Abstract
Camellia sinensis is an important economic crop worldwide since this plant is used to make one of the most popular non-alcoholic beverages, tea. Salinity together with drought pose a serious threat to the production and qualities of C. sinensis. However, the transcriptome dynamics occurring in response to drought stress and salt stress in tea plants are poorly understood at the molecular level. We reported the first large-coverage transcriptome datasets for C. sinensis under drought and salt stress using next-generation sequencing technology. Using a high-throughput Illumina sequencing platform, approximately 398.95 million high-quality paired-end reads were generated from young leaves of C. sinensis subjected to drought stress and salt stress, and these reads were used for de novo assembly. The transcripts with further processing and filtering yielded a set of 64,905 coding DNA sequences (CDSs) with an average length of 710 bp and an N50 of 933 bp. In total, 3936 and 3715 differentially expressed genes (DEGs) were identified from all analyzed time points of drought stress and salt stress, respectively. Identified in drought and salt stress were 2131 overlapping DEGs, and these are involved in galactosyltransferase activity, tetrapyrrole binding, and hydrolase activity, indicating that C. sinensis has a similar molecular response to these two stresses. We clustered the above DEGs from both sets into four clusters according to their expression dynamics, with the genes in each cluster showing enrichment for particular functional categories. We also found that under salt stress, most DEGs showed down-regulation at early time points and their expression levels were elevated after 48 h, whereas under drought stress most DEGs were down-regulated in all time points. The DEGs relative to pathways of osmotic product such as proline, sugar, and GABA were identified in C. sinensis. Noteworthy, among the identified DEGs are genes involved in the biosynthetic pathways of polyphenol and caffeine, providing evidence at the molecular level that salt and drought affect tea qualities. In addition, we analyzed the differential expression of transcription factors and revealed a large amount of crosstalk between the metabolic pathways of drought and salt stress. All findings suggest that gene expression exhibits rapid and coordinated changes during C. sinensis adaptations to drought stress and salt stress, and common themes in the response to both stresses were identified.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Abiotic stresses, especially salinity and drought, adversely affect plant growth, crop production, and geographical distribution (Boyer 1982; Allakhverdiev et al. 2000; Hasegawa et al. 2000; Zhu 2002; Arbona et al. 2013). Approximately 20% of the world cultivated land, and nearly 50% of irrigated land, is adversely affected by salinity (Anthony 1998; Tuteja 2007). Salinity, together with drought, continues to affect food security, economic sustainability, and biodiversity (Bahieldin et al. 2013). Given the increasing surface area of salty land and the growing water scarcity, a thorough understanding of the physiological and molecular bases for salt and drought tolerance in plants is required to develop salt- and drought-tolerant plants (Rus et al. 2005; Beritognolo et al. 2011).
Different abiotic stresses induce different (or common) biochemical and physiological effects on plant growth, development, and secondary metabolism. Drought restricts plant growth via photosynthetic decline, whereas osmotic stress imposes constraints on plant processes and interferes with nutrient availability as the soil dries. Salinity interferes with plant growth by physiological drought and ion toxicity (Zhu 2002); salt stress disrupts plant ion homeostasis, resulting in excess toxic Na+ in the cytoplasm, and a deficiency of essential ions such as K+ (Binzel and Ratajczak 2002). For plants to survive under these stress conditions, a complex mechanism must exist to sense and respond to the abiotic stresses (Bohnert et al. 1995). Responses usually involve complex networks controlling global gene expression, protein modification, and metabolite composition through specific factors (Urano et al. 2010). The DREB1/CBF pathway plays a key role in regulating ABA-independent gene expression in response to drought and cold stress; overexpression of DREB1/CBF can increase the tolerance of transgenic plants to cold temperatures, drought, and salt stress (Jaglo-Ottosen et al. 1998; Liu et al. 1998; Shinozaki and Yamaguchi-Shinozaki 2000; Goode and Organizers 2007). The salt overly sensitive (SOS) pathway, identified through the isolation and study of sos1, sos2, and sos3 mutants, is essential for defense response of salt stress (Zhu 2002). Abscisic acid (ABA), produced under water-deficit conditions, is important for plant tolerance to drought and high salinity. Though proline and betaine occur naturally in a wide variety of plants, exogenous application of such osmolytes mitigates the adverse effect of salt stress. Proline has been found to be more effective than betaine in mitigating salt stress-induced effects (Eiji et al. 2004; Hoque et al. 2007). Although several pathways involved in abiotic stress responses have been reported in Arabidopsis, little is known about the transcriptomic dynamics during prolonged salinity and drought stress in economically valuable crops.
Tea is one of the most prevalent non-alcoholic beverages that benefits human health due to the extensive presence of secondary metabolites in tea leaves, including polyphenols, theanine, and volatile oils (Rogers et al. 2008). The tea plant, Camellia sinensis, is an economically important horticultural crop. Appropriate soil and water conditions are required for tea plants to maintain balanced growth, which is fundamental to produce tea of superior quality and high yields. The increasing soil salinity and dry land are posing a serious threat to tea agriculture. Therefore, in-depth analyses of the physiological and molecular mechanisms involved in salt and drought tolerance in tea are required to develop tea plants that can tolerate high salt concentrations and severe drought conditions (Rus 2005; Beritognolo et al. 2011). However, a lack of genome information and genetic tools (such as tissue culture and transformation) remain a hurdle in tea research. Along with the development of next-generation sequencing (NGS) technologies, transcriptome sequencing using NGS technologies is a fast and cost-effective approach to generate genome-scale sequence resources (Liu et al. 2012; Kanak et al. 2012). NGS is not limited to detecting transcripts from known genomic sequences, and therefore is particularly attractive for studies in non-model organisms (Vera et al. 2008; Kristiansson et al. 2009; Meyer et al. 2009). NGS has also been used to study the transcriptome profiles of cold acclimation (CA) in tea plants, where a large number of genes involved in cold signal sensors or transduction, genes related to the stabilization of plasma membranes, osmosensing-responsive genes, and stress-responsive transcription factor genes were identified to be responsible for the CA process (Wang et al. 2013).
In this study, to investigate the molecular basis of tea plants under salt and drought stresses, we performed de novo transcriptome sequencing of tea under prolonged salt and drought stress using the NGS sequencing platform. The results provided the useful foundation for future genetic improvement of cultivated tea trees.
Materials and methods
Plant materials and RNA preparation
Tea plants of C. sinensis cv. Tieguanyin were cultivated in the Anxi County of the Fujian Province of China (east longitude 117°86, north altitude 24°90). One-year-old plants with similar growth and size (about 20 internodes) were used as source for the cuttings of C. sinensis and grown in 1/2 dose Hoagland nutrient solution for 3 days. All the selected cutting plants were divided into three groups, two of them exposed to 25% polyethylene glycol (PEG) (simulate drought-stress conditions) or 200 mM NaCl in 1/2 Hoagland nutrient solution for 24, 48, and 72 h after 3 days, respectively, the other group was exposed to 1/2 Hoagland nutrient solution alone for 0, 24, 48, and 72 h, respectively as control.
