Abstract
Antimicrobial peptides (AMPs) are small peptides existing in nature as an important part of the innate immune system in various organisms. Notably, the AMPs exhibit inhibitory effects against a wide spectrum of pathogens, showcasing potential applications in different fields such as food, agriculture, medicine. This review explores the application of AMPs in the food industry, emphasizing their crucial role in enhancing the safety and shelf life of food and how they offer a viable substitute for chemical preservatives with their biocompatible and natural attributes. It provides an overview of the recent advancements, ranging from conventional approaches of using natural AMPs derived from bacteria or other sources to the biocomputational design and usage of synthetic AMPs for food preservation. Recent innovations such as structural modifications of AMPs to improve safety and suitability as food preservatives have been discussed. Furthermore, the active packaging and creative fabrication strategies such as nano-formulation, biopolymeric peptides and casting films, for optimizing the efficacy and stability of these peptides in food systems are summarized. The overall focus is on the spectrum of applications, with special attention to potential challenges in the usage of AMPs in the food industry and strategies for their mitigation.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Outbreaks of foodborne diseases listed on FDA website reveal a dangerous trend. A total of 24 outbreaks are mentioned in 2023 resulting in illness or hospitalization (https://www.fda.gov/food/outbreaks-foodborne-illness/investigations-foodborne-illness-outbreaks). Some food microbes such as strains of Listeria monocytogenes, Salmonella, Escherichia coli are pathogenic and responsible for severe diseases; nevertheless, there are other microbes including but not limited to Brochothrix thermosphacta, Carnobacterium spp., Lactobacillus spp., Lactococcus spp., that may not cause illnesses, yet food spoilage caused by these microbes can be economically devastating. The actual amount of economic loss caused by food spoilage is unknown, although it is believed that 25% of food produced worldwide is lost owing to spoilage microbes (Bondi et al. 2014). Reducing food spoilage and FBDs is the main goal of using food preservation techniques, such as thermal or non-thermal treatments, food additives, and improved packaging. Artificial preservatives like sorbates, nitrites, and benzoates inhibit microbial activity, antioxidants like formaldehyde, butylated hydroxytoluene, and butylated hydroxyanisole are used to stop food from oxidizing. However, a growing number of individuals are looking for minimally processed foods free of artificial substances because they are worried about the safety of chemical preservatives.
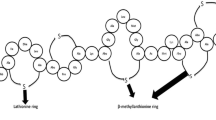
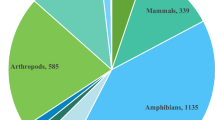
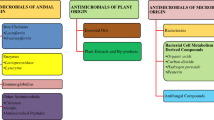
Antimicrobial peptides (AMPs) are a diverse class of molecules that all living things produce and are considered a part of innate immunity. These are sometimes also referred to as peptide antibiotics (Zasloff 2002). They are peptides that range in size from 7 to 100 amino acids. A lot of research interest has been generated by AMPs' capacity to fight bacteria that are resistant to drugs. Additionally, some AMPs have immune-modulatory qualities that support pathogen clearance in an indirect manner. They have found a role in diverse fields such as healthcare, agriculture, and the food industry. These peptides are classified based on their diverse sources, amino acid composition, activity, and structural attributes (Huan et al. 2020). Sources of AMPs encompass animals, plants, microorganisms, and recently synthetic AMPs. AMPs grouped on the basis of their amino acid composition include peptides rich in either proline, glycine, histidine, or tryptophan, while their activity can be characterized as either broad-spectrum or specific to certain pathogens. Structural classifications include α-helical, β-sheet, loop, and extended peptides. Mechanisms of action of AMPs include killing microbes by disrupting the cell membrane, inhibiting the production of proteins and nucleic acids, or binding to specific targets inside the cell, ultimately leading to cell death (Yonezawa et al. 1992; Graf et al. 2017; Boix-Lemonche et al. 2020).
Researchers are currently investigating the potential of AMPs to inhibit the growth of microbes, for extending the shelf life of various food products, and prevention of FBDs (Zhang et al. 2021; Baindara and Mandal 2022; Jha and Singh 2023). Nevertheless, there are still a number of technical obstacles that need to be addressed before AMPs may be used in food preservation. These include issues with large-scale synthesis, toxicity, haemolytic activity, stability, immunogenicity, and other possible drawbacks. Recent research investigations have focused on a number of solutions intended to alleviate the aforementioned limitations. These comprise rational design and modification of AMPs, biocomputational and bioinformatics analysis based in silico designing and active packaging systems (Agyei et al. 2018; Aguilera-Puga et al. 2023; Jordan et al. 2024).
