGTTCMTCGACAGCCAG-3′) and nosZ1622R (5′-CGSACCTTSTTGCCSTYGCG-3′) for the nosZ gene (Throback et al. 2004). The qPCR system (25.0 μL) was as follows: 12.5 μL SYBR Premix Ex Taq™ (Takara Biotech, Dalian, China), 0.5 μL of each primer (10 μM), 1.0 μL DNA (about 20 ng/μL), and 10.5 μL sterilized water. A tenfold serial dilution of the plasmids harboring the target functional genes was used to generate standard curves. The thermal cycling for the reaction of the three denitrification genes was performed according to a previous study (Liu et al. 2020b). Bio-Rad CFX Manager 3.1 (Bio-Rad, Hercules, CA) was used to analyze all qPCRs, and the specificity of the qPCR amplifications was checked based on melting curves.
2.4 16S rRNA gene sequencing and bioinformatics
The primer set 515F (5′-GTGYCAGCMGCCGCGGTAA-3′) and 806R (5′-GGACTACNVGGGTWTCTAAT-3′) was used to amplify the 16S rRNA gene (V4 region) to explore the soil bacterial communities (Walters et al. 2016). The amplification and purification of the 16S rRNA gene were detailed in our former study (Sun et al. 2018). The amplified products were purified, and sequencing was then performed using an Illumina NovaSeq platform (Shanghai Personalbio Technology Co., Ltd.). QIIME2 (Quantitative Insights Into Microbial Ecology, version 2019.7) was used to process the high-throughput raw sequencing data (Bolyen et al. 2019). DADA2 (Divisive Amplicon Denoising Algorithm) was used for quality filtering, denoising, joining, dechimeric sequences, and clustering of the sequencing data and then identified as amplicon sequence variants (ASVs) (Callahan et al. 2016). The classify-sklearn naïve Bayes taxonomy classifier in the feature-classifier plugin was used to assign the taxonomy of each ASV based on the Silva database (Release 132) (Bokulich et al. 2018). Diversity analyses (alpha- and beta-diversity) were calculated using the q2-diversity plugin (https://github.com/qiime2/q2-diversity). The annotation of the function of bacterial taxa was identified using the Functional Annotation of Prokaryotic Taxa (FAPROTAX) database (Louca et al. 2016). The sequencing data were uploaded to the National Center for Biotechnology Information (NCBI) under accession number PRJNA891031.
2.5 Statistical analysis
The variance analysis of soil properties, Shannon index, functional categories, and denitrifying gene abundances was carried out by one-way analysis of variance or with a nonparametric Kruskal‒Wallis test using IBM SPSS version 20. Principal coordinate analysis (PCoA) was performed to visualize the soil bacterial community structure. Based on the Bray–Curtis distance, permutational multivariate analysis of variance (PERMANOVA) was performed to examine the variations in bacterial communities at different soil depths and land-use types. The relationship between the bacterial communities and soil properties was determined using multivariate regression tree (MRT) analysis in the “mvpart” and “MVPARTwrap” packages. Keystone species were identified by the betweenness centrality value using the R package “igraph” (Chen et al. 2020; Csardi and Nepusz 2006). Spearman’s correlation analysis of the α-diversity, functional categories, denitrification gene abundance, and soil properties was performed using the R package “corrplot.” Heatmaps used for the relative abundance of functional categories under different land-use types with soil depth were generated using the R package “pheatmap.”
3 Results
3.1 Changes in soil properties with depth under different land-use types
The soil properties varied among the different land-use types (Table 1). At the 0–30 cm soil depth, WM treatment had the highest pH, SOM, TC, and NO3−–N contents than that in the VE and OR treatments. For example, the SOM and NO3−–N contents were 21.49 ± 1.80 g/kg and 24.92 ± 3.30 mg/kg in the WM treatment, 15.18 ± 3.51 g/kg and 6.10 ± 0.79 mg/kg in the OR treatment, and 15.69 ± 1.07 g/kg and 13.67 ± 3.44 mg/kg in the VE treatment, respectively. However, the changes in the soil properties with depth were inconsistent among the WM, VE, and OR treatments. In addition, the soil properties presented different trends with soil depth under each land-use type. Soil pH and the C:N ratio had an overall increasing tendency with soil depth, while SOM and TN decreased with soil depth under each land-use type. Notably, the NO3−–N content dramatically increased with soil depth in the OR treatment, from 6.10 ± 0.79 mg/kg at 0–30 cm to 256.31 ± 6.76 mg/kg at 200–300 cm, which was more than 20 times higher than that in the WM and VE treatments. The NO3−–N content first decreased with soil depth and then slightly increased in deeper soil layers in the WM and VE treatments.
