Abstract
The wide usage of neodymium (Nd) in industry, agriculture, and medicine has made it become an emerging pollutant in the environment. Increasing Nd pollution has potential hazards to plants, animals, and microorganisms. Thus, it is necessary to study the toxicity of Nd and the mechanism of Nd transportation and detoxification in microorganisms. Through genome-scale screening, we identified 70 yeast monogene deletion mutations sensitive to Nd ions. These genes are mainly involved in metabolism, transcription, protein synthesis, cell cycle, DNA processing, protein folding, modification, and cell transport processes. Furthermore, the regulatory networks of Nd toxicity were identified by using the protein interaction group analysis. These networks are associated with various signal pathways, including calcium ion transport, phosphate pathways, vesicular transport, and cell autophagy. In addition, the content of Nd ions in yeast was detected by an inductively coupled plasma mass spectrometry, and most of these Nd-sensitive mutants showed an increased intracellular Nd content. In all, our results provide the basis for understanding the molecular mechanisms of detoxifying Nd ions in yeast cells, which will be useful for future studies on Nd-related issues in the environment, agriculture, and human health.
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
Introduction
Rare earth elements include 17 rare earth elements, which are divided as light or heavy rare earth elements according to the atomic mass (Mowafy 2020; Pagano et al. 2019). Rare earth neodymium (Nd) is light rare earth, which is mainly present in the form of oxides, nitrate, and chloride in nature (Thornton and Burdette 2017). Nd is widely used in various fields such as medicine, agriculture, NdFB permanent magnet materials, electronics, and mechanical industries for its excellent performance (Gohda 2021). Moreover, Nd is also applied in non-ferrous metal materials, ceramic, glass materials, and rubber products (Bär et al. 2019; Zhang et al. 2007; Alarfaj et al. 2018).
It has been observed that the concentration of Nd is 0.0071 ~ 6.68 μg/L in the surface water of the Terengganu River Basin in Malaysia (Sultan and Shazili 2009). It is 771 μg/L Nd in streams draining from acid sulfate soils during high-water flow events in autumn in Finland (Åström 2001). In addition, the concentration of Nd is 10.8 μg/L in the surface water of an ex-mining pit lake in Malaysia (Khan et al. 2017), and it is 317 μg/L in a mining pit in Wisniowka (Poland) (Migaszewski et al. 2016). Previously, it was found that Nd3+ was a toxic element for the fungi Penicillium simplicissimum and Aspergillus japonicus. The combinations of Nd3+ and Sm3+ was the most toxic combination for the tested organisms and fungi (Bergsten-Torralba et al. 2020). The distribution of Nd was studied in the different species of mushrooms collected from unpolluted areas in the province of Ciudad Real, Central Spain, and the maximum absorption of Nd was 7.10 μg/L (Campos et al. 2009). In addition, the significant accumulation of Nd was detected in Bacillus cereus isolated from rare earth environments in India (Challaraj Emmanuel et al. 2011).
In addition, the positive effect of Nd has been observed on seed germination, root development, nutrient absorption, fruit weight, and fruit composition. Thus, it is widely used as a rare earth microfertilizer and plant growth regulator in agriculture (Wang et al. 2003; Ren et al. 2016). However, the wide application of Nd in various fields causes Nd to continuously enter the environment, which will result in soil and water pollution. Furthermore, Nd can be accumulated in the human body (Lemoine et al. 2019). The results of a previous study have shown that the content of Nd in the human blood, urine, and hair samples in the rare earth mining area of Baotou city of China has far exceeded that of the non-mining area (Bao et al. 2018). In addition, Nd can enter the human body through the respiratory system, digestive system, and skin, which will finally damage the lung, liver, central nervous system, and blood system (Zhuang et al. 2017; Wang et al. 2011). However, the mechanism of action and transport of Nd in living organisms have not been fully understood until recently.
As a fully sequencing eukaryotic cell, yeast Saccharomyces cerevisiae is a good experimental model for understanding eukaryotic systems, especially metal ion toxicity (Gerwien et al. 2018). Various studies on the genes and signal pathways related to metal ion stress have been investigated, and numerous signaling pathways and ion transporters involved in the metal ion stress have been found (Cyert and Philpott 2013; Bleackley and Macgillivray 2011). To fully understand the mechanism of Nd in eukaryotic cells, we studied the toxicity of Nd and screened the yeast double haploid gene missing library and analyzed the genes affected by Nd in this study. The results will provide the basis for understanding the molecular mechanisms of detoxifying Nd ions in yeast cells, which is useful for future studies on Nd-related issues in the environment, agriculture, and human health.
Material and methods
Yeast strains and culture
The diploid S. cerevisiae strains were derived from the BY4743 genetic background coming from a public collection. All gene mutant strains had KanMX4 genetic labels with G418 resistant, and the yeast library was purchased from Invitrogen Inc (USA). Yeast was grown at 30 °C in the YPD medium (1% yeast extract, 2% peptone, and 2% glucose, pH 5.6). Nd(NO3)3 and NaNO3 were purchased from Aladdin Co., Ltd (Shanghai, China).
Screening Nd-sensitive phenotype of yeast
Based on our preliminary experiments, we firstly tested Nd toxicity on fungi from 1 to 5 mM, then selected 3 to 4 mM during two days according to the method of a previous study (Luo et al. 2016). Finally, 3.7 mM Nd nitrate was used to screen the Nd-sensitive phenotype of yeast. All strains in the library were used for preliminary screening of Nd-sensitive mutations. We copied all strains to a medium containing or not containing 3.7 mM Nd nitrate by using 384 photographs. The tablet was incubated at 30 ℃ for 2 days, photographed, and used to analyze the growth of each mutant. Compared to the other surrounding mutants, it is considered Nd-sensitive if its colony size is reduced by more than 30% on the YPD media containing Nd nitrate based on the analysis of software and observation. Moreover, there is no change of colony size in YPD media containing no Nd.
In addition, the secondary screening was performed for Nd-sensitive mutants by continuous Nd dilution determination according to a previous method (Jiang et al. 2014). The sensitive mutants were streak cultured from the original library to the YPD medium (pH 5.6) and inoculated into the YPD medium for 12 h. A series of culture diluents (10–0 ~ 10–4) were put on the YPD tablets containing 0, 11.1 mM NaNO3, or 3.7 mM Nd(NO3)3, respectively. The 11.1 mM sodium nitrate was used for wild-type comparison to confirm that the mutant was sensitive to Nd3+ rather than NO3− (Luo et al. 2016). The dishes were cultured for 2 ~ 5 days at 30 ℃ and the secondary screening experiment was repeated three times.
Detecting the intracellular Nd concentration
For 3.7 mM Nd significantly inhibited the yeast growth, logarithmically growing yeast mutant cells and wild-type cells were treated with 1.85 mM Nd (NO3)3 for 4 h at 30 ℃. Then we collected fungi, measured and recorded its OD value (Zhao et al. 2013). An inductively coupled plasma mass spectrometer is used for detecting the intracellular Nd concentration. Three single colonies of each mutant were measured and wild type BY4743 was used as the control.
Test the missing strain genome by PCR
The yeast missing strain genome is extracted and the missing strain genome is used as a template for PCR test. Its upstream primer is designed within 100–300 bp upstream of the mutant gene open reading box. The downstream primer is designed at the internal sequence of the KanMX4 knockout box, which is about 1200 bp from the gene initiation codon. The primers used for testing the missing strain genome are listed in Supplementary Table 1.