AMPs derived from variety of sources as preservatives in different food types
In response to customer demands for safer and more natural food preservation techniques, AMPs present a possible substitute for conventional chemical preservatives. In addition to antimicrobial activity, AMPs have demonstrated antioxidant activity, which is of interest to the food industry (Lima et al. 2019). Some of the AMPs have the potential of being used as food additives and are classified as ‘Generally Regarded as Safe’ (GRAS) such as natural variants of Nisin (Nisin-A, Z, F, Q, U, U2, H, P) which are among the most important FDA approved AMPs as food preservative (Field et al. 2023). There are many similar approved/GRAS AMPs which include but not limited to Natamycin (Meena et al. 2021), Enterocin AS-48 (Dijksteel et al. 2021), Lactoferricin (Singh et al. 2023), ε-Polylysine and Pediocin PA-1 (Luz et al. 2018; Santos et al. 2018). Several AMPs have been evaluated for their potential to inhibit microbes in a number of food matrices, some of which are summarized as follows or in Table 1.
Milk and dairy products
Milk and its products are a vital source of nutrition, but they are susceptible to spoilage by microorganisms, leading to economic losses. Natural bio-preservatives, such as AMPs, offer a promising solution to preserve milk and its products without compromising their nutritional quality or safety (Mohanty et al. 2016). A novel AMP named LCWAP designed by screening amino acid sequences from the whey acidic protein (WAP) of large yellow croaker (Larimichthys crocea) exhibited a minimal inhibitory concentration (MIC) of 15.6 μg/mL against Staphylococcus aureus in milk and inhibited biofilm formation up to 94.3%, without showing cytotoxic effects on normal human hepatocytes (Yang et al. 2020). Similarly, a peptide Cerein 8A was found to reduce Listeria monocytogenes in milk by 3 log cycles over 14 days at 4 °C and decreased viable counts by 2 log cycles on contaminated cheese surfaces over 30 days (Bizani et al. 2008). Another study reported liposome-encapsulated Nisin and BLS P34 for preserving Minas frescal cheese by controlling L. monocytogenes in which these AMPs showed 88.9 and 100% entrapment efficiency in liposomes, respectively. These treatments significantly reduced the bacterial counts compared to the control during 21 days of storage of cheese at 7 °C (Malheiros et al. 2012).
Fruits and vegetables
AMPs have shown promise in inhibiting the growth of spoilage microorganisms and extending the shelf life of fruits and vegetables without the use of synthetic additives. Overall, the focus of research is shifting towards preserving fruits and vegetables in ways that are both effective and environmentally friendly. Arulrajah et al. (2021) studied antifungal peptides from kenaf seeds to extend the shelf life of tomato puree. Mixture of these peptides when applied to tomato puree significantly reduced fungal counts and delayed growth of Aspergillus niger and Fusarium sp. up to 14 and 23 days at 25 °C and 4 °C respectively. Shwaiki et al. (2020) examined Snakin-1, a peptide extracted from potato tubers, against Zygosaccharomyces bailii, a yeast responsible for spoilage in different beverages such as Fanta Orange (Coca-Cola, Ireland) (pH 3.1), SuperValu (Chilled 169 Cranberry Juice) (pH 2.7) and apple juice (CYPRINA) (pH 3.5), and observed a complete inhibition of Z. bailii in all juices at 200 µg/mL. A cationic antimicrobial peptide PAF56 inhibited spore formation, disrupted cell membranes, and was non-toxic to human red blood cells when it was evaluated in preservation of citrus fruits by targeting Penicillium digitatum, Penicillium italicum, and Geotrichum candidum, demonstrating its potential as an effective agent for postharvest citrus disease control (Wang et al. 2021). The antimicrobial peptide CB-M was studied by Yang et al. (2023) against Botrytis cinerea, a fungus causing Gray mold disease in cherry tomatoes, tomatoes, and grapes fruits. Results showed a strong dose dependent inhibitory effect on spore survival and mycelial growth of B. cinerea thus reducing the disease.
Meat, fish or sea food
Meat and fish are prized for their nutritional value and taste, but they are highly perishable due to microbial contamination and altered sensory properties because of lipid oxidation. Dang et al. (2020) found Musca domestica derived AMPs to increase membrane permeability and inhibition of bacterial growth in chilled pork at concentrations 0.4 to 0.8 mg/ml with negligible hemolytic activity against human erythrocytes. Nie et al. (2021) designed chimeric lysins by fusing Salmonella phage lysin with the peptide LeuA-P, which could effectively reduce the microbial counts in contaminated chilled chicken and extend its shelf life by seven days. Similar studies have reported the use of AMPs to minimize lipid oxidation and reduce the bacterial count in meat or meat products (Przybylski et al. 2016; Jabeen and Khanum 2017). AMPs have also been found to play a crucial role in preserving fish or sea food by inhibiting microbial growth (Shabir et al. 2018; Ning et al. 2019). Sm-A1, a peptide derived from turbot viscera, was successfully loaded into poly vinyl alcohol-chitosan hydrogel which could effectively protect the salmon muscle from the microbiological contamination and texture deterioration (Bi et al. 2020). Peng et al. (2018) investigated the effect of Nisin along with 1% chitosan as preservatives on the quality of Jumbo squid and reported a reduction in both microbial growth and nutritional loss.