3.2 Changes in the soil bacterial communities with depth under different land-use types
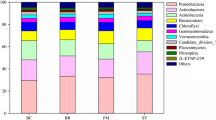
In total, 2,991,175 sequences were acquired from all 36 samples after quality screening was performed, which clustered into 2,795,348 ASVs. The bacterial community composition varied with different land-use types. In the 0–30 cm soil layer, six phyla Proteobacteria, Actinobacteria, Acidobacteria, Chloroflexi, Planctomycetes, and Gemmatimonadetes accounted for 85.11%, 86.37%, and 77.72% of the bacterial composition in the WM, OR, and VE treatments, respectively (Fig. 1). The relative abundance of Actinobacteria in the VE treatment was only 8.35%, which was much lower than that in the WM (21.51%) and OR (20.67%) treatments. Contrarily, the VE treatment harbored the highest relative abundances of Acidobacteria and Firmicutes, up to 20.33% and 13.12%, respectively. Additionally, the relative abundance of some phyla varied from the topsoil to deep soil layers (Fig. 1). For example, from 0–30 to 200–300 cm in the VE treatment, the relative abundance of Proteobacteria increased from 27.40 to 87.11%, and that of Bacteroidetes increased from 1.10 to 6.49%, while the relative abundances of Gemmatimonadetes, Chloroflexi, and Firmicutes overall decreased with soil depth, which had a similar trend in the WM and OR treatments.
According to the PCoA (Fig. 2a) and PERMANOVA (Table S1) results, there was no significant separation in the bacterial community structure in each soil layer across different land-use types. However, there were significant changes in the bacterial communities with soil depth, as the bacterial structure in the 0–100 cm soil layer was significantly different from that in the 100–200 cm and 200–300 cm layers (Table S1). In addition, the Shannon index, representing the bacterial α-diversity, had a similar trend with soil depth under each land-use type. For example, the Shannon index decreased with soil depth from 0–30 to 200–300 cm, except in the WM treatment, which showed a slight rebound at 200–300 cm compared with 100–200 cm (Fig. 2b). The WM treatment harbored significantly higher bacterial α-diversity than the OR and VE treatments in the 0–30 cm, 30–100 cm, and 200–300 cm soil layers. However, no obvious difference was found between the OR and VE treatments across all horizons, except 100–200 cm.
a Principal coordinate analysis (PCoA) plot of the soil bacterial communities at different soil depths under the three treatments. Each circle represents the same treatment; b bacterial α-diversity in different soil layers under the three treatments. Different letters indicate significant differences among treatments in the same soil layer (p < 0.05). WM, winter wheat‒summer maize field; OR, apple orchard field; VE, tomato vegetable field
3.3 Correlations between soil properties and bacterial communities
MRT analysis clarified the correlation between soil properties and the vertical distribution of the bacterial communities, which explained 81.81% of the bacterial community variation (Fig. 3). The C:N ratio divided the distribution of the bacterial communities into two major parts: one part mainly contained 0–100 cm samples with a C:N ratio < 35.00, and the other part contained 100–300 cm samples with a C:N ratio ≥ 35.00 (Fig. 3). In addition, the soil properties had different effects on the α-diversity in each soil layer (Table S2). For instance, α-diversity had a significant and positive correlation with SOM, TC, TN, and NO3−–N in the 0–30 cm soil layer (p < 0.05), but had a negative correlation with NO3−–N in the 30–100 cm soil horizon (p < 0.05) and TC in the 100–200 cm soil horizon (p < 0.05). Soil properties had no significant relationship with α-diversity in the 200–300 cm soil layer.
Multivariate regression tree (MRT) of bacterial communities in soil under the different land-use types. The land-use types and depth are in the rectangle, and the number of samples is in parentheses. The percentages under each split represent the percentage of difference explained by the split. WM, winter wheat‒summer maize field; OR, apple orchard field; VE, tomato vegetable field; SOM, soil organic matter; TC, total carbon
3.4 Keystone species and predictive functions under different land-use types
The top twenty ASVs were regarded as keystone species according to the betweenness centrality value (Table S3). The keystone species composition and relative abundance varied under the different land-use types. In the 0–30 cm soil horizon, the most predominant keystone species in the WM, OR, and VE treatments belonged to the Actinobacteria, Actinobacteria, and Chloroflexi phyla, which accounted for 0.49%, 1.37%, and 0.24%, respectively (Fig. S1). The keystone species, affiliated with Firmicutes, only existed in the VE treatment at 0–30 cm and 30–100 cm depths. Additionally, the relative abundance of keystone species varied with soil depth. For example, from 0–30 to 200–300 cm in the WM and OR treatments, the relative abundance of Rokubacteria in keystone species increased from 0 to 1.08% and from 0 to 0.39%, respectively. Moreover, Actinobacteria and Rokubacteria were dominant in the WM and OR treatments at 0–30 cm and 200–300 cm, respectively, while Chloroflexi and Actinobacteria were dominant in the VE treatment at 0–30 cm and 200–300 cm depths.