Gene function and localization analysis
The function and localization of the corresponding genes of Nd-sensitive mutants (http://www.yeastgenome.org), MIPS, BioGRID (http://www.thebiogrid.org), and FunSpec (http://funspec.med.utoronto.ca) were annotated using the yeast genome database.
Data analysis
Data was expressed as mean ± SEM; the statistical software SPSS (16.0) was used for analysis. The difference among the experimental groups was analyzed with the single factor variance analysis (LSD's test) analysis, and the significance level is set at P < 0.05.
Results
Sensitive mutants to Nd 3+ and the associated genes
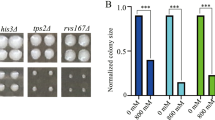
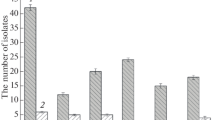
In this experiment, we firstly tested the sensitivity of diploid wild type BY4743 to Nd(NO3)3. After cultivating yeast cells with different concentration gradients of Nd(NO3)3 medium, we found that yeast cells had specific sensitivity at 3.7 mM Nd(NO3)3 (Fig. 1). Therefore, we used 3.7 mM Nd(NO3)3 to screen the diploid yeast mutant library, and we confirmed a total of 70 Nd sensitive mutants (Fig. 2). In addition, the genotype of these mutants was further tested by PCR (Supplementary Fig. 1).
Phenotypes of Nd-sensitive gene deletion mutants. Cells of the wild-type BY4743 and 70 gene deletion mutants were identified from the genome-scale screen, which were grown at 30℃ in liquid YPD overnight, serially diluted by 10 times and spotted on YPD plates with or without supplemented reagents as indicated, respectively. Plates were incubated for 1 ~ 5 days at 30℃
By functional analysis and subcellular localization of the derived genes through Saccharomyces Genome Database (SGD) and MIPS-related network resources, we found that the functional classification of these 70 genes includes 10 unknown function genes and 60 known function genes. In addition, 70 genes can be divided into the following 8 categories: A group, transcription (4 genes); B group, protein synthesis (5 genes); C group, protein fate (8 genes); D group, cell cycle and DNA processing (5 genes); E group, cellular transport, transport facilities and transport routes (30 genes); F group, metabolism (6 genes); G group, cell rescue, defense and virulence (2 genes); H group, unknown function genes (10 genes). In addition, the proteins encoded by these 60 genes are mainly localized in the nucleus (10 genes), vacuole (7 genes), and cytoplasm (18 genes). Some of them are present in the endoplasmic reticulum (ER) (6 genes), Golgi body (10 genes), mitochondrial (3 genes), endobody (1 genes), and plasma membrane (7 genes). These genes are listed in Table 1, and the functional notes for each gene are described in Table 2. Functional categories and localization of genes corresponding to sensitive phenotypes are shown in Fig. 3 and Table 3.
Functional categories and subcellular localization of the genes corresponding to the sensitive phenotypes and number of genes associated with each category. A Functional categories of the genes in 70 sensitive deletion mutants. The functional classifications are based on gene ontology and molecular function in SGD and MIPS, where A represents genes related to transcription, B for protein synthesis, C for protein fate, D for cell cycle and DNA processing, E for cellular transport, transport facilities, and transport routes, F for metabolism, G for cell rescue, defense and virulence, and H for uncharacterized or dubious ORFs. B Subcellular localization of proteins encoded by 70 genes identified from sensitive mutation
Four genes (BUD31, CRZ1, PHO4, and PHO2) in the A group participated in cell transcription. Five genes (RPS1A, RPL7A, MRT4, MSY1, and SMM1) in the B group were involved in regulating protein synthesis, in which RPS1A and RPL7A are assembly components of the ribosome. The eight genes (RBL2, VMA21, GDA1, SCJ1, SSZ1, OLA1, CNB1, and NCS2) in the C group were associated with protein fate (folding, modification, and determination). The D group contains five genes (SIF2, MCK1, RSC1, RAD57, and SAC7), which regulate cell cycle and DNA processing. The F group is a gene associated with metabolism, including TPS1, SUR1, and dfg5p.
We find that the largest category of Nd-sensitive genes is involved in cell transport, transport facilities, and transport routes. Seventeen genes (ENT3, COG5, COG6, PEP3, SRO7, DSS4, VPS9, ARF1, SYS1, ARL3, AKL1, LDB19, YPT6, ARL1, TRS85, MRL1, and SSO2) in the E group were involved in multiple processes of vesicular transport in the cell. There are 4 sensitive genes associated with Rab proteins (VPS9, YPT6, TRS85, and DSS4). The soluble N-ethylmaleimide-sensitive factor (NSF) attachment protein receptor (SNARE) family proteins are a key component of facilitating the specific fusion of transport vesicles with the destination membrane structure. In this study, we identified 3 SNARE proteins, including PEP3, SSO2, and SRO7, which mediate the fusion of the target membrane with transport membrane bubbles. Three genes (PHO84, PHO87, PHO86) are involved in regulating cellular phosphate transport pathways. In addition, the H group includes 10 unknown function genes and the function of these genes is still unknown until now days.
The content of intracellular Nd ions in yeast mutants
In this study, the yeast was further treated with 1.85 mM Nd. In the 70 missing mutants, compared to the wild type BY4743, the content of intracellular Nd ions was significantly increased in 30 mutants, and it was significantly decreased in 18 mutants. However, no significant difference was found in 22 mutants (Fig. 4). The intracellular Nd content in the other remaining 40 missing strains was near or lower than wild strains. It shows that these genes are not directly involved in affecting the function of intracellular Nd content. These genes may help to relieve Nd toxicity effects and further produce sensitive phenotypes during the loss of cell-related genes, such as repairing DNA damage under Nd stress.
Cellular Nd contents of 30 Nd-sensitive gene deletion mutants in response to Nd stress. Log-phase grown cells were treated with 1.85 mM Nd (NO3)3 for 4 h before they were collected for measurement of intracellular Nd content. The cellular Nd content of the wild type BY4743 is converted as an arbitrary unit of 1, and relative Nd content values of these mutants to that of the wild type are listed according to their functional categories. The value is the average of two independent assays for each strain
Discussion
Previously, it was found that Nd3+ was a toxic element for the fungi Penicillium simplicissimum and A. japonicus (Bergsten-Torralba et al. 2020). The mycelium cell membrane permeability of Fusarium oxysporum was increased when Nd concentrations ranged from 10 to 400 mg/L (Yufeng et al. 2007). In this study, we identified 70 Nd-sensitive gene deletion mutations from genome-scale genetic screening after fungi, they were treated with 3.7 mM Nd. By functional analysis and subcellular localization of the derived genes through SGD and MIPS-related network resources, we found that the functional classification of these 70 genes includes 60 known function genes and 10 unknown function genes. 70 genes can be divided into the following 8 categories: A ~ H groups.
