Abstract
This study aims at the application of a marine fungal consortium (Aspergillus sclerotiorum CRM 348 and Cryptococcus laurentii CRM 707) for the bioremediation of diesel oil-contaminated soil under microcosm conditions. The impact of biostimulation (BS) and/or bioaugmentation (BA) treatments on diesel-oil biodegradation, soil quality, and the structure of the microbial community were studied. The use of the fungal consortium together with nutrients (BA/BS) resulted in a TPH (Total Petroleum Hydrocarbon) degradation 42% higher than that obtained by natural attenuation (NA) within 120 days. For the same period, a 72 to 92% removal of short-chain alkanes (C12 to C19) was obtained by BA/BS, while only 3 to 65% removal was achieved by NA. BA/BS also showed high degradation efficiency of long-chain alkanes (C20 to C24) at 120 days, reaching 90 and 92% of degradation of icosane and heneicosane, respectively. In contrast, an increase in the levels of cyclosiloxanes (characterized as bacterial bioemulsifiers and biosurfactants) was observed in the soil treated by the consortium. Conversely, the NA presented a maximum of 37% of degradation of these alkane fractions. The 5-ringed PAH benzo(a)pyrene, was removed significantly better with the BA/BS treatment than with the NA (48 vs. 38 % of biodegradation, respectively). Metabarcoding analysis revealed that BA/BS caused a decrease in the soil microbial diversity with a concomitant increase in the abundance of specific microbial groups, including hydrocarbon-degrading (bacteria and fungi) and also an enhancement in soil microbial activity. Our results highlight the great potential of this consortium for soil treatment after diesel spills, as well as the relevance of the massive sequencing, enzymatic, microbiological and GC-HRMS analyses for a better understanding of diesel bioremediation.
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
Introduction
Environmental contamination with diesel oil and other petrochemicals endangers different ecosystems (Ugochukwu et al. 2013; Rakowska 2020). Diesel oil, which comprises a mixture of alkanes and aromatic compounds, is recognized worldwide as an important source of contamination by hydrocarbons (MacKinnon and Duncan 2013; Tribelli et al. 2018). Hydrocarbons have been associated with neurological damage, as well as with a higher prevalence of cancer and mutagenicity in petroleum workers (Latif et al. 2010; Johnston et al. 2019). In soil, accidental hydrocarbon spills can alter the physicochemical and biochemical properties, resulting in low fertility, changes in the microbiota, and ecological imbalance (Desforges et al. 2016; da Silva et al. 2020). The risks and negative effects of the transport, production, and commercialization of petroleum and its derivatives have driven the interest in the development of remediation techniques for these organic pollutants (Ugochukwu et al. 2013; Gaur et al. 2021).
Conventional remediation techniques using physical and chemical methods are economically impractical and often generate secondary contamination (Pandey et al. 2009; Simon and Joshi 2021). For this reason, bioremediation has been proposed as an efficient and eco-friendly strategy to mitigate environmental pollution (Tahri et al. 2013; Muthukumar et al. 2022b; Bhatt et al. 2022). Several studies have indicated bioremediation as a promising approach for the restoration of soils contaminated with hydrocarbons (Lu et al. 2012; Shahsavari et al. 2013; Guarino et al. 2017; Nwankwegu and Onwosi 2017; Poi et al. 2017; Kim et al. 2018). However, the possibility of applying bioremediation to treat soils contaminated with diesel oil is strictly related to microbial degradation capability. Therefore, it is important to identify microorganisms that are relevant for the degradation of the target molecules to develop bioremediation strategies (Prathiba 2014).
Several bacteria have been reported to be able to use diesel as the sole carbon source (Shukor et al. 2009; Gao et al. 2021; Vidal-Verdú et al. 2022; Liu et al. 2022), and many fungi can also act as hydrocarbon degraders (Aranda et al. 2017; Park et al. 2019; Asemoloye et al. 2020; Peidro-Guzmán et al. 2021). Nonetheless, the biodegradation of complex hydrocarbons, such as crude and diesel oils, requires the action of different metabolic pathways to achieve its complete mineralization (Garrido-Sanz et al. 2019). Ameen et al. (2016) reported that the degradation of diesel increased when five fungal strains were applied as a consortium, demonstrating the synergism among the species and indicating their ability to prosper when the energy source is limited to complex carbon sources. Similarly, Ebadi et al. (2021) reported that a bacterial consortium was more efficient in the degradation of crude oil than the isolated species, having reached 51.6% of degradation after seven days of cultivation.
According to Abed et al. (2014) and (Ho et al. 2020), the efficiency of bioremediation can be improved using the following methodologies: (i) biostimulation: this approach involves changes in the environmental conditions (e.g., nutrient concentration, pH, moisture content, and oxygen) in order to enhance the degradative potential of native and inoculated microorganisms; (ii) bioaugmentation: this approach implies the inoculation of degrading microorganisms in the environment/contaminated samples.
The effects on the soil microbial communities during biodegradation can help to expand the knowledge on this topic, providing subsidies for the improvement of the bioremediation processes (Sutton et al. 2013; Yadav et al. 2021). Nevertheless, due to the complexity of microbial activities in the soil, their influence on hydrocarbon degradation remains obscure (Kotoky et al., 2018). Thus, the combination of multidisciplinary and innovative approaches is necessary to elucidate and improve the understanding of these complex relationships.
In this context, the following objectives were established in this work: (i) to study the efficiency of a fungal consortium in the bioremediation of soil contaminated with diesel oil under microcosm conditions, (ii) to analyze how the different soil bioremediation approaches change soil quality, and (iii) to evaluate the dynamics of the microbial community during diesel degradation, as well as to assess the potential of the community's function.
Material and methods
Soil Characterization
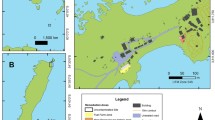
A soil sample collected at ‘Luiz de Queiroz’ College of Agriculture (ESALQ) located at University of São Paulo (Piracicaba, Brazil - coordinates −47° 38′ 57E, −22° 43′ 31S) was used in this study. The soil was classified as a sandy clay loam Ferralsol (Baxter 2007). This type of soil has no records of hydrocarbon contamination. It was collected at a depth of 0-20 cm and has the following characteristics: 201 g Kg-1 of clay, 11 g Kg-1 of silt, and 788 g Kg-1 of sand, density of 1.29 g cm3, 0.8% of total organic carbon. Prior to the experiments, the soil was air-dried and sieved on 4 mm mesh. The initial chemical parameters of the soil are shown in Table S1 (Supplementary material).
Experimental design: microcosms
The experiments were conducted using hermetically sealed glass flasks with a capacity of 1 L. The microcosms were assembled in triplicate with destructive repetitions, totalizing 75 experimental units. A total of 250 g of soil were added to each flask and the field capacity was adjusted to 60% with sterile distilled water. Subsequently, 20 g of diesel per Kg of soil (2%) was added. The diesel had been previously sterilized by filtration through a 0.22 μm membrane.
To assess the degradation of diesel oil, the following strategies were applied:
-
(C) Control: soil without diesel oil.
-
(T1) Natural attenuation: addition of 2% of diesel oil.
-
(T2) Biostimulation: addition of 2% of diesel oil and nutrients (nitrogen and phosphorus sources).
-
(T3) Bioaugmentation/biostimulation: addition of 2% of diesel oil, nutrients (nitrogen and phosphorus sources) and the marine fungal consortium.
-
(T4) Bioaugmentation: addition of 2% of diesel and the marine fungal consortium.
The nitrogen and phosphorus sources added to the biostimulation experiments (T2 and T3) were composed of urea (250 mg Kg-1 of soil) and K2HPO4 (100 mg Kg-1 of soil), respectively.
In this study, we used a microbial consortium composed of two fungi isolated from marine invertebrates: Aspergillus sclerotiorum CRM 348, which produces ligninolytic enzymes (Bonugli-Santos et al. 2010) and is able to degrade pyrene, benzo[a]pyrene, and diesel-oil (Passarini et al. 2011); and the yeast Cryptococcus laurentii CRM 707, which is able to degrade diesel-oil and produce lipases (Duarte et al. 2013) (Table 1). Additional data related to the microbial consortium used in the bioaugmentation treatments (T3 and T4) are shown in Table 1.
For the inoculum preparations: i) the filamentous fungus A. sclerotiorum CRM 348 was cultured in slant tubes with 2% Malt Agar (20 g L-1 of malt extract and 20 g L-1 of agar) for seven days at 28 °C. After this period, the spores (conidia) were removed and transferred to a 50 mL centrifuge tube and left for 12 h in saline solution (0.9% NaCl) in order to deplete the energy reserves; ii) the yeast C. laurentii CRM 707 was cultured in 2% Malt Broth and incubated for 24 h at 28 °C under the agitation of 160 rpm. Subsequently, the cells were centrifuged at 5,000 rpm for 10 min and washed with 20 mL of sterile saline (this procedure was repeated three times). The cells were kept for 12 h in sterile saline in order to deplete the energy reserves. Fungal spores and yeast cells were suspended in sterile saline to a final concentration of 106 CFU spores or cells Kg-1 of soil. These numbers of spores and cells were introduced in the bioaugmentation experiments at the moment of adjusting the field capacity (humidity) to 60%.