In total, 60 tea plants including 5 individuals for each time point of each stress type and control were collected. The second and third leaves under the seedling bud from PEG-stressed cutting plants and NaCl-stressed cutting plants as well as control plants were snap-frozen and stored at −80 °C until further processing. Total RNA from the above samples was isolated for transcriptome sequencing, using the previously described CTAB method (Shi et al. 2007). RNA integrity was tested using an Agilent 2100 Bioanalyzer with a minimum RNA Integrity Number (RIN) of 7. Equal amounts of total RNA from five samples for each time points of each stress and controls were pooled for cDNA preparation.
cDNA library construction and sequencing
Poly(A) RNA(mRNA)was isolated from 20 g of total RNA using Dynal oligo(dT) magnetic beads (Invitrogen) according to the manufacturer’s instructions. Following purification, mRNA was fragmented into smaller pieces at 70 °C for 5 min in the fragmentation buffer (Ambion), and reverse-transcribed to synthesize first strand cDNA using SuperScript III reverse transcriptase (Invitrogen) and N6 random hexamers (Takara). Subsequently, second strand cDNA was synthesized using RNase H and DNA polymerase. After end repair and ligation of adapters, products were enriched with PCR to prepare the final cDNA library. The integrity of cDNA libraries was analyzed by an Agilent 2100 Bioanalyzer. Ten paired-end cDNA libraries were constructed with 300 base pair (bp) insertions and then sequenced from both 5′ and 3′ ends using the Illumina HiSeq™ 2500 platform in the Center for Genomics and Biotechnology of Fujian Agriculture and Forestry University (Fuzhou, China) following the manufacturer’s instructions with 100 bp paired-end read length.
Data preprocessing, de novo assembly, and assessment
Raw data were initially processed using a custom script and NGS QC toolkit(Chen et al. 2015) to obtain clean reads by removing adapter sequences, empty reads, low-quality sequences (i.e., those containing more than 10% “N”s), and reads with more than 10% bases with Q value <20. Clean reads from ten samples were merged and assembled into unigenes using the Trinity package with default parameters (v2.0.6) (Grabherr et al. 2011; Haas et al. 2013).
To obtain high-quality unigenes and reduce the unavoidable redundancy produced by Trinity assembly, CD-Hit-est. software was used for clustering analysis (Li and Godzik 2006) with 90% identity and eight word length. We then annotated the protein coding sequences and obtained CDSs for C. sinensis using the “TransDecoder” module of the Trinity package with the “minimum protein length” parameter set to 100. All predicted CDSs were used for further analysis.
Transcriptome functional annotation
All non-redundant transcripts were annotated by searching against the NCBI non-redundant (NR), Pfam (version 28.0) (Robert et al. 2014) and Kyoto Encyclopedia of Genes and Genomes (KEGG: http://www.genome.jp/kegg/) databases by BLASTX with an E-value threshold of 10−6. To annotate Gene Ontology (GO: http//www.geneontology.org) terms describing biological processes, molecular functions, and cellular components of the predicted genes, we submitted the best BLASTX hits from the NR database for each transcript to the Blast2GO program (http://www.Blast2go.org) (Conesa et al. 2005). GO functional classification for all unigenes was performed using the Web Gene Ontology Annotation Plot (WEGO: http://wego.genomics.org.cn) software (Ye et al. 2006). For KEGG pathway annotation, we made sequence comparisons against the KEGG database (Kanehisa et al. 2004) using the BLASTP algorithm with an E-value threshold of 10−6.
Initial mapping of reads and gene expression analysis
We mapped RNA-seq reads from the control, salt-stressed, and drought-stressed samples onto the CDS assembly using the Bowtie2 (version 2.1.0) software (Langmead et al. 2009) with default parameters to estimate expression levels. Alignment output files from Bowtie were parsed using eXpress (version 1.5.0) (Roberts and Pachter 2013) to calculate the number of reads for each gene and standard FPKM values (numbers of fragments per kilobase per million mapped fragments on exon regions within a given gene) for each coding gene based on the final read assignments. The genes with FPKM ≥1 in samples from two or more time points were selected for further analysis. To compare expression levels of selected genes across all time points and reduce the bias in FPKM values due to highly expressed genes, the upper quartile normalization procedure (Bullard et al. 2010) was applied.
Identification and functional enrichment analysis of DEGs
To identify differentially expressed genes (DEGs) among different samples collected at different time points, we used the “abundance_estimates_to_matrix.pl,” “run_DE_analysis.pl” (edgeR), and “analyze_diff_expr.pl” modules of the Trinity package with default settings. The log2 fold changes >2 and a false discovery rate (FDR) significance score <0.001 were used as the threshold to identify significant differences in gene expression. The identified DEGs shared by all time points under both stresses were BLAST searched against PlantTFDB (plant transcription factor database) (version 3.0; http://planttfdb.cbi.pku.edu.cn/), clustered and used for GO and KEGG Orthology (KO) enrichment analysis.
Cluster analysis and functional enrichment
K-means clustering of transcript expression patterns based on FPKM values, obtained as described above, was performed using the K-Means/K-Medians Support Module (KMS) embedded in MultiExperiment Viewer 4.9 (http://www.tm4.org/mev). The optimal number of clusters (four clusters) was defined by the Figures of Merit (FOM) application (Yeung et al. 2001) within MEV4.9. The K-Means analysis was performed ten times, with each run generating four clusters using Euclidean distances. The final output generated four clusters in which all member genes clustered together in at least 80% of the K-Means runs (Li et al. 2010). Genes in each cluster were then classified according to GO terms and Fisher’s exact test was applied to test for enrichment of functional categories with Bonferroni’s correction (corrected P < 0.05).
Validation of DEGs by quantitative real-time PCR
To validate the reliability of the RNA-seq analyses without replication, ten candidate DEGs highly related to drought stress and salt stress were selected for quantitative RT-PCR (qRT-PCR) tests. Gene-specific primer pairs were designed using the Integrated DNA Technologies (IDT) software (http://www.idtdna.com/Primerquest/Home/Index) and are listed in Supplemental Table 1. After treatment with DNase I (Tiangen, China), 2 g of RNA was used for reverse transcription with the SuperScript VILO cDNA Synthesis Kit (Invitrogen) according to the manufacturer’s guidelines. Real-time qPCRs were performed using a Multicolor Real-Time PCR Detection System (Bio-Rad) and the conditions for all reactions were as follows: 95 °C for 30 s, 40 cycles of 95 °C for 5 s, followed by 60 °C for 30 s, and 95 °C for 10 s. Melting curve analysis was performed to confirm the PCR specificity. Three biological replicates were used for each gene. The relative expression levels of the selected DEGs, normalized to expression of the internal reference gene β-actin, were calculated using the 2−ΔΔCt method (Livak et al. 2001).
Results
Sequencing and de novo assembly of the C. sinensis transcriptome
To comprehensively cover the transcriptomic responses of C. sinensis to drought stress and salinity stress, ten cDNA libraries from the leaves of three PEG-stressed plant cuttings, three NaCl-stressed plant cuttings, and four no-stress controls were constructed. These libraries were then sequenced with an Illumina Hiseq 2500 platform. In total, 398.95 million 100 bp paired-end raw reads (~40.3 Gb sequence data) were generated (Table 1). After filtering the low-quality reads and trimming the adapter sequences, 370.39 million clean paired-end reads (~37.42 Gb) were acquired to perform de novo assembly using Trinity (Haas et al. 2013). To reduce redundancy and gain high-quality unigenes, 187,283 reconstructed transcripts were clustered into 137,477 unigenes with an N50 of 1085 bp and average length of 708 bp (Supplemental table 2) by CD-Hit (Li and Godzik 2006), then predicting the CDSs and removing those lower expressed CDSs and retain the CDSs with RPKM ≥1 in samples from two or more time points by custom PYTHON scripts. As a result, a high-quality dataset containing 64,905 CDs were recovered for C. sinensis with an N50 of 963 bp and average length of 730 bp (Table 2). The CDs number could be an overestimation due to allelic variation or alternative splicing. The size distribution for the assembled CDSs is shown in Fig. 1a. Those high-quality sequences were used for further analysis.
