Abstract
Exposure to environmental pollutants has been associated with alteration on relative levels of mitochondrial DNA copy number (mtDNAcn). However, the results obtained from epidemiological studies are inconsistent. This meta-analysis aimed to evaluate whether environmental pollutant exposure can modify the relative levels of mtDNAcn in humans. We performed a literature search using PubMed, Scopus, and Web of Science databases. We selected and reviewed original articles performed in humans that analyzed the relationship between environmental pollutant exposure and the relative levels of mtDNAcn; the selection of the included studies was based on inclusion and exclusion criteria. Only twenty-two studies fulfilled our inclusion criteria. A total of 6011 study participants were included in this systematic review and meta-analysis. We grouped the included studies into four main categories according to the type of environmental pollutant: (1) heavy metals, (2) polycyclic aromatic hydrocarbons (PAHs), (3) particulate matter (PM), and (4) cigarette smoking. Inconclusive results were observed in all categories; the pooled analysis shows a marginal increase of relative levels of mtDNAcn in response to environmental pollutant exposure. The trial sequential analysis and rate confidence in body evidence showed the need to perform new studies. Therefore, a large-scale cohort and mechanistic studies in this area are required to probe the possible use of relative levels of mtDNAcn as biomarkers linked to environmental pollution exposure.
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
Introduction
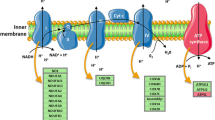
Mitochondria regulate several vital functions for maintaining cellular homeostasis and cell survival; some key processes are ATP production based on oxidative phosphorylation, cell signaling for controlling cell death by apoptosis, and others. Mitochondria possess their DNA; the mitochondrial DNA (mtDNA) is a circular double-stranded DNA molecule. Each mitochondrion contains several copies of mtDNA located in the mitochondrial matrix; mtDNA encodes 13 protein subunits implicated in oxidative phosphorylation, two rRNAs, and 22 tRNAs (Barshad et al. 2018). It is widely accepted that the mtDNA copy number (mtDNAcn) specifically variates according to cell or tissue types (Herbers et al. 2019).
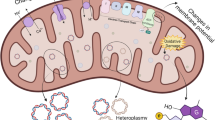
Toxicity starts with the interaction between the toxicant and its target molecule. It occurs when the ultimate toxicant reacts to an effective concentration at the target site. The target molecule has three key characteristics: (1) the capacity to react with the ultimate toxicant (2) that results in a disturbing function (3) which is mechanistically related to toxicity (Lehman-McKeeman and Armstrong 2022). In this direction, mitochondria could be considered a target molecule linked with or responsible for toxicity; it possesses several characteristics that increase the susceptibility to toxicity associated with environmental pollutant exposure. For example, mitochondria have double membrane rich in phospholipids that could attract lipophilic toxicants; mitochondria’s monooxygenases could participate in the bioactivation of environmental pollutants; metallic environmental pollutants could input cross calcium channels present in the mitochondria, and the gradient of protons in the intermembrane space, and the pH of the mitochondrial matrix could promote the accumulation of cationic metals and amphiphilic environmental pollutants (Meyer et al. 2013, 2017, 2018).
Consequently, several biomarkers have been suggested to evaluate human health linked to mitochondrial health. Early studies showed that deletions and mutations on mtDNA resulted in several complex diseases denominated as mitochondrial diseases. Interestedly, diagnosed individuals with the same mitochondrial disease show different levels of mtDNAcn; both increase and decrease in mtDNAcn were related with severe consequences in the clinical manifestations of mitochondrial disease (Bai et al. 2004). This observation indicated the possible use of mtDNAcn levels as a potential biomarker related to human health. In this context, a variation on mtDNAcn has been primarily studied as a possible predictor for the development of human diseases such as cancer, cardiovascular, kidney, liver, and neurodegenerative diseases (Castellani et al. 2020).
Several environmental pollutants can induce mitochondrial toxicity resulting in oxidative stress, altered mitochondrial ATP production, and disrupted mitochondrial-mediated cell signaling (Meyer et al. 2018). Moreover, early studies reported that environmental pollutants such as PAHs showed a high affinity to mtDNA compared to nuclear DNA (Allen and Coombs 1980). For all these reasons, the determination of changes on mtDNAcn has been monitored in response to environmental pollutant exposure. In vitro, in vivo, and epidemiological studies have reported a direct relationship between the alteration in mtDNAcn, and the exposure to environmental pollutants; nevertheless, the epidemiological studies based on environmental or occupational exposure to environmental pollutants showed contradictory results (Roubicek and Souza-Pinto 2017).
On the other hand, the implementation of systematic reviews focused on environmental issues allows assessment of the state of science concerning a particular topic. Moreover, the derived results are reproducible, transparent, formal, and accurate. Therefore, systematic reviews in environmental epidemiology are employed to make evidence-based decisions in the health environment (Rooney et al. 2014). Moreover, recently, a specific tool has been developed for performing a systematic review and meta-analyses in environmental epidemiology; this tool is used to evaluate the risks of bias, assess the quality of individual studies, and rate confidence in the body of evidence (NTP/OHAT 2019). Hence, this systematic review and meta-analysis aimed to determine the pooled effect of exposure to environmental pollutants on changes in mtDNAcn.
Materials and methods
To carry out this work, we followed the recommendations from the Handbook for Conducting a Literature-based Health Assessment Using OHAT Approach for Systematic Review and Evidence (NTP/OHAT 2019) and the Preferred Reporting Items for Systematic Reviews and Meta-Analyses (PRISMA) guidelines (Page et al. 2021). The PRISMA checklist is presented in supplementary data 4. A protocol was developed and registered (registration number CRD42020211134) in the international prospective register of systematic review (PROSPERO) database. A systematic search of the literature up to Oct 2020 was performed. The search was performed using PubMed, Scopus, and Web of Science databases, using the following keywords: “Environmental Pollutants” OR “Environmental Pollution” and “DNA, Mitochondrial.” We used the Medical Subject Headings (MeSH) and the most frequent synonyms. In addition, the search also included the review of the cited bibliographies to identify omitted articles.
Evidence searches
We followed the PECO (population, exposure, comparators, and outcomes) strategy for the reach question construction and evidence search (da Costa Santos et al. 2007). The critical elements of the PECO strategy are presented in Table 1.
Selection criteria
The selection criteria were as follows: (1) original articles (2) performed in humans (case–control, cross-sectional, or cohort studies) that (3) evaluated the association between mtDNAcn and the exposure to environmental pollutants. (4) The result of interest was the mtDNAcn expressed as the mean and standard error (or the data for their estimation).
The exclusion criteria were as follows: (1) review articles, letters, comments, correspondence, and conference reports; or (2) studies that did not provide enough data. In addition, we selected the most recent or most comprehensive manuscript if duplicate reports from a research group existed. Finally, if the authors only reported a graphic without descriptive data, we analyzed the reported graphic using the “WebPlotDigitizer—Extract data from plots, images, and maps” (version 4.0 PLOTCON 2017-Oakland, USA) for obtaining the descriptive data. Unselected articles that reported the effect of human exposure to environmental pollutants on mtDNAcn are presented in supplementary data 5.
Data extraction
For the included studies, we extracted the following data: family name of the first author, publication year, study design, geography localization, sample size, sex of participant, age, environmental pollutant, the employed method for exposure assessment, and biological matrix used for exposure assessment, confounding variables, and statistical methods. In addition, for all included studies, the information regarding the measure of mtDNAcn such as the method used for relative quantification, the mitochondrial and nuclear genes used in the assay, and the biological matrix used for the obtained DNA was extracted from each study. Two researchers independently selected the studies (CAR, MARV). Differences in study selection between researchers were infrequent and resolved by consensus.
Assessment of risk of bias
Two independent reviewers performed the quality assessment for each included study. The OHAT’s risk of bias tool for human and animal studies was used as a standardized method to quantify the risk of bias (NTP/OHAT 2019). The tool was designed for global selection-, confounding-, and detection-bias assessments. In brief, risk of bias questions addressed similarity between compared groups, confounder’s consideration (methodological measure and statistical treatment), pollutant exposure characterization (use of well-established methods; limits of detection and quantification reported and not near values), and outcome assessment (use of well-established methods, coefficient of variation reported, use of standards or internal controls, the blinding during the analysis). We rank studies by reported information in “Definitively low risk of bias,” “Probably low risk of bias,” “Probably high risk of bias,” or “Definitively high risk of bias.”