The microcosms containing the treatments and controls were incubated in the dark at room temperature and opened weekly to correct humidity with sterile distilled water, aeration, and to analyze microbial respiration. Samples from each of the flasks were collected after 15, 30, 60, 90, and 120 days for microbial and chemical characterization.
Microbial activity in microcosms
Enumeration of degrading microorganisms
The redox indicator method (2,3,5-triphenyltetrazolium) was applied to determine the most probable number (MPN) of microbial degraders using diesel oil as a carbon source. The preparation of the mineral medium with the redox indicator TTC followed the protocol proposed by Meyer et al. (2018). Firstly, the soil samples (1 g) were diluted (serial dilution). Aliquots of 100 μL of the respective dilutions were inoculated into 96-well plates containing 100 μL of the minimum mineral medium (KCl 0.7 g L-1, KH2PO4 2 g L-1, Na2HPO4 3 g L-1, NH4NO3 1 gL-1, MgSO4 4 mg L-1, FeSO4 0.2 mg L-1, MnCl2 0.2 mg L-1, CaCl2 0.2 mg L-1) and 10 μL of diesel oil in each well of the microplates. The plates were incubated at 28 °C for 14 days in the dark. Changes in the color of the medium indicated the presence of microorganisms that use diesel oil as carbon source and the results were compared with the table of MPN, according to the U.S. Food and Drug Administration protocol (U.S. Food and Drug Administration (FDA) 2001).
Enumeration of heterotrophic microorganisms
For the quantification of the heterotrophic microorganisms, the soil samples (1g) were serially diluted and aliquots of 100 μL of the respective dilutions were inoculated into 96-well plates containing 100 μL of BHI medium (brain heart infusion 8 g L-1, peptic digest of animal tissue 5 g L-1, pancreatic digest of casein 16 g L-1, sodium chloride 5 g L-1, glucose 2 g L-1, disodium hydrogen phosphate 2.5 g L-1). The microplates were incubated for 48 h at 28 °C. The turbidity of the medium indicates the presence of microorganisms. The results were compared with the table of most probable number, according to the U.S. Food and Drug Administration protocol (U.S. Food and Drug Administration (FDA) 2001).
Fluorescein diacetate (FDA) hydrolysis
FDA reflects the activity of lipase and esterase enzymes, which are associated with lipid degradation (Schnurer and Rosswall 1982). Fluorescein diacetate (FDA) hydrolysis was assessed according to Schumacher et al. (2015). Two grams of soil (fresh weight) were collected and added to 15 mL of phosphate buffer (60 mM; pH 7.6) and 0.2 mL of fluorescein diacetate solution (1000 μg mL-1). The mixture was incubated at 30 °C for 20 min and 140 rpm. After incubation, 0.2 mL of chloroform was added to the flask. The tubes were centrifuged at 25 °C for 5 min at 10,000 rpm. The supernatant was filtered and the absorbance of the filtrates was measured at 490 nm. Fluorescein concentration was determined applying a calibration curve (0-200 μg mL-1) of fluorescein sodium salt. Soil samples without the FDA solution were used as control.
Diesel oil degradation in soil microcosms
Total petroleum hydrocarbon (TPHs) analysis
Diesel degradation in the different treatments was assessed by gas chromatography according to the EPA methods (EPA 1986; Environmental Protection Agency 2007; US EPA 2007). The hydrocarbon extracted from the soil samples was injected in a gas chromatograph equipped with a flame ionization detector (FID) and capillary column VF-5MS (30 mx 0.53 mm x 1.5 μm). The concentration of the total petroleum hydrocarbons was determined by external standardization. Samples at 120 days of bioremediation were sent to the Company Eurofins (www.eurofins.com.br).
The value of TPH removal (%) was determined by Eq. (1):
Where T0 is the initial concentration of TPH in the soil (the moment of contamination with diesel) and T is the concentration of TPH in soil samples in the different treatments at each specific sampling day.
Non-target screening analysis with a GC-HRMS instrument
In parallel to TPH analysis, soil samples (from T0, T1, and T3) were also extracted for the non-target analysis of individual diesel components. Sample treatment consisted of a straightforward extraction of 3 g of soil with 3 mL of diethyl ether using a Digital sonifier (Branson 250, 30% Amplitude, 30 seconds) followed by centrifugation (5 min at 10.000 g). The supernatant was stored in a glass vial and the process was repeated three times to ensure complete extraction. QC samples were performed by pooling 50 μL of each sample extract. Blank samples were extracted following the same protocol, without the addition of soil samples.
Sample extract was directly injected into a GC-HRMS instrument, consisting of an Agilent 7890B gas chromatograph (Palo Alto, CA, USA) equipped with a PAL RSI autosampler (CTC Analytics AG, Zwingen, Switzerland), coupled to a quadrupole time-of-flight mass spectrometer (Xevo G2-XS QTOF, Waters Corporation, Manchester, UK), operating in APCI mode. Chromatographic separation was performed using a fused-silica GC (DB-5MS) capillary column of 30 m × 0.25 mm i.d. and a film thickness of 0.25 μm (J&W Scientific, Folson, CA, USA). Oven temperature was programmed as follows: from 50 °C, 30 °C/min to 100 °C (2.67 min); 10 °C/min to 300 °C (25 min); total runtime of 25 min. Splitless injections of 1 μL sample extract were performed at 250 °C. Helium was used as carrier gas at 1.4 mL/min. The interface temperature was set to 310 °C and the source temperature, to 150 °C using N2 as an auxiliary gas at 400 L h-1, a make-up gas at 350 mL min-1, and a cone gas at 150 L h-1. The APCI corona pin was operated at 1.6 μA with a cone voltage of 20 V. Xevo G2-XS QTOF MS was operated at a scan time of 0.4 s, acquiring the mass range m/z 50–650 with a resolution of 40,000 (FWHM) at m/z 614 in a data-independent acquisition (MSE) mode, acquiring a low energy full-scan function at 4eV collision energy and a high energy full-scan function using a ramp between 10-40 eV. External calibration was performed with heptacose (PFTBA), while internal calibration was performed every 30 seconds with octafluoronaphthalene. The samples were randomly injected, with a QC sample every 10 injections in order to control instrumental response variations. Blank samples were injected before and after the whole sequence, as well as a linear alkane solution, in order to control retention time variability.
Data treatment for GC-HRMS data
The data generated from the instrument were converted to mzXML using the MSconverter platform tool (ProteoWizard) and processed using MZmine 2.10 for peak detection, peak filtering, chromatogram construction, chromatogram deconvolution, isotopic peak grouping, chromatogram alignment, and gap filling. The following parameters were used for data processing: noise level at 5 x 105 for MS1 and 0 for MS2. The ADAP chromatogram builder was selected with the following parameter: a minimum group size in a number of scans of 4, a minimum height of 1x103, and m/z tolerance of 0.001 Da (or 10 ppm). Chromatogram deconvolution was set as follows: wavelets (ADAP) were used as the algorithm for peak recognition, m/z and RT range for MS2 scan pairing were 0.3 Da and 0.1 min, S/N threshold was 50, minimum feature height was 5 × 105, coefficient/area threshold was 90, peak duration range was 0.02–1.5 min, and RT wavelet range was 0.02–0.05. The chromatograms were then de-isotoped by isotopic peaks using a grouper algorithm with an m/z tolerance of 0.001 Da and an RT tolerance of 0.05 min. Peak alignment was carried out using a join aligner, with an m/z tolerance set at 0.001 Da, absolute RT tolerance at 0.05 min, and weight for m/z and RT at 30. The missing peaklist after the alignment was filled by gap filling of the same RT and m/z range gap filler module with an m/z tolerance of 0.001 Da. This resulted in a peaklist of 8350 features. Both peaklists were then exported to excel and used as input for MVA.
In addition, linear alkanes, as well as the selected PAHs and cyclosiloxanes, were searched in a targeted way to observe variations along the different treatments directly in joined tables of GC exported data, ensuring their identity by manually checking the spectral similarity with reference standards, accurate mass measurements for parent and fragment ions and kovats index (Table S3).
The total peak area of the alkanes, PAHs, and cyclosiloxanes detected was defined as the concentration of these compounds. The removal rate was calculated using eq. (2):
Where T0 refers to the total area of the peaks measured in the soil at the beginning of the experiment and T refers to the total area of the peaks measured in the soil samples in each treatment and at the specific sampling time.
Statistical analyses
All experiments were conducted in triplicate and the means and standard deviations (N-1) were calculated. The results were submitted to the Tukey’s means test using the statistical program (Minitab 17).
Illumina sequencing and diversity analyses
For the microbial community assessment, 0.5 g of microcosm soil (from the treatment with the best degradation response, natural attenuation, and controls) was removed at times 0, 30, 60 and 120 days. The DNeasy PowerSoil kit (Qiagen) was used for DNA extraction, and the extraction process was performed according to the manufacturer's instructions.