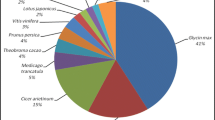
Characteristics of the C. sinensis transcriptome assembly and the homology search of CDSs against the NR database. a Size distribution of the assembled CDSs. b Effects of query sequence length on percentage of significant matches with a cutoff value of 1.0e−6. The proportion of sequences with matches in the NR database at NCBI is greater among the longer assembled sequences. c Similarity distribution of the best BLAST hits for each CDS. d E-value distribution of the top BLAST hits for each CDS. e Species distribution is shown as the percentage of the total homologous sequences
Functional annotation and classification of C. sinensis transcriptome
To predict the function of 64,905 transcripts, all transcript sequences were aligned against those from the NCBI NR database using BLASTP (Fig. 1a–e). Plant species giving top hits include Vitis vinifera with 11,246 (25.7%) hits, Coffea canephora with 2236 (5.1%) hits, Theobroma cacao with 2126 (4.9%) hits, Nelumbo nucifera with 1900 (4.3%) hits, and Sesamum indicum with 1824 (4.2%) hits (Fig. 1e). As reported in a previous study (Shi et al. 2011), the length of transcript sequences is crucial in determining the efficiency of BLAST searches. Our results showed that over 98% of the matching efficiency was observed for sequences longer than 1000 bp, whereas the matching efficiency dropped to about 86.4% for those ranging from 500 to 1000 bp, and to 59% for sequences between 400 and 500 bp; a lower matching efficiency (about 39.2%) was shown for sequences shorter than 400 bp (Fig. 1b). The similarity distribution of top hits from the NR database showed that 54.7% of mapped sequences had similarities higher than 80%, while a very small ratio of hits (0.025%) had similarities ranging between 30 and 40% (Fig. 1c). E-value distributions had a comparable pattern, with 45.4% of mapped sequences with high homologies (=<1e−100), and 54.6% of homologous sequences with E-values ranging between 1e−6 and 1e−100 (Fig. 1d). A total of 43,692 (67.3%) significant BLAST top hits were returned for GO annotation using Blast2GO (Table 2), of which 24,106 were successfully categorized into GO groups (S1 Fig). The distribution of GO terms for cellular components, molecular functions, and biological processes is shown in S1 Fig. As previously described (Zhang et al. 2015), for the cellular component category, genes involved in the “cell” (GO:0005623),“cell part” (GO:0044464) and “organelle part” (GO:0044422) were highly represented. For molecular function category, “catalytic” (GO:0003824) was the most enriched GO term, followed by “binding” (GO:0005488). For the biological processes category, the major subcategories were “metabolic process” (GO:0008152) and “cellular process” (GO:0009987). Of the 24,106 sequences annotated with GO terms, 7494 sequences were assigned with EC numbers (Table 2). To fully analyze EC numbers and biological pathways that are active in C. sinensis, we assigned all assembled sequences KO identifiers using the KEGG Automatic Annotation Server (KAAS) with the single-directional best hit information method, and subsequently mapped them to pathways and enzymes using the KEGG API. As a result, 23,371 sequences of 64,905 assembled sequences were assigned to 336 KEGG pathways (Supplemental Table 2). The most represented pathways by unique genes were ribosomes (129 members), biosynthesis of amino acids (102 members), and spliceosomes (102 members). The 64,905 assembled sequences were BLASTP searched against the Pfam database (version 28.0) with an E-value of 1e−6, of which 39,875 sequences were successfully assigned.
Mapping sequences to the C. sinensis transcriptome
Using the CDSs as reference, the FRKM values were analyzed to evaluate gene expression levels. The results showed that above 52% genes have FPKM values were distributed in the range of 1–100 for all time points (Fig. 2a). The correlation dendrogram in Fig. 2b illustrates expression patterns at the four analyzed time points under salt and drought stress. The results showed that gene expression patterns were clustered together under the salt-stressed and drought-stressed samples at 72 h, whereas the seedling plants at 24, 48 h for salt and drought stressed were clustered into one group, suggesting that to some extent the expression profiles of C. sinensis display the same patterns in response to drought and salt stress. In addition, the control samples collected at 0, 24, 48, and 72 h were clustered in a third group (Fig. 2b), demonstrating that gene expression levels were stable at the examined time points.
Transcriptomes characteristics of C. sinensis under drought stress and salt stress. a Number of genes expressed at each time point. b Hierarchical clustering of all samples by Spearman correlations based on FPKM (FPKM ≥1 in more than one time point). Heatmap with cluster dendrogram showing global relationships of the expressed genes between time points of drought stress and salt stress
In total, we identified 3936 and 3715 DEGs showing up- or down-regulation under drought stress and salt stress compared with the controls, respectively (Fig. 3a, b), while fewer DEGs (268) were identified in control samples (Control-0h, Control-24h, Control-48h, Control-72h) (Fig. 3c). Among these DEGs, 369 DEGs were present in all time points under drought stress (Fig. 3a), 207 DEGs were detected in all time points of salt stress (Fig. 3b), and 105 DEGs were shared under both salt stress and drought stress in all the detected time points. (Supplemental Fig. 2; Supplemental material 3). Noteworthy, no common DEGs were detected in all time points of control samples (Fig. 3c), indicating that only subtle changes in gene expressions in all time points under normal growth conditions were present; in contrast, great changes under drought stress and salt stress were experienced by the plants. Moreover, we found that 2131 DEGs (accounting for more than 67%) were shared by both stress conditions (Fig. 3d). This suggests that some mechanisms may be shared in the responses to drought and salt stress in tea. Out of all identified DEGs under drought stress and salt stress, 1306 and 1322 DEGs, respectively, were assigned GO annotations, for further analysis (Supplemental material 4, Supplemental material 5).
Venn diagrams showing unique and shared DEGs in the treated sample transcriptomes compared to the control samples (Control-0h) in C. sinensis with drought stress (PEG) and salt stress (NaCl). a DEGs of each time point of samples with drought stress. b DEGs of each time point of samples with salt stress. c DEG of each time point of samples without treatment. d DEGs shared by drought stress and salt stress in each time point and union set. The number of DEGs is detailed on the diagrams, all means the union set of all DEGs shared in drought stress and salt stress
GO category enrichment of DEGs under salt stress and drought stress
To speculate the function of the DEGs activated under drought stress or salt stress, we applied GO category enrichment analysis, using Fisher’s exact test with P value ≤0.05 as the cutoff. Using K-Mean clustering algorithm, we clustered all 3936 and 3715 DEGs into four main groups (designated P1-4 and N1-4, respectively) with highly similar temporal expression patterns (Fig. 4a, c; Supplemental material 4; Supplemental material 5). Four groups of genes with similar trends were detected under both stresses: (1) a continuously down-regulated trend was uncovered in genes of P1 and N1 in all time points, (2) in contrast to P1/N1, genes of P2 and N4 cluster were continuously up-regulated in all time points; (3) genes of P3 and N2 were most strongly expressed at 24 h, and recovered to the primary expression level at 48 h and 72 h; (4) genes of P4 and N3 showed a subtle up- and down-regulation at 24 and 48 h, respectively, and showed a great up-regulation at 72 h.