Statistical analysis
We calculated the standardized mean differences (SMDs) with 95% confidence intervals (95% CIs) of mtDNAcn in each study using a random-effects model. The random-effects analysis assumes that the true effect could vary between the studies and that the studies analyzed in the meta-analysis are a random sample of the effects that could be observed (Borenstein 2009). We considered the high versus low level of exposure for studies that reported categorical exposure. The I2 test was used to measure the proportion of variation between studies concerning total variation “heterogeneity.” For the I2 test, a value of 0% indicates no observed heterogeneity. The values between 25 and 50% indicate a low heterogeneity, and the values between 50 and 75% indicate a moderate heterogeneity, while the values ≥ 75% indicate a high heterogeneity (Higgins and Thompson 2002). In addition, the publication bias was evaluated by visual inspection of Egger’s publication bias plot, and the effect of small studies was tested using Egger’s test (Pustejovsky and Rodgers 2019). Statistical analyses were performed using the statistical package STATA version 16 (College Station, TX).
Trial sequential analysis
The meta-analyses are considered the best available evidence. However, the results derived from meta-analytic techniques could be influenced by statistical errors such as type 1 or 2. For this reason, it is essential to employ methods for ensuring that obtained results have a negligible risk of reporting false-positive or false-negative results. Trial sequential analysis (TSA) is used to decrease the false-positive or false-negative risk. TSA estimates the necessary number of events and participants to calculate a reliable statistical inference. In addition, the strength of evidence is measured using the accrued number of participants, observed difference of outcome measured in the included trials, and the impact of assortment. If the cumulative Z-curve crosses the TSA limit, it is inferred that the meta-analysis has enough evidence.
On the other hand, if the Z-curve does not cross the TSA limit, it is inferred that there is not enough evidence, and more trials are necessary (Shah and Smith 2020; Wetterslev et al. 2017). We examined the reliability and confidence of the available reports using TSA. We performed TSA analysis considering an overall 5% risk of a type I error and 20% of a type II error (power of 80%). The sequential trial analysis was performed using TSA version 0.9.5.10 β (“TSA”).
Rate the confidence in the body evidence
The OHAT method based on the Grading of Recommendations Assessment, Development and Evaluation “GRADE” was employed for rating the confidence in the body evidence (NTP/OHAT 2019). The GRADE tool starts evaluating the initial confidence rate considering the study design; human controlled trials are considered with a high initial confidence rating, and other studies such as case–control/cohorts or cross-sectional studies are evaluated with low-to-moderate initial confidence rate. After the first assessment, key factors can change the rate of confidence. Factors downgrading the confidence are the following: (1) risk of bias (the proportion of the included studies classified as T1 to T3 is considered); (2) imprecision (for continuous outcomes downrating is considered if the included studies reported sample sizes lower than 400); (3) publication bias (identified using funnel plots and Egger’s regression); (4) indirectness and applicability (epidemiological studies are not downgraded for directness regardless of the exposure level); (5) unexplained inconsistency (downgrading if significant and considerable heterogeneity is presented in meta-analysis).
Results
Literature search results
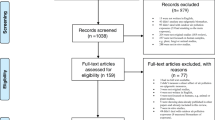
The electronic database searches yielded 465 records, and 69 records were identified from reviewing the referent list of the collected record. One hundred seven records were reviewed at the full-text level because of title and abstract screening. However, only twenty-two records fulfilled our inclusion criteria and were included in the meta-analysis (Fig. 1). Furthermore, in supplementary data 5, we summarized the unselected articles that evaluated the effect of human exposure to environmental pollutants on mtDNAcn.
Study characteristics
Table 2 summarizes the main characteristics of the included studies. Most included studies (≈ 82%) were performed under a cross-sectional design, and two studies reported results from prospective cohorts. In addition, a matched case–control or longitudinal design was performed by Linqing et al. (2016) and Hou et al. (2010), respectively. Fifty-five percent of the included studies in the meta-analysis captured both men and women; 32% of the studies only had men. Only one study did not report the sex of the included participants, and the remaining studies were performed on women.
Considering all age groups reported by the included studies, 95% of studies had adults, and only one study was performed on children. Seventy-three percent of the included studies measured the mtDNAcn using the ND1 as referent mitochondrial gene and β-globin as referent nuclear gene. The whole blood or isolated leukocytes was the most frequent (73.9%) biological matrix used by DNA isolation. Only 27% of the included studies did not conduct an adjustment for confounding factors. Other characteristics from included studies are shown in supplementary data 1.
Other characteristics such as recruitment strategy, inclusion, and exclusion criteria for the included participants, the method employed for exposure assessment, the instrument used (with limits of quantification or detection), and the standards employed are presented in supplementary data 1. The meta-analysis includes 6011 individuals exposed to environmental pollutants.
We grouped the included studies according to the type of environmental pollutant. We considered a conventional classification based on similar toxic mechanisms and physicochemical characteristics. We have grouped the included articles into four main groups. The heavy metal group includes metallic chemical elements characterized by their relatively high density and their toxic effect at low concentrations (Balali-Mood et al. 2021). Benzene and polycyclic aromatic hydrocarbons were grouped. Polycyclic aromatic hydrocarbons (PAHs, sometimes referred to as polynuclear aromatic hydrocarbons (PNAs)) are a large group of diverse organic compounds, which contain two or more fused aromatic rings. At the same time, benzene has only one aromatic ring (Fan and Lin 2011). Another group is comprised of articles that evaluate human exposure to particulate matter. Particle pollution is a general term for a mixture of solid and liquid droplets suspended in the air (Ukaogo et al. 2020). And finally, the fourth group is comprised of articles that evaluated the active exposure to cigarette smoke.
Effect of heavy metal exposure on the mtDNAcn
Alegría-Torres et al. (2020) characterized the concentrations of arsenic (As) and lead (Pb) in children from schools close to an oil refinery and thermoelectric plant. However, the author only reported the changes of mtDNAcn according to Pb blood concentration. Ameer et al. (2016) reported the changes of mtDNAcn according to inorganic As and its metabolites such as methylarsonic acid (MMA) and dimethylarsinic acid (DMA) in urine; the exposure to As occurred via the drinking water consumed by residents. Li et al. (2020a, b) included participants exposed to electronic waste; the reported results indicated that the concentrations of As, nickel (Ni), silver (Ag), lanthanum (La), and cerium (Ce) were higher in individuals exposed to electronic waste. Linqing et al. (2016) conducted a matched case–control study; they measured chromium (Cr), zinc (Zn), copper (Cu), cadmium (Cd), and selenium (Se) concentrations from blood in the exposed and control groups. Nevertheless, only the Cr concentration was higher in the exposed group.
The mtDNAcn showed an increase in highly exposed individuals to As, Cr, or electronic waste. An inverse effect was observed in the highly exposed individuals to Pb. Although the pooled analysis showed a marginal and insignificant increase in the mtDNAcn, the pooled standardized mean difference was 0.72-fold higher in highly exposed individuals (95% CIs: − 0.30 to 1.75; Z = 1.39; p = 0.17) (Fig. 2a). High and significant heterogeneity was observed (I2 = 96.7%, p < 0.0001).
Forest plot estimates for analyzing the changes in mtDNAcn induced by the exposure to a heavy metals, b benzene and polycyclic aromatic hydrocarbons, c particulate matter, and d cigarette smoking. The standardized mean differences (SMD) with the 95% CIs are presented. The size of the gray box is proportional to the weight assigned to each study, and horizontal lines represent the 95% CIs. The blue diamond indicates the overall effect size, the width of the blue diamond represents the 95% CIs, and the vertical dashed line represents the pooled error standard
Effect of benzene and polycyclic aromatic hydrocarbon exposure on the mtDNAcn
The changes in the mtDNAcn in response to PAH exposure were analyzed by Du et al. (2020), Duan et al. (2020), Ling et al. (2017), Pavanello et al. (2013), Shen et al. (2008), and Xu et al. (2018); and the effect to exposure to benzene on mtDNAcn was analyzed by Carugno et al. (2012), Li et al. (2020a, b), and Wong et al. (2017). However, the studies were performed on heterogeneous populations. For example, Du et al. (2020), Duan et al. (2020), and Pavanello et al. (2013) recruited coke-oven workers occupationally exposed to PAHs. The effect of occupational exposure to benzene on mtDNAcn was reported by Carugno et al. (2012), while other studies included individuals exposed to a complex mixture of fumes delivery from the combustion of solid fuels (Wong et al. 2017) or emit from hot asphalt (Xu et al. 2018). Only one study reported results from a cohort study (Ling et al. (2017)). The exposure assessment was performed measuring the PAH metabolite concentrations from the urine of the included individuals (Du et al. 2020; Ling et al. 2017; Pavanello et al. 2013; Xu et al. 2018) or quantifying the airborne PAHs from the breathing zone of everyone (Carugno et al. 2012; Duan et al. 2020; Li et al. 2020a, b; Shen et al. 2008; Wong et al. 2017).