The relative abundance of the functional categories of N cycling varied under different land-use types and exhibited various trends with soil depth (Fig. 4a). In the 0–30 cm soil layer, the relative abundance of the nitrification and aerobic ammonia oxidation functional categories was the lowest in the WM treatment, accounting for 8.60% and 6.19%, respectively. Additionally, the relative abundance of the denitrification functional category in the WM treatment was slightly higher than that in the OR and VE treatments. In the 30–100 cm soil layer, the OR treatment (7.40%) had a higher relative abundance of the denitrification functional category than that in the WM (3.72%) and VE (3.95%) treatments, but no significant difference was found in the 100–200 cm and 200–300 cm soil layers. The relative abundances of the nitrification, aerobic ammonia oxidation, and denitrification functional categories were significantly and negatively correlated with TC and the C:N ratio, while those of the aerobic ammonia oxidation and denitrification functional categories were positively correlated with SOM and TN (Fig. 4b). However, no significant correlation was found between denitrification and NO3−–N.
a The relative abundances of the nitrogen cycling functional categories in different soil layers under the WM, OR, and VE treatments. Different letters represent significant differences among treatments in the same soil layer (p < 0.05). WM, winter wheat‒summer maize field; OR, apple orchard field; VE, tomato vegetable field; b Spearman’s correlation between the nitrogen cycling functional categories and soil properties. The circle size represents the Spearman correlation coefficient. Red and blue represent positive correlation and negative correlation, respectively, and asterisks represent significant correlation. *: p < 0.05; **: p < 0.01; ***: p < 0.001
3.5 Changes in denitrifying genes with depth under different land-use types
The vertical distributions of nirK, nirS, and nosZ were explored under different land-use types (Fig. 5a). In the 0–30 cm soil layer, the nirK, nirS, and nosZ gene abundances in the WM treatment were the highest, reaching 2.51 × 108, 3.11 × 108, and 1.27 × 108 copies per g of dry soil, respectively, followed by those in the OR and VE treatments. From 30 to 300 cm, the OR treatment had the highest abundance of the nirK, nirS, and nosZ genes. In addition, the abundance of the nirK, nirS, and nosZ genes decreased with soil deepness in each land-use type, which decreased by approximately 2–4 orders of magnitude from 0–30 to 200–300 cm deep soil. In addition, the abundances of the nirK, nirS, and nosZ genes had significant and positive correlations with SOM and TN, but had negative correlations with the C:N ratio, TC, and pH (Fig. 5b). Unexpectedly, the soil NO3−–N content had no obvious relation with the denitrification genes.
a The absolute abundances (copies per g dry soil) of the nirK, nirS, and nosZ genes in the WM, OR, and VE treatments. Data were log-transformed. Different letters represent significant differences among treatments in the same soil layer (p < 0.05). WM, winter wheat‒summer maize rotation cropland; OR, apple orchard; VE, tomato vegetable field; b Spearman’s correlation between the denitrification gene abundances and soil properties. The circle size represents the Spearman correlation coefficient. Red and blue represent positive correlation and negative correlation, respectively, and asterisks represent significant correlation. *: p < 0.05; **: p < 0.01; ***: p < 0.001
4 Discussion
4.1 Land-use type effects on soil bacterial communities and denitrifying genes
As predicted, the soil bacterial communities were affected partly by land-use types, as indicated by their distinction from one another (Figs. 1 and 2), which is in line with earlier studies (Babin et al. 2019; Jimenez-Bueno et al. 2016). These distinctions were caused by multiple factors, such as fertilization, planting systems, and soil properties. In this study, only chemical fertilizer was applied in the WM treatment, while both chemical fertilizer and pig manure were applied in the OR and VE treatments. Vegetable fields rely heavily on manure fertilizer for regulating soil properties. Numerous studies have shown that the changes in soil properties caused by fertilization can induce differences in soil bacterial communities (Chai et al. 2021; Gu et al. 2019; Liu et al. 2020b). Taking the 0–30 cm soil depth as an example, the WM treatment harbored the most diverse bacterial communities compared with the OR and VE treatments, which is partly caused by the higher contents of SOM, TC, and NO3−–N, as they were significantly and positively correlated with the soil bacterial α-diversity (Table S2). Additionally, Francioli et al. found that the soil bacterial diversity with the combined application of chemical (NPK) and organic (cattle manure) fertilizer was higher than that with chemical fertilizer application alone (2016). However, a previous study reported that the quality and application of organic fertilizer also affect soil bacterial diversity (Hartmann et al. 2015). Therefore, low-quality organic fertilizer or improper application may lead to a decrease in bacterial diversity. In addition, Acosta-Martinez et al. demonstrated that the soil bacterial diversity in the rotation system was higher than that in the single cropping system because rooting systems and different crop residues in the rotation system had a stronger impact on soil bacterial diversity (2008). In contrast, the bacterial communities in the WM, OR, and VE treatments had no significant difference from each other (p > 0.05, Table S1), as all land-use types were geographically close to each other and existed under the same climatic conditions (Barberán et al. 2015). However, the dominant phyla had different relative abundances across these land-use types. Due to the heavy application of pig manure, VE treatment had the highest relative abundance of Firmicutes than that in the WM and OR treatments. Previous studies have reported that Firmicutes is the main phylum in organic fertilizer (Liu et al. 2022a; Semenov et al. 2021). This result indicated that manure-amended bacteria were abundant in these manure-amended soils. In addition, the relative abundance of Actinobacteria in the VE treatment was lower than that in the WM and OR treatments, which may be closely related to the cultivation cycle of greenhouse vegetables (Song et al. 2018). Previous research revealed that the relative abundance of Actinobacteria decreased with planting cycles in greenhouses (Song et al. 2018).