The DNA extracted from the samples was used in PCR reactions to amplify: the regions V3 and V4 of the 16S subunit of the bacterium rDNA, using primers 341F and 785 R (Klindworth et al. 2013); and the ITS region for fungal analysis, using primers ITS1F (Gardes and Bruns, 1993) and ITS2 (White et al. 1990). The reaction tubes contained 4 μL of Phusion™ High–Fidelity DNA Polymerase buffer, dNTPS (10 mM), 5 μL of each primer (1 pmol μL-1), and 2.5 μL of DNA (15 ng μL-1). The quality and size of the bands were verified by agarose gel electrophoresis. The PCR products were purified using the PureLink PCR purification Kit (Invitrogen®) and the triplicates were pooled in equal proportions. The pooled product was used to prepare the Illumina DNA library, and then the paired-end (2×300 bp) sequencing was performed by Macrogen Inc. (Seoul, Korea) using the MiSeqTM platform (Illumina, San Diego, USA).
The sequences obtained as described above were processed using QIIME (Quantitative Insights Into Microbial Ecology) (Caporaso et al. 2010) following a modified version of the protocols described in the Brazilian Microbiome Project (Pylro et al. 2014). All sequences were filtered for quality (Q25) and demultiplexed. The sequences were then dereplicated, filtered for chimeras and singletons, and clustered into 97% similarity OTUs using vsearch. The taxonomic affiliation was performed within QIIME using the algorithm uclust. A representative sequence from each OTU was classified according to the database Silva v.132 for 16S rRNA analysis and the database UNITE v8.0 for ITS sequences.
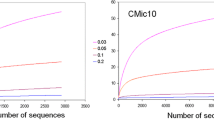
Subsequently, the 16S and ITS data were incorporated in the R environment using the phyloseq package, in which the data were rarefied to an even depth. The normalized data were used to quantify the alpha-diversity indexes on the Vegan package. The ANOVA test followed by Tukey HSD were performed on the agricolae package. The principal coordinates analysis was performed through Bray-Curtis distance matrixes in phyloseq. The differences among the communities were evaluated using PERMANOVA from the vegan package (Oksanen 2010). Taxa abundance plots were built by selecting the 10 most abundant taxa, which accounted for 42-82% of the bacterial community and 35-90% of the fungal community, while the other 10 genera represent less than 5% of these communities. The selected taxa were then normalized based on their relative abundance and plotted using the ggplot2 package. Heatmaps were constructed based on the abundance of the 100 most abundant OTUs grouped at family level using heatmap.2 function of the gplots package. Data is available at the MG-RAST server under the project number mgp92922.
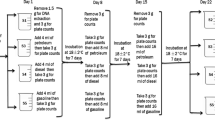
A summary of the methodology used in this study is shown in Fig. 1.
Results and Discussion
Diesel oil degradation (TPH and GC-HRMS analysis)
Results from the TPH analyses of the biostimulation (T2) and bioaugmentation (T4) strategies after 120 days of microcosm incubation did not show significant differences in relation to natural attenuation (3% and 10%, respectively). Nevertheless, 42% of the TPHs were depleted upon the combined application of the bioaugmentation/biostimulation strategies (T3) (Fig. 2). These results indicate the occurrence of a synergistic effect between nutrient amendments and consortium inoculation, ultimately resulting in higher diesel oil degradation. Since biostimulation alone was not effective at removing TPHs, the inoculated microorganisms appear to be decisive to increase the rate of hydrocarbon degradation. Although nutrients are considered relevant for the performance of microbial bioremediation (Roy et al. 2018), the results from the present study suggest that the fungal consortium should be genetically adapted to the microcosm conditions and could remain in this environment. Seklemova et al. (2001) and Bento et al. (2005) have found similar results on diesel oil degradation employing biostimulation. The authors have reported that after the addition of nutrients (nitrogen and phosphorus), no positive effects on the autochthonous microbiota were identified, which is in accordance with our findings. A possible explanation relates to the fact that the indigenous microbiota in these soils could not be naturally adapted to hydrocarbon degradation. This represents an important finding since accidental diesel spills are quite common in areas that have not been previously contaminated.
Total petroleum hydrocarbons (TPH) after 120 days of incubation in contaminated soil at room temperature, employing the different bioremediation strategies. (T1) Natural attenuation; (T2) Biostimulation; (T3) Biostimulation/Bioaugmentation; (T4) Bioaugmentation. The means followed by different letters indicate a significant difference among the treatments according to the Tukey's test (p <0.05) and the bars represent the standard deviation
In order to observe the most reduced hydrocarbon species, an analysis was performed using a GC-HRMS instrument. The results obtained from these data are consistent with TPH results, indicating a low individual degradation of linear alkanes (from dodecane to pentacosane) in natural attenuation (T1). In the case of short-chain alkanes (from C12 to C19) at 120 days, bioaugmentation/biostimulation showed a reduction of 72 to 92%, while treatment T1 removed 3 to 65% of these same compounds (Table 2). Possibly, the presence of short-chain alkanes was reduced in the soil of T1 over the 120 days due to their higher volatility (compared to the longer ones) (Koshlaf and Ball 2017), and not by the action of the autochthonous microbiota.
Bioaugmentation/biostimulation (T3) also showed a higher efficiency of long-chain alkane degradation (from C20 to C24), reaching 90 to 92% of icosane and heneicosane degradation, respectively. On the other hand, natural attenuation showed a maximum of 37% of degradation for these alkane fractions. Pentacosane was the alkane with the lowest rate of degradation by treatment T3, with no significant difference in relation to treatment T1 (Table 2). This possibly derives from its reduced bioavailability for the microorganisms caused by a lower solubility, considering that the solubility of alkanes decreases as the molecular weight increases (Rojo 2009).
In general terms, treatment T3 promoted a removal rate similar to that found by natural attenuation for the 4-ringed PAHs benzo(a)anthracene and pyrene (from 15 to 38%, respectively). For anthracene, a low molecular weight PAH with lower recalcitrance, natural attenuation showed a higher removal efficiency (about 92% of degradation), while treatment T3 degraded approximately 72% of this compound. Conversely, the 5-ringed PAH benzo(a)pyrene, which is considered the most toxic and recalcitrant PAH (Kuppusamy et al. 2017; Nzila and Musa 2020), was significantly more removed in treatment T3 in comparison to natural attenuation (48 vs. 38 % of biodegradation, respectively).
Non-target screening analysis with GC-HRMS
Apart from alkanes and PAHs, which were investigated in a targeted way, cyclosiloxanes were revealed as biostimulation/biodegradation markers (T3) after an untargeted approach (Table 3). It is known that cyclosiloxanes are added to diesel fuel (Amais et al. 2013) and used in petroleum refining industries as anti-foaming agents (Daâssi et al. 2021; Liu et al. 2021). In fact, the presence of these compounds in T0 samples (the moment of contamination with diesel) showed levels around 120% higher in comparison to the blank samples. However, after 120 days, the levels of D4 to D8 species increased significantly from 216 to 866% in treatment T3, with a constant increase in 30 days (128-314%) and 60 days (193-764%). The increase in cyclosiloxanes in T3 could be the result of microbial metabolism. Although little reported in the literature, some authors have identified a microbial production of cyclosiloxanes. Interestingly, Birdilla Selva Donio et al. (2018) reported a bioemulsifier produced by the halotolerant bacterium Haererehalobacter sp. JS1, which, after GC-MS analysis, was characterized as belonging to a siloxane-type compound (tetradecamethylcycloheptasiloxane, hexadecamethylcyclooctasiloxane, cyclononasiloxane, octadecamethyl, eicosamethylcyclodecasiloxane, cyclodecasiloxane, eicosamethyl and 1H,15H-hexadecamethyloctasiloxane). Additionally, Kumar et al. (2021) identified a new cyclic siloxane biosurfactant produced by the rhizobacterium Bacillus cereus BS14.
Biosurfactants are compounds that reduce the surface tension between two immiscible liquids, while an emulsifier helps the dispersion of droplets from one immiscible liquid into another and prevents coalescence (Ferreira et al. 2020). Although there are differences between surfactants and bioemulsifiers, both have properties that allow them to be used for the biodegradation of hydrocarbons, since these molecules play a specific role of binding tightly to dispersed hydrocarbons and oils, preventing them from merging together (Uzoigwe et al. 2015). Thus, the biodegradation of petroleum hydrocarbons can be significantly improved by increasing the solubility and bioavailability of these compounds to microorganisms.
Microbial activity in microcosms
Considering the MPN for heterotrophic and degraders in the different bioremediation strategies over time, small differences in the growth rate were found (Figs. 3A and B). In general, the number of microorganisms in the soil increased in the treatments in which nutrients were added (T2 and T3). In the case of heterotrophic microorganisms, the values obtained for biostimulation (T2) ranged from 5.4 to 6.0 log MPN g-1, whereas for bioaugmentation/biostimulation (T3) the values were between 5.1 and 5.2 log MPN g-1. For diesel-degrading microorganisms, the values for treatment T2 were between 5.1 and 5.2 log MPN g-1, while for treatment T3, the values remained at 4.8 log MPN g-1. In the control experiments, the lowest number of microorganisms were found: between 4.0 and 4.6 log MPN g-1 for the heterotrophic microorganisms, and 3.6-3.9 log MPN g-1 for the degraders.