A pooled analysis showed that the mtDNAcn was 0.008-fold higher in the highly exposed individuals than in the lowly exposed group (95% CIs: − 0.52 to 0.52; Z = 0.03; p = 0.97) (Fig. 2b), and significant heterogeneity was observed (I2 = 98.2%, p < 0.0001).
Effect of particulate matter exposure on the mtDNAcn
Studies performed to evaluate the effect of particulate matter on the mtDNAcn are listed below: Hou et al. (2013); Wang et al. (2020); Wong et al. (2017); Xu et al. (2017); Zhang et al. (2020). Wang et al. (2020) and Zhang et al. (2020) reported the results from two different cohorts that included only men; the included participants came through populations exposed to the highest or lowest levels of ambient air pollutant exposure (PM10, PM2.5, NO2, and CO). The occupational exposure to particulate matter was described in the truck drivers (Hou et al. 2013), welders (Xu et al. 2017), workers from an electric steel plant (Hou et al. 2010), and housewives (Wong et al. 2017). The pooled analysis showed an increase in the mtDNAcn in highly exposed individuals to the particulate matter. The overall estimated results were of 0.16-fold (95% CIs: − 0.1 to 0.43; Z = 1.18; p = 0.24); the linked heterogeneity was considered moderate (I2 = 74.1%, p = 0.002) (Fig. 2c).
Effect of cigarette smoking on the mtDNAcn
The effect of cigarette smoking on the mtDNAcn was evaluated by Bouhours-Nouet et al. (2005), Lee et al. (1998), Masayesva et al. (2006), and Tan et al. (2008). The increase in mtDNAcn was frequently observed in smokers. The pooled analysis showed an increase of 0.48-fold (95% CIs: 0.20–0.77; Z = 3.34; p = 0.001) higher in smokers than non-smokers. A low and insignificant heterogeneity was determined for the overall result (I2 = 34.9%, p = 0.2) (Fig. 2d). The obtained result clearly shows the importance of including cigarette smoking as a confounder exposure source.
Subgroup analysis for exploring the causes of heterogeneity
Unfortunately, we could not perform a subgroup analysis to explore the potential causes of heterogeneity because of the few included studies for heavy metal exposure and cigarette smoking. For polycyclic aromatic hydrocarbon exposure, we performed a subgroup analysis considering the classification according to NTP/OHAT risk of bias tool for exploring the causes of heterogeneity. Nevertheless, the subgroup analysis showed similar heterogeneity \({I}^{2}\)=92.7% and \({I}^{2}\)=97.9% for 1st tier and 2nd tier, respectively (see supplementary data 2, Fig. 1). Similarly, subgroup analysis for particulate matter exposure showed a low and insignificant heterogeneity (\({I}^{2}=\) 0%; p = 0.75) for 2nd tier. In the 1st tier, a high and significant heterogeneity was observed (\({I}^{2}=84.4\%\); p < 0.001) (see supplementary data 2, Fig. 2). Furthermore, a subgroup analysis considering the type of exposure (environmental or occupational) was performed; the heterogeneity for environmental exposure was low and insignificant (\({I}^{2}=\) 0%; p = 0.74). The subgroup of occupational exposure showed high and significant heterogeneity (\({I}^{2}=84.4\%\); p < 0.001) (see supplementary data 2, Fig. 3).
Egger’s funnel plot for measuring the risk of bias in studies included in the meta-analysis; circles indicate the observed standardized effect. The subsections represent the following information: a heavy metals, b benzene and polycyclic aromatic hydrocarbons, d particulate matter, and b cigarette smoking
In addition, we performed a subgroup analysis considering the type of tissue used for isolating mitochondrial and genomic DNA. Heterogeneity was low and non-significant for semen (\({I}^{2}=57.6\%; p=0.1\)), oral mucosa (\({I}^{2}=0\%; p=0.5\)), and other somatic tissues such as placenta or lung tissue (\({I}^{2}=0\%; p=0.6\)). However, high and significant heterogeneity was observed for isolated leukocytes (\({I}^{2}=94.2\%; p<0.001\)), whole blood (\({I}^{2}=97.4\%; p<0.001\)), and buffy coat (\({I}^{2}=85\%; p=0.01\)) (see supplementary data 2, Fig. 4).
Trial sequential analysis. The O’Brien-Fleming boundary line is displayed as the sloping red full lines with black diamonds, and the TSA line represents the required information size. The subsections describe the following information: a heavy metals, b benzene and polycyclic aromatic hydrocarbons, c particulate matter, and d cigarette smoking
Small-study effects and publication bias
Egger’s funnel plot was performed to assess the publication bias in the meta-analysis. The funnel plots showed symmetrical distribution for benzene and PAHs (Fig. 3b); the visual inspection of estimated Egger’s funnel plot for heavy metals, particulate matter, and cigarette smoking revealed an asymmetric distribution (Fig. 3a , c, and d, respectively). Moreover, we performed Egger’s regression test for assessing small-study effects. As shown in Table 3, the small-study effects could not be expected (p-value > 0.05). However, the estimated p-value (p = 0.051) for cigarette smoking thus provides weak evidence for the presence of small-study effects.
Trial sequential analysis
The obtained results of TSA are presented in Fig. 4. Figure 4a shows the TSA for heavy metals, Fig. 4b represents the TSA for benzene and PAHs, Fig. 4c shows the TSA for PM, and finally, Fig. 4d shows the TSA for cigarette smoking. The cumulative Z-curve does not cross the required information size in all cases. Furthermore, the cumulative Z-curve does not overpass the O’Brien-Fleming boundary lines. Accordingly, the results of TSA suggest that new studies are necessary to ensure the usefulness of mtDNAcn as a biomarker.
Risk of bias assessment
Table 4 summarizes the results from the risk of bias assessment. We employed the NTP/OHAT risk of bias tool to evaluate each included study’s quality. The first considered topic was the “selection bias.” Next, we assessed if the comparison groups were appropriate; many included studies reported the similarities in baseline characteristics between the analyzed groups. However, many studies that analyzed the effect of cigarette smoking on mtDNAcn did not report the similarities between the comparison groups.
The “confounding bias” topic was evaluated considering if the included studies thought the confounding and modifying variables in its design/statistical analysis. Fifty-six percent of the included studies did not adjust or control for other exposure. As previously mentioned, cigarette smoking could be considered a critical confounder. However, only one study considered cigarette smoking in the heavy metal group. For PM and PHA groups, 66% considered cigarette smoking as a confounder. A multivariate statistical approach was used to decrease the confounder effect linked to cigarette smoking in all cases. On the other hand, only 22% of the included studies considered the influence of confounding and modifying variables (key topic).
Finally, the “detection bias” topic was evaluated; a low risk of bias was considered when the included studies reported the following: the blinding in outcome analysis, the use of valid methods for measuring confounding variables, the use of a valid method, and techniques for exposure characterization (key topic) and outcome characterization (key issue); all included studies reported the use of valid, reliable, and sensitive methods enabled to measure the changes in the mtDNAcn in response to environmental pollutant exposure. The exposure characterization was realized by quantifying the environmental pollutant or its metabolites; the studies that evaluated the effect of cigarette smoking performed the exposure assessment using standardized questionnaires. Only three records performed the outcome measuring using blinding protocols. Sixty percent of the included studies analyzed the confounding variables employing sensitive methods of assessing substantial confounding or modifying variables.
Rate confidence in body evidence
Finally, we employed the OHAT method based on the Grading of Recommendations Assessment, Development and Evaluation “GRADE” to rate the confidence in the body evidence. This guide contemplates the strengths and weaknesses of the included articles; if the reported result showed strong confidence between pollutant exposure and the mtDNAcn levels, the obtained conclusion is considered high confidence. However, an unclear relationship between exposure to environmental pollutants and mtDNAcn is considered shallow confidence. In addition, the result of rate confidence indicates directly or indirectly the need to perform new research in a particular topic. For example, suppose the body evidence is considered with high confidence. In that case, it suggests that further research is improbable to modify the confidence in the relationship between exposure to environmental pollutants and mtDNAcn. On the contrary, if the rate of body evidence is considered very low, it suggests that new research is very likely to modify the relationship between exposure to environmental pollutants and outcome.
The GRADE assessment for heavy metals, benzene and PAHs, PM, and cigarette smoking is presented in supplementary data 3 Tables 1, 2, 3, and 4, respectively. Almost included studies were performed under a cross-sectional design. For this reason, the initial rate of confidence was considered low level. Imprecision was a common downgrading factor that decreased confidence. Low confidence in the body of evidence was the GRADE conclusion obtained for all included environmental pollutants. Larger-scale studies on this topic are required to find the role of mtDNAcn in response to environmental pollutant exposure.