Keystone species had various abundances and compositions across different land-use types. The keystone species belonging to Firmicutes existed only in the VE treatment in the 0–30 cm and 30–100 cm soil layers, which was attributed to Firmicutes as the dominant bacteria in manure fertilizer, and the application of manure fertilizer increased its abundance in manure-amended soils (Liu et al. 2022a). Additionally, a previous study reported that Firmicutes plays an important role in decomposing vegetable litter, which may be one of the reasons why Firmicutes exists only in the 0–100 cm soil layer of vegetable fields (Zhang et al. 2020). Furthermore, the N cycling functional categories revealed that the WM treatment had the lowest relative abundance of the nitrification and aerobic ammonia oxidation functional categories. This may be related to the lower N input, as N fertilizer applied to farmland soil could facilitate nitrification and the aerobic ammonia oxidation capacity (Chu et al. 2008). Rao et al. demonstrated that the changes in soil properties caused by fertilization also affect functional categories (2021). In this study, TC and the C:N ratio significantly affected nitrification, aerobic ammonia oxidation, and denitrification.
The WM treatment had higher abundances of denitrification genes than those in the OR and VE treatments in the 0–30 cm soil layer. A previous study also analyzed denitrification gene abundance under different land-use types and found that wheat‒maize fields had the highest denitrification gene abundance, which was attributed to sufficient NO3− promoting the growth of denitrifying bacteria (He et al. 2020). However, in this study, denitrification genes had no significant correlation with the NO3−–N content but had significant and positive relations with SOM and TN (Fig. 5b). It has been reported that rich soluble SOM in soil, especially water-soluble organic carbon, promotes the soil denitrification process (Surey et al. 2020). Additionally, other researches also found a positive relationship between TN and the abundance of most denitrification genes (Lindsay et al. 2010; Reverchon et al. 2015). The abundant SOM and TN in the WM treatment provided favorable conditions for denitrification.
4.2 Depth effect on soil bacterial communities and denitrifying genes
In this study, soil properties varied with soil depth. For instance, SOM and TN decreased with soil depth under each land-use type, which is in line with other studies (Emadi et al. 2008; Fetene and Amera 2018). Additionally, reduced root exudates and plant litter may result in a decrease in the SOM content with the soil profile (Kramer and Gleixner 2008). Notably, heavy amounts of NO3−–N accumulated in deep soil layers, posing a serious risk of N leaching (Cui et al. 2020). A previous study revealed that frequent irrigation and excessive N inputs led to severe soil nitrate leaching (Liu et al. 2022b).
Apart from soil properties, the influence of soil depth on bacterial richness and bacterial structures was also apparent (Figs. 1 and 2). Previous studies have shown that carbon and nitrogen sources are essential resources for cell growth and have crucial factors affecting bacterial diversity and community structure (Eilers et al. 2012; Seuradge et al. 2017). The NO3−–N content, SOM, and especially the C:N ratio had prominent effects on the vertical distribution of the bacterial communities in this study. Previous study demonstrated that the C:N ratio significantly influenced bacterial growth and reproduction (Gu et al. 2019). On another scale, plant inputs, such as root exudates and litter, may also partly reflect their influence on the variety of bacterial communities between topsoil and deep soil (Seuradge et al. 2017). Previous studies have shown that the various root system distributions account for the wide variations in carbon sources, which might be responsible for the stratification of microbes (Fierer et al. 2003; Goberna et al. 2005). In addition, a substantial portion of microorganisms existed in deep soil layers, and their potential capacities could not lower or even higher than those of microorganisms in surface soil (Fierer et al. 2003). Consistent with previous studies, the relative abundance of Proteobacteria increased with soil depth (Li et al. 2014; Liu et al. 2020a). Proteobacteria exists widely in various environments, including acidic and nutrient-poor conditions, and increases in labile substrate-abundant environments (Curtis et al. 2002; Goldfarb et al. 2011). Soil depth played a strong environmental factor that influenced the bacterial communities (Liu et al. 2020b; Schlatter et al. 2020). The relative abundance of Gemmatimonadetes decreased with soil depth, which was identified as one of the top nine phyla in soil and preferred soil environments (DeBruyn et al. 2011; He et al. 2017). In this study, the prominent keystone species in each soil layer were different in the WM, OR, and VE treatments. The differences in prominent keystone species with soil depth under different land-use types may reflect their viability and life history strategies in distinct habitats (Fierer 2017; Schlatter et al. 2020).