Four genes (BUD31, CRZ1, PHO4, and PHO2) in the A group participated in cell transcription. PHO4 is an alkaline helix-ring-helix (bHLH) transcription activator in yeast, which is dephosphorylated and transported from the cytoplasm to the nucleus. Then it improves the intracellular phosphate level with the transcription factor PHO2 (Zhou and O'Shea 2011). The sensitive phenotypes of PHO4 and PHO2 demonstrate the importance of intracellular phosphate levels against Nd stress. In addition, CRZ1 is a transcription factor of the calcium/calcium modulated neurophosphatase signaling pathway. CRZ1 is responsible for inducing the expression of cytocellular calcium pump genes PMC1 and PMR1, which can be used to decrease the Ca2+ concentration in the cytoplasm (Zhao et al. 2013). Thus, it suggests that the maintenance of calcium homeostasis could help cells adapt to Nd stress.
Five genes (RPS1A, RPL7A, MRT4, MSY1, and SMM1) in the B group were involved in regulating protein synthesis, in which RPS1A and RPL7A are assembly components of ribosome. RPS1A encodes ribosomal protein 10 of 40S small subunits and involves in the mature of small-subunit rRNA (Moteshareie et al. 2018). In addition, the RPL7A encodes the L7A protein of the ribosomal 60S large subunit and assembles the 60S subunit (Palumbo et al. 2017). MRT4 is involved in mRNA conversion and ribosome assembly (Sugiyama et al. 2011). Thus, the ribosomal function appears to play a key role in the sensitivity of yeast cells to Nd. MSY1 encodes the mitochondrial tyrosine-tRNA synthase and is involved in the assembly of RNA (Rodley et al. 2012). SMM1 modifies the urosine residue at 20 of cytoplasmic tRNAs location (Rinaldi et al. 2003). The sensitive phenotype of these mutations to Nd suggests that Nd exposure can lead to the decrease of inefficiency and accuracy of the translation.
Eight genes (RBL2, VMA21, GDA1, SCJ1, SSZ1, OLA1, CNB1, and NCS2) in the C group were associated with protein fate (folding, modification, and determination). It is noteworthy that the absence of regulatory subunit CNB1 encoding phosphestase makes yeast cells sensitive to Nd, which can coordinate cell response to development cues, environmental stimulation, and intracellular stress. It will finally influence cell proliferation, differentiation, death, and promoting cell survival (Connolly et al. 2018). In addition, VMA21 is an essential assembly companion for the vesicular ATP enzyme (V-ATPase). Loss of VMA21 reduces V-ATP enzyme activity and results in the increase of pH in the vacuole (Ramachandran et al. 2013). In the previous studies, V-ATPase was found against the toxic of many metal ions such as cadmium, cobalt, nickel, and calcium (Techo et al. 2020b). Other genes function in this group includes the molecular partners in folding β-tubulin, protein glycosylation, and tRNA mature genes (You et al. 2004; Klassen et al. 2015; Young et al. 2013). In all, the absence of these genes related to protein assembly, protein folding, and modification processes will make cells sensitive to Nd.
The D group contains five genes (SIF2, MCK1, RSC1, RAD57, and SAC7) that regulate cell cycle and DNA processing. As a member of the family of protein kinase GSK-3, MCK1 regulates RCN1, which can directly bind with caltunphosphaterase. In addition, MCK1 is involved in regulating the cell wall integrity pathway (Kassir et al. 2006), which is related to the cell sensitivity caused by the absence of caltunphosphaterase pathway genes CNB1 and CRZ1. It was previously found that the loss of MCK1 also made cells sensitive to lithium and cadmium stress (Lockshon et al. 2012; Jiang et al. 2014). The absence of GTPase-activated protein (GAP) of the core component of the SAC7-coded cell wall integrity (CWI) pathway makes yeast cells sensitive to Nd, which is similar to the result of lithium treatment. It suggests that the regulation of CWI may be associated with the Nd sensitivity of yeast cells. Moreover, the function of the remaining three genes is associated with histone modification, chromatin remodeling, and recombination repairing of DNA double-chain fracture (Chambers et al. 2012; Fung et al. 2009; Baek et al. 2016). The sensitive phenotype of gene deletion mutations supports the conclusion of previous studies that Nd can be genetically toxic and carcinogenic (Chen et al. 2020).
We find that the largest category of Nd-sensitive genes is involved in cell transport, transport facilities, and transport routes. Seventeen genes (ENT3, COG5, COG6, PEP3, SRO7, DSS4, VPS9, ARF1, SYS1, ARL3, AKL1, LDB19, YPT6, ARL1, TRS85, MRL1, and SSO2) in the E group were involved in multiple processes of vesicular transport in the cell. The vesicular transport process includes four steps: follicle formation, transport, tying, and fusion. Vesicular transport is a process mediated by intracellular localization through the combination of coated proteins, tethered factors, class Rab/Ypt small G proteins, and SNARE proteins (Guo et al. 2017). Six genes are related to the initiation of vesicles, and the absence of these genes makes cells sensitive to neodymium. During the assembly of the follicles, ARF firstly binds to the mass membrane and then forms vesicles through a series of ARF reactions to the cargo protein and the recruitment and assembly of the cap protein. The genes ARF1, ARL3, ARL1, and SYS1 are involved in the regulation of the ARF protein that is related to the vesicular initiation assembly. In addition, ARF1 belongs to the ARF protein and is involved in intracellular transport in Gorky. ARL1 belongs to the highly similar GTP enzyme ARL protein of ARF and is involved in the assembly of vesicles on the Golgi membrane (Yang and Rosenwald 2016). ARL3 is responsible for raising ARL1, and SYS1 transports Arf-like GTP enzyme Arl3p to the Golgi (Wang et al. 2017). ENT3 is also a protein associated with the assembly of vesicles, which is involved in Mesin recruitment and transport between the vesicular Golgi body and the inner body (Fang et al. 2010). Moreover, LDB19 is associated with the late maturation of the trans-Golgi network TGN that is rich in the meseshin cohesion protein complex-1 (AP-1) (Martínez-Márquez and Duncan 2018).
The F group is a gene TPS1 that is associated with metabolism. TPS1 is involved in alginose biosynthesis, which plays a role in stress response and inhibition of protein aggregation. It has been demonstrated that TPS1 is involved in cell resistance to nickel and arsenic as it is upregulated in nickel-resistant yeast mutants, and its deficiency grants sensitivity to As (III) (Johnson et al. 2016a; Bleackley and Macgillivray 2011). The biosynthesis of fatty acid (HTD2) and thiamine (THI20), as well as the metabolic regulatory process of glucose (REG1), exhibit sensitivity to Nd (Kastaniotis et al. 2004; French et al. 2011; Tabba et al. 2010). SUR1 is associated with the biosynthesis of scabelin (Tanaka and Tani 2018). SUR1 mutants are also highly sensitive to yttrium and calcium (Grosjean et al. 2018). dfg5p is the anchored membrane protein required for the formation of inositol (GPI)-bud to form cell wall biogenesis. The absence of dfg5p slows cell growth and cell wall synthesis (Vazquez et al. 2014). In addition, the absence sensitivity of SUR1 and DFG5 indicates that the damaged cell walls and membranes make cells sensitive to Nd.
