Abstract
Purpose
Implementing practices that maximize water use efficiency in arid zones is important as agriculture relies mostly on irrigation in these agroecosystems. The aim of this study was to identify the effect of combining different sowing irrigation methods (dry and wet) with contrasting tillage practices (conventional practices and conservation agriculture) on bacterial community structure in soils from the Norman E. Borlaug experimental field (CENEB), Yaqui Valley, Sonora.
Materials and methods
Soil samples were collected consecutively four times in the month after sowing from permanent and conventionally tilled beds under wet and dry sowing at CENEB. Soils were characterized and the bacterial communities were identified with Illumina MiSeq sequencing where the 16S rRNA gene was targeted.
Results and discussion
The bacterial community structure was affected by soil water content. Streptomyces and Balneimonas were enriched in dry sowed soil, the first in conventionally tilled beds and the latter in permanent beds, while Acinetobacter was enriched in wet sowed soil under conservation agriculture. Shannon index indicated that there was a high species diversity in both conventional practices and conservation agriculture.
Conclusions
Soil water content was the principal factor shaping the bacterial community structure. While the three most dominant phyla were affected by the difference in water content in the tillage practices under dry and wet sowing, it was at the genera level that these fluctuations were more obvious.
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
1 Introduction
Wheat (Triticum spp.), the third most important grain produced in the world, plays an important role in food security, as it is an affordable source of protein and carbohydrates (Balkovič et al. 2014). In Mexico, the leading wheat-producing states are in the north of the country, Sonora being the principal producer. More than 50% of wheat produced in Mexico is grown in the Yaqui Valley, Sonora, a region that is characterized by an arid climate (Verhulst et al. 2011), thus crop cultivation mainly depends on irrigation (Mondani et al. 2019). It is therefore important to adopt more sustainable agricultural practices in the region to ensure water use efficiency.
The International Maize and Wheat Improvement Center (CIMMYT, acronym in Spanish) has been experimenting with different agricultural practices to improve yields while maintaining sustainability of the agroecosystems for many years (Govaerts et al. 2006; Hobbs et al. 2008). Conservation agriculture (PB) is a sustainable alternative to conventional agricultural practices (CB), which rely heavily on tillage, a practice that causes physical soil disturbances that affect the mechanical structure and porosity of soil, leading to loss of nutrients and alteration in bacterial community structure (Wang et al. 2016; Anderson et al. 2017). Conservation agriculture involves the application of little or no tillage (minimize soil disturbance), retention of crop residue to form a permanent soil cover, and rotation of crops aiming to improve soil quality, reduce water runoff, improve water infiltration, and increase soil organic matter content, which ultimately increases plant nutrients (Hobbs 2007; Habig and Swanepoel 2015). A major advantage of PB is that it increases water-use efficiency while increasing yields and lowering management costs (Li et al. 2011; Verhulst et al. 2011).
Additionally, dry sowing as an alternative to wet sowing could increase water use efficiency. Conventionally in the Yaqui Valley, fields are irrigated about 2 to 3 weeks prior to crop sowing so that weeds that germinate can be controlled at sowing, allowing the crop to germinate in a weed-free field; this is referred to as wet sowing (Govaerts et al. 2009). In dry sowing, fields are irrigated 1 or 2 days after being sown, resulting in higher soil water content during germination compared to wet sowing (Mulvaney et al. 2014). Both sowing irrigation practices involve wetting and drying events of soil, which may have profound effects on bacterial activity and consequently on plant nutrient availability (Denej et al. 2001). Soil drying has a negative impact on matric and osmotic potential limiting the amount of water available to soil bacteria (Mavi and Marschner 2012). This can dehydrate bacteria and in extreme circumstances their activity may come to a halt (Muhr et al. 2010). Rewetting a soil stimulates microbial activity and increases the soil nutrient content, i.e., nitrogen and phosphorous, a phenomenon referred to as the “Birch effect” (Birch 1958; Thomson et al. 2010). Dry rewetting also alters the bacterial community structure as the changes in available C and nutrients enrich some bacterial groups, but not others (Fierer and Schimel 2002; Muhr et al. 2010).
For farmers that are adopting PB, sowing a dry crop residue-rich soil is easier than a moist soil. Therefore, CIMMYT has been investigating dry sowing under PB and conventional tillage practices. The difference in soil water content in dry sowing compared to wet sowing will affect the soil microbial population. Microbial activity will increase earlier in wet sowing than in dry sowing and thus stimulate plant growth–promoting bacteria earlier in wet than in dry sowing (Guo et al. 2013). Plant growth–promoting bacteria, such as Paenibacillus polymyxa, Achromobacter piechaudii ARV8, Pantoea agglomerans, and Azospirillum sp., help wheat crops cope with biotic and abiotic stress (Grover et al. 2011), so crop development might be favored in wet sowing than in dry sowing.
The aim of this study was to determine changes in bacterial population brought on by fluctuations in water content due to dry and wet sowing under PB (permanent beds) and CB (conventional tilled beds).
2 Materials and methods
2.1 Field experiment
Soil samples were taken from an experiment at the Norman Borlaug experimental station (CENEB) (lat. 27.33 N, long. 109.09 W, 38 masl) located in the Yaqui Valley, near Ciudad Obregon (Mexico) with an arid climate and a soil classified as a Hyposodic Vertisol (Calcaric, Chromic) (Verhulst et al. 2011).
The experiment was initiated in the winter of 2007–2008 and had a maize (Zea mays)–durum wheat (Triticum turgidum) rotation with maize in the summer and wheat in the winter. The experiment had two tillage treatments: PB, with permanents beds where only the furrows were reshaped each season but the soil on top of the beds is not tilled, and CB, with conventionally tilled beds that were tilled and remade each season. Crop residues were left on the soil surface in PB and incorporated through tillage in CB. The tillage treatments have been in place since 1996 and were divided in treatments with dry and wet sowing. Sub-plots had six different N management treatments, of which the treatment fertilized with 240 kg urea-N ha−1 (30% applied pre-planting and 70% at first node) was selected for this study. This resulted in four treatments (n = 4), including PB and CB, each with a wet and dry sowing practice, with three replicated plots (n = 3). As such, 12 plots were sampled (n = 12) (Fig. S1, Table S1). Each plot contained four raised beds of 0.75 m width and 10 m length (total plot size 30 m2), with two wheat rows sown on the bed at 24-cm spacing.