Count of the heterotrophic microorganisms (a) and diesel degraders (b) by the MPN over time employing the different diesel oil bioremediation strategies. (C) Control; (T1) Natural attenuation; (T2) Biostimulation; (T3) Bioaugmentation/Biostimulation; (T4) Bioaugmentation. The means followed by different letters indicate a significant among the between treatments according to the Tukey's test (p <0.05) and the bars represent the standard deviation
In general, during the 120 days of the bioremediation, an increase in the soil microbial community abundance was not evidenced. Nonetheless, for the treatments with biostimulation (T2) and bioaugmentation/biostimulation (T3), an enhanced microbial activity was found during the period of microcosm incubation (Fig. 4). According to Miethe et al. (1994), the microbial counts of an environment do not always correlate with biological activity, because many microorganisms may be dormant or uncultured under laboratory conditions. Additionally, pollutant degradation is not always necessarily related to microbial growth rates (Montgomery et al. 2010).
FDA activity over time employing the different diesel oil bioremediation strategies. (C) Control; (T1) Natural attenuation; (T2) Biostimulation; (T3) Bioaugmentation/Biostimulation; (T4) Bioaugmentation. The means followed by different letters indicate a significant difference among the treatments according to the Tukey's test (p <0.05) and the bars represent the standard deviation
The positive correlation between FDA and microbial biomass has been used to indicate microbial activity in soils. In addition, FDA reflects the activity of lipase and esterase enzymes, which are associated with lipid degradation (Schnurer and Rosswall 1982). In the treatment in which the microorganisms and nutrients were added (bioaugmentation/biostimulation, T3) (Fig. 4, times 15, 30, and 60 days), a significant difference in the activity of these enzymes in relation to the other treatments was observed (0.28; 0.23; and 0.33 μg of fluorescein/g-1 of soil, respectively). Additionally, an increase was observed in these enzymatic activities over time, and after 90 days of incubation, the biostimulation (T2) treatment reached enzymatic values close to the bioaugmentation/biostimulation (T3) treatment (0.5 μg of fluorescein g-1 of soil). These data suggest that biostimulation may have caused an increase in the activity of the indigenous degrading microbiota, although this was not reflected in terms of the degradation of TPHs (Fig. 2), alkanes and PAHs (Table 2). The lack of hydrocarbon degradation in the biostimulation treatment could be related to the delay in the natural selection of microorganisms with greater degradation capacity in relation to the treatment in which the consortium was added, involving higher FDA values at the beginning of the process. Yuan et al. (2018) reported a higher FDA activity in the treatment with a microbial consortium, along with the degradation rate of crude oil in culture conditions. Martínez Álvarez et al. (2017) also found enhanced FDA activity in biostimulated soils (nitrogen and phosphorus) in comparison to the control.
An increase in microbial activity in the biostimulation (T2) and bioaugmentation/biostimulation (T3) treatments was observed over time, whereas in natural attenuation (T1) and bioaugmentation (T4) the overall microbial activity in the soils remained low during the 120 days. Although the microbial activity of the biostimulated soil (T2) showed high microbial activity, hydrocarbon degradation did not occur. Therefore, it is reasonable to believe that the autochthonous microbiota of the soil, even with biostimulation and enhanced microbial activity, did not present a metabolic support for hydrocarbon biodegradation.
These results highlighted the efficiency of the consortium composed of two marine-derived fungi (A. sclerotiorum and C. laurentii) in degrading diesel oil from contaminated soils. In the study reported by Huarte-Bonnet et al. (2018), the expression of genes responsible for aromatic hydrocarbon oxidation (cytochrome P450 - CYP53 family) was significantly induced when Aspergillus niger was cultured in the presence of phenanthrene. Filamentous fungi from the genus Aspergillus have high proportions of CYP genes in their genomes, a phenomenon that may be related to new physiological processes (Chen et al. 2014). Additionally, Schwarz et al. (2018) reported that representatives of Basidiomycota were dominant in the fungal community obtained from phenanthrene-contaminated soil, including the genus Cryptococcus. Members of this yeast genus have also been reported as biosurfactants (Kamyabi et al., 2017) and lipase producers (Aarthy et al., 2018), and pyrene degraders (Kamyabi et al. 2017).
Diversity and composition of the fungal and bacterial communities
All samples were properly assessed, as evidenced by the good coverage indices obtained, which were higher than 99% (Table S2). Table S2 presents the microbial richness expressed by the Chao1 index. Both the T1 and T3 treatments revealed changes in fungal/bacterial richness. A decreasing trend in fungal richness can be observed over the period of bioremediation for treatments T1 and T3, being more pronounced in treatment T3. Bacterial richness increased in treatment T1 and decreased in treatment T3 in comparison to the control.
The results of the Shannon and Simpson indices suggest that microbial diversity decreased in response to the addition of diesel in the soil (Table S2). Bell et al. (2013), Jung et al. (2016) and Sutton et al. (2013) reported similar results. Apparently, the bioaugmentation/biostimulation (T3) treatment inhibited some autochthonous fungal and bacterial microbiota over time or promoted the propagation of specific groups of microorganisms, making them more pronouncedly represented in all treatments.
In the first 30 days of bioremediation there was a drop in bacterial diversity, mainly in treatment T3. The analyses at 60 and 120 days revealed that diversity increased, but did not reach the initial values. In natural attenuation (T1), a lower fungal and bacterial diversity occurred throughout the bioremediation period. However, a lower diversity of both fungi and bacteria was observed in treatment T3 (bioaugmentation/biostimulation), concomitantly with the greater diesel degradation rate. These results may indicate an increase in the populations responsible for hydrocarbon metabolism during soil incubation, which could have promoted an inhibition in the native microbiota. Although hydrocarbons provide carbon sources for many microorganisms, they may be toxic to others and their use requires specific adaptations of the microbiota (Bell et al. 2013).
Data from this study indicate that the degradation of hydrocarbons by the soil microbial community may not be directly connected to higher richness and diversity, but rather to the selection of specific groups of degrading microorganisms. Additionally, the toxicity of diesel oil and the metabolites derived from the microbial degradation could have affected microbial community composition, causing a decrease in richness and diversity.
PCoA analyses clearly showed the separation and variation in the fungal community composition in the different treatments, explaining 95% of the total variability (Fig. 5A). It was possible to verify that samples within the same treatment were clustered together, showing the role of bioaugmentation/biostimulation (T3) in the soil fungal community structures. Regarding the bacterial community (Fig. 5B), there were contrasting separations of diversity, mainly in treatment T3 in relation to the control and natural attenuation (T1), explaining a total of 76.4% of the variability. The PCoA plot also shows different trajectories of the bacterial community in different treatments over time. In treatment T3, the abundance of each group varied according to the successional time. The data from the PCoA analyses indicate that the diesel oil degradation processes were performed by distinct fungal/bacterial groups, which may have had a determining role in the bioremediation process.
Principal coordinate analysis (PCoA) based on the profiles of the fungal (a) and bacterial (b) communities found in the soils under the different bioremediation treatments at the genus level. (C) Control; (T1) Natural attenuation and (T3) Bioaugmentation/Biostimulation. The values in the axis indicate the percentage of variance explained by the principal coordinate
Figure 6A shows the heatmap comparing the influence of the different bioremediation treatments (T1 and T3) in relation to the control, considering the different incubation times of the 100 most abundant fungal OTUs at the family level. Based on this data, two distinct clusters appeared: the first one composed of soils from the different incubation times of treatment T3, and the second composed of natural attenuation (T1) and control (C), although the control soil was distinctly segregated from treatment T1. The grouping pattern of the different treatments was consistent with PCoA (Figs. 5A and B), indicating that the soils from treatment T3 presented a fungal community considerably distinct from the other treatments. The fungal community was drastically affected by diesel addition. In the control, we can see a high abundance of fungi from the families Pleosporaceae and Psathyrellaceae, in addition to unidentified families. In natural attenuation (T1), there was an increase in the families Chaetomiaceae, Hypocreaceae, and Nectriaceae. Regarding the bioaugmentation/biostimulation (T3) treatment, there was a pronounced increase in the families Trichocomaceae and Microascaceae (Fig. 7). The increase in the family Trichocomaceae was already expected, since the spores of the fungus A. sclerotiorum CRM 348 were inoculated in this treatment.