The absolute abundances of the nirK, nirS, and nosZ genes also decreased with soil deepness under these three land-use types (Fig. 5a), which is consistent with other studies (Chen et al. 2018; Wang et al. 2021). This was also reflected by the denitrification functional categories (Fig. 4a). Although NO3−–N, an important substrate for denitrification, has a significant effect on the denitrification potential (Wang et al. 2018), the denitrifier abundance still decreased at 100–300 cm under the OR treatment, which had more than 30–40 times the NO3−–N content of the surface soil. One study noted that denitrifiers are mostly heterotrophic and grow in an environment rich in organic carbon (Gao et al. 2015), indicating that a lower carbon source inhibits the growth of denitrifiers in deep soil. Recent studies have noted that carbon availability is the major hindering factor for denitrification activity in deep soil (Chen et al. 2018; Zhou et al. 2016). Carbon amendments have been considered to sharpen denitrification capacity in deep vadose zone soils (Chen et al. 2018).
5 Conclusion
In this study, the orchard had serious nitrate accumulation at 200–300 cm compared with the other two land-use types, due to the large amount of fertilizer input and the irrigation pattern. In addition, the vertical distribution of the soil bacterial communities under different land-use types was affected by the soil properties, and the C:N ratio was the major driving factor. Land-use types also remarkably influenced the soil bacterial diversity, with the crop field having the highest bacterial diversity in most soil layers. Moreover, the denitrification gene abundance decreased with soil depth under these three land-use types, and the lower denitrification activity partly caused the accumulation of nitration in the orchard soil. This study provided insights into the status quo of the soil nitrate, bacterial community, and denitrification activity in deep soil under three typical land-use types.
Data availability
The data that support the findings of this study are available from the corresponding author upon reasonable request.
References
Acosta-Martinez V, Dowd S, Sun Y, Allen V (2008) Tag-encoded pyrosequencing analysis of bacterial diversity in a single soil type as affected by management and land use. Soil Biol Biochem 40:2762–2770
Babin D, Deubel A, Jacquiod S, Sorensen SJ, Geistlinger J, Grosch R, Smalla K (2019) Impact of long-term agricultural management practices on soil prokaryotic communities. Soil Biol Biochem 129:17–28
Banerjee S, Schlaeppi K, van der Heijden MGA (2018) Keystone taxa as drivers of microbiome structure and functioning. Nat Rev Microbiol 16:567–576
Barberán A, Ladau J, Leff JW, Pollard KS, Menninger HL, Dunn RR, Fierer N (2015) Continental-scale distributions of dust-associated bacteria and fungi. Proc Natl Acad Sci USA 112:5756–5761
Bokulich NA, Kaehler BD, Rideout JR, Dillon M, Bolyen E, Knight R, Huttley GA, Caporaso JG (2018) Optimizing taxonomic classification of marker-gene amplicon sequences with QIIME 2 ’ s q2-feature-classifier plugin. Microbiome 6:90
Bolyen E et al (2019) Reproducible, interactive, scalable and extensible microbiome data science using QIIME 2. Nat Biotechnol 37:852–857
Boot CM, Hall EK, Denef K, Baron JS (2016) Long-term reactive nitrogen loading alters soil carbon and microbial community properties in a subalpine forest ecosystem. Soil Biol Biochem 92:211–220
Bowles TM, Acosta-Martinez V, Calderon F, Jackson LE (2014) Soil enzyme activities, microbial communities, and carbon and nitrogen availability in organic agroecosystems across an intensively-managed agricultural landscape. Soil Biol Biochem 68:252–262
Braker G, Matthies D, Hannig M, Brandt FB, Brenzinger K, Groengroeft A (2015) Impact of land use management and soil properties on denitrifier communities of namibian savannas. Microb Ecol 70:981–992
Callahan BJ, McMurdie PJ, Rosen MJ, Han AW, Johnson AJA, Holmes SP (2016) DADA2: high-resolution sample inference from Illumina amplicon data. Nat Methods 13:581–583
Chai X, Yang Y, Wang X, Hao P, Wang L, Wu T, Zhang X, Xu X, Han Z, Wang Y (2021) Spatial variation of the soil bacterial community in major apple producing regions of China. J Appl Microbiol 130:1294–1306
Chen J, Wang P, Wang C, Wang X, Miao L, Liu S, Yuan Q, Sun S (2020) Fungal community demonstrates stronger dispersal limitation and less network connectivity than bacterial community in sediments along a large river. Environ Microbiol 22:832–849