The PB and CB under wet sowing were first irrigated on 6 November, approximately 2 weeks before the wheat cultivar CIRNO C2008 was sown on 23 November 2015 (Fig. S1). Wheat cultivar CIRNO C2008 was also sown in the PB and CB under dry sowing on 23 November 2015 and first irrigated on 25 November.
2.2 Soil sampling
Soil samples were collected from 12 plots (dry and wet sowing in the PB and CB treatments in triplicate) on the day the seeds were sown, i.e., 23 November, and additionally on 2, 8, and 16 December 2015 as described below. Bulk soil was collected randomly in 10 sampling points from the 0–7 cm top layer of the 12 plots and pooled separately so that 12 soil samples were obtained on each sampling day. A 50-g subsample of each soil sample was weighed and dried at 105 °C to determine soil water content. Overall, a total of 48 samples (12 on each sampling day (n = 4)) were obtained and each sample was air dried before transportation to the Laboratory of Ecology (Cinvestav, Mexico City). Soil samples were characterized separately and extracted for DNA separately as described below.
2.3 Soil characterization
The soil samples were characterized physicochemically. A glass electrode was used to determine pH in a soil–water suspension (Thomas 1996) and electrolytic conductivity (EC) was determined using the saturated soil-paste extract method described by Rhoades et al. (1989). The method described by Cassel and Nielsen (1986) was used to determine the water holding capacity (WHC) of the soil samples Soil water content was determined by the filter-paper method (Fawcett and Collis-George 1967) nitrogen (TN). The hydrometer method was used to determine soil texture (Bouyoucos 1962), total organic carbon (TOC) was measured following Tiessen and Moir (1993) dry oxidation method, and the Kjeldahl acid digestion described by Bremner (1996) was used to determine total.
2.4 Metagenomic DNA extraction and PCR amplification of the bacterial gene
Fulvic and humic acids were removed from a 0.5-g sub-sample of soil as described by Ceja-Navarro et al. (2010). Three methods (enzymatic, chemical, and thermal shock lysis) were used for DNA extraction and described in Chávez-Romero et al. (2016). Each extraction method was done in triplicate; hence, 1.5 g soil was extracted for DNA per method per sample. The DNA extraction samples obtained with each method (n = 3) were pooled, amplified, and sent for analysis. As such, 4.5 g soil was extracted for DNA from each sample of each plot (n = 3). Overall, 13.5 g soil was extracted per treatment per sampling day.
The variable V3 and V4 regions of the 16S rRNA gene were amplified using the following region of specific primers with overhang adapters (Klindworth et al. 2013): forward (5′-TCGTCGGCAGCGTCAGATGTGTATAAGAGACAG-3′); reverse (5′-GTCTCGTGGGCTCGGAGATGTGTATAAGAGACAG-3′). The conditions of the PCR cycles used were as indicated by Navarro-Noya et al. (2013). Amplicons of approximately 500 bp were purified using the Ultra Clean PCR Clean-up kit (MO BIO Laboratories, CA) as recommended by the manufacturer and quantified in a NanoDrop™ 3300 (Thermo Scientific NanoDrop) using Quant-iT™ PicoGreen® dsDNA (Invitrogen, Carlsbad, USA). The libraries were prepared, normalized and pooled, and subsequently sequenced on the Illumina MiSeq system by Macrogen, Inc. (DNA Sequencing Service, Seoul, Korea).
2.5 Analysis of Illumina sequencing data
The QIIME pipeline version 1.9.1 was used to process the raw bacterial sequences (Caporaso et al. 2010b). Sequences were demultiplexed and filtered for low-quality reads with a Phred score <19 removed. Operational taxonomic unit (OTU) clusters were generated at a similarity threshold of 97% (OTU97) using Uclust by running open reference OTU picking pipeline to compare reads against Greengenes v13_8 database (Edgar 2010). The representative sequence for each OTU97 was aligned with Python nearest alignment space termination (PyNAST) version 1.2.2 and those that did not align at a sequence identity of >75% were removed (Caporaso et al. 2010a). The taxonomic composition was assigned with ribosomal database project using the naïve Bayesian rRNA classifier (http://edp.cme.msu.edu/classifier/classifier.jps) for a representative set of sequences selected for each OTU97 (Wang et al. 2007). The taxonomic assignments were used to construct biological observations matrices (BIOM). The alpha diversity indices were calculated with BIOM tables rarefied at 3000 reads per sample (Kuczynski et al. 2011). The raw sequences were deposited in the National Center for Biotechnology Information (NCBI) database Sequence Read Archive (SRA) under the BioProject PRJNA542494.
2.6 Statistical analysis
All statistical analyses were done in R (R Development Core Team 2013). A non-parametric test (t2way test in the WRS2 package “A collection of robust statistical methods”) (Mair 2018) was used to determine the effect of the tillage practice (PB and CB) and sowing irrigation practice (wet and dry sowing) and their interaction on soil characteristics. Abundance of the different bacterial phyla and genera was explored separately with a principal component analysis (PCA) and a constrained analysis of principal coordinates (CAP) was used to explore the effect of treatment on the bacterial phyla and genera and soil characteristics. A permutational multivariate analysis of variance (PERMANOVA) using distance matrices test (adonis, method Bray–Curtis, argument strata) was used to determine the effect of tillage practice and sowing irrigation on the bacterial phyla and genera. The PCA, CAP, and adonis tests were done with the vegan package (Oksanen et al. 2018). Heatmaps were constructed with the pheatmap package (Kolde 2018). The random forest algorithm was used to evaluate the effect of the tillage practices and sowing irrigation on soil characteristics, alpha diversity, and bacterial phyla and genera over time (Breiman et .al 2018).
3 Results
3.1 Soil physicochemical properties
Water content, and total C and N showed highly significant differences between the treatments over time (Table S2). Tillage practices (PB or CB) (F = 8.41, p = 0.008) and sowing irrigation (wet or dry sowing) (F = 8.84, p = 0.007) significantly affected EC (Table 1, S3). Other soil characteristics were not affected significantly by sowing irrigation or tillage practices.
3.2 Bacterial community structure
The number of reads was filtered and rarefied at 2996 to include all 48 samples in the analysis, generating a total of 143,808 reads whose sequences resulted in 42,050 OTUs. The rarefication curves showed that analyzing more sequences would generate a limited number of new OTUs (Fig. S2). The Chao1 (F = 4.20, p = 0.008), Shannon (F = 18.48, p < 0.001), and Simpson indexes (F = 4.09, p = 0.009) were highly significant different between the treatments on the different sampling days (Table S4). Tillage practices had a significant effect on the Shannon (F = 9.18, p = 0.006) and Simpson (F = 10.01, p = 0.004) index, but not wet or dry sowing (Table S5).