Dynamic progression of C. sinensis transcriptome under drought stress and salt stress. a and c showing the dynamic expression of the C. sinensis transcriptome under drought stress (P1–4) and salt stress (N1–4), respectively, by K-means clustering. b and d showing the functional enrichment among the clusters. e Functional enrichment among the DEGs shared in drought stress and salt stress
Furthermore, the groups of genes in each cluster showed confident enrichments for particular functional categories (corrected P < 0.05) (Fig. 4b, d). In general, similar functional categories were also enriched under both stress conditions. Under both stress conditions, continuously down-regulated genes (P1 and N1) were suggested to be required for developmental process, metabolic process, response to stimulus, and catalytic activity, whereas continuously up-regulated genes (P2 and N3) were predicted to be involved in catalytic activity, developmental processes, transporter activity, response to stress, and secondary metabolism were greatly enriched. Consistently, GO category enrichment analysis for 2131 DEGs shared by drought stress and salinity stress showed that these DEGs were mainly involved in “hydrolase activity,” “transferase activity,” “tetrapyrrole binding,” and “oxidoreductase complex” (Fig. 4d), in which most of them belong to continuously down-regulated genes and continuously up-regulated genes. However, the functional categories of the early stresses response genes were observed to be different between drought stressed and salinity stressed plants. In the early drought stressed stages (24 h, cluster P3), the up-regulated genes were mainly involved in “response to stimulus,” “anatomical structure morphogenesis,” “transporter activity,” and “kinase activity,” while genes whose expression peaked during the early salinity stressed stages (24 h, cluster N2) were “DNA binding,” “catalytic activity,” “carbohydrate binding,” “receptor activity,” “response to stress,” and “kinase activity.” These results provide support for molecular mechanisms being generally common in response to both drought-stressed and salinity-stressed plants in spite of differences at the early response stage (24 h).
Moreover, we performed the comparative analysis of GO category enrichment between DEGs under salt stress and drought stress (Fig. 4e; Supplemental material 6; Supplemental material 7). The results showed that the DEGs of both of stresses were commonly involved in 13 GO categories. DEGs belonging to seven categories including catalytic activity, biotic stimulus, ion binding, glutamate metabolic process, glutamate biosynthetic process, carboxylyase activity, and cofactor binding were present under salt stress but absent under drought stress. In contrast, four GO categories including copper ion binding, UDP-galactorsyltransferase activity, cofactor transporter activity, and NAD transporter activity only contained the DEGs from drought stress. These results also supported that some mechanisms may be shared in the responses to drought and salt stress in tea, and further suggest variation of molecular response to drought and salt stress in tea plants.
Identification of transcription factors in response to salt stress and drought stress
Transcriptional control of the expression of stress-responsive genes is a crucial part of the plant response to stresses (Singh et al. 2002). Dynamics of the expression of transcription factors (TFs) under salinity and drought stress were extensively determined in our RNA-seq data. Based on the K-Means clustering algorithm, the expression patterns of differentially expressed TFs could be clustered into three groups for both of the stress tolerance in C. sinensis (K1–K3 for drought stress and J1–J3 for salt stress) (Fig. 5a, b; Supplemental material 8; Supplemental material 9). Under drought stress, more than half of these genes (54%) were expressed at highest levels at 72 h (J1), 19% were expressed at highest levels at 0 h (J2), and the remaining 27% showed a peak of expression at 24 h (J3). Moreover, the TF family-specific expression trends were also analyzed (Fig. 5c). Members of the bHLH family, PPR family, and homeobox domain (HB) of transcriptional regulators were highly expressed in the leaf of C. sinensis under drought stress after 72 h treatment; these are known transcriptional activators in abscisic acid signaling in Arabidopsis thaliana (Abe et al. 2003). Members of the MYB, ERF, ZFWD4, and ABRF gene families were highly expressed in the leaves of C. sinensis under drought stress for 24 h, these transcription factors have been reported to play roles in controlling the expression of genes in response to stress at early stages in model plants (Abe et al. 1997; Terol et al. 2001; Sakuma et al. 2002).
Dynamics of transcription factor accumulation profiles. a and b showing the dendrogram of the transcription factors (TF); 476 and 395 significantly differentially expressed TFs from each time point of drought stress and salt stress clustered into three lineages respectively (J1–3 in drought stress and K1–3 in salt stress) using MEV4.9. c and d show the distribution of TF families among J1–3 and K1–3, respectively
Under salinity stress, the TFs within the K1 cluster presented down-regulation at all stressed time points; TFs within K2 cluster were expressed at the highest levels at the 24 h time point, and the TFs within K3 cluster had the highest levels at 72 h (Fig. 5b). The analysis of expression of TF family-specific trends showed that the K1 cluster included members of ARF, HB, TIR-NBS-LRR, and SBP families, K2 cluster included DNAJ and C2H2, and K3 cluster included PKF, MYB, ERF, WRKY, MIKC, and LOB (Fig. 5d). These TF families are known as key regulators of gene expression in response to abiotic stresses (Yamaguchi-Shinozaki and Shinozaki 2005).
Analysis of drought stress and salt stress related metabolic pathways using DEGs
KEGG pathway enrichment analyses were performed to identify the candidate pathway/s involved in stress response using KOBAS 2.0 (Xie et al. 2011). Genes with KO numbers within 3936 DEGs under drought stress were enriched in 18 KEGG pathways (Fig. 6a; Supplemental material 10) with pathways such as starch and sucrose metabolism, plant hormone signal transduction, photosynthesis-antenna proteins, photosynthesis, phenylpropanoid biosynthesis, glutathione metabolism, and cysteine and methionine metabolism being overrepresented during drought stress. Genes with KO numbers within 3715 DEGs under salt stress were enriched in 23 KEGG pathways (Fig. 6b; Supplemental material 11), in which phenylpropanoid biosynthesis, phenylalanine metabolism, starch and sucrose metabolism, plant hormone signal transduction, photosynthesis, and cysteine and methionine metabolism pathways were overrepresented. Furthermore, enriched by genes with KO numbers within the pathways such as starch and sucrose metabolism, plant hormone signal transduction, photosynthesis, starch and sucrose metabolism, photosynthesis-antenna proteins, photosynthesis, galactose metabolism, and cysteine and methionine metabolism with a lower rich factor were overrepresented in the 2131 DEGs shared by drought stress and salt stress (Fig. 6c; Supplemental material 12).
Scatterplot of enriched KEGG pathways for differentially expressed genes in drought stress and salt stress. The DEGs from each time point of drought stress (a), salt stress (b), and shared in drought stress and salt stress (c). Rich factor is the ratio of the differentially expressed gene number to the total gene number in a certain pathway. The size and color of dot represent the gene number and the range of the FDR value, respectively
Validation of transcriptome expression patterns by qRT-PCR analysis
To validate the gene expression results based on RNA-seq, ten candidate DEGs associated with salt- and drought-related processes were selected for qRT-PCR assays, the detected expression patterns were consistent for all ten genes, confirming the reliability of the RNA-seq results (Fig. 7).
qRT-PCR verification of ten selected DEGs. Comparison of RNA-seq data (blue bar) with qRT-PCR data (red line). The normalized expression level (FPKM) of RNA-seq is indicated on the left y-axis. The relative qRT-PCR expression level is shown on the right y-axis. Actin was used as the internal control. Both methods showed similar gene expression trends (color figure online)
Discussion
C. sinensis is an economically important crop, greatly affected by environmental abiotic stresses such as drought and high salinity, which are critical factors affecting its quality. Although cold stress (Vinay and Sudesh 2009) and temperature stress (Wu et al. 2015) in C. sinensis have already been reported in previous studies, drought and salt stress remain poorly understood in C. sinensis. Much of the work addressing the mechanisms of plant response to drought and salt stresses focused on Arabidopsis or rice, where many regulatory networks are known to participate in these processes (Zhu 2002). In this study, to understand the molecular mechanisms of C. sinensis response to both stresses, we performed a time-course RNA-seq analysis for C. sinensis for salt and drought stress. Due to the high costs associated with RNA-seq, we pooled five individuals for one library construction to avoid the bias of individual samples. However, pooling biological replicate samples causes loss of group variance, and thus we cannot determine the statistical significance of DEGs in our study. These large-coverage transcriptome datasets based on RNA-seq provided the clue for estimating the potential gene function, but wet-lab experiment would be necessary to further verify gene expression. Therefore, ten candidate DEGs were selected for qRT-PCR tests; the results showed that the gene expression levels identified by RNA-seq were consistent with qRT-PCR, confirming the reliability of RNA-seq analysis for gene expression.