During vesicular transport, there is a small molecule GTP binding protein Rab, which allows the transport vesicular to be anchored on the appropriate target membrane (Li et al. 2019). There are 4 sensitive genes associated with Rab proteins (VPS9, YPT6, TRS85, and DSS4). YPT6 is the GTP enzyme in the Rab family, which is required for retrograde and reverse transport of ER in Gorky. Its absence or dysfunction can cause temperature sensitivity of cell growth (Yang and Rosenwald 2016). VPS9 is a guanine nucleotide exchange factor (GEF) in yeast S. cerevisiae. Membrane transport in the lysosomal pathway can be regulated by activating the Rab5 GTPases (Li et al. 2019). TRS85 is a specialized subunit of TRAPP complex III, belonging to the polyunit Rab family Ypt1p guanine nucleotide exchange factor. The absence of this gene will inhibit non-specific autophagy (Zou et al. 2015). DSS4 is a guanine nucleotide dissociation stimulating factor of the Rab protein Sec4p, which is necessary for normal delivery and transporting Gorky posterior vesicles to target mass membrane polarization growth sites (Itzen et al. 2007).
The SNARE family proteins are a key component of facilitating the specific fusion of transport vesicles with the destination membrane structure. In this study, we identified 3 SNARE proteins, namely PEP3, SSO2, and SRO7, which mediate the fusion of the target membrane with transport membrane bubbles. PEP3 is a component of the CORVET membrane-bound complex (the membrane retention complex), and it promotes the binding of transport vesicles to the vacuole vesicular SNARE protein. It is found that PEP3 overexpression strains exhibit higher V-ATPase activity (Ding et al. 2015). In addition, Sec4 and its dual-effect proteins SRO7 and t-SNARE Sec9p form a complex. It is used to butt and fuse the Golgi body vesicles with the plasma membrane (Rossi et al. 2020). SSO2 is the plasma membrane t-SNARE, and it is directly involved in the fusion of the plasma membrane secretion of vesicles (Yamamoto et al. 2018). Another important factor during vesicular transport is the tethered complex, which interacts with vesicular to promote the formation of the SNARE complex and transports vesicular to the destination. COG5 and COG6 are essential components for encoding the oligomeric Gorky complex, which are conserved in eukaryotes and play a role in protein transport. Moreover, COG5 and COG6 mediate the fusion of the transport vesicular to the Golgi region chamber (Wang et al. 2017).
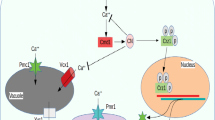
In addition, three genes (PHO84, PHO87, PHO86) are involved in regulating cellular phosphate transport pathways. PHO84 is a high-affinity inorganic phosphate (Pi) transporter and a low-affinity manganese transporter, which is regulated by Pho4p and Spt7p. The mutations of PHO84 increase the resistance to arsenates, and the mature endoplasmic reticulum secretion requires PHO86p. The low-affinity phosphate transport system PHO87 uptakes Pi at high phosphate levels (Mouillon and Persson 2006). After metals enter the yeast, cells can reduce metal toxicity through phosphate and polyphosphate cation buffer. The previous studies have shown that most Zn2+ can bind to a very rich polyphosphate in the vacuole (MacDiarmid et al. 2000). In addition, three genes (MID1, PMR1, and VCX1) are involved in the calcium ion homeostasis systems in cells. MID1 is involved in the mass membrane voltage control (Voltage-gated) Ca2+ channel, which is necessary for the internal flow of Ca2+. The PMR1 is a calcium pump, which pumps Ca2+ into the endoplasmic network and Golgi body. VCX1 is a H+/Ca2+ exchange protein localized on the vacuole membrane, which will transport calcium ions from the cytoplasm to the vacuole (Zhao et al. 2013).
Rare-earth ions have similar effects as Ca2+ in the cells (Kolouchova et al. 2016). Various reports suggest that yeast uses Ca2+ to mediate signals in response to various environmental stimuli. High concentrations of metal exposure transmit signals within the cell through a second messenger (such as Ca2+), which enable cells to take the necessary steps for living. It has been found that high concentrations of Cd emit a hazard signal through a sudden increase of cytoplasm Ca2+ (Ruta et al. 2014). These results imply the importance of a calcium steady state in combating neodymium toxicity. Moreover, VMA1 and VMA2 are two subunits of the V1 outer membrane domain of the V-ATP enzyme, and the vacuole H+-ATPase (V-ATPase) plays a key role in maintaining intracellular pH levels. In addition, V-ATPase maintains intracellular homeostasis by pumping excess protons from the cytoplasm into the vacuole (Hirata et al. 1990).
In the 70 Nd-sensitive gene deletion mutations from genome-scale genetic screening. The proteins encoded by these genes are involved in a number of important cell processes, mainly including transcription, protein synthesis, protein destination, cell cycle, DNA processing, metabolism, and cell transport. The largest number of genes are involved in the process of cell transport, which are associated with 30 genes in 70 Nd-sensitive deletion mutants. It shows that the cell transport process is a key mechanism of Nd ion detoxification in yeast cells. Our results are consistent with the results of previous studies on metal ions (Delorme-Axford and Klionsky 2018; Jiang et al. 2014; Zhao et al. 2013; Grosjean et al. 2018). The functions of these genes are related to cell wall protection, biological synthesis of ribosomes, mRNA/tRNA modification, transcription regulation, V-ATP enzymes, signal transduction, protein transport, and autophagy, which is helpful to cope with Nd toxicity.
Under heavy metal stress, yeast precipitates heavy metals through cell walls, which are the main deposition sites of Pb2+, Cd2+, and Zn2+ (Belde et al. 1988; Suman et al. 2014). It has been demonstrated that the cell wall integrity (CWI) pathway is involved in dealing with cadmium and arsenate-induced cell wall stress (Techo et al. 2020a). The yeast that loss of subunit SUR1 mannose-mannitol phosphate neuramide (MIPC) synthase is sensitive to Nd, and the biosynthesis of MIPC is necessary for maintaining normal cell wall function (Tanaka and Tani 2018). In addition, we found that GTPase-activating protein SAC7 of the cell wall integrity pathway and GPI-anchored membrane protein Dfg5 make yeast sensitive to Nd, which are also necessary for normal biosynthesis of cell walls (Lockshon et al. 2012). Our previous studies found that Nd is mainly bound or deposited in cell wall (Shi et al. 2021). Thus, maintaining the cell wall intact plays a positive role in protecting yeast cells against Nd toxicity.
Rare earth Nd has a certain mutagenic effect on DNA. A certain dose of Nd can cause damage to genetic substances at both DNA level and chromosome level, and it shows the genetic toxic effect (Chen et al. 2020). In this study, we indirectly confirmed the genetic toxicity of Nd by genes related to DNA repair and cell cycle in the sensitive phenotypes. The remodeling complex RSC1 of RAD57 and RSC chromatin is involved in the repair of double-stranded DNA broken (Chambers et al. 2012; Fung et al. 2009). It has been found that Nd treatment causes the DNA chain breaking in juvenile rainbow trout (Hanana et al. 2021). Thus, the repairing of DNA damage is used by yeast to protect against Nd stress. Gda1 plays an important role in the S phase before subdivision (Wang et al. 2015). The subunits containing the WD40 repeat sequence of the Set3c histone deacetylase (HDAC) complex are involved in regulating spore production (Ryu et al. 2020). The deficiency of FUN26 nucleoside/nucleobase transporters, which recycle nucleoside and bases from the vacuole and recycle them to the cytoplasm pool, will produce a serious fault of spore formation (Boswell-Casteel et al. 2018). Thus, the spore-producing genes are also important for the survival of yeast cells under Nd stress during cell reproduction.