Hierarchically clustered heatmap of the distribution of the top 100 fungal OTUs from the different communities at the family level (a) and bacterial OTUs at the class level (b). (C) Control; (T1) Natural attenuation and (T3) Bioaugmentation/Biostimulation. The abundance z-score of each OTU is color-coded according to the legend above the heatmaps
In the heatmap of the bacteria (Fig. 6B), the separation of the community into two clusters was also observed: the first group was composed of treatment T3 (times 30 and 60 days), and the second was composed of the treatments C and T1 (30, 60, and 120 days) and T3 (120 days), although T3 (120 days) was separated from the others, indicating less similarity. The control soil (C) presented a high abundance of Bacilli, Verrucomicrobiae, and subgroups 5 and 25. These bacterial classes decreased in both treatments T1 and T3. In the natural attenuation (T1 30 days), there was an increase in the representatives of the classes Negativicutes, Methanomicrobia, Bacteriodia, Alphaproteobacteria, and Gammaproteobacteria in relation to the control. On the 60th day of incubation, the most abundant taxonomic groups were Chlamydiae, Leptospirae, Methanomicrobia, and Acidobacteria, very differently from the 120th day of incubation, which showed a greater abundance of Saccharimonadia, Anaerolineae, Blastocatellia, and Leptospirae. In treatment T3, there was a reduction in Anaerolineae, Bacilli, Verrucomicrobiae, Blastocatellia, Bacterioidia, and subgroups 5 and 25 on the 30th day, whereas an increase in Negativicutes, Clostridia, and Gammaproteobacteria occurred. On the 60th day of incubation, there was an increase in representatives of the classes Clostridia, Alphaproteobacteria, Deltaproteobacteria, Actinobacteria, and interestingly, a decrease in Gammaproteobacteria. On the 120th day of incubation, the bacterial community of treatment T3 markedly differed in comparison to the 30th and 60th days. Deltaproteobacteria and Actinobacteria were the most abundant classes, whereas Gammaproteobacteria continued to decline. Furthermore, there was an increase, although less pronounced, in the classes Anaerolineae, Acidobacteria, Methanomicrobia, and Alphaproteobacteria. It is evident that treatment T3 stimulated the dominance of certain classes, mainly Deltaproteobacteria, Gammaproteobacteria, and Actinobacteria. Although Gammaproteobacteria was the most dominant class at the beginning of the bioremediation process, lower proportions of that class were verified over time, accompanied by higher proportions of Actinobacteria and Deltaproteobacteria. These data corroborate the findings of a previous study reported by Yergeau et al. (2012), in which an increase in Actinobacteria and lower proportions of Gammaproteobacteria during bioremediation was verified. Several families of hydrocarbon-degradation enzymes, including alkane monooxygenases, ring-hydroxylating dioxygenases, and cytochrome P450 oxidase, are distributed in Proteobacteria and Actinobacteria (Fuentes et al. 2014).
The patterns of fungal and bacterial abundance at the genus level (10 most abundant) during bioremediation were also investigated (Figs. 7A and B). These genera represent between 42-82% of the bacterial community and 35-90% of the fungal community, while the other 10 genera Srepresent less than 5% of these communities. The fungal community of the control soil (C) had a greater abundance of unidentified fungi (65%), followed by representatives of the genus Gibberella (18.3%), while in natural attenuation (T1) the representatives of the genus Fusarium were the most abundant (53 to 62%), followed by unidentified fungi (19 to 33%) at all incubation times. The fungal community present in the bioaugmentation/biostimulation (T3) treatment differed considerably from the initial soil community, as well as from the fungal community of natural attenuation (T1). The abundance of several groups was negatively affected (Giberella, Humicola, Fusarium, Penicillium, and Bionectria) in response to the treatment, in addition to an evident decrease in fungal diversity, probably due to the induction in the growth of other fungal and bacterial species. In treatment T3, an increase in representatives of the genus Pseudollescheria (71.6 to 90%), Scedosporium (5 to 13.3%), and Aspergillus (4.1%) was observed in relation to the control (C) and natural attenuation (T1). It is important to highlight that Scedosporium is the anamorphic state of the genus Pseudallescheria (Gilgado et al. 2008). These results suggest that the fungal inoculum remained in the soil, although over time fungi from the genus Aspergillus decreased in abundance, reaching close to zero on the 120th day of incubation (Fig. 7A). This data corroborates those presented in Fig. 6A, which showed a greater abundance of the classes Trichocomaceae and Microascaceae in treatment T3. Greco et al. (2019) reported a significantly higher degradation of polycyclic aromatic hydrocarbons in an experiment with the addition of a fungal consortium composed of Pseudallescheria boydii, Talaromyces amestolkiae, and Sordaria fimicola. In the study conducted by Morales et al. (2017), the complete genome of one representative of Scedosporium apiospermum was characterized and a group of gene families related to hydrocarbon degradation was found (e.g. dioxygenases and cytochrome P450). Zafra et al. (2016) used a functional metagenomic approach to investigate the effects of bioaugmentation (with a microbial consortium) and biostimulation on the bioremediation of hydrocarbon-contaminated soil. The authors reported changes in the abundance of aromatic genes, including those related to dioxygenases, laccases, protocatechuate, salicylate, and benzoate-degrading enzymes.
Figure 7B illustrates the relative abundance of the bacterial community at the genus level for each treatment at different incubation times. Diesel seems to affect the general diversity and the dominance of certain phyla, such as Proteobacteria, which was found in higher abundance after the application of the pollutant. In the control soil (C), the most abundant organisms were uncultured bacteria (70%), followed by Bacillus (18.3%) and Sphingomonas (8.3%). Soil bacterial community during natural attenuation (T1) differed significantly from the control, with less abundance of uncultured bacterium and Bacillus. In addition, after the exposure of the soil to diesel, representatives of the genera Acinetobacter, Pseudomonas, and Cupriavidus showed a notable increase. In the bioaugmentation/biostimulation (T3) treatment, the bacterial community in the first 30 days of bioremediation was dominated by representatives of the genera Pseudomonas (46.6%) and Acinetobacter (40%). In contrast, the abundance of Bacillus, Sphingomonas, and uncultured bacterium decreased dramatically in this period in comparison to the control soil (C). These results indicate that bacteria of these genera may have a relevant function in the degradation of diesel. The genus Pseudomonas has been largely studied and proven to be efficient in the degradation of hydrocarbons. Previous studies have reported the presence of the monooxygenase genes alkB (Muriel-Millán et al., 2019) and almA (Muriel-Millán et al. 2019), as well as surfactant production (Muriel-Millán et al. 2019; Elumalai et al. 2021; Muthukumar et al. 2022a) in the genus Pseudomonas. Roy et al. (2018) used bioaugmentation/biostimulation for the degradation of hydrocarbons from the oil of refinery sludge and reported an increase in the population of Pseudomonas accompanied by 75% of TPH degradation in 120 days of bioremediation. Similarly, members of the genus Acinetobacter are often regarded as hydrocarbon degraders (Tiralerdpanich et al. 2018; Elumalai et al. 2021; Parthipan et al. 2022) and as carriers of the gene almA (Throne-Holst et al. 2007).
After 60 days of bioremediation, although Pseudomonas (30%) and Acinetobacter (25%) were still the most abundant groups in treatment T3, their abundance decreased in comparison to the 30th day. Concomitantly, there was an increase in the abundance of Caulobacter (16.7%), Rhodococcus (11.6%), and Anaeromyxobacter (3.6%). The bacterial population of the soil under the bioaugmentation/biostimulation (T3) treatment changed dramatically over the incubation time (120 days). This soil showed a marked decrease in the genera Pseudomonas (1.6%) and Acinetobacter (1.6%). Similarly, Yergeau and collaborators (2012) showed that the genus Pseudomonas was predominant in a diesel-contaminated soil during the initial bioremediation period, but representatives of this genus decreased when the residual soil hydrocarbons were depleted. In the present study, the decrease in the abundance of Pseudomonas was associated with an enrichment of representatives of the genera Rhodococcus (31.6%), Caulobacter (25%), and Anaeromyxobacter (13.3%). These data corroborate the information shown in Fig. 7B, in which an increase in the groups Proteobacteria and Actinobacteria in treatment T3 was observed. The role of Rhodococcus in hydrocarbon degradation is well known. In the study reported by Gielnik et al. (2019), representatives of the genus Rhodococcus were dominant in the bacterial community associated with the highest diesel degradation rate. The increase in the population of this bacterium over the bioremediation treatment may be related to the particular genetic characteristics of this genus, since most strains have the gene alkB (Brzeszcz and Kaszycki 2018), which encodes the enzyme alkane hydroxylase, responsible for the degradation of linear and branched alkanes. Many of these strains have multiple copies of the gene alkB (Orro et al. 2015) or homologous genes in their genomes (Nie et al. 2014), which can effectively enhance the range of alkane degradation (Amouric et al. 2010), resulting in high bioremediation efficiency.
Conclusions
Results from this study indicate that the marine fungal consortium played a decisive role in the degradation of diesel oil in the contaminated soil. The bioaugmentation/biostimulation strategy caused a decrease in the total soil microbial diversity and an increase in the abundance of specific microbial groups. The changes in the structure of the fungal and bacterial communities can explain the reduction in TPHs, PAHs, and alkanes in the bioaugmentation/biostimulation treatment. In addition, the higher presence of cyclosiloxanes in the bioaugmentation/biostimulation treatment may be related to their biosynthesis by the microbial community, acting as a biosurfactant/bioemulsifier and enhancing the bioavailability of hydrocarbons. Furthermore, the use of a combined approach (massive sequencing, enzymatic activity, microbiological analyses, and advanced methods like GC-HRMS) reinforces the importance of a polyphasic study to improve the understanding of organic pollutant mitigation in soils, providing a better overview of this complex system.