Discussion
In the present study, we investigated the impact of environmental pollutants on the changes in relative levels of mtDNAcn. Although the results showed conflicting results in all included studies for each environmental pollutant, the TSA demonstrated the need to draw up and perform new studies focusing on clarifying the relationship between environmental or occupational exposure to environmental pollutants and the changes mtDNAcn. Finally, the analysis of rate confidence in body evidence showed that the available information provides weak and insufficient information.
Environmental pollution has been associated with several human diseases and exercises influence over global mortality. According to the World Health Organization, air pollution, water pollution, and poisoning are significant environmental risks (WHO 2019). Induction of oxidative stress is considered a common molecular mechanism through which environmental pollutants induce/promote/potential the development of human diseases. Classical biomarkers of oxidative stress such as antioxidant status (analyzed measuring the activity/or relative mRNA expression of key antioxidant enzymes, or total GSH level, or GSH/GSSG ratio), oxidative status (estimated by measuring of ROS levels), or oxidative damage markers (such as lipid peroxidation, carbonylated proteins or oxidative adducts in DNA) have been used for evaluating the damage induced by exposure to environmental pollutants (Ghezzi 2020; Münzel and Daiber 2018). Recently, new molecular biomarkers such as miRNAs and lncRNAs have been analyzed to identify critical patterns of expression associated with exposure to environmental pollutants (Kotsyfakis and Patelarou 2019; Miguel et al. 2020). Therefore, the changes in the mtDNAcn in response to environmental pollutant exposure have been analyzed with contradictory results. For this reason, we performed a systematic review and meta-analysis following the recommendations of the NTP/OHAT 2019; the NTP/OHAT 2019 indicates the guidelines for researching, selecting, extracting, synthesizing the evidence, and rating the level of confidence in the body evidence. The result could be used in environmental sciences to support state-of-science or regulatory evaluation (Rooney et al. 2014).
Exploring sources of heterogeneity
In the present meta-analysis, the standardized mean differences were performed for evaluating the changes in mtDNAcn according to the level of environmental pollutant exposure. Four pooled analyses ((1) heavy metals, (2) cigarette smoking, (3) benzene and polycyclic aromatic hydrocarbons, and (4) particulate matter) were performed in the present meta-analysis. Nevertheless, some of the statistical results lack statistical significance, and some suffer from excessive heterogeneity. Subgroup analysis was not performed for cigarette smoking and heavy metals because of the small number of the included studies. The minimum number of studies for performing a subgroup analysis is five studies (Spineli and Pandis 2020). Nevertheless, the heterogeneity for cigarette smoking is considered low and insignificant; low levels of heterogeneity are associated with common variation of the included studies; the studies that analyzed the effect of cigarette smoking were performed in individuals with self-report of cigarette consumption. Moreover, almost included studies were considered with a similar risk of bias. In this line, the risk of bias estimated for each included study is regarded as a critical point directly associated with the heterogeneity. The inclusion of low-quality studies can potentially cause a low-quality conclusion. For this reason, we performed a subgroup analysis considering the final score of risk of bias according to the NTP/OHAT risk of bias tool. However, only the subgroup analysis of PM showed a reduction in heterogeneity; and surprising studies with probably low risk of bias showed low heterogeneity compared to studies with definitely low risk of bias. The assessment of environmental pollutant exposure is considered a high source of heterogeneity (Gong et al. 2022); nevertheless, the present study almost included studies that used methodologies that allow confidence in the environmental pollutant exposure. Moreover, heterogeneity in the meta-analysis is influenced by variations in individual human responses to environmental pollutant exposure. It is widely known that the variation in the outcome is associated with genetic polymorphism, age, and sex (Aldridge et al. 2003). Another point of consideration is that Egger’s funnel plot for cigarette smoking, heavy metals, and PM showed an asymmetrical distribution; the asymmetric distribution indicates the lack of negative results and leads to an overestimation of pooled results (Lin 2019). However, Egger’s regression shows non-evidence of small studies effects indicating that the obtained pooled results have been estimated based on studies with significant and insignificant results (Schwarzer et al. 2015).
Impact of tissue on the measuring of mtDNAcn
The whole blood was the most frequent biological matrix used for estimating the mtDNAcn because of accessible collection and relatively minimally invasive techniques. However, whole blood is a heterogeneous mix composing for 55% of blood plasma and 45% of the formed elements (approximately 98% of the formed elements are composed of erythrocytes, and the remaining 2% are covered for leukocytes and platelets together); typical results from leukocytes differential count are described below: neutrophils (50–70%), eosinophils (2–4%), basophils (0.5–1%), lymphocytes (25–45%), and monocytes (3–8%) (Marieb and Hoehn 2010). Platelets are more abundant than leukocytes (proportion 40:1); they contain mtDNA but lack nuclear DNA. Considering that the estimation of mtDNAcn is frequently calculated as mtDNA/nuclear DNA ratio, the number of platelets in whole blood could be used as an adjust factor from mtDNAcn for each studied individual (Hurtado-Roca et al. 2016). Typically, the use of buffy coat as a biological matrix results in lower levels of mtDNAcn than whole blood; reduction is attributed to a decrease in the high influence exercise for the number of platelets in the sample (Andreu et al. 2009). Another biological matrix used for determining the levels of mtDNAcn was the isolated peripheral blood mononuclear cells (PBMCs) using density gradients. However, PBMCs and platelets show similar relative density resulting in an overestimation of mtDNAcn attributed to platelet contamination (Urata et al. 2008). Platelet elimination before PBMCs obtention decreases the risk of overestimating mtDNAcn linked to platelet contamination (Banas et al. 2004).
Rausser et al. (2021) (using fluorescence-activated cell sorting) have reported other critical issues for consideration when blood is used for measuring the mtDNAcn: (1) mtDNAcn varies according to specific cell phenotype; for example, lymphocytes B show higher levels of mtDNAcn than neutrophils; (2) mtDNAcn displayed substantial weekly variation in the same individual (changes does not attribute to genetics, constitutional, or behavior factors); (3) serum lipid levels (that including triglycerides, total cholesterol, and low- or high-density lipoproteins) were correlated with substantial changes in mtDNAcn; (4) a sexual dimorphism for mtDNAcn is observed (men showed higher mtDNAcn in comparison with women); and (5) mtDNAcn increases according to age (for each decade an increase ~ 10% is observed in monocytes and naïve T cells); the above-cited results are based in isolated PBMCs. In whole blood or buffy coat, the increase of age was negatively associated with mtDNAcn; a marked hedging effect is observed in persons older than 50 years (Mengel-From et al. 2014; Verhoeven et al. 2018). However, a recent cohort study has shown that reducing mtDNAcn attributed to age could be explicitly accounted for by the platelet count and differential percentage in leukocytes (Moore et al. 2018). Neither of the included studies (using whole blood, buffy coat, or isolated PBMCs) in the meta-analysis considered a statistical adjustment using the platelet count. Only two included studies use the total leucocyte count (Alegría-Torres et al. 2020) or percentage of neutrophils (Hou et al. 2010) as statistical adjusters. Considering the current level of evidence on isolated mtDNAcn from blood, we suggest using PBMCs isolated with a previous step of platelet elimination with statistical adjustment that includes biological sex, neutrophil/lymphocyte ratio, age, and serum lipid levels (triglycerides and total cholesterol) for future studies. For example, suppose the authors used whole blood or buffy coat. In that case, they could include a statistical adjustment by biological sex, platelet count, differential percentage in leukocytes, age, and serum lipid levels (triglycerides and total cholesterol).
Other constraints to consider include that mtDNAcn isolated from blood could not always reflect a genuine change in mtDNAcn for another tissue or organ. Correlation analysis of mtDNAcn from different human tissues (that including blood, cerebellum, cerebrum, cortex, kidney, large intestine, liver, myocardial muscle, ovary, small intestine, skin, and skeletal muscle) showed that the mtDNAcn level from blood does not correlate with measured mtDNAcn levels for another organ/tissue (Wachsmuth et al. 2016). While most xenobiotics cross several barriers from the point of contact to target tissue/organ, a bloodstream step is required. For this reason, the toxicokinetic associated with a particular environmental pollutant could be considered before selecting the blood as an excellent biological matrix for the isolation of mtDNA.