Cereal crops and their products
AMPs have been found to be effective in protection of various cereal crops or their products. Unclean conditions in storage godowns make food grain unfit for human consumption due to microbial deterioration. AMPs LR14 were studied for the prevention of wheat grain spoilage under storage which prevented fungal growth even after a prolonged storage for more than 2 years (Gupta and Srivastava 2014). Furthermore, transgenic technology has been used for heterogenous expression of AMPs in crops like rice and maize which provides them with resistance against phytopathogens (Noonan et al. 2017; Tang et al. 2023). Additionally, studies have reported the efficacy of AMPs in improving shelf life of cereal derived food products such as cakes and noodles (Xiao and Niu 2015; Luz et al. 2018; Lu et al. 2022).
Challenges of the AMPs applications
The application of AMPs in general, and particularly for food preservation, has many challenges. These include high production costs, possible toxicity for eukaryotic cells, a lack of stability, antigenicity and development of resistance, that need to be addressed. The final fate of the AMPs, including their absorption, distribution, metabolism, excretion, and toxicity (ADMET), are other aspects to be looked into for efficient therapeutic efficiency (Okella et al. 2022).
The major challenge that synthetic AMPs face in their application in food preservation is the expensive cost of synthesis, which impedes the large-scale production and usage of these peptides. The poor stability of AMPs is another major challenge due to their susceptibility to the action of proteases or other environmental factors such as temperature and pH. AMPs stability has also been associated with geometrical features such as ovality, lipophilicity, radius of gyration, and polar surface area. The thermostability of AMPs is very important in food production since thermal treatments are generally used during food processing (Al-sahlany et al. 2020).
Another major problem with applications of AMPs is their possible toxicity to the host cells. Due to hydrophobic structure with positive charges, AMPs have both advantage of broad-spectrum bioactivity and problem of potential biotoxicity (Wei and Zhang 2022). Although the nonspecific mechanism of action of AMPs is a major advantage in terms of broad-spectrum activity encompassing a wide range of microorganisms, there may be chances of uptake of these AMPs by off-target cells instead or in addition to the target cells. It may result in undesired effects on eukaryotic cells, leading to toxicity or hemolytic activity. Natural AMPs with cationic charges can interact with negative ions on the surface of the cell membrane, and then form oligomers to destroy cells (Spaller et al. 2013). These potential toxicities may be reduced by change in amino acid composition of peptides or by attachment with biomaterials (Wei and Zhang 2022). In addition, certain microbiota that are part of the digestive system may uptake these AMPs and develop resistance against the peptides, resulting in a risk of dissemination of the resistance through horizontal gene transfer to pathogens (Crits-Christoph et al. 2022; Tajer et al. 2024).
Further, there are many elements in foods such as proteases and peptidases that can react with or interact with AMPs, thereby reducing their bioactivity (Udenigwe and Fogliano 2017). Also, reactions can occur between amino groups on peptides with carbonyl groups on reducing sugars present in foods, resulting in inactive form of peptides (Lund and Ray 2017). The need for studies on the behavior of AMPs within complex food systems is imperative since the effect of some food processing parameters on AMP activity, such as temperature and pH, is often studied separately.
Improved design or modification of AMPs
There are many strategies for rationally designing AMPs for improved efficiency against microbes, reduced toxicity, and increased stability (Rai et al. 2016; Giacometti and Buretić-Tomljanović 2017; Agyei et al. 2018). Important concerns in applications of AMPs and designing strategies to mitigate these concerns are shown as Fig. 1.
The effective functioning of AMPs requires adherence to appropriate design principles. It involves creating the AMP units, or AMP moiety, to be available for the best possible performance. Even the size and aggregation of the newly created peptides affect the functional characteristics of the AMPs. In order to achieve the interaction of the proper domain, the hydrophilic and hydrophobic regions of a composite AMP assembly should be oriented appropriately, e.g., the systematic evolution of ligands by exponential enrichment (SELEX) forms. Certain peptides may operate better even when combined with single-stranded oligonucleotide versions of nucleic acids (Lee et al. 2022). The biological activity of AMPs has been demonstrated to be enhanced via hybridization, which involves covalently joining two or more peptide segments to reap the benefits of each fragment (Tian et al. 2019).
The multiple hurdle concept is another approach wherein combinations of natural antimicrobials with nonthermal processing technologies such as ultrasound, pulsed-electric field, high pressure, and ozone treatment have shown potential synergistic effects. Applying these nonthermal techniques may cause bacterial cell membranes to deteriorate or make them more vulnerable to AMPs, which would increase the lethality of these peptides (Molinos et al. 2008).