Chen S, Wang F, Zhang Y, Qin S, Wei S, Wang S, Hu C, Liu B (2018) Organic carbon availability limiting microbial denitrification in the deep vadose zone. Environ Microbiol 20:980–992
Chu H, Fujii T, Morimoto S, Lin X, Yagi K (2008) Population size and specific nitrification potential of soil ammonia-oxidizing bacteria under long-term fertilizer management. Soil Biol Biochem 40:1960–1963
Csardi G, Nepusz T (2006) The igraph software package for complex network research. Int J Complex Syst 1695:1–9
Cui M, Zeng L, Qin W, Feng J (2020) Measures for reducing nitrate leaching in orchards: a review. Environ Pollut 263:114553
Curtis P, Nakatsu CH, Konopka A (2002) Aciduric Proteobacteria isolated from pH 2.9 soil. Arch Microbiol 178:65–70
DeBruyn JM, Nixon LT, Fawaz MN, Johnson AM, Radosevich M (2011) Global biogeography and quantitative seasonal dynamics of Gemmatimonadetes in soil. Appl Environ Microbiol 77:6295–6300
Ding L, Zhou J, Li Q, Tang J, Chen X (2022) Effects of land-use type and flooding on the soil microbial community and functional genes in reservoir riparian zones. Microb Ecol 83:393–407
Eilers KG, Debenport S, Anderson S, Fierer N (2012) Digging deeper to find unique microbial communities: the strong effect of depth on the structure of bacterial and archaeal communities in soil. Soil Biol Biochem 50:58–65
Emadi M, Emadi M, Baghernejad M, Fathi H, Saffari M (2008) Effect of land use change on selected soil physical and chemical properties in North Highlands of Iran. J Appl Sci 8:496–502
Fan Z, Lin S, Zhang X, Jiang Z, Yang K, Jian D, Chen Y, Li J, Chen Q, Wang J (2014) Conventional flooding irrigation causes an overuse of nitrogen fertilizer and low nitrogen use efficiency in intensively used solar greenhouse vegetable production. Agric Water Manag 144:11–19
Fetene EM, Amera MY (2018) The effects of land use types and soil depth on soil properties of Agedit watershed, Northwest Ethiopia. Ethiop J Sci Technol 11:39–56
Fierer N (2017) Embracing the unknown: disentangling the complexities of the soil microbiome. Nat Rev Microbiol 15:579–590
Fierer N, Lauber CL, Ramirez KS, Zaneveld J, Bradford MA, Knight R (2012) Comparative metagenomic, phylogenetic and physiological analyses of soil microbial communities across nitrogen gradients. ISME J 6:1007–1017
Fierer N, Schimel JP, Holden PA (2003) Variations in microbial community composition through two soil depth profiles. Soil Biol Biochem 35:167–176
Francioli D, Schulz E, Lentendu G, Wubet T, Buscot F, Reitz T (2016) Mineral vs. organic amendments: microbial community structure, activity and abundance of agriculturally relevant microbes are driven by long-term fertilization strategies. Front Microbiol 7:1446
Gao J, Wang S, Li Z, Wang L, Chen Z, Zhou J (2021) High nitrate accumulation in the vadose zone after land-use change from croplands to orchards. Environ Sci Technol 55:5782–5790
Gao W, Yang H, Kou L, Li S (2015) Effects of nitrogen deposition and fertilization on N transformations in forest soils: a review. J Soils Sediments 15:863–879
Goberna M, Insam H, Klammer S, Pascual JA, Sanchez J (2005) Microbial community structure at different depths in disturbed and undisturbed semiarid Mediterranean forest soils. Microb Ecol 50:315–326
Goldfarb KC, Karaoz U, Hanson CA, Santee CA, Bradford MA, Treseder KK, Wallenstein MD, Brodie EL (2011) Differential growth responses of soil bacterial taxa to carbon substrates of varying chemical recalcitrance. Front Microbiol 2:94
Gu Y, Wei Y, Xiang Q, Zhao K, Yu X, Zhang X, Li C, Chen Q, Xiao H, Zhang X (2019) C: N ratio shaped both taxonomic and functional structure of microbial communities in livestock and poultry breeding wastewater treatment reactor. Sci Total Environ 651:625–633
Hartmann M, Frey B, Mayer J, Maeder P, Widmer F (2015) Distinct soil microbial diversity under long-term organic and conventional farming. ISME J 9:1177–1194
He H, Miao Y, Zhang L, Chen Y, Gan Y, Liu N, Dong L, Dai J, Chen W (2020) The structure and diversity of nitrogen functional groups from different cropping systems in Yellow River Delta. Microorganisms 8:424
He S, Guo L, Niu M, Miao F, Jiao S, Hu T, Long M (2017) Ecological diversity and co-occurrence patterns of bacterial community through soil profile in response to long-term switchgrass cultivation. Sci Rep 7:3608