Autophagy is a highly conservative catabolic pathway, which is essential for metal stress response. Moreover, autophagy plays a role in maintaining intracellular homeostasis and preventing cell damage from heavy metals. When cells are exposed to Cd, SEC17 expression is induced, which encodes the membrane protein required for vesicular transport and autophagy (Muthukumar and Nachiappan 2013). Autophagy also acts on yeast resisting nickel stress, and a number of nickel-related studies show that loss of autophagy-related genes will lead to the sensitivity to nickel, such as YVH1 (encoding autophophy structures), VPS30, and VPS38 (encoding the phosphatidylinositol 3-kinase complexes I and II) (Luo et al. 2016). In this study, there are some mutations associated with autophagy. Some studies have also found that rare-earth compounds Nd oxide–induced cancer cell autophagy (Chen et al. 2005). Thus, autophagy plays a potential role in yeast resistance to Nd stress. Cytoplasmic-vacuole targeting (Cvt) pathway is a special biosynthetic form of yeast selective autophagy (Torggler et al. 2017). It has been found that knocking out several genes involved in Cvt and autophagy pathways increased the sensitivity of yeast cerevisiae to Zn2+. Microscopic analysis shows that Zn2+ partially inhibits the fusion of Cvt vesicles with liquid follicles (Dziedzic and Caplan 2011). In addition, three autophagy-related genes are involved in Cvt pathways. YPT6 and ARL1 are critical for retrograde transport of vesicles from the interior to the trans-Golgi network (TGN) and help form autophagy. The mutations of these genes will lead to severe protein dislocation and growth defects (Yang and Rosenwald 2016). TRS85 regulates Endo-Golgi transport and is necessary for membrane amplification in the autophagy and Cvt pathway (Zou et al. 2015). Moreover, loss of the subunits of cytosol-binding complex causes yeast sensitivity to Nd, and the subunits of the cytosol-binding complex also showed sensitive phenotypic in the previous genome-wide screening under Al, Pb, Y, and Cr stress (Tun et al. 2014; Johnson et al. 2016b; Du et al. 2015; Grosjean et al. 2018). It is also found that the abnormal COG complex function leads to Cvt pathway defects and non-selective autophagy (Wang et al. 2017). In our study, we also find that the Cvt pathway plays a central role in autophagy against Nd poisoning in yeast cells.
Conclusion
Through genome-scale screening, we identified 70 yeast monogene deletion mutations sensitive to Nd ions. These genes are involved in metabolism, transcription, protein synthesis, cell cycle, DNA processing, protein folding, modification, and cell transport processes. Furthermore, these data are mapped to the protein interaction group to identify the regulatory networks of Nd toxicity. These networks are associated with various signal pathways, mainly including calcium ion transport, phosphate pathways, vesicular transport, and cell autophagy. In addition, the content of Nd ions in yeast was detected by an inductively coupled plasma mass spectrometry, and most of these Nd-sensitive mutants showed an increased intracellular Nd content. In all, our results provide the basis for understanding the molecular mechanisms of detoxifying Nd ions in yeast cells, which will be useful for future studies on Nd-related issues in the environment, agriculture, and human health.
Availability of data and materials
The data obtained during and/or analyzed during the current study are available from the corresponding author on reasonable request.
References
Alarfaj MA, Hasen AA, Al-Yami SS (2018) Traumatic subhyaloid macular hemorrhage with complete resolution following neodymium-doped yttrium aluminium garnet laser. Am J Ophthalmol Case Rep 9:85–87. https://doi.org/10.1016/j.ajoc.2018.01.020
Åström M (2001) Abundance and fractionation patterns of rare earth elements in streams affected by acid sulphate soils. Chem Geol 175(3):249–258. https://doi.org/10.1016/S0009-2541(00)00294-1
Baek SH, Kwon EY, Kim SY, Hahn JS (2016) GSF2 deletion increases lactic acid production by alleviating glucose repression in Saccharomyces cerevisiae. Sci Rep 6:34812. https://doi.org/10.1038/srep34812
Bao TM, Tian Y, Wang LX, Ting WU, Li-Na LU, Hong-Yu MA, Wang L (2018) Lanthanum, cerium, praseodymium, and neodymium levels in residential environment of rare earth ore area in Baotou City. J Environ Occup Med 35(2):158–162
Bär F, Berger L, Jauer L, Kurtuldu G, Schäublin R, Schleifenbaum JH, Löffler JF (2019) Laser additive manufacturing of biodegradable magnesium alloy WE43: a detailed microstructure analysis. Acta Biomater 98:36–49. https://doi.org/10.1016/j.actbio.2019.05.056
Belde PJM, Kessels BGF, Moelans IM, Borst-Pauwels GWFH (1988) Cd2+ uptake, Cd2+ binding and loss of cell K+ by a Cd-sensitive and a Cd-resistant strain of Saccharomyces cerevisiae. FEMS Microbiol Lett 49(3):493–498
Bergsten-Torralba LR, Magalhães DP, Giese EC, Nascimento CRS, Pinho JVA, Buss DF (2020) Toxicity of three rare earth elements, and their combinations to algae, microcrustaceans, and fungi. Ecotoxicol Environ Saf 201:110795. https://doi.org/10.1016/j.ecoenv.2020.110795
Bleackley MR, Macgillivray RT (2011) Transition metal homeostasis from yeast to human disease. Biometals 24(5):785–809. https://doi.org/10.1007/s10534-011-9451-4
Boswell-Casteel RC, Johnson JM, Hays FA (2018) Functional characterization of the Saccharomyces cerevisiae Equilibrative Nucleoside Transporter 1 (ScENT1). Molecules (Basel, Switzerland) 23(4):732. https://doi.org/10.3390/molecules23040732
Campos JA, Tejera NA, Sánchez CJ (2009) Substrate role in the accumulation of heavy metals in sporocarps of wild fungi. Biometals 22(5):835–841. https://doi.org/10.1007/s10534-009-9230-7
Challaraj Emmanuel ES, Vignesh V, Anandkumar B, Maruthamuthu S (2011) Bioaccumulation of cerium and neodymium by Bacillus cereus isolated from rare earth environments of Chavara and Manavalakurichi. India Indian J of Microbiol 51(4):488–495. https://doi.org/10.1007/s12088-011-0111-8
Chambers AL, Brownlee PM, Durley SC, Beacham T, Kent NA, Downs JA (2012) The two different isoforms of the RSC chromatin remodeling complex play distinct roles in DNA damage responses. PLoS ONE 7(2):e32016. https://doi.org/10.1371/journal.pone.0032016