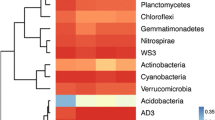
Overall, phylotypes belonged to 37 phyla with the most abundant being Proteobacteria (relative abundance 33.38 ± 4.66%, mostly Alphaproteobacteria 12.12 ± 1.66%), followed by Acidobacteria (19.69 ± 3.11%) and Actinobacteria (11.96 ± 2.55%). These three phyla were clustered together, i.e., they were affected in the same way by tillage practices and sowing irrigation (Fig. 1a).
Heatmap with relative abundance (%) of the bacterial (a) phyla and (b) genera in wet (Wet) and dry sowed soil (Dry) with conservation agriculture (PB) and conventional agricultural practices (CB) at CIMMYT’s Norman E. Borlaug experimental station (CENEB) near Ciudad Obregón (Sonora, Mexico). Soil was sampled on 23 November (d1), 2 December (d2), 8 December (d3), and 16 December 2015 (d4). Boolean values were used to cluster rows (bacterial groups) and columns (different treatments on different sampling days)
Steroidobacter (relative abundance 1.80 ± 0.56%) was the most abundant genus followed by Bacillus (1.66 ± 0.73%), Kaistobacter (1.46 ± 0.51%), Flavisolibacter (1.27 ± 0.42), Acinetobacter (1.08 ± 1.72), and Rubrobacter (1.07 ± 0.35), the 50 most abundant genera with relative abundance >0.10% (Fig. 1b). These most abundant bacterial genera generally responded in the same way to tillage practices and wet or dry sowing.
3.3 Effect of sowing irrigation and tillage practices on the bacterial community
Wet sowing or dry sowing had a significant effect on the bacterial community structure in both PB and CB (F ≥ 2.36, p ≤ 0.015) (Table 2). The PCA separated the wet from the dry sowing in both CB and PB, but the separation was more accentuated when the 50 most abundant genera were considered than when the bacterial phyla were used (Fig. 2). In CB, Pedobacter and Streptomyces were enriched in dry sowing and Pseudonocardia in wet sowing. In PB, Balneimonas and Thermomonas were enriched in dry sowing and Methylibium in wet sowing.
Principal component analysis (PCA) with the relative abundance (%) of the bacterial phyla in (a) soil with conventional agriculture practices (CB) soil and (b) with conservation agriculture (PB), and with the 50 most abundant bacterial genera in soil in (c) CB and (d) PB. Wet sowed soil sampled on 23 November (○), 2 December (□), 8 December (◊), and 16 December (∆), and in dry sowed soil on 23 November (●), 2 December (■), 8 December (♦), and 16 December (▲) at CIMMYT’s Norman E. Borlaug experimental station (CENEB) near Ciudad Obregón (Sonora, Mexico). Soil was sampled in 2015
Tillage practices had a highly significant effect on the bacterial community structure in wet sowing considering the bacterial phyla and 50 most abundant genera (F ≥ 4.42, p ≤ 0.003), but not in dry sowing (Table 2). The PCA separated CB from PB in the wet sowing, but not in the dry sowing (Fig. 3). In the wet sowing, the separation between the PB and CB treatments was more accentuated when the 50 most abundant genera were considered than when the bacterial phyla were used. In wet sowing, Cellvibrio, Flavisolibacter, Pontibacter, and Proteobacteria were enriched in CB and Actinobacteria, Nitrospirae, Rhodoplanes, and [Thermi] in PB.
Principal component analysis (PCA) with the relative abundance (%) of the bacterial phyla in (a) wet sowed soil and (b) dry sowed soil, and with the 50 most abundant bacterial genera in (c) wet sowed soil and (d) dry sowed soil. Soil was sampled in conventional agricultural practices (CB) on 23 November (○), 2 December (□), 8 December (◊), and 16 December (∆), and in conservation agriculture (PB) on 23 November (●), 2 December (■), 8 December (♦), and 16 December (▲ permanent beds at CIMMYT’s Norman E. Borlaug experimental station (CENEB) near Ciudad Obregón (Sonora, Mexico). Soil was sampled in 2015
On each of the sampling days, the PCA with the bacterial phyla and genera separated the different treatments (Figs. S3 and S4). The effect of sowing irrigation, i.e., wet versus dry sowing, on the bacterial community structure considering the different bacterial phyla (p ≤ 0.009) and genera (p ≤ 0.022) was significantly different, except on 2 December 2015. The effect of tillage practices on the bacterial phyla was only significant on 23 November and 8 December, and on 16 December 2015 considering the 50 most abundant genera (Table S6).
Considering the mean of the four sampling days, the PERMANOVA test indicated that the bacterial phyla (F = 6.11, p < 0.001) and genera (F = 2.33, p = 0.013) were affected significantly by wet or dry sowing. Tillage practices had a significant effect on the bacterial community structure considering the different genera (F = 2.33, p = 0.013), but not when considering the bacterial phyla (F = 2.48, p = 0.081) (Table S6). Considering the mean of the four sampling days, the relative abundance of Acidobacteria, Cyanobacteria, Elusimicrobia, Nitrosovibrio, Nitrospirae, and Virgisporangium was higher in soil with wet sowing compared to dry sowing, while that of Bacteroidetes, Balneimonas, Kaistobacter, and Proteobacteria and WS3 was lower (Fig. S5). The relative abundance of Verrucomicrobia was highly positive correlated with soil water content (p < 0.01) and Streptomyces negatively with total N content (p < 0.001) (Fig. S6).
On each sampling day, the CAP separated the different treatments considering the bacterial phyla and genera (Fig. S7 and S8). The function capscale (CAP) in the vegan package in R is a constrained version of metric scaling, a.k.a. principal coordinates analysis, which is based on the Euclidean distance, but can be used, and is more useful, with other dissimilarity measures (Oksanen et al. 2018). The capscale function (vegan) was used to constrain the physicochemical parameters analyzed and the means of the different bacterial phyla and genera relative abundances of the combined sampling days (Bulgarelli et al. 2015). The CAP analysis showed a clear shift with wet sowing compared to dry sowing in both PB and CB and variations between the plots were smaller in wet sowing than in dry sowing (Fig. 4). The relative abundance of Actinobacteria, Bacteroidetes, and Proteobacteria was higher in soil under dry sowing compared to wet sowing and that of Acidobacteria was lower (Fig. 4a). The relative abundance of Acinetobacter and Stenotrophomonas decreased generally in soil with dry sowing compared to wet sowing, and that of Bacillus and Steroidobacter increased (Fig. 4b).