Semen, oral mucosa, and somatic tissues (lung and placenta) were other tissues used to measure the mtDNAcn in the included studies. At the same time, a low and insignificant heterogeneity was linked to the use of these tissues. There are some issues to consider. As is well known, the number of mitochondria changes in accordance with tissue-specific energetic demands directly linked to the rate of oxidative phosphorylation activity. For example, the muscle, heart, and liver showed higher levels of mitochondria than the kidney and lung (D’Erchia et al. 2015). The use of lung tissue could be associated with low levels of mtDNAcn and marginal changes linked to environmental pollution exposure. Furthermore, the use of semen for measuring mtDNAcn is limited because mammalian spermatozoids are characterized by a low number of mitochondria (22 to 75) with low mtDNAcn (for example, human spermatozoa show a range of 2.8 to 226 mtDNAcn for each spermatozoon) (Song and Lewis 2008). In addition, semen contains ≈15% of nonspermatozoal cells (including immature germ cells, epithelial cells, and leukocytes) (Fedder 1996), which could cause a wrong estimation of real changes in spermatic mtDNAcn associated with environmental pollutant exposure. The selection of mitochondrial gen for measuring mtDNAcn in spermatozoids is another crucial topic. A study performed in isolated spermatozoids from horse semen showed that the use of cytochrome b (cytb) and ND4 (mitochondrially encoded NADH dehydrogenase 4) genes results in the overestimation of mtDNAcn in comparison with the use of ND1 (mitochondrially encoded NADH dehydrogenase 1) gen. The authors suggested that the localization of cytb and ND4 into mtDNA (near zones of origin of heavy strands replication and the origin of light strand replication) results in a high deletion rate of cytb and ND4. Furthermore, this fact is associated with an overestimation of mtDNAcn (Darr et al. 2017). We suggest using isolated spermatozoids to measure the impact of environmental pollutant exposure on spermatic mtDNAcn. If the authors use semen, we suggest performing a count of nonspermatozoal cells for statistical adjustment.
Moreover, the number of spermatozoa in both sceneries could be used as another statistical adjuster. There is no evidence of changes on mtDNAcn according to the class of epithelium cells (keratinocyte differentiation). However, an experimental study showed that the number and dimension of mitochondria decrease in the upper cell layer (Simpson et al. 2021). For this reason, the loss of mitochondria (from basal cells to differentiated cells) and the loss of cell nuclei (from differentiated cells to karyolytic cells) through the differentiation cycle of epithelial cells could be considered when the stratified squamous epithelium from oral mucosa is used for measuring the mtDNAcn.
The potential use of mtDNAcn as pollution biomarker
It is widely accepted that the ideal biomarker for characterizing the environmental pollution exposure would potentially have the following characteristics: (1) the use of biomarker has no ethical issues; (2) the obtention and analysis of biomarker are distinguished primarily by its minimal invasion and reliable results; (3) the biomarker is specific for a particular pollutant; (4) the changes in biomarker levels reflect reversible subclinical/clinical effects and predict or monitor patient response to intervention/treatment medical (Gil and Pla 2001). Therefore, mtDNAcn has shown an easy obtention without ethical implications. The mtDNAcn measurements use the latest technology and instruments (that include quantitative real-time PCR: qPCR; digital PCR: dPCR, and genotyping microarray probe intensities and DNA sequencing read counts such as whole-exome sequencing: WES, and whole-genome sequencing: WGS). Comparison of these techniques showed that WGS could be considered the gold standard for measuring the mtDNAcn compared to the current gold standard qPCR and dPCR. Furthermore, the authors showed a significant variation in mtDNAcn levels associated with the employed method for isolating DNA (Longchamps et al. 2020). The changes in mtDNAcn are not environmental pollutant-specific; several environmental pollutants induce change in the mtDNAcn (see supplementary table 5 for visualizing other studies). Moreover, the association between the changes in mtDNAcn levels and clinical effects has yet to be clarified. For example, an increase in mtDNAcn has been associated with the development of cardiovascular diseases (Ashar et al. 2017), and leukemia, myeloproliferative, and spleen diseases (Hägg et al. 2021), while neurodegenerative diseases are associated with a reduction in mtDNAcn (Yang et al. 2021). Changes in birth weight and cognitive/locomotor development are frequent clinical events studied about changes in placental mtDNAcn induced by air pollution; nevertheless, contradictory results are reported through published reports (Li et al. 2019). Another studied clinical event is the changes in blood pressure in response to air pollution and its association with the mtDNAcn. The reported results showed that subjects exposed to similar levels of air pollution showed a differential increment in blood pressure associated with mtDNAcn levels. Subjects with the highest mtDNAcn showed a minor increase in blood pressure (in response to air pollution) in comparison with the lowest mtDNAcn subjects (Zhong et al. 2016). Unfortunately, the currently available information is not able to state the facts between a clinic event and the changes in mtDNAcn induced by the exposure to environmental pollutants in a clear manner. Moreover, the biological effect of modifications in mtDNAcn levels is unclear. Early reports suggested that an increase in the mtDNAcn could be an indicator of mitochondrial biogenesis and/or a mechanism for compensating defects in the functionality of mitochondrial complexes that results in a decrease from the bioenergetic capacity of mitochondrial (Miyako et al. 1997; Renis et al. 1989). However, correlation analysis between mtDNAcn and mitochondrial content in skeletal muscle from healthy young subjects showed that mtDNAcn levels are not linked to mitochondrial content (Larsen et al. 2012). A similar result showed that VDAC1 levels (a biomarker of mitochondrial mass) are not correlated to mtDNAcn levels in human skeletal muscle and hippocampus (Brinckmann et al. 2010). In addition, the increase of mtDNAcn by transfection of transcriptional mtDNA factors (such as TFAM or Twinkle) in mice does not change the levels of mitochondrial complexes (biomarkers of mitochondrial activity) (Ikeda et al. 2015). For these reasons, an increase of mtDNAcn could not be considered a good standard to indirectly measure the mitochondrial status of biogenesis, content, or bioenergetic activity. Conversely, a reduction of mtDNAcn could be a positive indicator of mitochondria’s bioenergetic adaptation or biogenesis. For example, an increase in activity/relative mRNA levels and protein content from citrate synthase, cytochrome c oxidase subunit IV, and cytochrome c (biomarkers of bioenergetic functionality of mitochondrial) was observed in human skeletal muscle from subjects under short-term, aerobic exercise training. But mitochondrial fission regulator DRP-1 (dynamin-related protein 1) and mitochondrial fusion regulator Mfn (mitofusins) showed an increase in the protein content and relative mRNA levels, respectively. Surprisingly, a reduction in mtDNAcn levels was observed (Egan et al. 2013). Isolated human fibroblasts exposed to cigarette smoke condensate (at 80 μg/mL by 72 h) showed an increase in biomarkers of mitochondrial mass and biogenesis such as citrate synthase, the subunit A of succinate dehydrogenase, and nuclear respiratory factor 1; nevertheless, the mtDNAcn levels were reduced (Giordano et al. 2015). As can be seen, there is a need for performing new studies that show the molecular, cellular, biochemical, and biological repercussions linked to change in mtDNAcn levels.
A possible causal link between environmental pollutant exposure, oxidative stress, and mtDNAcn
Mitochondria are considered a subcellular target for many environmental pollutants; while the exact mechanism of environmental pollutant-induced mitochondrial toxicity is not fully known, it is considered to occur either as a consequence of the oxidative stress induction that results in mitochondrial dysfunction (Meyer et al. 2018). ROS regulates several cellular signaling pathways in normal physiological conditions, including nuclear-mitochondrial communication (Moldogazieva et al. 2018). It has been observed that under an oxidative stress state, the mtDNAcn increases in a dose-time-dependent manner (Lee et al. 2000). This fact could be explicated by activating the Keap1-Nrf2 pathway; the increase in cytoplasmic ROS promotes the oxidation of Keap1, abolishing its repressive function over Nrf2. Then, Nrf2 is translocated to the nucleus and promotes transcription of antioxidant response element and other genes (Deshmukh et al. 2017). Both nuclear transcriptional factors Nrf2 and Nrf1 are implicating in the biogenomic coordination between nuclear and mitochondrial genomes and the regulation of mitochondrial biogenesis (Goffart and Wiesner 2003). Moreover, the mtDNAcn levels could be regulated by Nrf1 and Nrf2; both factors can recognize consensus-binding sites in mitochondrial transcription factor A (mtTFA) (Virbasius and Scarpulla 1994); mtTNFA has been directly linked to regulation of the mtDNAcn in mammalian cells (Ekstrand et al. 2004). Considering that several environmental pollutants can induce an overactivation of transcriptional factors, including the Nrf1 and Nrf2 (Jennings et al. 2013; Venugopal and Jaiswal 1998), it is possible that the increase in mtDNAcn could be considered the result of environmental pollutant–induced activation of Nrf1 and Nrf2. However, more mechanistic studies are required to verify this proposed theory.