The similarity and divergence of DEGs under drought and salt stress in C. sinensis
In this study, 3936 DEGs and 3715 were identified in C. sinensis under salt stress and drought stress, respectively (Fig. 3d). Comparative analysis of GO enrichments between the two stresses suggested that the two stresses contained the DEGs commonly involved in more than half of GO categories (13/20 under salt stress and 13/17 under drought stress) (Fig. 4e). The similarity of the DEGs between drought and salt stresses in C. sinensis indicated that C. sinensis has similar response mechanisms under these two stresses. Moreover, 2131 DEGs were identified to be shared under drought and salt stresses in C. sinensis (Fig. 3d). KEGG pathway enrichment results showed that C. sinensis responses to drought and salt stress overlapped in starch and sucrose metabolism, plant hormone signal transduction, cysteine and methionine metabolism, and glutathione metabolism (Fig. 6c), findings that are consistent with previous observations showing an overlap of the responses in Arabidopsis (Wong et al. 2006), rice (Rabbani et al. 2003), and maize (Sun et al. 2015) to drought and salt stress.
Drought and salt stress generally have shared DEGs; however, the analysis of different DEGs between salt and drought stress may help discover the different molecular mechanisms. Under drought stress, the GO analysis showed that the DEGs were relative to developmental growth, single-organism developmental processes, macromolecular complex binding, and molecular transducer activity in C. sinensis. Under salt stress, the DEGs were relative to lipid metabolic process, carbohydrate binding, homeostatic processes, regulation of biological quality, transferase activity, cellular homeostasis, and cell death in C. sinensis (Fig. 4a, b). In plants, prevention of osmotic stress caused by drought depends upon minimizing stomatal and cuticular water loss and maximizing water uptake, while osmotic adjustment appears to play a major role in maintaining osmotic homeostasis during salt stress (Chinnusamy et al. 2002). Therefore, it is understandable that the DEGs relative to homeostatic processes, cellular homeostasis were present under salt stress in C. sinensis . Based on the presence of these general and specific abiotic stress tolerance mechanisms, it is reasonable to expect C. sinensis to have multiple stress perception and signal transduction pathways, which may have crosstalk at various steps in the pathways.
In this study, the result of gene expression pattern and its ontology and KEGG pathway enrichment show that, under salt stress, the expression of genes participating in the process of “homeostatic process”, “cellular homeostasis” were down-regulated at all analyzed time points, while genes participating in the process of “catalytic activity”, “kinase activity”, “transferase activity”, “response to stress” and “receptor activity” were up-regulated either at the initial stage (24 h) or at the all four stages (24, 48, and 72 h) (Fig. 4c, d). These results indicated that ion transporters may be phosphorylated by protein kinases to reestablish cellular homeostasis in salt-treated C. sinensis.
Under drought stress, we identified 25 genes involved in both cysteine and methionine metabolism, and in the glutathione metabolism pathway, with a further 33 and 36 genes being identified in starch and sucrose metabolism and plant hormone signal transduction, respectively (Fig. 6a). These results indicated that C. sinensis may regulate carbohydrate metabolic processes and developmental processes through hormones (such as ABA) and secondary metabolites in response to drought stress.
A large body of evidence has been gathered to support the idea that many abiotic stresses act in a cooperative manner, and nature has developed diverse signaling pathways for combating and tolerating them. These pathways act in cooperation to alleviate stress (Mahajan and Tuteja 2005). The correlation dendrogram illustrates expression patterns of salt and drought stress at the four points, the two stresses show a divergent expression pattern at 24 h, while a similar pattern was exhibited at 48 and 72 h (Fig. 2b); this result indicated that C. sinensis may have a slightly different mechanism in the response to salt and drought stress at the initial stage (24 h), but a signal transduction crosstalk may exist at the mid (48 h) and longtime stage (72 h).
The molecular response relative to metabolism associated drought and salt response in C. sinensis
Under water-deficit and other stress regimes, proline functions as a mediator of osmotic adjustment, a compatible solute to protect enzymes, a stabilizer of macromolecules, and as a store for carbon and nitrogen for use (Delauney and Verma 2002; Ashraf and Foolad 2007). As predicted, the DEGs related to the regulation of osmotic pressure were identified, such as the proline pathway genes Delta-1-pyrroline-5-carboxylate synthase (P5CS) (contig ID: m.60374 in Supplemental material 4), and two proline transporters (contig ID: m.60225 and m.60227) were observed to be unregulated in C. sinensis. Noteworthy, these genes were all present at higher expression levels under salt stress than under drought stress at the end of stress treatment (72 h), suggesting that salt stress has a more prolonged effect in regulating the pathway of proline in C. sinensis. Moreover, the GO categories of glutamate metabolisms including glutamate metabolic process and glutamate biosynthetic process were presented under salt stress but absent in drought stress (Fig. 4e). Glutamate appears to be the main precursor in stress-induced proline accumulation, which has been reported to occur in response to salt stress in tomato, potato, and Chinese Mitten Crab (Berteli et al. 2008; Teixeira and Fidalgo 2009; Wang et al. 2012). These results further supported that salt stress has a more prolonged effect than drought stress in regulating the pathway of proline in C. sinensis.
Under stress conditions, sugars act as osmoprotectants to help maintain osmotic balance and stabilize macromolecules (Suresh and A 1998). In this study, we identified 13 DEGs associated with sugar metabolism (Supplemental material 5). Remarkably, nine of these DEGs are sugar transporters, including five SWEETs (SUGARS WILL EVENTUALLY BE EXPORTED TRANSPORTER) and four other types of sugar transporters. All of them were up-regulated at the transcriptome level. To our knowledge, SWEETs have not been linked to drought and salt stress yet, while AtSWEET16 was shown to increase freezing tolerance in Arabidopsis. Additional work is required to evaluate the role of SWEETs in recapturing carbon during stress. These results suggest that disaccharide transport tends toward accumulation processes under salt and drought conditions to increase substance solubility and weaken osmotic pressure in C. sinensis.
Similarly, accumulation of amino acids has been observed in many studies on plants exposed to abiotic stress (Sanchez et al. 2008; Usadel et al. 2008; Lugan et al. 2010). The non-protein amino acid c-aminobutyric acid accumulates rapidly to high levels under multiple stress conditions (Rhodes et al. 1986; Bown and Shelp 1999; Kempa et al. 2008; Renault et al. 2010). In this study, the DEGs relative to GABA metabolism and transport were identified in C. sinensis (Supplemental material 5). Glutamate decarboxylase (GAD) (contigs ID: m.60581) was unregulated 2.3–4.8-fold and 1.8–5.6-fold in drought and salt-treated C. sinensis, and GABA transporters (m.41679 and m.41682) had higher expression levels under salt treatment than drought treatment in C. sinensis. GABA is mainly synthesized from glutamate by GAD in the cytosol, and is subsequently transported to the mitochondria in plants. Salt stress enhances the activity of enzymes involved in GABA metabolism in model plants (Renault et al. 2010). It is possible that GABA is accumulated under both salt and drought stress but is transported more actively under salt than under drought stress in C. sinensis.