Canonical analysis of principal coordinates (CAP) with the mean of the relative abundance (%) of (a) the bacterial phyla and (b) the 50 most abundant bacterial genera and physicochemical characteristics of the soil sampled on 23 November, 2 December, 8 December, and 16 December 2015. Wet sowed soil with conventional agricultural practices (CB) (□) and conservation agriculture (PB) (○), and dry sowed soil with CB (■) and PB (●) at CIMMYT’s Norman E. Borlaug experimental station (CENEB) near Ciudad Obregón (Sonora, Mexico)
3.4 Effect of sampling time on the bacterial community
A PCA with the different bacterial phyla and genera did not clearly separate the different sampling days in the different treatments (Fig. S9 and S10). However, considering the mean relative abundance of the different bacterial phyla and genera of the different tillage practices, the PCA separated the different sampling days (Fig. 5). The relative abundance of Acidobacteria, Bacillus, Elusimicrobia, Nitrospirae, Rubrobacter, and Steroidobacter was higher on the first sampling day while that of Acinetobacter and Proteobacteria was higher on the last sampling day. The PERMANOVA analysis indicated that the composition of the bacterial phyla (F = 6.05, p < 0.001) and genera (F = 2.74, p = 0.003) was highly significantly different between the sampling days (Table S6). A CAP analysis with the different bacterial phyla and genera and soil characteristics did not clearly separate the different sampling days in the different treatments (Figs. S11 and S12). However, considering the mean relative abundance of the different bacterial phyla and genera of the different tillage practices, the CAP analysis separated the different sampling days (Fig. 6).
Principal component analysis (PCA) with the mean relative abundance (%) of (a) the bacterial phyla and (b) the 50 most abundant bacterial genera in wet sowed and dry sowed soil with conservation agriculture and conventional practices on 23 November (□), 2 December (■), 8 December (○), and 16 December (●) 2015 at CIMMYT’s Norman E. Borlaug experimental station (CENEB) near Ciudad Obregón (Sonora, Mexico)
Canonical analysis of principal coordinates (CAP) with the mean relative abundance (%) of (a) the bacterial phyla and (b) the 50 most abundant bacterial genera and physicochemical characteristics in wet sowed and dry sowed soil with conservation agriculture and conventional practices on 23 November (□), 2 December (■), 8 December (○), and 16 December (●) 2015 at CIMMYT’s Norman E. Borlaug experimental station (CENEB) near Ciudad Obregón (Sonora, Mexico)
4 Discussion
The bacterial community was determined in the bulk soil. The first sampling day was before plant emergence, while seedlings with minimal rhizosphere development were present on the other sampling days. As such, the effect of the crop on the bacterial community should be absent or minimal. It is possible that the summer crop, i.e., maize, affected the bacterial community at the beginning of our sampling period, as suggested by Alvey et al. (2003). Since a maize–wheat crop rotation has been applied for over 10 years in both agricultural practices studied here, the effect of plant species (root exudates) on bacterial community structure to be similar in all samples (Benitez et al. 2017).
4.1 Changes in the soil physicochemical characteristics due to wet or dry sowing, tillage practices, and sampling time
There was more than 2 weeks’ difference between application of the first irrigation in wet sowed soils, i.e., 6 November, and dry sowed soils, i.e., 25 November. Hence, the water content was higher in wet sowed soil than in dry sowed on the first sampling day, 23 November. Electrolytic conductivity is susceptible to fluctuations in soil water content (Zhang and Wienhold 2002). An increase in water content during rewetting causes an increase in water potential and diffusion of soluble substrates. These soluble ions tend to leach to deeper layers thus causing a decrease in EC, whereas soil drying, hence a decrease in water availability, avoided this, i.e., soluble salts accumulated (USDA 2011). As such, EC was generally higher in wet sowed than in dry sowed soil.
Soil drainage, which leaches out soil minerals, is another factor that affects EC (Grisso et al. 2009). Tillage breaks up aggregates, decreasing pore size, ultimately decreasing infiltration. Additionally, the use of heavy machinery in CB compacts soil, which lowers also infiltration (Manyiwa and Dikinya 2014). Minimum tillage and crop residue cover improve aggregate stability thereby preventing run-off and improving water infiltration (Verhulst et al. 2011). Improved water infiltration in PB increased drainage and the leaching out of minerals resulting in a decrease in EC (Stagnari et al. 2009).
4.2 The bacterial community structure
The Shannon index indicated that the bacterial diversity was similar in conservation and conventional agricultural practices as well as in dry and wet sowing. The Shannon index in this study was close to the value (H′ = 11.21) reported in organic farming under similar climatic conditions where rice straw, water hyacinth, wood chips, organic waste, clay, chicken, and cow manure were amended to soil (Köberl et al. 2011). High and diverse organic matter availability will increase bacterial diversity (Thiele-Bruhn et al. 2012).
Numerous studies have shown that soil pH helps shape soil bacterial diversity (Wang et al. 2012), and extreme soil alkalinity or acidity might reduce bacterial diversity (Lauber et al. 2009). The pH in soil of CENEB was slightly alkaline as in the study of Köberl et al. (2011), so its effect on the bacterial diversity was limited.
The bacterial community was dominated by Proteobacteria, Acidobacteria, and Actinobacteria, often described as the most abundant bacterial phyla in arable soil. For instance, Zhao et al. (2014) found also that these three phyla were dominant in arable soils with a similar wheat–maize rotation. Wang et al. (2016) also reported that Proteobacteria, Actinobacteria, and Acidobacteria were the most abundant phyla in conventional and conservation tillage practices in arid to semi-arid zones. Arid regions are prone to extremely long periods of dry weather before rewetting, so these three bacterial phyla seemed to be well adapted these conditions (Blazewicz et al. 2014). It made them also well adapted to the arid conditions and sporadic rewetting events at CENEB (Evans and Wallenstein 2012). None of the bacterial genera was clearly dominant. The most abundant bacterial genera have often been described in arable soil. Moreno-Espíndola et al. (2018) reported a high relative abundance of Bacillus, Flavisolibacter, Kaistobacter, and Steroidobacter in soils under a conventional agricultural and an organic milpa system in the central highlands of Mexico.