Jimenez-Bueno NG, Valenzuela-Encinas C, Marsch R, Ortiz-Gutierrez D, Verhulst N, Govaerts B, Dendooven L, Navarro-Noya YE (2016) Bacterial indicator taxa in soils under different long-term agricultural management. J Appl Microbiol 120:921–933
Ju X, Kou C, Zhang F, Christie P (2006) Nitrogen balance and groundwater nitrate contamination: comparison among three intensive cropping systems on the North China Plain. Environ Pollut 143:117–125
Kramer C, Gleixner G (2008) Soil organic matter in soil depth profiles: distinct carbon preferences of microbial groups during carbon transformation. Soil Biol Biochem 40:425–433
Kramer SB, Reganold JP, Glover JD, Bohannan BJM, Mooney HA (2006) Reduced nitrate leaching and enhanced denitrifier activity and efficiency in organically fertilized soils. Proc Natl Acad Sci U S A 103:4522–4527
Lauber CL, Strickland MS, Bradford MA, Fierer N (2008) The influence of soil properties on the structure of bacterial and fungal communities across land-use types. Soil Biol Biochem 40:2407–2415
Li C, Yan K, Tang L, Jia Z, Li Y (2014) Change in deep soil microbial communities due to long-term fertilization. Soil Biol Biochem 75:264–272
Li H, Zhang Y, Wang T, Feng S, Ren Q, Cui Z, Cao C (2019) Responses of soil denitrifying bacterial communities carrying nirS, nirK, and nosZ genes to revegetation of moving sand dunes. Ecol Indic 107:105541
Lian J, Wang H, Deng Y, Xu M, Liu S, Zhou B, Jangid K, Duan Y (2022) Impact of long-term application of manure and inorganic fertilizers on common soil bacteria in different soil types. Agric Ecosyst Environ 337:108044
Lindsay EA, Colloff MJ, Gibb NL, Wakelin SA (2010) The abundance of microbial functional genes in grassy woodlands is influenced more by soil nutrient enrichment than by recent weed invasion or livestock exclusion. Appl Environ Microbiol 76:5547–5555
Liu C, Yan J, Huang Q, Liu H, Qiao C, Li R, Shen B, Shen Q (2022a) The addition of sawdust reduced the emission of nitrous oxide in pig manure composting by altering the bacterial community structure and functions. Environ Sci Pollut Res 29:3733–3742
Liu G, Chen L, Deng Q, Shi X, Lock TR, Kallenbach RL, Yuan Z (2020a) Vertical changes in bacterial community composition down to a depth of 20 m on the degraded Loess Plateau in China. Land Degrad Dev 31:1300–1313
Liu M, Min L, Wu L, Pei H, Shen Y (2022b) Evaluating nitrate transport and accumulation in the deep vadose zone of the intensive agricultural region. North China Plain Sci Total Environ 825:153894
Liu M, Zhang W, Wang X, Wang F, Dong W, Hu C, Liu B, Sun R (2020b) Nitrogen leaching greatly impacts bacterial community and denitrifiers abundance in subsoil under long-term fertilization. Agric Ecosyst Environ 294:106885
Louca S, Parfrey LW, Doebeli M (2016) Decoupling function and taxonomy in the global ocean microbiome. Science 353:1272–1277
Lv X, Ma B, Sun L, Cai Y, Chang SX (2022) Long-term nitrogen fertilization, but not short-term tillage reversal, affects bacterial community structure and function in a no-till soil. J Soils Sediments 22:630–639
Merloti LF, Mendes LW, Pedrinho A, de Souza LF, Ferrari BM, Tsai SM (2019) Forest-to-agriculture conversion in Amazon drives soil microbial communities and N-cycle. Soil Biol Biochem 137:107567
Muhammad I, Yang L, Ahmad S, Zeeshan M, Farooq S, Ali I, Khan A, Zhou X (2022) Irrigation and nitrogen fertilization alter soil bacterial communities, soil enzyme activities, and nutrient availability in maize crop. Front Microbiol 13:833758
Rao D, Meng F, Yan X, Zhang M, Yao X, Kim KS, Zhao J, Qiu Q, Xie F, Zhang W (2021) Changes in soil microbial activity, bacterial community composition an d function in a long-term continuous soybean cropping system after corn insertion and fertilization. Front Microbiol 12:638326
Reverchon F, Bai SH, Liu X, Blumfield TJ (2015) Tree plantation systems influence nitrogen retention and the abundance of nitrogen functional genes in the Solomon Islands. Front Microbiol 6:1439
Romdhane S, Spor A, Banerjee S, Breuil M-C, Bru D, Chabbi A, Hallin S, van der Heijden MGA, Saghai A, Philippot L (2022) Land-use intensification differentially affects bacterial, fungal and protist communities and decreases microbiome network complexity. Environ Microbiome 17:1
Schlatter DC, Kahl K, Carlson B, Huggins DR, Paulitz T (2020) Soil acidification modifies soil depth-microbiome relationships in a no-till wheat cropping system. Soil Biol Biochem 149:107939
Semenov MV, Krasnov GS, Semenov VM, Ksenofontova N, Zinyakova NB, van Bruggen AHC (2021) Does fresh farmyard manure introduce surviving microbes into soil or activate soil-borne microbiota? J Environ Manag 294:113018