Data Availability
There is no additional data.
References
Aarthy M, Puhazhselvan P, Aparna R et al (2018) Growth associated degradation of aliphatic-aromatic copolyesters by Cryptococcus sp. MTCC 5455. Polym Degrad Stab 152:20–28. https://doi.org/10.1016/j.polymdegradstab.2018.03.021
Abed RMM, Al-Sabahi J, Al-Maqrashi F et al (2014) Characterization of hydrocarbon-degrading bacteria isolated from oil-contaminated sediments in the Sultanate of Oman and evaluation of bioaugmentation and biostimulation approaches in microcosm experiments. Int Biodeterior Biodegrad 89:58–66. https://doi.org/10.1016/j.ibiod.2014.01.006
Amais RS, Donati GL, Schiavo D, Nóbrega JA (2013) A simple dilute-and-shoot procedure for determination in diesel and biodiesel by microwave-induced plasma optical emission spectrometry. Microchem J 106:318–322. https://doi.org/10.1016/J.MICROC.2012.09.001
Ameen F, Moslem M, Hadi S, Al-Sabri AE (2016) Biodegradation of diesel fuel hydrocarbons by mangrove fungi from Red Sea Coast of Saudi Arabia. Saudi J Biol Sci 23:211–218. https://doi.org/10.1016/j.sjbs.2015.04.005
Amouric A, Quéméneur M, Grossi V et al (2010) Identification of different alkane hydroxylase systems in Rhodococcus ruber strain SP2B, an hexane-degrading actinomycete. J Appl Microbiol 108:1903–1916. https://doi.org/10.1111/j.1365-2672.2009.04592.x
Aranda E, Godoy P, Reina R et al (2017) Isolation of Ascomycota fungi with capability to transform PAHs: Insights into the biodegradation mechanisms of Penicillium oxalicum. Int Biodeterior Biodegrad 122:141–150. https://doi.org/10.1016/j.ibiod.2017.05.015
Asemoloye MD, Tosi S, Daccò C et al (2020) Hydrocarbon Degradation and Enzyme Activities of Aspergillus oryzae and Mucor irregularis Isolated from Nigerian Crude Oil-Polluted Sites. Microorganisms 8:1–19. https://doi.org/10.3390/MICROORGANISMS8121912
Baxter S (2007) World Reference Base for Soil Resources. World Soil Resources Report 103 . Rome: Food and Agriculture Organization of the United Nations (2006), pp. 132, US$22.00 (paperback). ISBN 92-5-10511-4
Bell TH, Yergeau E, Maynard C et al (2013) Predictable bacterial composition and hydrocarbon degradation in Arctic soils following diesel and nutrient disturbance. ISME J 7:1200–1210. https://doi.org/10.1038/ismej.2013.1
Bento FM, Camargo FAO, Okeke BC, Frankenberger WT (2005) Comparative bioremediation of soils contaminated with diesel oil by natural attenuation, biostimulation and bioaugmentation. Bioresour Technol 96:1049–1055. https://doi.org/10.1016/j.biortech.2004.09.008
Bhatt P, Bhandari G, Bhatt K, Simsek H (2022) Microalgae-based removal of pollutants from wastewaters: Occurrence, toxicity and circular economy. Chemosphere 306:135576. https://doi.org/10.1016/J.CHEMOSPHERE.2022.135576
Birdilla Selva Donio M, Chelladurai Karthikeyan S, Michaelbabu M et al (2018) Haererehalobacter sp. JS1, a bioemulsifier producing halophilic bacterium isolated from Indian solar salt works. J Basic Microbiol 58:597–608. https://doi.org/10.1002/JOBM.201800056
Bonugli-Santos RC, Durrant LR, da Silva M, Sette LD (2010) Production of laccase, manganese peroxidase and lignin peroxidase by Brazilian marine-derived fungi. Enzym Microb Technol 46:32–37. https://doi.org/10.1016/j.enzmictec.2009.07.014
Brzeszcz J, Kaszycki P (2018) Aerobic bacteria degrading both n-alkanes and aromatic hydrocarbons: an undervalued strategy for metabolic diversity and flexibility. Biodegradation 29:359–407
Caporaso JG, Kuczynski J, Stombaugh J et al (2010) correspondence QIIME allows analysis of high- throughput community sequencing data Intensity normalization improves color calling in SOLiD sequencing. Nat Publ Group 7:335–336. https://doi.org/10.1038/nmeth0510-335
Chen W, Lee MK, Jefcoate C et al (2014) Fungal cytochrome P450 monooxygenases: Their distribution, structure, functions, family expansion, and evolutionary origin. Genome Biol Evol 6:1620–1634. https://doi.org/10.1093/gbe/evu132
da Silva IGS, de Almeida FCG, da Rocha e Silva NMP et al (2020) Soil bioremediation: Overview of technologies and trends. Energies 13
Daâssi D, Nasraoui-Hajaji A, Bawasir S et al (2021) Biodegradation of C20 carbon clusters from Diesel Fuel by Coriolopsis gallica: optimization, metabolic pathway, phytotoxicity. 3 Biotech 11:1–21. https://doi.org/10.1007/S13205-021-02769-W/FIGURES/5
Desforges JPW, Sonne C, Levin M et al (2016) Immunotoxic effects of environmental pollutants in marine mammals. Environ Int 86:126–139
Duarte AWF, Dayo-Owoyemi I, Nobre FS et al (2013) Taxonomic assessment and enzymes production by yeasts isolated from marine and terrestrial Antarctic samples. Extremophiles 17:1023–1035. https://doi.org/10.1007/s00792-013-0584-y
Ebadi A, Ghavidel A, Khoshkholgh Sima NA et al (2021) New strategy to increase oil biodegradation efficiency by selecting isolates with diverse functionality and no antagonistic interactions for bacterial consortia. J Environ Chem Eng 9:106315. https://doi.org/10.1016/J.JECE.2021.106315
Elumalai P, Parthipan P, Huang M et al (2021) Enhanced biodegradation of hydrophobic organic pollutants by the bacterial consortium: Impact of enzymes and biosurfactants. Environ Pollut 289. https://doi.org/10.1016/j.envpol.2021.117956
Environmental Protection Agency (2007) Ultrasonic Extraction. EPA Method 3550C:1–17
EPA (1986) EPA Method 8100: PAH analysis. https://www.epa.gov/sites/production/files/2015-12/documents/8310.pdf. Accessed Jun 2023
Ferreira INS, Rodríguez DM, Campos-Takaki GM, Andrade RF da S (2020) Biosurfactant and bioemulsifier as promising molecules produced by Mucor hiemalis isolated from Caatinga soil. Electron J Biotechnol 47:51–58. https://doi.org/10.1016/J.EJBT.2020.06.006
Fuentes S, Méndez V, Aguila P, Seeger M (2014) Bioremediation of petroleum hydrocarbons: Catabolic genes, microbial communities, and applications. Appl Microbiol Biotechnol 98:4781–4794
Gao J, Ming J, Xu M et al (2021) Isolation and characterization of a high-efficiency marine diesel oil-degrading bacterium. Pet Sci 18:641–653. https://doi.org/10.1007/S12182-020-00540-Z/TABLES/7
Gardes M, Bruns TD (1993) ITS primers with enhanced specificity for basidiomycetes - application to the identification of mycorrhizae and rusts. Mol Ecol 2:113–118. https://doi.org/10.1111/j.1365-294X.1993.tb00005.x
Garrido-Sanz D, Redondo-Nieto M, Guirado M et al (2019) Metagenomic Insights into the Bacterial Functions of a Diesel-Degrading Consortium for the Rhizoremediation of Diesel-Polluted Soil. Genes 10. https://doi.org/10.3390/GENES10060456
Gaur VK, Gupta S, Pandey A (2021) Evolution in mitigation approaches for petroleum oil-polluted environment: recent advances and future directions. Environmental Science and Pollution Research 2021 29:41 29:61821–61837. https://doi.org/10.1007/S11356-021-16047-Y
Gielnik A, Pechaud Y, Huguenot D et al (2019) Bacterial seeding potential of digestate in bioremediation of diesel contaminated soil. Int Biodeterior Biodegrad 143. https://doi.org/10.1016/j.ibiod.2019.06.003
Gilgado F, Cano J, Gené J et al (2008) Molecular and phenotypic data supporting distinct species statuses for Scedosporium apiospermum and Pseudallescheria boydii and the proposed new species Scedosporium dehoogii. J Clin Microbiol 46:766–771. https://doi.org/10.1128/JCM.01122-07
Greco G, Di Piazza S, Cecchi G et al (2019) Mycoremediation of Oily Slime Containing a Polycyclic Aromatic Hydrocarbon Mixture. Waste Biomass Valorization 10:3821–3831. https://doi.org/10.1007/s12649-019-00802-x
Guarino C, Spada V, Sciarrillo R (2017) Assessment of three approaches of bioremediation (Natural Attenuation, Landfarming and Bioagumentation - Assistited Landfarming) for a petroleum hydrocarbons contaminated soil. Chemosphere 170:10–16. https://doi.org/10.1016/J.CHEMOSPHERE.2016.11.165