Environmental pollutant’s affinity for mtDNA
Another point to consider is the significant variation in the affinity to mtDNA over nuclear DNA. For example, early reports showed that PAHs have a high affinity to produce adducts in mtDNA in comparison with nuclear DNA (Allen and Coombs 1980); it results in a large adducts formation in mtDNA approximately 40- to 90-fold higher in comparison with nuclear DNA (Backer and Weinstein 1980); on the other hand, a differential affinity for mtDNA or nuclear DNA has not been proven for heavy metals; nevertheless, these are associated with the induction of oxidative stress and the formation of oxidative adducts in DNA (8-OHdG mainly), and decrease the expression of key enzymes involved in DNA repair mechanism (OGG1 mainly) (Al Bakheet et al. 2013); similar mechanisms are linked to particulate matter exposure; the exposure of human endothelial cells to PM results in an increase in mitochondrial ROS, and the disrupt of key mitochondrial functions such as regulation of both mitochondrial biogenesis and fission (Wang et al. 2020). These facts could explain the non-systematic effect of environmental pollutant exposure on mtDNAcn.
Limitations
In the present work, we analyzed the effect of exposure to environmental pollutants in changes on mtDNAcn levels. We performed a comprehensive search strategy in the most appropriate databases. And the literature selection, risk of bias evaluation, and rate confidence in the body of evidence were performed employing the OHAT’s tools because these tools were developed to be used in systematic review and meta-analysis with environmental epidemiology implications. There are limitations associated with our study: (1) the obtained result derivatives of subgroups that contain heterogeneous environmental pollutants; (2) some included studies do not perform a complete characterization of exposure to environmental pollutants; (3) most included studies were performed using a cross-sectional design that increases the risk of bias; (4) the obtained results do not represent a dose–response effect because the pooled analysis was carried out considered only the lowest level of exposure versus highest level. The mentioned limitations should be considered when interpreting the obtained results.
Conclusions
The systematic review and meta-analysis show evidence that exposure to environmental pollutants induces marginal changes on mtDNAcn levels. Nevertheless, the data obtained in this study also suggest the need to perform new studies with mechanistic or cohort designs are required to elucidate the biological effect associated with the changes in mtDNAcn. And its possible use as a biomarker linked to environmental pollution exposure.
Data availability
All the data and material pertinent to this manuscript are included and have been reviewed by all authors.
References
Al Bakheet SA, Attafi IM, Maayah ZH et al (2013) Effect of long-term human exposure to environmental heavy metals on the expression of detoxification and DNA repair genes. Environ Pollut 181:226–232. https://doi.org/10.1016/j.envpol.2013.06.014
Aldridge JE, Gibbons JA, Flaherty MM et al (2003) Heterogeneity of toxicant response: sources of human variability. Toxicol Sci 76:3–20. https://doi.org/10.1093/toxsci/kfg204
Alegría-Torres JA, Pérez-Rodríguez RY, García-Torres L et al (2020) Exposure to arsenic and lead in children from Salamanca México, effects on telomeric lengthening and mitochondrial DNA. Environ Sci Pollut Res Int 27:6420–6428. https://doi.org/10.1007/s11356-019-07108-4
Allen JA, Coombs MM (1980) Covalent binding of polycyclic aromatic compounds to mitochondrial and nuclear DNA. Nature 287:244–245. https://doi.org/10.1038/287244a0
Ameer SS, Xu Y, Engström K et al (2016) Exposure to inorganic arsenic is associated with increased mitochondrial DNA Copy Number and longer telomere length in peripheral blood. Front Cell Dev Biol 4:87. https://doi.org/10.3389/fcell.2016.00087
Andreu AL, Martinez R, Marti R, García-Arumí E (2009) Quantification of mitochondrial DNA copy number: pre-analytical factors. Mitochondrion 9:242–246. https://doi.org/10.1016/j.mito.2009.02.006
Ashar FN, Zhang Y, Longchamps RJ et al (2017) Association of mitochondrial DNA copy number with cardiovascular disease. JAMA Cardiol 2:1247–1255. https://doi.org/10.1001/jamacardio.2017.3683
Backer JM, Weinstein IB (1980) Mitochondrial DNA is a major cellular target for a dihydrodiol-epoxide derivative of benzo[a]pyrene. Science 209:297–299. https://doi.org/10.1126/science.6770466
Bai R-K, Perng C-L, Hsu C-H, Wong L-JC (2004) Quantitative PCR analysis of mitochondrial DNA content in patients with mitochondrial disease. Ann N Y Acad Sci 1011:304–309. https://doi.org/10.1007/978-3-662-41088-2_29
Balali-Mood M, Naseri K, Tahergorabi Z et al (2021) Toxic mechanisms of five heavy metals: mercury, lead, chromium, cadmium, and arsenic. Front Pharmacol 12:227. https://doi.org/10.3389/fphar.2021.643972
Banas B, Kost BP, Goebel FD (2004) Platelets, a typical source of error in real-time PCR quantification of mitochondrial DNA content in human peripheral blood cells. Eur J Med Res 9:371–377
Barshad G, Marom S, Cohen T, Mishmar D (2018) Mitochondrial DNA transcription and its regulation: an evolutionary perspective. Trends Genet TIG 34:682–692. https://doi.org/10.1016/j.tig.2018.05.009
Borenstein M (ed) (2009) Introduction to meta-analysis. John Wiley & Sons, Chichester, U.K
Bouhours-Nouet N, May-Panloup P, Coutant R et al (2005) Maternal smoking is associated with mitochondrial DNA depletion and respiratory chain complex III deficiency in placenta. Am J Physiol-Endocrinol Metab 288:E171–E177. https://doi.org/10.1152/ajpendo.00260.2003
Brinckmann A, Weiss C, Wilbert F et al (2010) Regionalized pathology correlates with augmentation of mtDNA copy numbers in a patient with myoclonic epilepsy with ragged-red fibers (MERRF-syndrome). PLoS ONE 5:e13513. https://doi.org/10.1371/journal.pone.0013513
Carugno M, Pesatori AC, Dioni L et al (2012) Increased mitochondrial DNA copy number in occupations associated with low-dose benzene exposure. Environ Health Perspect 120:210–215. https://doi.org/10.1289/ehp.1103979
Castellani CA, Longchamps RJ, Sun J et al (2020) Thinking outside the nucleus: mitochondrial DNA copy number in health and disease. Mitochondrion 53:214–223. https://doi.org/10.1016/j.mito.2020.06.004
da Costa Santos CM, de Mattos Pimenta CA, Nobre MRC (2007) The PICO strategy for the research question construction and evidence search. Rev Lat Am Enfermagem 15:508–511. https://doi.org/10.1590/s0104-11692007000300023
Darr CR, Moraes LE, Connon RE et al (2017) The relationship between mitochondrial DNA copy number and stallion sperm function. Theriogenology 94:94–99. https://doi.org/10.1016/j.theriogenology.2017.02.015
D’Erchia AM, Atlante A, Gadaleta G et al (2015) Tissue-specific mtDNA abundance from exome data and its correlation with mitochondrial transcription, mass and respiratory activity. Mitochondrion 20:13–21. https://doi.org/10.1016/j.mito.2014.10.005
Deshmukh P, Unni S, Krishnappa G, Padmanabhan B (2017) The Keap1-Nrf2 pathway: promising therapeutic target to counteract ROS-mediated damage in cancers and neurodegenerative diseases. Biophys Rev 9:41–56. https://doi.org/10.1007/s12551-016-0244-4
Du J, Pan B, Cao X et al (2020) Urinary polycyclic aromatic hydrocarbon metabolites, peripheral blood mitochondrial DNA copy number, and neurobehavioral function in coke oven workers. Chemosphere 261:127628. https://doi.org/10.1016/j.chemosphere.2020.127628
Duan X, Yang Y, Zhang H et al (2020) Polycyclic aromatic hydrocarbon exposure, miRNA genetic variations, and associated leukocyte mitochondrial DNA copy number: a cross-sectional study in China. Chemosphere 246:125773. https://doi.org/10.1016/j.chemosphere.2019.125773