Chen Y, Yang L, Feng C, Wen LP (2005) Nano neodymium oxide induces massive vacuolization and autophagic cell death in non-small cell lung cancer NCI-H460 cells. Biochem Biophys Res Commun 337(1):52–60. https://doi.org/10.1016/j.bbrc.2005.09.018
Chen Y, Zhu W, Shu F, Fan Y, Yang N, Wu T, Ji L, Xie W, Bade R, Jiang S, Liu X, Shao G, Wu G, Jia X (2020) Nd(2)O(3) Nanoparticles induce toxicity and cardiac/cerebrovascular abnormality in zebrafish embryos via the apoptosis pathway. Int J Nanomed 15:387–400. https://doi.org/10.2147/ijn.s220785
Connolly S, Quasi-Woode D, Waldron L, Eberly C, Waters K, Muller EM, Kingsbury TJ (2018) Calcineurin regulatory subunit calcium-binding domains differentially contribute to calcineurin signaling in Saccharomyces cerevisiae. Genetics 209(3):801–813. https://doi.org/10.1534/genetics.118.300911
Cyert MS, Philpott CC (2013) Regulation of cation balance in Saccharomyces cerevisiae. Genetics 193(3):677–713. https://doi.org/10.1534/genetics.112.147207
Delorme-Axford E, Klionsky DJ (2018) Transcriptional and post-transcriptional regulation of autophagy in the yeast Saccharomyces cerevisiae. J Biol Chem 293(15):5396–5403. https://doi.org/10.1074/jbc.R117.804641
Ding J, Holzwarth G, Bradford CS, Cooley B, Yoshinaga AS, Patton-Vogt J, Abeliovich H, Penner MH, Bakalinsky AT (2015) PEP3 overexpression shortens lag phase but does not alter growth rate in Saccharomyces cerevisiae exposed to acetic acid stress. Appl Microbiol Biotechnol 99(20):8667–8680. https://doi.org/10.1007/s00253-015-6708-9
Du J, Cao C, Jiang L (2015) Genome-scale genetic screen of lead ion-sensitive gene deletion mutations in Saccharomyces cerevisiae. Gene 563(2):155–159. https://doi.org/10.1016/j.gene.2015.03.018
Dziedzic SA, Caplan AB (2011) Identification of autophagy genes participating in zinc-induced necrotic cell death in Saccharomyces cerevisiae. Autophagy 7(5):490–500. https://doi.org/10.4161/auto.7.5.14872
Fang P, Li X, Wang J, Niu L, Teng M (2010) Structural basis for the specificity of the GAE domain of yGGA2 for its accessory proteins Ent3 and Ent5. Biochem 49(36):7949–7955. https://doi.org/10.1021/bi1010255
French JB, Begley TP, Ealick SE (2011) Structure of trifunctional THI20 from yeast. Acta Crystallogr D Biol Crystallogr 67(Pt 9):784–791. https://doi.org/10.1107/s0907444911024814
Fung CW, Mozlin AM, Symington LS (2009) Suppression of the double-strand-break-repair defect of the Saccharomyces cerevisiae rad57 mutant. Genetics 181(4):1195–1206. https://doi.org/10.1534/genetics.109.100842
Gerwien F, Skrahina V, Kasper L, Hube B, Brunke S (2018) Metals in fungal virulence. FEMS Microbiol Rev 42(1):fux050. https://doi.org/10.1093/femsre/fux050
Gohda Y (2021) First-principles determination of intergranular atomic arrangements and magnetic properties in rare-earth permanent magnets. Sci Technol Adv Mater 22(1):113–123. https://doi.org/10.1080/14686996.2021.1877092
Grosjean N, Gross EM, Le Jean M, Blaudez D (2018) Global deletome profile of Saccharomyces cerevisiae exposed to the technology-critical element yttrium. Front Microbiol 9:2005. https://doi.org/10.3389/fmicb.2018.02005
Guo Y, Yang F, Tang X (2017) An overview of protein secretion in yeast and animal cells. Methods in Mol Biol (clifton, NJ) 1662:1–17. https://doi.org/10.1007/978-1-4939-7262-3_1
Hanana H, Kleinert C, Gagné F (2021) Toxicity of representative mixture of five rare earth elements in juvenile rainbow trout (Oncorhynchus mykiss) juveniles. Environ Sci Pollut Res Int 28(22):28263–28274. https://doi.org/10.1007/s11356-020-12218-5
Hirata R, Ohsumk Y, Nakano A, Kawasaki H, Suzuki K, Anraku Y (1990) Molecular structure of a gene, VMA1, encoding the catalytic subunit of H(+)-translocating adenosine triphosphatase from vacuolar membranes of Saccharomyces cerevisiae. J Biol Chem 265(12):6726–6733
Itzen A, Rak A, Goody RS (2007) Sec2 is a highly efficient exchange factor for the Rab protein Sec4. J Mol Biol 365(5):1359–1367. https://doi.org/10.1016/j.jmb.2006.10.096
Jiang L, Cao C, Zhang L, Lin W, Xia J, Xu H, Zhang Y (2014) Cadmium-induced activation of high osmolarity glycerol pathway through its Sln1 branch is dependent on the MAP kinase kinase kinase Ssk2, but not its paralog Ssk22, in budding yeast. FEMS Yeast Res 14(8):1263–1272. https://doi.org/10.1111/1567-1364.12220
Johnson AJ, Veljanoski F, O’Doherty PJ, Zaman MS, Petersingham G, Bailey TD, Münch G, Kersaitis C, Wu MJ (2016a) Molecular insight into arsenic toxicity via the genome-wide deletion mutant screening of Saccharomyces cerevisiae. Metallomics Integr Biometal Sci 8(2):228–235. https://doi.org/10.1039/c5mt00261c
Johnson AJ, Veljanoski F, O’Doherty PJ, Zaman MS, Petersingham G, Bailey TD, Münch G, Kersaitis C, Wu MJ (2016b) Revelation of molecular basis for chromium toxicity by phenotypes of Saccharomyces cerevisiae gene deletion mutants. Metallomics Integr Biometal Sci 8(5):542–550. https://doi.org/10.1039/c6mt00039h
Kassir Y, Rubin-Bejerano I, Mandel-Gutfreund Y (2006) The Saccharomyces cerevisiae GSK-3 beta homologs. Curr Drug Targets 7(11):1455–1465. https://doi.org/10.2174/1389450110607011455
Kastaniotis AJ, Autio KJ, Sormunen RT, Hiltunen JK (2004) Htd2p/Yhr067p is a yeast 3-hydroxyacyl-ACP dehydratase essential for mitochondrial function and morphology. Mol Microbiol 53(5):1407–1421. https://doi.org/10.1111/j.1365-2958.2004.04191.x
Khan AM, Bakar NKA, Bakar AFA, Ashraf MA (2017) Chemical speciation and bioavailability of rare earth elements (REEs) in the ecosystem: a review. Environ Sci Pollut Res Int 24(29):22764–22789. https://doi.org/10.1007/s11356-016-7427-1
Klassen R, Grunewald P, Thüring KL, Eichler C, Helm M, Schaffrath R (2015) Loss of anticodon wobble uridine modifications affects tRNA(Lys) function and protein levels in Saccharomyces cerevisiae. PLoS ONE 10(3):e0119261. https://doi.org/10.1371/journal.pone.0119261
Kolouchova I, Sigler K, Zimola M, Rezanka T, Matatkova O, Masak J (2016) Influencing fatty acid composition of yeasts by lanthanides. World J Microbiol Biotechnol 32(8):126. https://doi.org/10.1007/s11274-016-2093-5