4.3 Effect of wet or dry sowing on the bacterial community
Since our team has extensively investigated the effect of different agricultural practices on dynamics of C and N in arable soil (Ramirez-Villanueva et al. 2015; Chavez-Romero et al. 2016), the focus of this study was strictly to investigate how the difference in water content as a result of different irrigation strategies explained affected the bacterial community structures.
Water content is one of the major factors shaping the bacterial community structure in arid soils (Lennon et al. 2012). In our study, for instance, the relative abundance of Verrucomicrobia was directly affected by fluctuations in water availability and it increased as soil water content increased. Maestre et al. (2015) found the same in their study on the effects of drying and rewetting in drylands. Prolonged dry conditions inhibit normal functioning of the soil microorganisms and they become mostly inactive, but they have developed different mechanisms to survive (Fierer and Schimel 2002). For instance, some members of Bacillus form spores while some strains of Cyanobacteria and Pseudomonas secrete mucilage, extracellular polymeric substances, to protect themselves against drought (Costa et al. 2018). When a dry soil is wetted, microbial activity resumes (Muhr et al. 2010). Organic material that was previously physically protected is released providing microorganisms with an easily decomposable organic C substrate, while microorganisms that died under dry conditions further increase the available organic material (Mavi and Marschner 2012). Copiotrophs that favor nutrient-rich environments are enriched, while oligotrophs that favor nutrient deficient conditions are replaced (Blazewicz et al. 2014; Naylor and Coleman-Derr 2018). Under prolonged wet conditions, O2 diffusion is restricted and facultative or obligated anaerobes flourish (Yan et al. 2015). Nitrification and CH4 oxidation, both aerobic processes, are inhibited and denitrification and methanogenesis, anaerobic processes, are favored (Lennon et al. 2012). Microorganisms involved in the latter processes are enriched, while those involved in the first cease to be active (Moreno-Espíndola et al. 2018).
Water was applied in the wet sowing treatment on 6 November. Microbial activity was reactivated, and the availability of easily decomposable organic material subsequently changed the bacterial community structure (Miller et al. 2005). The relative abundance of Acidobacteria, Bacillus, and Rubrobacter was higher on the first sampling day (23 November) compared to the other sampling days. Acidobacteria respond opportunistically to rewetting. Their relative abundance decreases during desiccation, but rapidly increases when the soil is rewetted (Maestre et al. 2015). The phylum also responded opportunistically in this study. Members of Bacillus form spores to protect against abiotic stress factors, such as desiccation, remaining in a dormant state with little or no metabolic activity until conditions for growth improve, such as increased water and nutrient availability (Setlow 2014). After irrigation was applied in wet sowing, the number of spores (relative abundance of Bacillus) was high due to the germination of their dormant spores and their enrichment as conditions were favorable on the first sampling day. Many species of the genus Rubrobacter are known to resist desiccation as they are mostly mesophilic, moderately thermophilic, or thermophilic (Chen et al. 2018). Phylotypes belonging to Rubrobacter responded favorably to the increased water content after irrigation was applied in wet sowed soil and were abundant on the first sampling day.
In the dry sowing treatment, the microbial activity was still low on the first sampling day (23 November), so the bacterial community structure was different from that in the wet sowing treatment. The second sampling took place 1 week after the dry sowing treatment was irrigated and an increase in microbial activity occurred. However, this change in microbial activity occurred 22 days later than in the wet sowing. The changes in the bacterial community structure followed the same pattern in the dry as in the wet sowing treatments, but later. Consequently, the bacterial community structures were different between the wet and dry sowing. For instance, Bacteroidetes and Proteobacteria exhibited copiotrophic characteristics as previously reported by Fierer et al. (2007). They were enriched in the dry sowed soil, compared to wet sowed soil, where the availability of decomposable organic matter increased after rewetting. Acidobacteria and Acinetobacter, both oligotrophs (Fierer et al. 2007), were more abundant in wet sowed soil than in dry sowed soil. In the wet sowed soil, the easily decomposable organic material was decomposed earlier than in the dry sowed soil. Consequently, oligotrophs were enriched earlier in the wet sowed soil than in dry sowed soil.
In this study, the timing of rewetting the soil had a strong effect on the bacterial community independent of the tillage practices applied. The timing of moistening the soil had such a strong impact on the bacterial community that considering all sampling days or averaging them did not alter its highly significant effect. The difference in bacterial community structure between wet and dry sowing also lasted at least a month. For instance, Acinetobacter was more abundant on the fourth sampling day (16 December) compared to the other sampling days. This suggests that the genus can survive lower water availability. Marasco et al. (2012) found Acinetobacter in cultivated desert soil where water was a major limiting factor and confirmed that the genus can survive dry conditions with limited availability of water by forming spores.
4.4 Effect of tillage practices on the bacterial community
The bacterial community structure was different between PB and CB treatment in wet sowing. In the wet sowing, water was applied on 6 November and the first sampling took place 17 days later. The effect of wetting the soil, i.e., a flush in microbial activity as the organic material released was mineralized, had ceased after 17 days. As such, differences between PB and CB depended nearly solely on differences in soil conditions due to differences in the treatment of crop residue in the field. For instance, the amount of organic material available in PB and CB was different as it was left on the soil surface in the first and incorporated in the second. Members of Actinobacteria have been reported to be copiotrophic (Leff et al. 2015) and should have thrived better in tilled soil where organic matter would have been more accessible than in permanent beds with no tillage. However, our results showed that this phylum was more abundant in wet sowed soil under PB than CB so they could not be characterized as a copiotroph in this study. Fierer et al. (2007) also found that the abundance of Actinobacteria did not change as expected by C availability. In the dry sowing treatment, no significant effect of tillage practices on the bacterial community was detected. On the first sampling day, i.e., 23 November 2015, the soil was still dry, so microbial activity was low in both PB and CB and the differences in bacterial community minimal. The soil was rewetted on 25 November, so the effect on the bacterial community structure of wetting the soil was still strong on the second sampling day, i.e., 7 days later. As such, the effect of increased water content masked a possible effect of tillage practices on the bacterial community, i.e., applying water trumped the effect of the different tillage practices in this study. It was only after the effect of water application on the short-term microbial activity had ceased that the effect of the tillage practices became evident in the microbial community in the dry sowing treatment.