Egan B, O’Connor PL, Zierath JR, O’Gorman DJ (2013) Time course analysis reveals gene-specific transcript and protein kinetics of adaptation to short-term aerobic exercise training in human skeletal muscle. PLoS ONE 8:e74098. https://doi.org/10.1371/journal.pone.0074098
Ekstrand MI, Falkenberg M, Rantanen A et al (2004) Mitochondrial transcription factor A regulates mtDNA copy number in mammals. Hum Mol Genet 13:935–944. https://doi.org/10.1093/hmg/ddh109
Fan Z, Lin L (2011) Exposure science: contaminant mixtures. In: Nriagu J (ed) Encyclopedia of environmental health, 2nd edn. Elsevier, Oxford, pp 805–815
Fedder J (1996) Nonsperm cells in human semen: with special reference to seminal leukocytes and their possible influence on fertility. Arch Androl 36:41–65. https://doi.org/10.3109/01485019608987883
Ghezzi P (2020) Environmental risk factors and their footprints in vivo – a proposal for the classification of oxidative stress biomarkers. Redox Biol 34:101442. https://doi.org/10.1016/j.redox.2020.101442
Gil F, Pla A (2001) Biomarkers as biological indicators of xenobiotic exposure. J Appl Toxicol JAT 21:245–255. https://doi.org/10.1002/jat.769
Giordano L, Deceglie S, d’Adamo P et al (2015) Cigarette toxicity triggers Leber’s hereditary optic neuropathy by affecting mtDNA copy number, oxidative phosphorylation and ROS detoxification pathways. Cell Death Dis 6:e2021. https://doi.org/10.1038/cddis.2015.364
Goffart S, Wiesner RJ (2003) Regulation and coordination of nuclear gene expression during mitochondrial biogenesis. Exp Physiol 88:33–40. https://doi.org/10.1113/eph8802500
Gong C, Wang J, Bai Z et al (2022) Maternal exposure to ambient PM2.5 and term birth weight: A systematic review and meta-analysis of effect estimates. Sci Total Environ 807:150744. https://doi.org/10.1016/j.scitotenv.2021.150744
Hägg S, Jylhävä J, Wang Y et al (2021) Deciphering the genetic and epidemiological landscape of mitochondrial DNA abundance. Hum Genet 140:849–861. https://doi.org/10.1007/s00439-020-02249-w
Herbers E, Kekäläinen NJ, Hangas A et al (2019) Tissue specific differences in mitochondrial DNA maintenance and expression. Mitochondrion 44:85–92. https://doi.org/10.1016/j.mito.2018.01.004
Higgins JPT, Thompson SG (2002) Quantifying heterogeneity in a meta-analysis. Stat Med 21:1539–1558. https://doi.org/10.1002/sim.1186
Hou L, Zhang X, Dioni L et al (2013) Inhalable particulate matter and mitochondrial DNA copy number in highly exposed individuals in Beijing, China: a repeated-measure study. Part Fibre Toxicol 10:17. https://doi.org/10.1186/1743-8977-10-17
Hou L, Zhu Z-Z, Zhang X et al (2010) Airborne particulate matter and mitochondrial damage: a cross-sectional study. Environ Health Glob Access Sci Source 9:48. https://doi.org/10.1186/1476-069X-9-48
Hurtado-Roca Y, Ledesma M, Gonzalez-Lazaro M et al (2016) Adjusting MtDNA quantification in whole blood for peripheral blood platelet and leukocyte counts. PLoS ONE 11:e0163770. https://doi.org/10.1371/journal.pone.0163770
Ikeda M, Ide T, Fujino T et al (2015) Overexpression of TFAM or Twinkle increases mtDNA copy number and facilitates cardioprotection associated with limited mitochondrial oxidative stress. PLoS ONE 10:e0119687. https://doi.org/10.1371/journal.pone.0119687
Jennings P, Limonciel A, Felice L, Leonard MO (2013) An overview of transcriptional regulation in response to toxicological insult. Arch Toxicol 87:49–72. https://doi.org/10.1007/s00204-012-0919-y
Kotsyfakis M, Patelarou E (2019) MicroRNAs as biomarkers of harmful environmental and occupational exposures: a systematic review. Biomark Biochem Indic Expo Response Susceptibility Chem 24:623–630. https://doi.org/10.1080/1354750X.2019.1652348
Larsen S, Nielsen J, Hansen CN et al (2012) Biomarkers of mitochondrial content in skeletal muscle of healthy young human subjects. J Physiol 590:3349–3360. https://doi.org/10.1113/jphysiol.2012.230185
Lee H-C, Lu C-Y, Fahn H-J, Wei Y-H (1998) Aging- and smoking-associated alteration in the relative content of mitochondrial DNA in human lung. FEBS Lett 441:292–296. https://doi.org/10.1016/S0014-5793(98)01564-6
Lee HC, Yin PH, Lu CY et al (2000) Increase of mitochondria and mitochondrial DNA in response to oxidative stress in human cells. Biochem J 348:425–432
Lehman-McKeeman LD, Armstrong LE (2022) Chapter 2 - Biochemical and molecular basis of toxicity. In: Haschek WM, Rousseaux CG, Wallig MA, Bolon B (eds) Haschek and Rousseaux’s handbook of toxicologic pathology (fourth edition). Academic Press, pp 15–49
Li Z, Fu J, Li Z et al (2019) Air pollution and placental mitochondrial DNA copy number: mechanistic insights and epidemiological challenges. Environ Pollut 255:113266. https://doi.org/10.1016/j.envpol.2019.113266
Li A, Sun Y, Wang T et al (2020a) Effects of micronucleus frequencies and mitochondrial DNA copy numbers among benzene-exposed workers in China. Environ Mol Mutagen 61:355–360. https://doi.org/10.1002/em.22354
Li Z, Liu H, Qian Y et al (2020b) Influence of metals from e-waste dismantling on telomerelength and mitochondrial DNA copy number in people living near recycling sites. Environ Int 140:105769. https://doi.org/10.1016/j.envint.2020.105769
Lin L (2019) Graphical augmentations to sample-size-based funnel plot in meta-analysis. Res Synth Methods 10:376–388. https://doi.org/10.1002/jrsm.1340
Ling X, Zhang G, Sun L et al (2017) Polycyclic aromatic hydrocarbons exposure decreased sperm mitochondrial DNA copy number: a cross-sectional study (MARHCS) in Chongqing, China. Environ Pollut Barking Essex 1987 220:680–687. https://doi.org/10.1016/j.envpol.2016.10.026
Linqing Y, Bo X, Xueqin Y et al (2016) Mitochondrial DNA hypomethylation in chrome plating workers. Toxicol Lett 243:1–6. https://doi.org/10.1016/j.toxlet.2015.11.031
Longchamps RJ, Castellani CA, Yang SY et al (2020) Evaluation of mitochondrial DNA copy number estimation techniques. PLoS ONE 15:e0228166. https://doi.org/10.1371/journal.pone.0228166
Marieb EN, Hoehn K (2010) Human anatomy & physiology. Benjamin Cummings, San Francisco
Masayesva BG, Mambo E, Taylor RJ et al (2006) Mitochondrial DNA content increase in response to cigarette smoking. Cancer Epidemiol Biomark Prev Publ Am Assoc Cancer Res Cosponsored Am Soc Prev Oncol 15:19–24. https://doi.org/10.1158/1055-9965.EPI-05-0210
Mengel-From J, Thinggaard M, Dalgård C et al (2014) Mitochondrial DNA copy number in peripheral blood cells declines with age and is associated with general health among elderly. Hum Genet 133:1149–1159. https://doi.org/10.1007/s00439-014-1458-9
Meyer JN, Hartman JH, Mello DF (2018) Mitochondrial toxicity. Toxicol Sci 162:15–23. https://doi.org/10.1093/toxsci/kfy008
Meyer JN, Leung MCK, Rooney JP et al (2013) Mitochondria as a target of environmental toxicants. Toxicol Sci 134:1–17. https://doi.org/10.1093/toxsci/kft102
Meyer JN, Leuthner TC, Luz AL (2017) Mitochondrial fusion, fission, and mitochondrial toxicity. Toxicology 391:42–53. https://doi.org/10.1016/j.tox.2017.07.019
Miguel V, Lamas S, Espinosa-Diez C (2020) Role of non-coding-RNAs in response to environmental stressors and consequences on human health. Redox Biol 37:101580. https://doi.org/10.1016/j.redox.2020.101580
Miyako K, Kai Y, Irie T et al (1997) The content of intracellular mitochondrial DNA is decreased by 1-methyl-4-phenylpyridinium ion (MPP+). J Biol Chem 272:9605–9608. https://doi.org/10.1074/jbc.272.15.9605
Moldogazieva NT, Mokhosoev IM, Feldman NB, Lutsenko SV (2018) ROS and RNS signalling: adaptive redox switches through oxidative/nitrosative protein modifications. Free Radic Res 52:507–543. https://doi.org/10.1080/10715762.2018.1457217
Moore AZ, Ding J, Tuke MA et al (2018) Influence of cell distribution and diabetes status on the association between mitochondrial DNA copy number and aging phenotypes in the InCHIANTI study. Aging Cell 17. https://doi.org/10.1111/acel.12683
Münzel T, Daiber A (2018) Environmental stressors and their impact on health and disease with focus on oxidative stress. Antioxid Redox Signal 28:735–740. https://doi.org/10.1089/ars.2017.7488