Lemoine A, Mamann N, Larroquet M, Tounian P, Irtan S, Lemale J (2019) Ingestion of neodymium magnet spheres: three case studies. Arch Pediatr 26(3):179–181. https://doi.org/10.1016/j.arcped.2019.02.001
Li W, Wu Z, Liang Y (2019) Vrl1 relies on its VPS9-domain to play a role in autophagy in Saccharomyces cerevisiae. Cell Biol Int 43(8):875–889. https://doi.org/10.1002/cbin.11156
Lockshon D, Olsen CP, Brett CL, Chertov A, Merz AJ, Lorenz DA, Van Gilst MR, Kennedy BK (2012) Rho signaling participates in membrane fluidity homeostasis. PLoS ONE 7(10):e45049. https://doi.org/10.1371/journal.pone.0045049
Luo C, Cao C, Jiang L (2016) The endosomal sorting complex required for transport (ESCRT) is required for the sensitivity of yeast cells to nickel ions in Saccharomyces cerevisiae. FEMS Yeast Res 16(3):fow028. https://doi.org/10.1093/femsyr/fow028
MacDiarmid CW, Gaither LA, Eide D (2000) Zinc transporters that regulate vacuolar zinc storage in Saccharomyces cerevisiae. EMBO J 19(12):2845–2855. https://doi.org/10.1093/emboj/19.12.2845
Martínez-Márquez JY, Duncan MC (2018) Investigation of Ldb19/Art1 localization and function at the late Golgi. PLoS ONE 13(11):e0206944. https://doi.org/10.1371/journal.pone.0206944
Migaszewski ZM, Gałuszka A, Dołęgowska S (2016) Rare earth and trace element signatures for assessing an impact of rock mining and processing on the environment: Wiśniówka case study, south-central Poland. Environ Sci Pollut Res Int 23(24):24943–24959. https://doi.org/10.1007/s11356-016-7713-y
Moteshareie H, Hajikarimlou M, Mulet Indrayanti A, Burnside D, Paula Dias A, Lettl C, Ahmed D, Omidi K, Kazmirchuk T, Puchacz N, Zare N, Takallou S, Naing T, Hernández RB, Willmore WG, Babu M, McKay B, Samanfar B, Holcik M, Golshani A (2018) Heavy metal sensitivities of gene deletion strains for ITT1 and RPS1A connect their activities to the expression of URE2, a key gene involved in metal detoxification in yeast. PLoS ONE 13(9):e0198704. https://doi.org/10.1371/journal.pone.0198704
Mouillon JM, Persson BL (2006) New aspects on phosphate sensing and signalling in Saccharomyces cerevisiae. FEMS Yeast Res 6(2):171–176. https://doi.org/10.1111/j.1567-1364.2006.00036.x
Mowafy AM (2020) Biological leaching of rare earth elements. World J Microbiol Biotechnol 36(4):61. https://doi.org/10.1007/s11274-020-02838-x
Muthukumar K, Nachiappan V (2013) Phosphatidylethanolamine from phosphatidylserine decarboxylase2 is essential for autophagy under cadmium stress in Saccharomyces cerevisiae. Cell Biochem Biophys 67(3):1353–1363. https://doi.org/10.1007/s12013-013-9667-8
Pagano G, Thomas PJ, Di Nunzio A, Trifuoggi M (2019) Human exposures to rare earth elements: Present knowledge and research prospects. Environ Res 171:493–500. https://doi.org/10.1016/j.envres.2019.02.004
Palumbo RJ, Fuchs G, Lutz S, Curcio MJ (2017) Paralog-specific functions of RPL7A and RPL7B mediated by ribosomal protein or snoRNA dosage in Saccharomyces cerevisiae. G3 (Bethesda Md) 7(2):591–606. https://doi.org/10.1534/g3.116.035931
Ramachandran N, Munteanu I, Wang P, Ruggieri A, Rilstone JJ, Israelian N, Naranian T, Paroutis P, Guo R, Ren ZP, Nishino I, Chabrol B, Pellissier JF, Minetti C, Udd B, Fardeau M, Tailor CS, Mahuran DJ, Kissel JT, Kalimo H, Levy N, Manolson MF, Ackerley CA, Minassian BA (2013) VMA21 deficiency prevents vacuolar ATPase assembly and causes autophagic vacuolar myopathy. Acta Neuropathol 125(3):439–457. https://doi.org/10.1007/s00401-012-1073-6
Ren Y, Ren X, Ma J, Yan L (2016) Effects of mixed rare earth fertilizer on yield and nutrient quality of leafy vegetables during different seasons. J Rare Earths 34(6):638–643. https://doi.org/10.1016/S1002-0721(16)60073-X
Rinaldi T, Gambadoro A, Francisci S, Frontali L (2003) Nucleo-mitochondrial interactions in Saccharomyces cerevisiae: characterization of a nuclear gene suppressing a defect in mitochondrial tRNA(Asp) processing. Gene 303:63–68. https://doi.org/10.1016/s0378-1119(02)01154-x
Rodley CD, Grand RS, Gehlen LR, Greyling G, Jones MB, O’Sullivan JM (2012) Mitochondrial-nuclear DNA interactions contribute to the regulation of nuclear transcript levels as part of the inter-organelle communication system. PLoS ONE 7(1):e30943. https://doi.org/10.1371/journal.pone.0030943
Rossi G, Lepore D, Kenner L, Czuchra AB, Plooster M, Frost A, Munson M, Brennwald P (2020) Exocyst structural changes associated with activation of tethering downstream of Rho/Cdc42 GTPases. J Cell Biol 219(2):e201904161. https://doi.org/10.1083/jcb.201904161
Ruta LL, Popa VC, Nicolau I, Danet AF, Iordache V, Neagoe AD, Farcasanu IC (2014) Calcium signaling mediates the response to cadmium toxicity in Saccharomyces cerevisiae cells. FEBS Lett 588(17):3202–3212. https://doi.org/10.1016/j.febslet.2014.07.001
Ryu HY, Zhao D, Li J, Su D, Hochstrasser M (2020) Histone sumoylation promotes Set3 histone-deacetylase complex-mediated transcriptional regulation. Nucleic Acids Res 48(21):12151–12168. https://doi.org/10.1093/nar/gkaa1093
Shi K, Liu C, Liu D, Lyu K, Chen J, Wang X (2021) The accumulation and effect of rare earth element neodymium on the root of rice seedlings. Environ Sci Pollut Res Int 28(35):48656-48665. https://doi.org/10.1007/s11356-021-14072-5
Sugiyama M, Nugroho S, Iida N, Sakai T, Kaneko Y, Harashima S (2011) Genetic interactions of ribosome maturation factors Yvh1 and Mrt4 influence mRNA decay, glycogen accumulation, and the expression of early meiotic genes in Saccharomyces cerevisiae. J Biochem 150(1):103–111. https://doi.org/10.1093/jb/mvr040
Sultan K, Shazili NA (2009) Rare earth elements in tropical surface water, soil and sediments of the Terengganu River Basin, Malaysia. J Rare Earths 27(6):1072–1078. https://doi.org/10.1016/S1002-0721(08)60391-9
Suman J, Kotrba P, Macek T (2014) Putative P1B-type ATPase from the bacterium Achromobacter xylosoxidans A8 alters Pb2+/Zn2+/Cd2+-resistance and accumulation in Saccharomyces cerevisiae. Biochem Biophys Acta 5:1338–1343. https://doi.org/10.1016/j.bbamem.2014.01.023