Ho MT, Li MSM, McDowell T et al (2020) Characterization and genomic analysis of a diesel-degrading bacterium, Acinetobacter calcoaceticus CA16, isolated from Canadian soil. BMC Biotechnol 20. https://doi.org/10.1186/s12896-020-00632-z
Huarte-Bonnet C, Kumar S, Saparrat MCN et al (2018) Insights into Hydrocarbon Assimilation by Eurotialean and Hypocrealean Fungi: Roles for CYP52 and CYP53 Clans of Cytochrome P450 Genes. Appl Biochem Biotechnol 184:1047–1060. https://doi.org/10.1007/s12010-017-2608-z
Johnston JE, Lim E, Roh H (2019) Impact of upstream oil extraction and environmental public health: A review of the evidence. Sci Total Environ 657:187–199. https://doi.org/10.1016/J.SCITOTENV.2018.11.483
Jung J, Philippot L, Park W (2016) Metagenomic and functional analyses of the consequences of reduction of bacterial diversity on soil functions and bioremediation in diesel-contaminated microcosms. Sci Rep 6. https://doi.org/10.1038/srep23012
Kamyabi A, Nouri H, Moghimi H (2017) Synergistic Effect of Sarocladium sp. and Cryptococcus sp. Co-Culture on Crude Oil Biodegradation and Biosurfactant Production. Appl Biochem Biotechnol 182:324–334. https://doi.org/10.1007/s12010-016-2329-8
Kim J, Lee AH, Chang W (2018) Enhanced bioremediation of nutrient-amended, petroleum hydrocarbon-contaminated soils over a cold-climate winter: The rate and extent of hydrocarbon biodegradation and microbial response in a pilot-scale biopile subjected to natural seasonal freeze-thaw temperatures. Sci Total Environ 612:903–913. https://doi.org/10.1016/J.SCITOTENV.2017.08.227
Klindworth A, Pruesse E, Schweer T et al (2013) Evaluation of general 16S ribosomal RNA gene PCR primers for classical and next-generation sequencing-based diversity studies. Nucleic Acids Res 41. https://doi.org/10.1093/nar/gks808
Koshlaf E, Ball AS (2017) Soil bioremediation approaches for petroleum hydrocarbon polluted environments. AIMS Microbiology 3:25–49. https://doi.org/10.3934/MICROBIOL.2017.1.25
Kotoky R, Rajkumari J, Pandey P (2018) The rhizosphere microbiome: Significance in rhizoremediation of polyaromatic hydrocarbon contaminated soil. J Environ Manag 217:858–870
Kumar S, Dheeman S, Dubey RC et al (2021) Cyclic siloxane biosurfactant-producing Bacillus cereus BS14 biocontrols charcoal rot pathogen Macrophomina phaseolina and induces growth promotion in Vigna mungo L. Arch Microbiol 203:5043–5054. https://doi.org/10.1007/S00203-021-02492-3
Kuppusamy S, Thavamani P, Venkateswarlu K et al (2017) Remediation approaches for polycyclic aromatic hydrocarbons (PAHs) contaminated soils: Technological constraints, emerging trends and future directions. Chemosphere 168:944–968. https://doi.org/10.1016/J.CHEMOSPHERE.2016.10.115
Latif IK, Karim AJ, Zuki ABZ et al (2010) Pulmonary modulation of benzo[a]pyrene-induced hemato-and hepatotoxicity in broilers. Poult Sci 89:1379–1388. https://doi.org/10.3382/ps.2009-00622
Liu N, Sun H, Xu L, Cai Y (2021) Methylsiloxanes in petroleum refinery facility: Their sources, emissions, environmental distributions and occupational exposure. Environ Int 152:106471. https://doi.org/10.1016/J.ENVINT.2021.106471
Liu S, Sun R, Cai M et al (2022) Petroleum spill bioremediation by an indigenous constructed bacterial consortium in marine environments. Ecotoxicol Environ Saf 241:113769. https://doi.org/10.1016/J.ECOENV.2022.113769
Lu XY, Li B, Zhang T, Fang HHP (2012) Enhanced anoxic bioremediation of PAHs-contaminated sediment. Bioresour Technol 104:51–58. https://doi.org/10.1016/j.biortech.2011.10.011
MacKinnon G, Duncan HJ (2013) Phytotoxicity of branched cyclohexanes found in the volatile fraction of diesel fuel on germination of selected grass species. Chemosphere 90:952–957. https://doi.org/10.1016/j.chemosphere.2012.06.038
Martínez Álvarez LM, Ruberto LAM, Lo Balbo A, Mac Cormack WP (2017) Bioremediation of hydrocarbon-contaminated soils in cold regions: Development of a pre-optimized biostimulation biopile-scale field assay in Antarctica. Sci Total Environ 590–591:194–203. https://doi.org/10.1016/j.scitotenv.2017.02.204
Meyer DD, Beker SA, Heck K et al (2018) Simulation of a surface spill of different diesel/biodiesel mixtures in an ultisol, using natural attenuation and bioaugmentation/biostimulation. An Acad Bras Cienc 90:2741–2752. https://doi.org/10.1590/0001-3765201820170268
Miethe D, Riis V, Babel W (1994) The relationship between the microbial activity of the autochthonous microorganisms of pristine and contaminated soils and their potential for the degradation of mineral oil hydrocarbons. Acta Biotechnol 14:131–140. https://doi.org/10.1002/abio.370140204
Montgomery MT, Boyd TJ, Osburn CL, Smith DC (2010) PAH mineralization and bacterial organotolerance in surface sediments of the Charleston Harbor estuary. Biodegradation 21:257–266. https://doi.org/10.1007/s10532-009-9298-3
Morales LT, González-García LN, Orozco MC et al (2017) The genomic study of an environmental isolate of Scedosporium apiospermum shows its metabolic potential to degrade hydrocarbons. Stand Genomic Sci 12. https://doi.org/10.1186/s40793-017-0287-6
Muriel-Millán LF, Rodríguez-Mejía JL, Godoy-Lozano EE et al (2019) Functional and Genomic Characterization of a Pseudomonas aeruginosa Strain Isolated From the Southwestern Gulf of Mexico Reveals an Enhanced Adaptation for Long-Chain Alkane Degradation. Front Mar Sci 6. https://doi.org/10.3389/fmars.2019.00572
Muthukumar B, Al Salhi MS, Narenkumar J et al (2022a) Characterization of two novel strains of Pseudomonas aeruginosa on biodegradation of crude oil and its enzyme activities. Environ Pollut 304. https://doi.org/10.1016/j.envpol.2022.119223
Muthukumar B, Parthipan P, AlSalhi MS et al (2022b) Characterization of bacterial community in oil-contaminated soil and its biodegradation efficiency of high molecular weight (C40) hydrocarbon. Chemosphere 289:133168. https://doi.org/10.1016/j.chemosphere.2021.133168
Nie Y, Chi CQ, Fang H et al (2014) Diverse alkane hydroxylase genes in microorganisms and environments. Sci Rep 4. https://doi.org/10.1038/srep04968
Nwankwegu AS, Onwosi CO (2017) Bioremediation of gasoline contaminated agricultural soil by bioaugmentation. Environ Technol Innov 7:1–11. https://doi.org/10.1016/J.ETI.2016.11.003
Nzila A, Musa MM (2020) Current Status of and Future Perspectives in Bacterial Degradation of Benzo[a]pyrene. Int J Environ Res Public Health 18:1–24. https://doi.org/10.3390/IJERPH18010262
Oksanen P (2010) Vegan 1.17-0
Orro A, Cappelletti M, D’Ursi P et al (2015) Genome and phenotype microarray analyses of Rhodococcus sp. BCP1 and Rhodococcus opacus R7: Genetic determinants and metabolic abilities with environmental relevance. PLoS One 10. https://doi.org/10.1371/journal.pone.0139467
Pandey J, Chauhan A, Jain RK (2009) Integrative approaches for assessing the ecological sustainability of in situ bioremediation. FEMS Microbiol Rev 33:324–375
Park H, Min B, Jang Y et al (2019) Comprehensive genomic and transcriptomic analysis of polycyclic aromatic hydrocarbon degradation by a mycoremediation fungus, Dentipellis sp. KUC8613. Appl Microbiol Biotechnol 103:8145–8155. https://doi.org/10.1007/S00253-019-10089-6
Parthipan P, Cheng L, Dhandapani P et al (2022) Impact of biosurfactant and iron nanoparticles on biodegradation of polyaromatic hydrocarbons (PAHs). Environ Pollut 306. https://doi.org/10.1016/j.envpol.2022.119384
Passarini MRZ, Sette LD, Rodrigues MVN (2011) Improved extraction method to evaluate the degradation of selected PAHs by marine fungi grown in fermentative medium. J Braz Chem Soc 22:564–570. https://doi.org/10.1590/S0103-50532011000300022
Peidro-Guzmán H, Pérez-Llano Y, González-Abradelo D et al (2021) Transcriptomic analysis of polyaromatic hydrocarbon degradation by the halophilic fungus Aspergillus sydowii at hypersaline conditions. Environ Microbiol 23:3435–3459. https://doi.org/10.1111/1462-2920.15166