National Toxicology Program (2019) Handbook for conducting a literature-based health assessment using OHAT approach for systematic review and evidence integration. https://ntp.niehs.nih.gov/whatwestudy/assessments/noncancer/handbook/index.html. Accessed 21 Oct 2020
Organization WH (2019) World health statistics 2019: monitoring health for the SDGs, sustainable development goals. World Health Organization
Page MJ, McKenzie JE, Bossuyt PM et al (2021) The PRISMA 2020 statement: an updated guideline for reporting systematic reviews. BMJ 372:n71. https://doi.org/10.1136/bmj.n71
Pavanello S, Dioni L, Hoxha M et al (2013) Mitochondrial DNA copy number and exposure to polycyclic aromatic hydrocarbons. Cancer Epidemiol Biomark Prev Publ Am Assoc Cancer Res Cosponsored Am Soc Prev Oncol 22:1722–1729. https://doi.org/10.1158/1055-9965.EPI-13-0118
Pustejovsky JE, Rodgers MA (2019) Testing for funnel plot asymmetry of standardized mean differences. Res Synth Methods 10:57–71. https://doi.org/10.1002/jrsm.1332
Rausser S, Trumpff C, McGill MA, et al (2021) Mitochondrial phenotypes in purified human immune cell subtypes and cell mixtures
Renis M, Cantatore P, Loguercio Polosa P et al (1989) Content of mitochondrial DNA and of three mitochondrial RNAs in developing and adult rat cerebellum. J Neurochem 52:750–754. https://doi.org/10.1111/j.1471-4159.1989.tb02518.x
Rooney AA, Boyles AL, Wolfe MS et al (2014) Systematic review and evidence integration for literature-based environmental health science assessments. Environ Health Perspect 122:711–718. https://doi.org/10.1289/ehp.1307972
Roubicek DA, de Souza-Pinto NC (2017) Mitochondria and mitochondrial DNA as relevant targets for environmental contaminants. Toxicology 391:100–108. https://doi.org/10.1016/j.tox.2017.06.012
Schwarzer G, Carpenter J, Rücker G (2015) Small-study effects in meta-analysis. In: Schwarzer G, Carpenter JR, Rücker G (eds) Meta-analysis with R. Springer International Publishing, pp 107–141
Shah A, Smith AF (2020) Trial sequential analysis: adding a new dimension to meta-analysis. Anaesthesia 75:15–20. https://doi.org/10.1111/anae.14705
Shen M, Zhang L, Bonner MR et al (2008) Association between mitochondrial DNA copy number, blood cell counts, and occupational benzene exposure. Environ Mol Mutagen 49:453–457. https://doi.org/10.1002/em.20402
Simpson CL, Tokito MK, Uppala R et al (2021) NIX initiates mitochondrial fragmentation via DRP1 to drive epidermal differentiation. Cell Rep 34:108689. https://doi.org/10.1016/j.celrep.2021.108689
Song GJ, Lewis V (2008) Mitochondrial DNA integrity and copy number in sperm from infertile men. Fertil Steril 90:2238–2244. https://doi.org/10.1016/j.fertnstert.2007.10.059
Spineli LM, Pandis N (2020) Exploring heterogeneity in meta-analysis: Meta-regression analysis. Am J Orthod Dentofacial Orthop 158:623–625. https://doi.org/10.1016/j.ajodo.2020.07.002
Tan D, Goerlitz DS, Dumitrescu RG et al (2008) Associations between cigarette smoking and mitochondrial DNA abnormalities in buccal cells. Carcinogenesis 29:1170–1177. https://doi.org/10.1093/carcin/bgn034
Ukaogo PO, Ewuzie U, Onwuka CV (2020) 21 - Environmental pollution: causes, effects, and the remedies. In: Chowdhary P, Raj A, Verma D, Akhter Y (eds) Microorganisms for sustainable environment and health. Elsevier, pp 419–429
Urata M, Koga-Wada Y, Kayamori Y, Kang D (2008) Platelet contamination causes large variation as well as overestimation of mitochondrial DNA content of peripheral blood mononuclear cells. Ann Clin Biochem 45:513–514. https://doi.org/10.1258/acb.2008.008008
Venugopal R, Jaiswal AK (1998) Nrf2 and Nrf1 in association with Jun proteins regulate antioxidant response element-mediated expression and coordinated induction of genes encoding detoxifying enzymes. Oncogene 17:3145–3156. https://doi.org/10.1038/sj.onc.1202237
Verhoeven JE, Révész D, Picard M et al (2018) Depression, telomeres and mitochondrial DNA: between- and within-person associations from a 10-year longitudinal study. Mol Psychiatry 23:850–857. https://doi.org/10.1038/mp.2017.48
Virbasius JV, Scarpulla RC (1994) Activation of the human mitochondrial transcription factor A gene by nuclear respiratory factors: a potential regulatory link between nuclear and mitochondrial gene expression in organelle biogenesis. Proc Natl Acad Sci U S A 91:1309–1313. https://doi.org/10.1073/pnas.91.4.1309
Wachsmuth M, Hübner A, Li M et al (2016) Age-related and heteroplasmy-related variation in human mtDNA copy number. PLOS Genet 12:e1005939. https://doi.org/10.1371/journal.pgen.1005939
Wang M, Zhao J, Wang Y et al (2020) Genome-wide DNA methylation analysis reveals significant impact of long-term ambient air pollution exposure on biological functions related to mitochondria and immune response. Environ Pollut Barking Essex 1987 264:114707. https://doi.org/10.1016/j.envpol.2020.114707
Wetterslev J, Jakobsen JC, Gluud C (2017) Trial sequential analysis in systematic reviews with meta-analysis. BMC Med Res Methodol 17:39. https://doi.org/10.1186/s12874-017-0315-7
Wong JYY, Hu W, Downward GS et al (2017) Personal exposure to fine particulate matter and benzo[a]pyrene from indoor air pollution and leukocyte mitochondrial DNA copy number in rural China. Carcinogenesis 38:893–899. https://doi.org/10.1093/carcin/bgx068
Xu Y, Li H, Hedmer M et al (2017) Occupational exposure to particles and mitochondrial DNA - relevance for blood pressure. Environ Health 16:22. https://doi.org/10.1186/s12940-017-0234-4
Xu Y, Lindh CH, Jönsson BAG et al (2018) Occupational exposure to asphalt mixture during road paving is related to increased mitochondria DNA copy number: a cross-sectional study. Environ Health Glob Access Sci Source 17:29. https://doi.org/10.1186/s12940-018-0375-0
Yang L, Xia B, Yang X et al (2016) Mitochondrial DNA hypomethylation in chrome plating workers. Toxicol Lett 243:1–6. https://doi.org/10.1016/j.toxlet.2015.11.031
Zhang G, Jiang F, Chen Q et al (2020) Associations of ambient air pollutant exposure with seminal plasma MDA, sperm mtDNA copy number, and mtDNA integrity. Environ Int 136:105483. https://doi.org/10.1016/j.envint.2020.105483
Zhong J, Cayir A, Trevisi L et al (2016) Traffic-related air pollution, blood pressure, and adaptive response of mitochondrial abundance. Circulation 133:378–387. https://doi.org/10.1161/CIRCULATIONAHA.115.018802
Acknowledgements
The authors would like to thank MPS. Fabiola Carolina Olguin Higuera from Facultad de ciencias políticas y sociales for her support and cooperation on external access to UNAM database collection. During the study, Cristian Aviles-Ramirez (CVU:1043502) was a grant recipient of the Consejo Nacional de Ciencia y Tecnología (CONACyT) of Mexico.
Author information
Authors and Affiliations
Contributions
Conceptualization: Marco Antonio Ramírez-Vargas. Investigation: Marco Antonio Ramírez-Vargas and Cristian Avilés-Ramírez. Formal analysis: Eugenia Flores-Alfaro and Mónica Ramírez. Visualization: Cristian Avilés-Ramírez and Gerardo Huerta-Beristain. Writing–original draft: Ma. Elena Moreno-Godínez and Marco Antonio Ramírez-Vargas. Writing–review and editing: Isela Parra-Rojas and Matthew R. Bonner.
Corresponding author
Ethics declarations
Ethics approval
Not applicable.
Consent to participate
Not applicable.
Consent for publication
Not applicable.
Competing interests
The authors declare no competing interests.
Additional information
Responsible Editor: Robert Duran
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Avilés-Ramírez, C., Moreno-Godínez, M.E., Bonner, M.R. et al. Effects of exposure to environmental pollutants on mitochondrial DNA copy number: a meta-analysis. Environ Sci Pollut Res 29, 43588–43606 (2022). https://doi.org/10.1007/s11356-022-19967-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11356-022-19967-5