Nevertheless, tea qualities were apparently affected by stress as the plants responded to osmotic pressure by changing their metabolic products. Polyphenol and caffeine are considered to be key components for tea qualities. Both polyphenol and caffeine are the important active ingredients in tea. In this study, we observed that two DEGs (m.62690 and m.62689) encoding caffeine synthase decreased their expression under salt stress and under drought stress. Moreover, a DEG (m.25702) encoding polyphenol oxidase dramatically increased up to 91-fold under both stresses, demonstrating that the metabolism of polyphenol is regulated by salt and drought stress in C. sinensis. The recognizable brown color of tea is developed by enzymatic browning by PPO during the fermentation process in tea production (Subramanian et al. 1999). Therefore, these results provide the evidence at the molecular level that stress affects tea quality. Further experiments such as characterizing the spatio-temporal expression, enzymatic activity assays, and physiological assays would be necessary to evaluate the extent of the effect of salt and drought stress in C. sinensis.
Transcription factors involved in drought- and salt-stress responses in C. sinensis
Both salinity and drought stress have a strong impact on gene expression leading to changes of stress-associated metabolite levels (Krasensky and Jonak 2012). Distinct TFs family genes, such as bHLH, WRKY, MYB, NAM, CBF, and NBS, have been documented to be functional as early responders to environmental signals, mediating plant adaptation to various stresses, such as drought and salt stress, and overexpression of these TFs enhanced drought or salt tolerance of transgenic plants (Seki et al. 2002; Takasaki et al. 2010). In this study, we detected 77, 79, and 50 DEGs in bHLH, MYB, and NAC family, respectively (Fig. 5a), that were either at initial stage (24 h) or at later stage (72 h) up-regulated in drought stressed C. sinensis (Fig. 5c), while a higher proportion of DEGs in those families (87% in bHLH family, 88% in MYB family, 75% in NAC family) were up-regulated in salt-stressed C. sinensis (Fig. 5b, d). In addition, all TIR-NBS-LRR DEGs were up-regulated in both drought- and salt-stressed C. sinensis (Fig. 5a–d). Previous research found that 80% of DEGs in the bHLH family and MYB family, 60% of DEGs in the WRKY family, and 33% of DEGs in the NAC family were up-regulated in cold-stressed C. sinensis (Wang et al. 2013). Functional characterization of bHLH TFs indicated that these TF family members act as critical regulators in various physiological processes, such as stomata development (Pires and Dolan 2010), and chloroplast development (Monte et al. 2004); OsbHLH148 was shown to be involved in mediating drought tolerance by modulating jasmonate signaling (Seo et al. 2011), TabHLH1 is critical in mediating plant adaptation to osmotic stresses largely through its function in the modulation of the ABA-dependent-signaling pathway (Seo et al. 2011). Thus, in C. sinensis, it is apparent that various regulatory mechanisms may act to adapt to drought, salt, and cold stress, and crosstalk pathways exist among the responses to these stresses.
In nature, stress does not generally come in isolation and many different stresses act at the same time. In response to those crosstalking stress signals, nature has developed diverse signaling pathways for combating and tolerating them. These pathways act in cooperation to alleviate stress (Mahajan and Tuteja 2005). The components of drought and salt stress exhibit crosstalk, and these stress conditions could both lead to cell dehydration, osmotic imbalance, and associated oxidative stress (Mahajan and Tuteja 2005; Wong et al. 2006).
Conclusions
In this study, we presented an original transcriptome sequencing analysis of RNA from C. sinensis leaves under a prolonged time-course for drought stress and salt stress. A total of ~18.7 Gb reads was generated and assembled into 64,905 CDSs, of which 67.3% were annotated with gene descriptions, while the remaining 32.7% were unknown. Identified from all analyzed time points of drought and salt stress were 3936 and 3715 DEGs, respectively; 2131 overlapping DEGs were further identified in drought stress and salt stress indicating that C. sinensis has a similar molecular response to these two stresses. Two DEGs (m.62690 and m.62689) encoding caffeine synthase decreased their expression under salt stress and under drought stress, and a DEG (m.25702) encoding polyphenol oxidase dramatically increased under both stresses, demonstrating that the metabolism of polyphenol is regulated by salt and drought stress in C. sinensis. The expression patterns of ten unigenes were analyzed by qRT-PCR to verify the RNA-seq results. Our studies provide valuable insight into C. sinensis responses to drought and salt stress, and the identified DEGs provide important resources for further functional analyses and potential transgenic modifications to develop drought and salt-tolerant C. sinensis.
References
Abe H, Urao T, Ito T et al (2003) Arabidopsis AtMYC2 (bHLH) and AtMYB2 (MYB) function as transcriptional activators in abscisic acid signaling. Plant Cell 15(1):63–78. doi:10.1105/tpc.006130
Abe H, Yamaguchi-Shinozaki K, Urao T et al (1997) Role of Arabidopsis MYC and MYB homologs in drought- and abscisic acid-regulated gene expression. Plant Cell 9(10):1859–1868. doi:10.1105/tpc.9.10.1859
Allakhverdiev SI, Sakamoto A, Nishiyama Y et al (2000) Ionic and osmotic effects of NaCl-induced inactivation of photosystems I and II in Synechococcus sp. Plant Physiol 123(3):1047–1056. doi:10.1104/pp.123.3.1047
Anthony Y (1998) Predicting the interaction between the effects of salinity and climate change on crop plants. Sci Hortic 78(1–4):159–174. doi:10.1016/S0304-4238(98)00193-9
Arbona V, Manzi M, Cd O, Gómez-Cadenas A (2013) Metabolomics as a tool to investigate abiotic stress tolerance in plants. Int J Mol Sci 14(3):4885–4911. doi:10.3390/ijms14034885
Ashraf M, Foolad MR (2007) Roles of glycine betaine and proline in improving plant abiotic stress resistance. Environ Exp Bot 59(2):206–216. doi:10.1016/j.envexpbot.2005.12.006
Bahieldin A, Atef A, Sabir JS (2013) Analysis of the barley leaf transcriptome under salinity stress using mRNA-Seq. C R Biol 35(6):1915–1924. doi:10.1016/j.crvi.2015.03.010
Beritognolo I, Harfouche A et al (2011) Comparative study of transcriptional and physiological responses to salinity stress in two contrasting Populus alba L. genotypes. Tree Physiol 31(12):1335–1355. doi:10.1093/treephys/tpr083
Berteli F, Corrales E, Guerrero C et al (2008) Salt stress increases ferredoxin-dependent glutamate synthase activity and protein level in the leaves of tomato. Physiol Plantarum 93(2):259–264. doi:10.1111/j.1399-3054.1995.tb02226.x
Binzel M, Ratajczak R (2002) Function of membrane transport systems under salinity: tonoplast. Springer, Netherlands
Bohnert HJ, Nelson DE, Jensen RG (1995) Adaptations to environmental stresses. Plant Cell 7(7):1099–1111. doi:10.1105/tpc.7.7.1099
Boyer JS (1982) Plant productivity and environment. Science 218(4571):443–448. doi:10.1126/science.218.4571.443
Bown AW, Shelp BJ (1999) Metabolism and functions of gamma-aminobutyric acid. Plant Physiol 115(1):1–5. doi:10.1104/pp.115.1.1
Bullard JH, Purdom E, Hansen KD, Dudoit S (2010) Evaluation of statistical methods for normalization and differential expression in mRNA-Seq experiments. BMC Bioinformatics 11:94. doi:10.1186/1471-2105-11-94
Chen C, Khaleel SS, Huang H, Wu CH (2015) NGS QC toolkit: a platform for quality control of next-generation sequencing data. Source Code Biol Med 7(2):1–5. doi:10.1186/1751-0473-9-8