It is worth pointing out that although no screening was done for specific traits. Bacterial genera, such as Acinetobacter, Bacillus, Pseudomonas, and Streptomyces, abundant in this study, have been identified as plant growth–promoting bacteria. They stimulate plant growth and increase their resistance to biotic and abiotic stresses (Beneduzi et al. 2012). They may secrete antibiotics and other metabolites to suppress the activity and growth of plant pathogens (Beneduzi et al. 2012). Plant growth–promoting bacteria might produce also hormones, such as auxins and cytokinins, and other substances that aid in the uptake of nutrients (Dias et al. 2015).
5 Conclusions
This study revealed that soil water content was important in shaping the bacterial community structure, regardless of the tillage practice applied, i.e., conventional practices or conservation agriculture. In fact, the most abundant bacterial groups identified in this study were affected more by wet and dry sowing than by tillage practices and the effect of tillage practices on bacterial structure was only significant in wet sowed soil. The changes in relative abundance over time were due to the sudden increase in soil water content on the different irrigation days in dry and wet sowing.
References
Alvey S, Yang CH, Buerkert A, Crowley DE (2003) Cereal/legume rotation effects on rhizosphere bacterial community structure in West African soils. Biol Fertil Soils 37:73–82
Anderson C, Beare M, Buckley HL, Lear G (2017) Bacterial and fungal communities respond differently to varying tillage depth in agricultural soils. PeerJ 5:e3930
Balkovič J, van der Velde M, Skalský R, Xiong W, Folberth C, Khabarov N, Smirnov A, Mueller ND, Obersteiner M (2014) Global wheat production potentials and management flexibility under the representative concentration pathways. Glob Planet Chang 122:107–121
Beneduzi A, Ambrosini A, Passaglia LM (2012) Plant growth-promoting rhizobacteria (PGPR): their potential as antagonists and biocontrol agents. Genet Mol Biol 35:1044–1051
Benitez MS, Osborne SL, Lehman RM (2017) Previous crop and rotation history effects on maize seedling health and associated rhizosphere microbiome. Sci Rep-UK 7:15709
Birch HF (1958) The effect of soil drying on humus decomposition and nitrogen availability. Plant Soil 10:9–31
Blazewicz SJ, Schwartz E, Firestone MK (2014) Growth and death of bacteria and fungi underlie rainfall-induced carbon dioxide pulses from seasonally dried soil. Ecology 95:162–1172
Bouyoucos GJ (1962) Hydrometer method improved for making particle size analyses of soils. Agron J 54:464–465
Breiman L, Cutler A, (Fortran original) Liaw A, Wiener M (R port) (2018) Breiman and Cutler’s random forests for classification and regression. Package ‘randomForest’. Version: 4.6–14
Bremner JM (1996) Nitrogen-total. In: Sparks DL (ed) Methods of soil analysis: chemical methods part 3. ASA, SSSA, Madison, pp 1085–1122
Bulgarelli D, Garrido-Oter R, Münch PC, Weiman A, Dröge J, Pan Y, McHardy AC, Schulze-Lefert P (2015) Structure and function of the bacterial root microbiota in wild and domesticated barley. Cell Host Microbe 17:392–403
Caporaso JG, Bittinger K, Bushman FD, Desantis TZ, Andersen GL, Knight R (2010a) PyNAST: a flexible tool for aligning sequences to a template alignment. Bioinformatics 26:266–267
Caporaso JG, Kuczynski J, Stombaugh J, Bittinger K, Bushman FD, Costello EK, Fierer N, Peña AG, Goodrich JK, Ji G, Huttley GA, Kelley ST, Knights D, Koening JE, Ley RE, Lozupone CA, McDonald D, Muegge BD, Pirrung M, Reeder J, Sevinsky JR, Turnbaugh PJ, Walters WA, Widmann J, Yatsunenko T, Zaneveld J, Knight R (2010b) QIIME allows analysis of high-throughput community sequencing data. Nat Methods 7:335–336
Cassel DK, Nielsen DR (1986) Field capacity and available water capacity. In: Klute A, Campbell GS, Jackson RD, Mortland MM, Nielsen DR (eds) Methods of soil analysis. Part 1. Physical, mineralogical methods. ASA, SSSA, Madison, pp 901–924
Ceja-Navarro JA, Rivera-Orduña FN, Patiño-Zúñiga L, Villa-Sanjurjo A, Crossa J, Govaerts B, Dendooven L (2010) Phylogenetic and multivariate analyses to determine the effects of different tillage and residue management practices on soil bacterial communities. Appl Environ Microb 76:3685–3691
Chávez-Romero Y, Navarro-Noya Y, Reynoso-Martínez S, Sarria-Guzmán Y, Govaerts B, Verhulst N, Dendooven L, Luna-Guido M (2016) 16S metagenomics reveals changes in the soil bacterial community driven by soil organic C, N-fertilizer and tillage-crop residue management. Soil Till Res 159:1–8
Chen RW, Wang KX, Wang FZ, He YQ, Long LJ, Tien XP (2018) Rubrobacter indicoceani sp. nov., a new marine actinobacterium isolated from Indian Ocean sediment. Int J Syst Evol Micr 68:3487–3493
Costa OYA, Raaijmakers JM, Kuramae EE (2018) Microbial extracellular polymeric substances: ecological function and impact on soil aggregation. Front Microbiol 9:1636
Denej K, Six J, Bossuyt H, Frey SD, Elliott ET, Merckx R, Paustian K (2001) Influence of dry–wet cycles on the interrelationship between aggregate, particulate organic matter, and microbial community dynamics. Soil Biol Biochem 33:1599–1611
Dias T, Dukes A, Antunes PM (2015) Accounting for soil biotic effects on soil health and crop productivity in the design of crop rotations. J Sci Food Agri 95:447–454
Edgar RC (2010) Search and clustering orders of magnitude faster than BLAST. Bioinformatics 26:2460–2461
Evans SE, Wallenstein MD (2012) Soil microbial community response to drying and rewetting stress: does historical precipitation regime matter? Biogeochemistry 109:101–116
Fawcett RG, Collis-George N (1967) A filter paper method for determining the moisture characteristic of soil. Aust J Exp Agr Anim Husb 7:162–167