Various strategies like cyclization, capping, lipidation, glycosylation, dimerization/multimerization, dendrimerization, phosphorylation, acetylation have been explored for increasing the stability of the AMPs (Rounds and Straus 2020; Bitencourt et al. 2023; Kumari et al. 2023; Mironov et al. 2024). Cyclization (Liu et al. 2017) and peptide stapling (Selvarajan et al. 2023) have been employed to enhance the stability of the AMPs against proteolytic degradation. Another way to improve AMP stability against proteases is to cap the AMPs. Attaching an acyl group or hydrophobic end modification are some of the N- or C-terminal capping methods to improve the stability of AMPs (Zhong et al. 2019). Dimerization or multimerization is obtained by incorporating the peptide side chains or peptide branches in the α- or ε-amino groups of certain amino acids (Santos-Filho et al. 2021). Using this strategy, more peptide monomers are grouped, which eventually boosts the stability of the whole unit by providing protection against proteolytic degradation due to steric hindrance (Bitencourt et al. 2023).
To create AMPs that are both highly active and less toxic, different approaches have been used, which include truncating the original peptides while preserving the active component and site-directed alteration of the amino acid residues (Huan et al. 2020).
In silico analysis for AMP prediction and discovery
AMPs in their initial days of discovery were isolated from different natural sources, but now a days, synthetic AMPs are also in application. These molecules are well known for their diverse modes of action against pathogens, which reflects their potential in terms of therapeutic and food preservative usage. However, a few limitations have also been identified with the applications of AMPs (Lombardi et al. 2015; Zhao et al. 2016; Oshiro et al. 2019), as mentioned earlier. These limitations can be overcome through in silico analysis using a combination of bioinformatics or computational models with the aim of improving the performance, biocompatibility, and safety of AMPs. The classical methods, such as isolation and identification of AMPs, are very time-consuming and laborious. For the practical use of AMPs as food preservatives, it is important to reduce the experimental time spent identifying AMPs. A number of computational models have been developed for the effective design and discovery of AMPs through in silico analysis. These models have helped in the rapid and accurate prediction of peptides that can be chemically synthesized and investigated for their suitability for use as food preservatives. In general, these predictive computational models are broadly based on empirical methods and machine learning (ML) approaches (Porto et al. 2012).
The empirical methods used for AMP design are qualitative in nature. These models are governed by rules or patterns of antimicrobial activity. Practical issues may arise for empirical approaches because of their low accuracy and complex analysis. In contrast, the ML models have demonstrated their usefulness in well-structured screening and prediction of AMP sequences. The models used on such platforms make final inferences using comprehensive training data (Jia et al. 2015). One popular machine learning approach for creating and optimizing AMPs is the quantitative structure activity relationship, or QSAR model (Mitchell 2014). This model uses physicochemical parameters and amino acid sequences to envisage the biological action of AMPs (Hilpert et al. 2008).
Interestingly, the combination of the predictive and generative machine learning (ML) models is opined to be best suited for the designing of AMPs (Aguilera-Puga et al. 2023). The ML platforms use various algorithms like support vector machine (SVM), K-nearest neighbor (KNN), random forest (RF), and neural network (NN). The SVM algorithms consider factors like amino acid composition, physiochemical properties, and structural features of AMPs. These can also be used to forecast peptide membrane activity with high predictive value. The KNN algorithms rely on pattern recognition methods. Xiao et al. (2013) demonstrated the use of the fuzzy K-nearest neighbor (F-KNN) method and multi-label classifier which finally classify AMPs into different types based on activity. The RF algorithm-based methods predict AMPs using the distribution pattern of properties of peptides acquired from available AMPs and non-AMPs sequences (Lawrence et al. 2021), while the NN method can be used to identify the pattern in a sequence database and model the structure–activity relationship of AMPs (Wang et al. 2024).
In addition to the popular ML methods, de novo computational prediction platforms have helped generate a variety of peptide sequences with different amino acid compositions, interaction structures, and modes of action (Hiss et al. 2010). These platforms use information on amino acid frequency and position preferences which offers insights into the structure and amphipathicity of peptides (Porto et al. 2012).
An evolutionary/genetic algorithm-based approach has been utilized recently using fitness functions like activity descriptors and data gathered from databases to identify a variety of AMPs (Torres and de La Fuente-Nunez 2019). Using these algorithms in conjunction with other models like NNs and computational tools such as molecular docking, the output potential of AMP prediction can be further enhanced. Table 2 provides details on several models and tools for in silico investigation of AMPs.
Workflow for in silico analysis
Adoption of a prudent AMPs analytical workflow or pipeline is necessary for a thorough assessment of the antibacterial potential, physicochemical characteristics, and biocompatibility of peptides in order to design and identify appropriate AMPs with a broad range of activity. A typical workflow for AMP development or in silico prediction may have four stages of analysis (Melo et al. 2021; Aguilera-Puga et al. 2023). Gathering experimental data on amino acid composition, structure, biological activity, and physiochemical characteristics is the initial stage. In the second stage, the information is converted to a digital format that supports ML so that it may be analyzed. The third step comprises the actual use of ML models with selection of algorithms having high accuracy and reliability. In the fourth phase, the best AMP candidate is identified, which can be validated for activity. Various in vitro biocompatibility analysis parameters like hemolysis prediction, water solubility prediction, toxicity prediction, peptide half-life prediction, and protease susceptibility analysis can also be included (Melo et al. 2021). One of the important aspects of in silico analysis is antigenicity prediction, as designed AMPs should be free from antigenicity. Various safety analysis databases and tools to design AMP are given in Table 3.