Tabba S, Mangat S, McCartney R, Schmidt MC (2010) PP1 phosphatase-binding motif in Reg1 protein of Saccharomyces cerevisiae is required for interaction with both the PP1 phosphatase Glc7 and the Snf1 protein kinase. Cell Signal 22(7):1013–1021. https://doi.org/10.1016/j.cellsig.2010.02.003
Tanaka S, Tani M (2018) Mannosylinositol phosphorylceramides and ergosterol coodinately maintain cell wall integrity in the yeast Saccharomyces cerevisiae. FEBS J 285(13):2405–2427. https://doi.org/10.1111/febs.14509
Techo T, Jindarungrueng S, Tatip S, Limcharoensuk T, Pokethitiyook P, Kruatrachue M, Auesukaree C (2020b) Vacuolar H(+) -ATPase is involved in preventing heavy metal-induced oxidative stress in Saccharomyces cerevisiae. Environ Microbiol 22(6):2403–2418. https://doi.org/10.1111/1462-2920.15022
Techo T, Charoenpuntaweesin S, Auesukaree C (2020a) Involvement of the cell wall integrity pathway of Saccharomyces cerevisiae in protection against cadmium and arsenate stresses. Appl Environ Microbiol 86(21):e01339-20. doi:https://doi.org/10.1128/aem.01339-20
Thornton BF, Burdette SC (2017) The Neodymium Neologism. Nat Chem 9(2):194. https://doi.org/10.1038/nchem.2722
Torggler R, Papinski D, Kraft C (2017) Assays to monitor autophagy in Saccharomyces cerevisiae. Cells 6(3):23. doi:https://doi.org/10.3390/cells6030023
Tun NM, O’Doherty PJ, Chen ZH, Wu XY, Bailey TD, Kersaitis C, Wu MJ (2014) Identification of aluminium transport-related genes via genome-wide phenotypic screening of Saccharomyces cerevisiae. Metallomics Integr Biometal Sci 6(8):1558–1564. https://doi.org/10.1039/c4mt00116h
Vazquez HM, Vionnet C, Roubaty C, Conzelmann A (2014) Cdc1 removes the ethanolamine phosphate of the first mannose of GPI anchors and thereby facilitates the integration of GPI proteins into the yeast cell wall. Mol Biol Cell 25(21):3375–3388. https://doi.org/10.1091/mbc.E14-06-1033
Wang D, Wang C, Ye S, Qi H, Zhao G (2003) Effects of spraying rare earths on contents of rare Earth elements and effective components in tea. J Agric Food Chem 51(23):6731–6735. https://doi.org/10.1021/jf0303417
Wang Y, Jin H, Deng S, Chen Y, Yu Y (2011) Effects of neodymium on growth and physiological characteristics of Microcystis aeruginosa. J Rare Earths 29(4):388–395. https://doi.org/10.1016/S1002-0721(10)60466-8
Wang Q, Liu C, Tang C, Guo H, Liu Y, Wang L, Zhao H, Shang Y, Wen Y, Lin Y, Zhou T, Zhou Z, Dong W, Hu Z, Guo X, Sha J, Li W (2015) Yeast model identifies ENTPD6 as a potential non-obstructive azoospermia pathogenic gene. Sci Rep 5:11762. https://doi.org/10.1038/srep11762
Wang IH, Chen YJ, Hsu JW, Lee FJ (2017) The Arl3 and Arl1 GTPases co-operate with Cog8 to regulate selective autophagy via Atg9 trafficking. Traffic (copenhagen, Denmark) 18(9):580–589. https://doi.org/10.1111/tra.12498
Yamamoto T, Yamamoto D, Rokugawa K, Yoshimura K, Imura Y, Yoshimura E, Suzuki M (2018) Decreased aluminium tolerance in the growth of Saccharomyces cerevisiae with SSO2 gene disruption. Biometals 31(2):203–215. https://doi.org/10.1007/s10534-017-0069-z
Yang S, Rosenwald AG (2016) Autophagy in Saccharomyces cerevisiae requires the monomeric GTP-binding proteins Arl1 and Ypt6. Autophagy 12(10):1721–1737. https://doi.org/10.1080/15548627.2016.1196316
You L, Gillilan R, Huffaker TC (2004) Model for the yeast cofactor A-beta-tubulin complex based on computational docking and mutagensis. J Mol Biol 341(5):1343–1354. https://doi.org/10.1016/j.jmb.2004.06.081
Young CL, Raden DL, Robinson AS (2013) Analysis of ER resident proteins in Saccharomyces cerevisiae: implementation of H/KDEL retrieval sequences. Traffic (copenhagen, Denmark) 14(4):365–381. https://doi.org/10.1111/tra.12041
Yufeng Z, Lifen Y, Kaoshan C, Liang D (2007) Effects of neodymium on growth, pectinase activity and mycelium permeability of Fusarium oxysporum. J Rare Earths 25(1):100–105. https://doi.org/10.1016/S1002-0721(07)60053-2
Zhang H, Han H, Su C, Zhang H, Hou Z, Song Q (2007) Application of stereology on microstructure of neodymium-doped yttrium aluminum garnet (Nd:YAG) transparent ceramics. Mater Sci Eng, A 445–446:180–185. https://doi.org/10.1016/j.msea.2006.09.030
Zhao Y, Du J, Zhao G, Jiang L (2013) Activation of calcineurin is mainly responsible for the calcium sensitivity of gene deletion mutations in the genome of budding yeast. Genomics 101(1):49–56. https://doi.org/10.1016/j.ygeno.2012.09.005
Zhou X, O’Shea EK (2011) Integrated approaches reveal determinants of genome-wide binding and function of the transcription factor Pho4. Mol Cell 42(6):826–836. https://doi.org/10.1016/j.molcel.2011.05.025
Zhuang M, Wang L, Wu G, Wang K, Jiang X, Liu T, Xiao P, Yu L, Jiang Y, Song J, Zhang J, Zhou J, Zhao J, Chu Z (2017) Health risk assessment of rare earth elements in cereals from mining area in Shandong. China Sci Rep 7(1):9772. https://doi.org/10.1038/s41598-017-10256-7
Zou S, Liu Y, Zhang C, Yu S, Liang Y (2015) Bet3 participates in autophagy through GTPase Ypt1 in Saccharomyces cerevisiae. Cell Biol Int 39(4):466–474. https://doi.org/10.1002/cbin.10416
Funding
This study was supported by the Major Research Plan of the Shandong Natural Science Foundation (ZR2020ZD19) and the National Natural Science Foundation of China (30900071).
Major Research plan of the Shandong Natural Science Foundation,ZR2020ZD19,Xue Wang,National Natural Science Foundation of China,30900071,Xue Wang
Author information
Authors and Affiliations
Contributions
X Wang: conceptualization, supervision; CK Liu, KL Lyu, and KL Shi: methodology; CK Liu and DW Liu: writing–original draft preparation; KL Shi and CK Liu: investigation; X Wang: writing–reviewing and editing.
Ethics declarations
Ethics approval
Not applicable. Animals were not used in this study.
Consent to participate
Not applicable.
Consent to publication
Not applicable.
Competing interests
Not applicable.
Additional information
Responsible Editor: Robert Duran.
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Liu, C., Shi, K., Lyu, K. et al. The toxicity of neodymium and genome-scale genetic screen of neodymium-sensitive gene deletion mutations in the yeast Saccharomyces cerevisiae. Environ Sci Pollut Res 29, 41439–41454 (2022). https://doi.org/10.1007/s11356-021-18100-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11356-021-18100-2