Chinnusamy V, Stevenson B, Lee BH, Zhu JK (2002) Screening for gene regulation mutants by bioluminescence imaging. Sci STKE 2002(140):l10. doi:10.1126/stke.2002.140.pl10
Conesa A, Götz S, García-Gómez JM et al (2005) Blast2GO: a universal tool for annotation, visualization and analysis in functional genomics research. Bioinformatics 21(18):3674–3676. doi:10.1093/bioinformatics/bti610
Delauney AJ, Verma DPS (2002) Proline biosynthesis and osmoregulation in plants. Plant J 4(2):215–223. doi:10.1046/j.1365-313X.1993.04020215.x
Eiji O, Yuki M, Yasuaki S et al (2004) Effects of exogenous application of proline and betaine on the growth of tobacco cultured cells under saline conditions. Soil Science and Plant Nutrition 50(8):1301–1305. doi:10.1080/00380768.2004.10408608
Goode JA,Organizers DC (2007) Improving plant drought, salt and freezing tolerance by gene transfer of a single stress-inducible transcription factor. John Wiley & Sons, Ltd. doi:10.1002/9780470515778.ch13
Grabherr MG, Haas BJ, Yassour M et al (2011) Full-length transcriptome assembly from RNA-Seq data without a reference genome. Nat Biotechnol 29(7):644–652. doi:10.1038/nbt.1883
Haas BJ, Papanicolaou A, Yassour M et al (2013) De novo transcript sequence reconstruction from RNA-seq using the Trinity platform for reference generation and analysis. Nat Protoc 8(8):1494–1512. doi:10.1038/nprot.2013.084
Hasegawa PM, Bressan RA, Zhu JK, Bohnert HJ (2000) Plant cellular and molecular responses to high salinity. Annu Rev Plant Physiol Plant Mol Biol 51:463–499. doi:10.1146/annurev.arplant.51.1.463
Hoque MA, Okuma E, Banu MN et al (2007) Exogenous proline mitigates the detrimental effects of salt stress more than exogenous betaine by increasing antioxidant enzyme activities. J Plant Physiol 164(5):553–561. doi:10.1016/j.jplph.2006.03.010
Jaglo-Ottosen KR, Gilmour SJ, Zarka DG et al (1998) Arabidopsis CBF1 overexpression induces COR genes and enhances freezing tolerance. Science 280(5360):104–106. doi:10.1126/science.280.5360.104
Kanak V, Arpita G, Vinay K et al (2012) De novo transcriptome sequencing in L. to identify genes involved in the biosynthesis of diosgenin. Plant Genome 6(2):1–11. doi:10.3835/plantgenome2012.08.0021
Kanehisa M, Goto S, Kawashima S et al (2004) The KEGG resource for deciphering the genome. Nucleic Acids Res 32(Database issue):277–280. doi:10.1093/nar/gkh063
Kempa S, Krasensky J, Dal Santo S et al (2008) A central role of abscisic acid in stress-regulated carbohydrate metabolism. PLoS One 3(12):e3935. doi:10.1371/journal.pone.0003935
Krasensky J, Jonak C (2012) Drought, salt, and temperature stress-induced metabolic rearrangements and regulatory networks. J Exp Bot 63(4):1593–1608. doi:10.1093/jxb/err460
Kristiansson E, Asker N, Förlin L, Larsson DG (2009) Characterization of the Zoarces viviparus liver transcriptome using massively parallel pyrosequencing. BMC Genomics 10(1):345. doi:10.1186/1471-2164-10-345
Langmead B, Trapnell C, Pop M, Salzberg SL (2009) Ultrafast and memory-efficient alignment of short DNA sequences to the human genome. Genome Biol 10(3):1–10. doi:10.1186/gb-2009-10-3-r25
Li P, Ponnala L, Gandotra N et al (2010) The developmental dynamics of the maize leaf transcriptome. Nat Genet 42(12):1060–1067. doi:10.1038/ng.703
Li W, Godzik A (2006) Cd-hit: a fast program for clustering and comparing large sets of protein or nucleotide sequences. Bioinformatics 22(13):1658–1659. doi:10.1093/bioinformatics/btl158
Liu M, Qiao G, Jiang J et al (2012) Transcriptome sequencing and de novo analysis for ma bamboo (Dendrocalamus latiflorus Munro) using the Illumina platform. PLoS One 7(10):136–136. doi:10.1371/journal.pone.0046766
Liu Q, Kasuga M, Sakuma Y et al (1998) Two transcription factors, DREB1 and DREB2, with an EREBP/AP2 DNA binding domain separate two cellular signal transduction pathways in drought- and low-temperature-responsive gene expression, respectively, in Arabidopsis. Plant Cell 10(8):1391–1406. doi:10.1105/tpc.10.8.1391
Livak KJ, Schmittgen TD et al (2001) Analysis of relative gene expression data using real-time quantitative PCR and the 2-ΔΔCt method. Methods 25:402–408
Lugan R, Niogret MF, Leport L et al (2010) Metabolome and water homeostasis analysis of Thellungiella salsuginea suggests that dehydration tolerance is a key response to osmotic stress in this halophyte. Plant J 64(2):215. doi:10.1111/j.1365-313X.2010.04323.x
Mahajan S, Tuteja N (2005) Cold, salinity and drought stresses: an overview. Arch Biochem Biophys 444(2):139–158. doi:10.1016/j.abb.2005.10.018
Meyer E, Aglyamova GV, Wang S et al (2009) Sequencing and de novo analysis of a coral larval transcriptome using 454 GSFlx. BMC Genomics 10(1):219. doi:10.1186/1471-2164-10-219
Monte E, Tepperman JM, Al-Sady B et al (2004) The phytochrome-interacting transcription factor, PIF3, acts early, selectively, and positively in light-induced chloroplast development. Proc Natl Acad Sci U S A 101(46):16091–16098. doi:10.1073/pnas.0407107101
Pires N, Dolan L (2010) Origin and diversification of basic-helix-loop-helix proteins in plants. Mol Biol Evol 27(27):862–874. doi:10.1093/molbev/msp288
Rabbani MA, Maruyama K, Abe H et al (2003) Monitoring expression profiles of rice genes under cold, drought, and high-salinity stresses and abscisic acid application using cDNA microarray and RNA gel-blot analyses. Plant Physiol 133(4):1755–1767. doi:10.1104/pp.103.025742
Renault H, Roussel V, El Amrani A et al (2010) The Arabidopsis pop2-1 mutant reveals the involvement of GABA transaminase in salt stress tolerance. BMC Plant Biol 10(1):20. doi:10.1186/1471-2229-10-20
Rhodes D, Handa S, Bressan RA (1986) Metabolic changes associated with adaptation of plant cells to water stress. Plant Physiol 82(4):890–903. doi:10.1104/pp.82.4.890
Robert DF, Bateman A, Clements J et al (2014) Pfam: the protein families database. Nucleic Acids Res 42(Database issue):D222–D230. doi:10.1093/nar/gkt1223
Roberts A, Pachter L (2013) Streaming fragment assignment for real-time analysis of sequencing experiments. Nat Methods 10(1):71–73. doi:10.1038/nmeth.2251
Rogers PJ, Smith JE, Heatherley SV, Pleydell-Pearce CW (2008) Time for tea: mood, blood pressure and cognitive performance effects of caffeine and theanine administered alone and together. Psychopharmacology 195(4):569–577. doi:10.1007/s00213-007-0938-1
Rus AM, Bressan RA, Hasegawa PM (2005) Unraveling salt tolerance in crops. Nat Genet 37(10):1029–1030. doi:10.1038/ng1005-1029
Sakuma Y, Liu Q, Dubouzet JG et al (2002) DNA-binding specificity of the ERF/AP2 domain of Arabidopsis DREBs, transcription factors involved in dehydration- and cold-inducible gene expression. Biochem Biophys Res Commun 290(3):998–1009. doi:10.1006/bbrc.2001.6299
Sanchez DH, Siahpoosh MR, Roessner U et al (2008) Plant metabolomics reveals conserved and divergent metabolic responses to salinity. Physiol Plant 132(2):209–219. doi:10.1111/j.1399-3054.2007.00993.x
Seki M, Narusaka M, Ishida J et al (2002) Monitoring the expression profiles of 7000 Arabidopsis genes under drought, cold and high-salinity stresses using a full-length cDNA microarray. Plant J 31(31):279–292. doi:10.1046/j.1365-313X.2002.01359.x
Seo JS, Joo J, Kim MJ et al (2011) OsbHLH148, a basic helix-loop-helix protein, interacts with OsJAZ proteins in a jasmonate signaling pathway leading to drought tolerance in rice. Plant J 65(6):907–921. doi:10.1111/j.1365-313X.2010.04477.x
Shi CY,Wan XC,Jiang CJ, et al. (2007) Method for high-quality total RNA isolation from tea plants [Camellia sinensis (L.) O. Kuntze)]. Journal of Anhui Agricultural University.