Active food packaging and creative fabrication of AMPs
Food packaging is an important aspect of today’s food industry, as it plays a major role in safeguarding the quality of products and the safety of consumers. In smart packaging, functionality beyond the standard function of serving as a physical barrier between the product and its surroundings is included. Various types of packaging on the basis of the engineered functionality of the package include ergonomic packaging, informative packaging, responsive packaging, and active packaging (Singh and Heldman 2001; Brockgreitens and Abbas 2016).
Active packaging is achieved by incorporating antimicrobial agents into the packaging material and ensuring a controlled release over an extended period to preserve the food quality. The mechanism of packaging of AMPs can be summed up by three interconnected sections, which include the incorporation system, which comprises the addition of AMPs into the packaging material; the release system, which ensures the diffusion of peptides from the surface of the package to the food; and the interaction and inhibition system, involving the interaction and subsequently destruction of microbes present in the food (Sultana et al. 2021).
For the purpose of enhancing efficacy and stability in a range of applications, including food packaging, AMPs are manufactured or encapsulated in a variety of ways, as described as follows.
Nanoencapsulation
One of the most common methods used for the fabrication of AMPs for active packaging of food is nanoencapsulation. The two primary stages in nanoencapsulation are incorporating antibacterial agents into their appropriate carriers and reducing their size to the nanoscale. The stability of AMPs is generally improved by nanoencapsulation, which also offers controlled release, high adsorption, defense against environmental variables, and less unfavorable interactions with food ingredients. Additionally, it gives the loaded particles a larger surface area than the bigger particles for higher efficacy. The efficiency of AMPs has been improved by their integration into nanoparticles such as lactoferrin, chitosan, polycaprolactone, polyethylene oxide, liposomes, poly (lactic-co-glycolic acid) commonly known as PLGA and even extending to binary or ternary complexes of such nanoparticles in order to preserve the physiochemical properties of food items (Duarte and Picone 2022). There are a variety of nanoencapsulation systems, including nanofibers, liposomes, nanoemulsions.
Nanofibers can be synthesized using artificial or natural polymers. These possess a large surface area of contact, which makes it possible to load more than one molecule at once. They are produced by various methods, such as electrospinning, self-assembly, laser spinning, and melt spinning, of which electrospinning is the most common. AMP-loaded nanoparticles, when electrospun into nanofibers such as polyethylene oxide, have demonstrated increased effectiveness against foodborne bacteria without producing any effect on sensory characteristics (Cui et al. 2017). AMP-conjugated nanotubes or nanocomposites have also been studied with promising results (Hemmati et al. 2021).
Liposomes are spherical nanocarriers based on lipids that are frequently utilized in delivery systems due to their biocompatibility, stability, and ease of synthesis. Liposome-encapsulated AMPs are also widely used in the food industry. AMPs such as Nisin have shown higher antimicrobial activity when loaded into liposomes as compared to when directly applied to the food (Malheiros et al. 2012). Furthermore, liposomes have demonstrated stabilization of the trapped materials against different environments and chemical changes such as pH, temperature, enzymatic modifications. (Mozafari et al. 2008).
Nano-emulsions are also one of the lipid-based nanoencapsulation systems that have been employed in active food packaging. A variety of biopolymers are employed to create emulsions for the inclusion of AMPs (Imran et al. 2012).
Biopolymeric AMPs
Another approach to using AMPs in food preservation is their integration into biopolymers for increased effectiveness. Biopolymers can be synthesized in vitro in cell-free systems, e.g., dextran, or derived from nature, e.g., alginate or hyaluronic acid. Being biocompatible and biodegradable in nature, they can be used as edible films or for packaging food materials (Baranwal et al. 2022). One of the most commonly used biopolymers is chitosan. Its antifungal, antibacterial, and antioxidant qualities are widely recognized. It has shown a synergistic effect in combination with AMPs against the pathogens, along with other characteristics such as decreased moisture content and higher oxygen barrier effectiveness (Luo et al. 2023). Many approaches have been used for the incorporation of AMPs into biopolymers. One of them involves mixing polymer with AMP using click chemistry and then embedding it into another polymer (Chiloeches et al. 2023). It creates a robust packaging material that would prevent leaching out of the components. Combining biopolymer derivatives with antimicrobial peptide dendrimers (AMPDs) is another strategy that targets both the inner and outer membranes of bacteria (Jordan et al. 2024). Controlled release due to retention of the antimicrobial compounds in polymer matrices is also achieved by the use of nanofillers (Jamróz et al. 2019).