Fierer N, Schimel JP (2002) Effects of drying-rewetting frequency on soil carbon and nitrogen transformations. Soil Biol Biochem 34:777–787
Fierer N, Bradford MA, Jackson RB (2007) Toward an ecological classification of soil bacteria. Ecology 88:1354–1364
Govaerts B, Sayre KD, Deckers J (2006) A minimum data set for soil quality assessment of wheat and maize cropping in the highlands of Mexico. Soil Till Res 87:163–174
Govaerts B, Verhulst N, Sayre K, Kienle F, Flores D, Limon-Ortega A (2009) Implementing conservation agriculture concepts for irrigated wheat based systems in Northwest Mexico: a dynamic process towards sustainable production. In: Rattan RK, Mathur VC, Sharma AR, Manjaiah KM, Datta SP, Chakraborty A, Chakraborty D, Datta KK (eds) 4th world congress on conservation agriculture: innovations for improving efficiency, equity and environment. M/s Print Process, New Delhi, pp 136–145
Grisso R, Alley MM, Wysor WG, Holshouser D, Thomason W (2009) Precision farming tools: soil electrical conductivity. VA Coop Ext 442-508:1–6
Grover M, Ali SZ, Sandhya V, Rasul A, Venkateswarlu B (2011) Role of microorganisms in adaptation of agriculture crops to abiotic stresses. World J Microb Biot 27:1231–1240
Guo Q, Wang W, Liu Y, Wen X, Liao Y (2013) Effects of soil moisture and temperature on soil microbial ecological environment and soil respiration in rainfed wheat field. Res Crop 14:374–381
Habig J, Swanepoel CM (2015) Effects of conservation agriculture and fertilization on soil microbial diversity and activity. Environments 2:358–384
Hobbs P (2007) Conservation agriculture: what is it and why is it important for future sustainable food production? J Agr Sci 145:127–137
Hobbs PR, Sayre K, Gupta R (2008) The role of conservation agriculture in sustainable agriculture. Philos Trans R Soc B 363:543–555
Klindworth A, Pruesse E, Schweer T, Peplies J, Quast C, Horn M, Glöckner FO (2013) Evaluation of general 16S ribosomal RNA gene PCR primers for classical and next-generation sequencing-based diversity studies. Nucleic Acids Res 41:e1
Köberl M, Müller H, Ramadan EM, Berg G (2011) Desert farming benefits from microbial potential in arid soils and promotes diversity and plant health. PLoS One 6:e24452
Kolde KR (2018) Pretty Heatmaps, package ‘pheatmap’ version: 1.0.10
Kuczynski J, Stombaugh J, Walters WA, González A, Caporaso JG, Knight R (2011) Using QIIME to analyze 16S rRNA gene sequences from microbial communities. Curr Protoc Bioinformatics 36:10.7.1–10.7.20. https://doi.org/10.1002/0471250953.bi1007s36
Lauber CL, Hamady M, Knight R, Fierer N (2009) Pyrosequencing-based assessment of soil pH as a predictor of soil bacterial community structure at the continental scale. Appl Environ Microb 75:5111–5120
Leff JW, Jones SE, Prober SM, Barberán A, Borer ET, Firn JL, Harpole WS, Hobbie SE, Hofmockel KS, Knops JMH, McCulley RL, La Pierre K, Risch AC, Seabloom EW, Schütz M, Steenbock C, Stevens CJ, Fierer N (2015) Consistent responses of soil microbial communities to elevated nutrient inputs in grasslands across the globe. Proc Natl Acad Sci USA 112:10967–10972
Lennon JT, Aanderud ZT, Lehmkuhl BK, Schoolmaster DR Jr (2012) Mapping the niche space of soil microorganisms using taxonomy and traits. Ecology 93:1867–1879
Li L, Huang G, Zhang R, Bill B, Guangdi L, Kwong YC (2011) Benefits of conservation agriculture on soil and water conservation and its progress in China. Agric Sci China 10:850–859
Maestre FT, Delgado-Baquerizo M, Jeffries TC, Eldridge DJ, Ochoa V, Gozalo B, Quero JL, García-Gómez M, Gallardo A, Ulrich W, Bowker MA, Arrendondo T, Barraza-Zepeda C, Bran D, Florentino A, Gaitán J, Gutiérrez JR, Huber-Sannwald E, Jankju M, Mau RL, Miriti M, Naseri K, Ospina A, Stavi I, Wang D, Woods NN, Yuan X, Zaady E, Singh BK (2015) Increasing aridity reduces soil microbial diversity and abundance in global drylands. Proc Natl Acad Sci USA 112:15684–15689
Mair P (2018) Wilcox, package ‘WRS2 a collection of robust statistical methods’, Version 0.10–0
Manyiwa T, Dikinya O (2014) Impact of tillage types on compaction and physical properties of soils of Sebele farms in Botswana. Plant Soil Environ 33:124–132
Marasco R, Rolli E, Ettoumi B, Vigani G, Mapelli F, Borin S, Abou-Habid AF, El-Behairy UA, Sorlini C, Cherif A, Zocchi G, Daffonchio D (2012) A drought resistance-promoting microbiome is selected by root system under desert farming. PLoS One 7:e48479
Mavi MS, Marschner P (2012) Drying and wetting in saline and saline-sodic soils—effects on microbial activity, biomass and dissolved organic carbon. Plant Soil 355:51–62
Miller AE, Schimel JP, Meixner T, Sickman JO, Melack JM (2005) Episodic rewetting enhances carbon and nitrogen release from chaparral soils. Soil Biol Biochem 37:2195–2204
Mondani F, Khani K, Honarmand SJ, Saedidi M (2019) Evaluating effects of plant growth-promoting rhizobacteria on the radiation use efficiency and yield of soybean (Glycine max) under water deficit stress condition. Agric Water Manage 213:707–713
Moreno-Espíndola IP, Ferrara-Guerrero MJ, Luna-Guido ML, Ramírez-Villanueva DA, De León-Lorenzana AS, Gómez-Acata S, González-Terreros E, Ramírez-Barajas B, Navarro-Noya YE, Sánchez-Rodríguez LM, Fuentes-Ponce M, Macedas-Jímenez JU, Dendooven L (2018) The bacterial community structure and microbial activity in a traditional organic milpa farming system under different soil moisture conditions. Front Microbiol 9:2737
Muhr J, Franke J, Borken W (2010) Drying-rewetting events reduce C and N losses from a Norway spruce forest floor. Soil Biol Biochem 42:1303–1312
Mulvaney MJ, Verhulst N, Herrera JM, Mezzalama M, Govaerts B (2014) Improved wheat performance with seed treatments under dry sowing on permanent raised beds. Field Crop Res 164:189–198