Shi CY, Yang H, Wei CL et al (2011) Deep sequencing of the Camellia sinensis transcriptome revealed candidate genes for major metabolic pathways of tea-specific compounds. BMC Genomics 12(1):131. doi:10.1186/1471-2164-12-131
Shinozaki K, Yamaguchi-Shinozaki K (2000) Molecular responses to dehydration and low temperature: differences and cross-talk between two stress signaling pathways. Curr Opin Plant Biol 3(3):217–223. doi:10.1016/S1369-5266(00)80068-0
Singh K, Foley RC, Oñate-Sánchez L (2002) Transcription factors in plant defense and stress responses. Curr Opin Plant Biol 5(5):430–436. doi:10.1016/S1369-5266(02)00289-3
Subramanian N, Venkatesh P, Ganguli S, Sinkar VP (1999) Role of polyphenol oxidase and peroxidase in the generation of black tea theaflavins. J Agric Food Chem 47(7):2571. doi:10.1021/jf981042y
Sun C, Gao X, Fu J et al (2015) Metabolic response of maize ( Zea mays L.) plants to combined drought and salt stress. Plant Soil 388(1):99–117. doi:10.1007/s11104-014-2309-0
Suresh I, Allan C (1998) Products of proline catabolism can induce osmotically regulated genes in rice. Plant Physiol 116(1):203–211. doi:10.1104/pp.116.1.203
Takasaki H, Maruyama K, Kidokoro S et al (2010) The abiotic stress-responsive NAC-type transcription factor OsNAC5 regulates stress-inducible genes and stress tolerance in rice. Mol Gen Genomics 284(3):173–183. doi:10.1007/s00438-010-0557-0
Teixeira J, Fidalgo F (2009) Salt stress affects glutamine synthetase activity and mRNA accumulation on potato plants in an organ-dependent manner. Plant Physiol Bioch 47(9):807. doi:10.1016/j.plaphy.2009.05.002
Terol J, Bargues M, Pérez-Alonso M (2001) ZFWD: a novel subfamily of plant proteins containing a C3H zinc finger and seven WD40 repeats. Gene 260(1–2):45–53. doi:10.1016/S0378-1119(00)00446-7
Tuteja N (2007) Mechanisms of high salinity tolerance in plants. Methods Enzymol 428:419–438. doi:10.1016/S0076-6879(07)28024-3
Urano K, Kurihara Y, Seki M, Shinozaki K (2010) ‘Omics’ analyses of regulatory networks in plant abiotic stress responses. Curr Opin Plant Biol 13(2):132–138. doi:10.1016/j.pbi.2009.12.006
Usadel B, Bläsing OE, Gibon Y et al (2008) Multilevel genomic analysis of the response of transcripts, enzyme activities and metabolites in Arabidopsis rosettes to a progressive decrease of temperature in the non-freezing range. Plant Cell Environ 31(4):518–547. doi:10.1111/j.1365-3040.2007.01763.x
Vera JC, Wheat CW, Fescemyer HW et al (2008) Rapid transcriptome characterization for a nonmodel organism using 454 pyrosequencing. Mol Ecol 17(7):1636–1647. doi:10.1111/j.1365-294X.2008.03666.x
Vinay K, Sudesh K (2009) Proline and betaine provide protection to antioxidant and methylglyoxal detoxification systems during cold stress in Camellia sinensis (L.) O. Kuntze. Acta Physiol Plant 31(2):261–269. doi:10.1007/s11738-008-0227-6
Wang XC, Zhao QY, Ma CL et al (2013) Global transcriptome profiles of Camellia sinensis during cold acclimation. BMC Genomics 14(1):1–15. doi:10.1186/1471-2164-14-415
Wang Y, Li E, Yu N et al (2012) Characterization and expression of glutamate dehydrogenase in response to acute salinity stress in the Chinese mitten crab, Eriocheir sinensis. PLoS One 7(5):e37316. doi:10.1371/journal.pone.0037316
Wong CE, Li Y, Labbe A et al (2006) Transcriptional profiling implicates novel interactions between abiotic stress and hormonal responses in Thellungiella, a close relative of Arabidopsis. Plant Physiol 140(140):1437–1450. doi:10.1104/pp.105.070508
Wu ZJ, Li XH, Liu ZW et al (2015) Transcriptome-based discovery of AP2/ERF transcription factors related to temperature stress in tea plant (Camellia sinensis). Funct Integr Genomics 15(6):741. doi:10.1007/s10142-015-0457-9
Xie C, Mao X, Huang J et al (2011) KOBAS 2.0: a web server for annotation and identification of enriched pathways and diseases. Nucleic Acids Res 39(Web Server issue):W316–W322. doi:10.1093/nar/gkr483
Yamaguchi-Shinozaki K, Shinozaki K (2005) Organization of cis-acting regulatory elements in osmotic- and cold-stress-responsive promoters. Trends Plant Sci 10(2):88–94. doi:10.1016/j.tplants.2004.12.012
Ye J, Fang L, Zheng H et al (2006) WEGO: a web tool for plotting GO annotations. Nucleic Acids Res 34(Web Server issue):W293–W297. doi:10.1093/nar/gkl031
Yeung KY, Haynor DR, Ruzzo WL (2001) Validating clustering for gene expression data. Bioinformatics 17(4):309–318. doi:10.1093/bioinformatics/17.4.309
Zhang HB, Xia EH, Huang H et al (2015) De novo transcriptome assembly of the wild relative of tea tree (Camellia taliensis) and comparative analysis with tea transcriptome identified putative genes associated with tea quality and stress response. BMC Genomics 16:298. doi:10.1186/s12864-015-1494-4
Zhu JK (2002) Salt and drought stress signal transduction in plants. Annu Rev Plant Biol 53(53):247–273. doi:10.1146/annurev.arplant.53.091401.143329
Acknowledgements
This project was supported by grants from the 863 program (2013AA102604) and the funding from the Fujian Agriculture and Forestry University.
Data archiving statement
The Illumina sequencing datasets are available from the European Nucleotide Archive database (ENA; http://www.ebi.ac.uk/ena) under project number accession PRJEB11522. The experiment accession numbers of cDNA libraries obtained from the control tea plants for 0, 24, 48, and 72 h in cultivation nutrient solution only are ERX1180353, ERX1180350, ERX1180351, and ERX1180352, respectively. The samples from plant cuttings exposed to salt stress or drought stress for 24, 48, and 72 h were ERX1180347, ERX1180348, ERX1180349, ERX1180344, ERX1180345, and ERX1180346, respectively.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Rights and permissions
About this article
Cite this article
Zhang, Q., Cai, M., Yu, X. et al. Transcriptome dynamics of Camellia sinensis in response to continuous salinity and drought stress. Tree Genetics & Genomes 13, 78 (2017). https://doi.org/10.1007/s11295-017-1161-9
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1007/s11295-017-1161-9