Casting films
The fabrication of antimicrobial films integrating AMPs using the casting approach has been documented (Meira et al. 2017). The use of nanofillers results in films being mechanically resistant, with enhanced thermal performance and good water-resistant qualities. AMP encapsulation has also been combined with techniques like biopolymer immobilization to form biodegradable films that showed improved barrier properties against food-borne pathogens (Imran et al. 2012). Jamróz et al. (2021) demonstrated the effectiveness of active double-layered furcellaran/gelatin hydrolysate films containing Ala-Tyr peptide for fish preservation.
Cold plasma technology
Cold plasma technology is a novel environmental-friendly approach for activating polymers that employs the use of naturally or laboratory-produced plasma, which contains a mixture of ions, photons, reactive species for synthesizing polymers or modifying their surface for conjugating AMPs (Jordá-Vilaplana et al. 2014). This technology has been used for the immobilization of biologically active functional substances such as Nisin, Lysozyme, and Vanillin into the packaging material. Polymers used in food packaging, such as polyethylene terephthalate, effectively prevented the formation of biofilms when they were surface-plasma activated and subsequently conjugated to immobilized peptides (Agrillo et al. 2019).
Conclusion and future perspectives
Although antimicrobial peptides offer themselves as an ideal substitute to chemical additives for preventing food spoilage and foodborne infections, their application in this sector is not without limitations. Ideally AMPs to be used for food preservation should be chemically well designed for high effectiveness, nontoxic to native host cells and normal microbiota, stable and cost-effective. As a result, biocompatibility testing is required prior to employing AMPs as food preservatives. Furthermore, a comprehensive approach is required to establish particular conditions that must replicate those present during food processing and storage to determine AMPs applicability, stability, safety, biocompatibility, and potential interactions with food components. Numerous online tools including those based on ML can be utilized to investigate the various properties required for an effective and safe AMP. These are quite fast and accurate, potentially saving time and money as compared to lengthy experimental studies for AMP discovery and prediction. With the addition of new AMPs to the database, ML predictions are constantly improving, which will pave the way for the discovery of more effective and safe peptides in the future.
Data availability
No datasets were generated or analysed during the current study.
References
Agrawal P, Raghava GPS (2018) Prediction of antimicrobial potential of a chemically modified peptide from its tertiary structure. Front Microbiol 9:418508. https://doi.org/10.3389/fmicb.2018.02551
Agrillo B, Balestrieri M, Gogliettino M, Palmieri G, Moretta R, Proroga YT, Rea I, Cornacchia A, Capuano F, Smaldone G, De Stefano L (2019) Functionalized polymeric materials with bio-derived antimicrobial peptides for “active” packaging. Int J Mol Sci 20(3):601. https://doi.org/10.3390/ijms20030601
Aguilera-Puga MD, Cancelarich NL, Marani MM, de la Fuente-Nunez C, Plisson F (2023) Accelerating the discovery and design of antimicrobial peptides with artificial intelligence. Computational drug discovery and design, New York. Springer, NY, pp 329–352
Agyei D, Tsopmo A, Udenigwe CC (2018) Bioinformatics and peptidomics approaches to the discovery and analysis of food-derived bioactive peptides. Anal Bioanal Chem 410:3463–3472. https://doi.org/10.1007/s00216-018-0974-1
Al-sahlany STG, Altemimi AB, Al-Manhel AJ, Niamah AK, Lakhssassi N, Ibrahim SA (2020) Purification of bioactive peptide with antimicrobial properties produced by Saccharomyces cerevisiae. Food 9:324–325. https://doi.org/10.3390/foods9030324
Arulrajah B, Muhialdin BJ, Qoms MS, Zarei M, Hussin ASM, Hasan H, Saari N (2021) Production of cationic antifungal peptides from kenaf seed protein as natural bio preservatives to prolong the shelf-life of tomato puree. Int J Food Microbiol 359:109418. https://doi.org/10.1016/j.ijfoodmicro.2021.109418