Navarro-Noya YE, Gómez-Acata S, Montoya-Ciriaco N, Rojas-Valdez A, Suárez-Arriaga MC, Valenzuela-Encinas C, Jiménez-Bueno N, Verhulst N, Govaerts B, Dendooven L (2013) Relative impacts of tillage, residue management and crop-rotation on soil bacterial communities in a semi-arid agroecosystem. Soil Biol Biochem 65:86–95
Naylor D, Coleman-Derr D (2018) Drought stress and root-associated bacterial communities. Front Plant Sci 8:2223
Oksanen J, Blanchet FG, Friendly M, Kindt R, Legendre P, McGlinn D, Minchin PR, O'Hara RB, Simpson GL, Solymos P, Stevens MHH, Szoecs E, Wagner H (2018) Vegan package in R. community ecology package. Version:2.5-3
R Development Core Team (2013) R: A language and environment for statistical computing. Version:2.6.2, R Foundation for Statistical Computing, Vienna, Austria. URL http://www.r-project.org/
Ramirez-Villanueva DA, Bello-López JM, Navarro-Noya YE, Luna-Guido M, Verhulst N, Govaerts B, Dendooven L (2015) Bacterial community structure in maize residue amended soil with contrasting management practices. Appl Soil Ecol 90:49–59
Rhoades JD, Manteghi NA, Shouse PJ, Alves WJ (1989) Soil electrical conductivity and soil salinity: new formulations and calibrations. Soil Sci Soc Am J:433–439
Setlow P (2014) Germination of spores of Bacillus species: what we know and do not know. J Bacteriol 196:1297–1305
Stagnari F, Ramazzotti S, Pisante M (2009) Conservation agriculture: a different approach for crop production through sustainable soil and water management: a review. In: Lichtfouse E (ed) Organic farming, pest control and remediation of soil pollutants. Sustain Agr Rev. Springer, Dordrecht, pp 55–83
Thiele-Bruhn S, Bloem J, de Vries FT, Kalbitz K, Wagg C (2012) Linking soil biodiversity and agricultural soil management. Curr Opin Env Sust 4:523–528
Thomas GW (1996) Soil pH and soil acidity. In: Sparks DK (ed) Methods of soil analysis: chemical methods. Part 3. ASA, SSSA, Madison, pp 475–490
Thomson BC, Ostle NJ, McNamara NP, Whiteley AS, Griffiths RI (2010) Effects of sieving, drying and rewetting upon soil bacterial community structure and respiration rates. J Microbiol Meth 83:69–73
Tiessen H, Moir JO (1993) Total and organic carbon. In: Carter MR, Gregorich EG (eds) Soil sampling and methods of analysis, Can Soc Soil Sci, 2nd edn. CRC Press, Boca Raton, pp 187–200
USDA, NRCS (2011) Soil quality indicators. NRCS information sheets https://wwwnrcsusdagov/wps/PA_NRCSConsumption/download?cid=nrcs142p2_053136&ext=pdf Assessed 12 Jan 2019
Verhulst N, Kienle F, Sayre KD, Deckers J, Raes D, Limon-Ortega A, Tijerina-Chavez L, Govaerts B (2011) Soil quality as affected by tillage-residue management in a wheat–maize irrigated bed planting system. Plant Soil 340:453–466
Wang Q, Garrity GM, Tiedje JM, Cole JR (2007) Naïve Bayesian classifier for rapid assignment of rRNA sequences into the new bacterial taxonomy. Appl Environ Microb 73:5261–5267
Wang BZ, Zhang CX, Liu JL, Zen XW, Li FR, Wu YC, Lin XG, Xiong ZQ, Xu J, Jia ZJ (2012) Microbial community changes along a land-use gradient of desert soil origin. Pedosphere 22:593–603
Wang Z, Liu L, Chen Q, Wen X, Lia Y (2016) Conservation tillage increases soil bacterial diversity in the dryland of northern China. Agron Sustain Dev 36:28
Yan N, Marschner P, Cao W, Zuo C, Qin W (2015) Influence of salinity and water content on soil microorganisms. Int Soil Water Conserv Res 3:316–323
Zhang R, Wienhold BJ (2002) The effect of soil moisture on mineral nitrogen, soil electrical conductivity, and pH. Nutr Cycl Agroecosys 63:251–254
Zhao J, Zhang R, Xue C, Xun W, Sun L, Xu Y, Shen Q (2014) Pyrosequencing reveals contrasting soil bacterial diversity and community structure of two main winter wheat cropping systems in China. Microb Ecol 67:443–453
Acknowledgments
We thank “Centro Internacional de Mejoramiento de Maíz y Trigo (CIMMYT)” via CRP WHEAT for allowing us to sample soils from their experimental field in Sonora.
Funding
This research was funded by “Centro de Investigación y de Estudios Avanzados del Instituto Politécnico Nacional (CINVESTAV-IPN-Mexico)” and project “Infraestructura 205945 from Consejo Nacional de Ciencia y Tecnología (CONACyT, Mexico),” and “Centro Internacional de Mejoramiento de Maíz y Trigo (CIMMYT)” via CRP WHEAT. The research forms part of the strategic research for MasAgro Productor, supported by “Secretaría de Agricultura y Desarrollo Rural (SADER).” L.P. received a doctoral grant from CONACyT.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Disclosure of potential conflicts of interest
The authors hereby declare that they have no conflicts of interest.
Research involving human participants and/or animals
No studies with human participants or animals were performed by any of the authors to obtain data for this article.
Informed consent
Not applicable.
Additional information
Responsible editor: Zhihong Xu
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
ESM 1
(PDF 83 kb)
ESM 2
(PDF 52 kb)
ESM 3
(PDF 148 kb)
ESM 4
(PDF 152 kb)
ESM 5
(PDF 92 kb)
ESM 6
(PDF 141 kb)
ESM 7
(PDF 388 kb)
ESM 8
(PDF 77.2 kb)
ESM 9
(PDF 73.6 kb)
ESM 10
(PDF 84.2 kb)
ESM 11
(PDF 69.6 kb)
ESM 12
(PDF 71.4 kb)
ESM 13
(PDF 74 kb)
ESM 14
(PDF 313 kb)
ESM 15
(PDF 77 kb)
ESM 16
(PDF 198 kb)
ESM 17
(PDF 379 kb)
ESM 18
(PDF 73 kb)
Rights and permissions
About this article
Cite this article
Prince, L., Verhulst, N., Govaerts, B. et al. Wet or dry sowing had a larger effect on the soil bacterial community composition than tillage practices in an arid irrigated agro-ecosystem. J Soils Sediments 20, 3316–3329 (2020). https://doi.org/10.1007/s11368-020-02626-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11368-020-02626-y