Poi G, Aburto-Medina A, Mok PC et al (2017) Large scale bioaugmentation of soil contaminated with petroleum hydrocarbons using a mixed microbial consortium. Ecol Eng 102:64–71. https://doi.org/10.1016/J.ECOLENG.2017.01.048
Prathiba J (2014) Molecular identification of the isolated diesel degrading bacteria and optimization studies. J Biochem Technol 5:727–730
Pylro VS, Roesch LFW, Ortega JM et al (2014) Brazilian Microbiome Project: Revealing the Unexplored Microbial Diversity-Challenges and Prospects. Microb Ecol 67:237–241. https://doi.org/10.1007/s00248-013-0302-4
Rakowska J (2020) Remediation of diesel-contaminated soil enhanced with firefighting foam application. Scientific Reports 2020 10:1 10:1–9. https://doi.org/10.1038/s41598-020-65660-3
Rojo F (2009) Degradation of alkanes by bacteria: Minireview. Environ Microbiol 11:2477–2490
Roy A, Dutta A, Pal S et al (2018) Biostimulation and bioaugmentation of native microbial community accelerated bioremediation of oil refinery sludge. Bioresour Technol 253:22–32. https://doi.org/10.1016/j.biortech.2018.01.004
Schnurer J, Rosswall T (1982) Fluorescein diacetate hydrolysis as a measure of total microbial activity in soil and litter. Appl Environ Microbiol 43:1256–1261. https://doi.org/10.1128/aem.43.6.1256-1261.1982
Schumacher TE, Eynard A, Chintala R (2015) Rapid cost-effective analysis of microbial activity in soils using modified fluorescein diacetate method. Environ Sci Pollut Res 22:4759–4762. https://doi.org/10.1007/s11356-014-3922-4
Schwarz A, Adetutu EM, Juhasz AL et al (2018) Microbial Degradation of Phenanthrene in Pristine and Contaminated Sandy Soils. Microb Ecol 75:888–902. https://doi.org/10.1007/s00248-017-1094-8
Seklemova E, Pavlova A, Kovacheva K (2001) Biostimulation-based bioremediation of diesel fuel: Field demonstration. Biodegradation 12:311–316. https://doi.org/10.1023/A:1014356223118
Shahsavari E, Adetutu EM, Anderson PA, Ball AS (2013) Plant residues - A low cost, effective bioremediation treatment for petrogenic hydrocarbon-contaminated soil. Sci Total Environ 443:766–774. https://doi.org/10.1016/j.scitotenv.2012.11.029
Shukor MY, Dahalan FA, Jusoh AZ et al (2009) Characterization of a diesel-degrading strain isolated from a hydrocarbon-contaminated site. J Environ Biol 30:145–150
Simon M, Joshi H (2021) A review on green technologies for the rejuvenation of polluted surface water bodies: Field-scale feasibility, challenges, and future perspectives. J Environ Chem Eng 9:105763. https://doi.org/10.1016/J.JECE.2021.105763
Sutton NB, Maphosa F, Morillo JA et al (2013) Impact of long-term diesel contamination on soil microbial community structure. Appl Environ Microbiol 79:619–630. https://doi.org/10.1128/AEM.02747-12
Tahri N, Bahafid W, Sayel H, El Ghachtouli N (2013) Biodegradation: Involved Microorganisms and Genetically Engineered Microorganisms. In: Biodegradation - Life of Science
Throne-Holst M, Wentzel A, Ellingsen TE et al (2007) Identification of novel genes involved in long-chain n-alkane degradation by Acinetobacter sp. strain DSM 17874. Appl Environ Microbiol 73:3327–3332. https://doi.org/10.1128/AEM.00064-07
Tiralerdpanich P, Sonthiphand P, Luepromchai E et al (2018) Potential microbial consortium involved in the biodegradation of diesel, hexadecane and phenanthrene in mangrove sediment explored by metagenomics analysis. Mar Pollut Bull 133:595–605. https://doi.org/10.1016/j.marpolbul.2018.06.015
Tribelli PM, Rossi L, Ricardi MM et al (2018) Microaerophilic alkane degradation in Pseudomonas extremaustralis: a transcriptomic and physiological approach. J Ind Microbiol Biotechnol 45:15–23. https://doi.org/10.1007/s10295-017-1987-z
U.S. Food and Drug Administration (FDA) (2001) Microbiological Methods & Bacteriological Analytical Manual (BAM). US Department of Health and Human Services 10–12
Ugochukwu SC, Uche A, Ifeanyi O (2013) Preliminary phytochemical screening of different solvent extracts of stem bark and roots of Dennetia tripetala. Asian J Plant Sci Res 3:10–13
US EPA (2007) Method 8015C Nonhalogenated organics by gas chromatography. In: SW-846 OnLine Test Methods for Evaluating Solid Waste, Physical/Chemical Methods. https://www.epa.gov/sites/default/files/2015-12/documents/8015c.pdf. Accessed Aug 2023
Uzoigwe C, Burgess JG, Ennis CJ, Rahman PKSM (2015) Bioemulsifiers are not biosurfactants and require different screening approaches. Front Microbiol 6:245. https://doi.org/10.3389/FMICB.2015.00245/BIBTEX
Vidal-Verdú À, Gómez-Martínez D, Latorre-Pérez A, et al (2022) The car tank lid bacteriome: a reservoir of bacteria with potential in bioremediation of fuel. Biofilms and Microbiomes 2022 8:1 8:1–12. https://doi.org/10.1038/s41522-022-00299-8
White TJ, Bruns T, Lee S, Taylor J (1990) Amplification and direct sequencing of fungal ribosomal rna genes for phylogenetics. PCR Protocols 315–322. https://doi.org/10.1016/B978-0-12-372180-8.50042-1
Yadav R, Rajput V, Dharne M (2021) Functional metagenomic landscape of polluted river reveals potential genes involved in degradation of xenobiotic pollutants. Environ Res 192. https://doi.org/10.1016/j.envres.2020.110332
Yergeau E, Sanschagrin S, Beaumier D, Greer CW (2012) Metagenomic analysis of the bioremediation of diesel-contaminated canadian high arctic soils. PLoS One 7. https://doi.org/10.1371/journal.pone.0030058
Yuan X, Zhang X, Chen X et al (2018) Synergistic degradation of crude oil by indigenous bacterial consortium and exogenous fungus Scedosporium boydii. Bioresour Technol 264:190–197. https://doi.org/10.1016/j.biortech.2018.05.072
Zafra G, Taylor TD, Absalón AE, Cortés-Espinosa DV (2016) Comparative metagenomic analysis of PAH degradation in soil by a mixed microbial consortium. J Hazard Mater 318:702–710. https://doi.org/10.1016/j.jhazmat.2016.07.060
Acknowledgments
PG and LLS thank the Fundação de Amparo à Pesquisa do Estado de São Paulo (FAPESP) for the Postdoctoral Fellowships (#2016/17488-4 and #2019/11823-4; 2018/10734-5, respectively). LDS thanks Conselho Nacional de Desenvolvimento Tecnológico for the Research Productivity Fellowship (#303218/2019-3).
Funding
This research was financially supported by the Fundação de Amparo à Pesquisa do Estado de São Paulo (FAPESP) (grant #2016/07957-7) and by Conselho Nacional de Desenvolvimento Tecnológico (CNPq) (grant #407986/2018-6).
Author information
Authors and Affiliations
Contributions
Patricia Giovanella: Conceptualization, Methodology, Investigation, Formal analysis, and Writing-original draft preparation; Rodrigo Gouvêa Taketani: Formal analysis, Writing - Review and Editing; Ruben Gil-Solsona: Formal analysis, Investigation, Writing - Review and Editing; Luiz Leonardo Saldanha: Formal analysis; Samantha Beatríz Esparza Naranjo: Investigation; Juan V. Sancho: Formal analysis; Tania Portolés: Formal analysis; Fernando Dini Andreote: Reviewing and Editing; Sara Rodríguez-Mozaz: Supervision, Reviewing and Editing; Damià Barceló: Supervision, Resources Provision, Reviewing and Editing; Lara Durães Sette: Conceptualization, Supervision, Funding acquisition, Reviewing and Editing.
Corresponding author
Ethics declarations
Ethical Approval
Not applicable.
Consent to Participate
Not applicable.
Consent to Publish
Not applicable.
Competing Interests
The authors declare no competing interests.
Additional information
Responsible Editor: Robert Duran
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
ESM 1
(DOCX 25.9 kb)
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Giovanella, P., Taketani, R.G., Gil-Solsona, R. et al. A comprehensive study on diesel oil bioremediation under microcosm conditions using a combined microbiological, enzymatic, mass spectrometry, and metabarcoding approach. Environ Sci Pollut Res 30, 101250–101266 (2023). https://doi.org/10.1007/s11356-023-29474-w
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11356-023-29474-w