Baindara P, Mandal SM (2022) Plant-derived antimicrobial peptides: novel preservatives for the food industry. Foods 11(16):2415. https://doi.org/10.3390/foods11162415
Baranwal J, Barse B, Fais A, Delogu GL, Kumar A (2022) Biopolymer: a sustainable material for food and medical applications. Polymers 14(5):983. https://doi.org/10.3390/polym14050983
Bi J, Tian C, Jiang J, Zhang GL, Hao H, Hou HM (2020) Antibacterial activity and potential application in food packaging of peptides derived from turbot viscera hydrolysate. J Agric Food Chem 68(37):9968–9977. https://doi.org/10.1021/acs.jafc.0c03146
Bitencourt NV, Righetto GM, Camargo ILBC, de Godoy MO, Guido RVC, Oliva G, Santos-Filho NA, Cilli EM (2023) Effects of dimerization, dendrimerization, and chirality in p-BthTX-I peptide analogs on the antibacterial activity and enzymatic inhibition of the SARS-CoV-2 PLpro protein. Pharmaceutics 15(2):436. https://doi.org/10.3390/pharmaceutics15020436
Bizani D, Morrissy JA, Dominguez AP, Brandelli A (2008) Inhibition of Listeria monocytogenes in dairy products using the bacteriocin-like peptide cerein 8A. Int J Food Microbiol 121(2):229–233. https://doi.org/10.1016/j.ijfoodmicro.2007.11.016
Boix-Lemonche G, Lekka M, Skerlavaj B (2020) A rapid fluorescence-based microplate assay to investigate the interaction of membrane active antimicrobial peptides with whole gram-positive bacteria. Antibiotics 9:92. https://doi.org/10.3390/antibiotics9020092
Bondi M, Messi P, Halami PM, Papadopoulou C, De Niederhausern S (2014) Emerging microbial concerns in food safety and new control measures. BioMed Res Int. https://doi.org/10.1155/2014/251512
Bournez C, Riool M, de Boer L, Cordfunke RA, de Best L, van Leeuwen R, Drijfhout JW, Zaat SA, van Westen GJ (2023) CalcAMP: A new machine learning model for the accurate prediction of antimicrobial activity of peptides. Antibiotics (Basel) 12(4):725. https://doi.org/10.3390/antibiotics12040725
Brockgreitens J, Abbas A (2016) Responsive food packaging: recent progress and technological prospects. Compr Rev Food Sci Food Saf 15(1):3–15. https://doi.org/10.1111/1541-4337.12174
Chaudhary K, Kumar R, Singh S, Tuknait A, Gautam A, Mathur D, Anand P, Varshney GC, Raghava GP (2016) A web server and mobile app for computing hemolytic potency of peptides. Sci Rep 6(1):22843. https://doi.org/10.1038/srep22843
Chiloeches A, Zágora J, Plachá D, Torres MD, de la Fuente-Nunez C, López-Fabal F, Gil-Romero Y, Fernández-García R, Fernández-García M, Echeverría C, Muñoz-Bonilla A (2023) Synergistic combination of antimicrobial peptides and cationic polyitaconates in multifunctional PLA fibers. ACS Appl Bio Mater 6(11):4805–4813. https://doi.org/10.1021/acsabm.3c00576
Crits-Christoph A, Hallowell HA, Koutouvalis K, Suez J (2022) Good microbes, bad genes? The dissemination of antimicrobial resistance in the human microbiome. Gut Microbes 14(1):2055944. https://doi.org/10.1080/19490976.2022.2055944
Cui H, Wu J, Li C, Lin L (2017) Improving anti-listeria activity of cheese packaging via nanofiber containing nisin-loaded nanoparticles. LWT 81:233–242. https://doi.org/10.1016/j.lwt.2017.04.003
Dang X, Zheng X, Wang Y, Wang L, Ye L, Jiang J (2020) Antimicrobial peptides from the edible insect Musca domestica and their preservation effect on chilled pork. J Food Process Preserv 44(3):14369. https://doi.org/10.1111/jfpp.14369
Dijksteel GS, Ulrich MM, Middelkoop E, Boekema BK (2021) Lessons learned from clinical trials using antimicrobial peptides (AMPs). Front Microbiol 12:616979. https://doi.org/10.3389/fmicb.2021.616979
Dong B, Wang Y, Cui G, Wang Y, Lin Y, Su Z, Zhao G (2024) In vitro antimicrobial activity of the novel antimicrobial peptide mytimacin-4 and its influence on the microbial community and quality of pork during refrigerated storage. Food Control 163:110486. https://doi.org/10.1016/j.foodcont.2024.110486
Duarte LG, Picone CS (2022) Antimicrobial activity of lactoferrin-chitosan-gellan nanoparticles and their influence on strawberry preservation. Food Res Int 159:111586. https://doi.org/10.1016/j.foodres.2022.111586
El-Saadony MT, Abd El-Hack ME, Swelum AA, Al-Sultan SI, El-Ghareeb WR, Hussein EO, Ba-Awadh HA, Akl BA, Nader MM (2021) Enhancing quality and safety of raw buffalo meat using the bioactive peptides of pea and red kidney bean under refrigeration conditions. Ital J Anim Sci 20(1):762–776. https://doi.org/10.1080/1828051X.2021.1926346