Seuradge BJ, Oelbermann M, Neufeld JD (2017) Depth-dependent influence of different land-use systems on bacterial biogeography. FEMS Microbiol Ecol 93:fiw239
Song Y, Xu M, Li X, Bian Y, Wang F, Yang X, Gu C, Jiang X (2018) Long-term plastic greenhouse cultivation changes soil microbial community structures: a case study. J Agric Food Chem 66:8941–8948
Sun R, Guo X, Wang D, Chu H (2015) Effects of long-term application of chemical and organic fertilizers on the abundance of microbial communities involved in the nitrogen cycle. Appl Soil Ecol 95:171–178
Sun R, Li W, Dong W, Tian Y, Hu C, Liu B (2018) Tillage changes vertical distribution of soil bacterial and fungal communities. Front Microbiol 9:699
Surey R, Lippold E, Heilek S, Sauheitl L, Henjes S, Horn MA, Mueller CW, Merbach I, Kaiser K, Boettcher J, Mikutta R (2020) Differences in labile soil organic matter explain potential denitrification and denitrifying communities in a long-term fertilization experiment. Appl Soil Ecol 153:103630
Throback IN, Enwall K, Jarvis A, Hallin S (2004) Reassessing PCR primers targeting nirS, nirK and nosZ genes for community surveys of denitrifying bacteria with DGGE. FEMS Microbiol Ecol 49:401–417
Walters W, Hyde ER, Berg-Lyons D, Ackermann G, Humphrey G, Parada A, Gilbert JA, Jansson JK, Caporaso JG, Fuhrman JA, Apprill A, Knight R (2016) Improved bacterial 16S rRNA gene (V4 and V4–5) and fungal internal transcribed spacer marker gene primers for microbial community surveys. mSystems 1:e00009
Wang F, Chen S, Qin S, Sun R, Zhang Y, Wang S, Hu C, Hu H, Liu B (2021) Long-term nitrogen fertilization alters microbial community structure and denitrifier abundance in the deep vadose zone. J Soils Sediments 21:2394–2403
Wang F, Chen S, Wang Y, Zhang Y, Hu C, Liu B (2018) Long-term nitrogen fertilization elevates the activity and abundance of nitrifying and denitrifying microbial communities in an upland soil: implications for nitrogen loss from intensive agricultural systems. Front Microbiol 9:2424
Wang R, Liu Z, Yao Z, Lei Y (2014) Modeling the risk of nitrate leaching and nitrate runoff loss from intensive farmland in the Baiyangdian Basin of the North China Plain. Environ Earth Sci 72:3143–3157
Wang Y, Chen P, Wang F, Han W, Qiao M, Dong W, Hu C, Zhu D, Chu H, Zhu Y (2022) The ecological clusters of soil organisms drive the ecosystem multifunctionality under long-term fertilization. Environ Int 161:107133
Wang Y, Ying H, Yin Y, Zheng H, Cui Z (2019) Estimating soil nitrate leaching of nitrogen fertilizer from global meta-analysis. Sci Total Environ 657:96–102
Wheeler DC, Nolan BT, Flory AR, DellaValle CT, Ward MH (2015) Modeling groundwater nitrate concentrations in private wells in Iowa. Sci Total Environ 536:481–488
Xun W, Liu Y, Li W, Ren Y, Xiong W, Xu Z, Zhang N, Miao Y, Shen Q, Zhang R (2021) Specialized metabolic functions of keystone taxa sustain soil microbiome stability. Microbiome 9:35
Zhang Q, Han Y, Chen W, Guo Y, Wu M, Wang Y, Li H (2022) Soil type and pH mediated arable soil bacterial compositional variation across geographic distance in North China Plain. Appl Soil Ecol 169:104220
Zhang Z, Han X, Yan J, Zou W, Wang E, Lu X, Chen X (2020) Keystone microbiomes revealed by 14 years of field restoration of the degraded agricultural soil under distinct vegetation scenarios. Front Microbiol 11:1915
Zhou J, Gu B, Schlesinger WH, Ju X (2016) Significant accumulation of nitrate in Chinese semi-humid croplands. Sci Rep 6:25088
Zhou J, Jiang X, Wei D, Zhao B, Ma M, Chen S, Cao F, Shen D, Guan D, Li J (2017) Consistent effects of nitrogen fertilization on soil bacterial communities in black soils for two crop seasons in China. Sci Rep 7:3267
Funding
This work was financially supported by the National Natural Science Foundation of China (No. 41930865, No. 42077358, No. 41877425) and the Strategic Priority Research Program of the Chinese Academy of Sciences (No. XDB40020204).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Responsible editor: Hang-Wei Hu
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Han, W., Wang, F., Zhang, L. et al. Variations of soil bacterial community and denitrifier abundance with depth under different land-use types. J Soils Sediments 23, 1889–1900 (2023). https://doi.org/10.1007/s11368-023-03428-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11368-023-03428-8