Field D, Fernandez de Ullivarri M, Ross RP, Hill C (2023) After a century of nisin research-where are we now? FEMS Microbiol Rev 47(3):fuad023. https://doi.org/10.1093/femsre/fuad023
Fingerhut LC, Miller DJ, Strugnell JM, Daly NL, Cooke IR (2020) ampir: an R package for fast genome-wide prediction of antimicrobial peptides. Bioinformatics 36(21):5262–5263. https://doi.org/10.1093/bioinformatics/btaa653
Gasteiger E, Hoogland C, Gattiker A, Wilkins MR, Appel RD, Bairoch A (2005) Protein identification and analysis tools on the Expasy server. In: Walker JM (ed) The proteomics protocols handbook. Humana Press, Totowa, pp 571–607
Giacometti J, Buretić-Tomljanović A (2017) Peptidomics as a tool for characterizing bioactive milk peptides. Food Chem 230:91–98. https://doi.org/10.1016/j.foodchem.2017.03.016
Graf M, Mardirossian M, Nguyen F, Seefeldt AC, Guichard G, Scocchi M, Innis CA, Wilson DN (2017) Proline-rich antimicrobial peptides targeting protein synthesis. Nat Prod Rep 34(7):702–711. https://doi.org/10.1039/C7NP00020K
Grande MJ, Lucas R, Abriouel H, Omar NB, Maqueda M, Martínez-Bueno M, Martínez-Cañamero M, Valdivia E, Gálvez A (2005) Control of Alicyclobacillus acidoterrestris in fruit juices by enterocin AS-48. Int J Food Microbiol 104(3):289–297. https://doi.org/10.1016/j.ijfoodmicro.2005.03.010
Gupta S, Ansari HR, Gautam A, Open-Source Drug Discovery Consortium, Raghava GP (2013) Identification of B-cell epitopes in an antigen for inducing specific class of antibodies. Biol Direct 8:1–15. https://doi.org/10.1186/1745-6150-8-27
Gupta S, Kapoor P, Chaudhary K, Gautam A, Kumar R, Open-Source Drug Discovery Consortium, Raghava GP (2013) In silico approach for predicting toxicity of peptides and proteins. PLoS ONE 8(9):73957. https://doi.org/10.1371/journal.pone.0073957
Gupta R, Srivastava S (2014) Antifungal effect of antimicrobial peptides (AMPs LR14) derived from Lactobacillus plantarum strain LR/14 and their applications in prevention of grain spoilage. Food Microbiol 42:1–7. https://doi.org/10.1016/j.fm.2014.02.005
Hazam PK, Selvaraj SP, Negi A, Lin WC, Chen JY (2024) Use of natural peptide TP4 as a food preservative prevents contamination by fungal pathogens. Food Chem 455:139874. https://doi.org/10.1016/j.foodchem.2024.139874
Hemmati F, Bahrami A, Esfanjani AF, Hosseini H, McClements DJ, Williams L (2021) Electrospun antimicrobial materials: advanced packaging materials for food applications. Trends Food Sci Technol 111:520–533. https://doi.org/10.1016/j.tifs.2021.03.014
Hilpert K, Fjell CD, Cherkasov A (2008) Short linear cationic antimicrobial peptides: screening, optimizing, and prediction. In: Otvos L (ed) Peptide-based drug design. Springer, Berlin, pp 127–159
Hiss JA, Hartenfeller M, Schneider G (2010) Concepts and applications of “natural computing” techniques in de novo drug and peptide design. Curr Pharm Des 16:1656–1665. https://doi.org/10.2174/138161210791164009
Hou J, Li YQ, Wang ZS, Sun GJ, M HZ, (2017) Applicative effect of glycinin basic polypeptide in fresh wet noodles and antifungal characteristics. LWT 83:267–274. https://doi.org/10.1016/j.lwt.2017.05.028
Huan Y, Kong Q, Mou H, Yi H (2020) Antimicrobial peptides: classification, design, application and research progress in multiple fields. Front Microbiol 11:582779. https://doi.org/10.3389/fmicb.2020.582779
Huang RH, Xiang Y, Liu XZ, Zhang Y, Hu Z, Wang DC (2002) Two novel antifungal peptides distinct with a five-disulfide motif from the bark of Eucommia ulmoides Oliv. FEBS Lett 521:87–90. https://doi.org/10.1016/s0014-5793(02)02829-6
Imran M, Revol-Junelles AM, René N, Jamshidian M, Akhtar MJ, Arab-Tehrany E, Jacquot M, Desobry S (2012) Microstructure and physico-chemical evaluation of nano-emulsion-based antimicrobial peptides embedded in bioactive packaging films. Food Hydrocoll 29(2):407–419. https://doi.org/10.1016/j.foodhyd.2012.04.010
Jabeen U, Khanum A (2017) Isolation and characterization of potential food preservative peptide from Momordica charantia L. Arab J Chem 10:3982–3989. https://doi.org/10.1016/j.arabjc.2014.06.009
Jamróz E, Kulawik P, Kopel P (2019) The effect of nanofillers on the functional properties of biopolymer-based films: a review. Polymers 11:1–42. https://doi.org/10.3390/polym11040675
Jamróz E, Kulawik P, Tkaczewska J, Guzik P, Zając M, Juszczak L, Krzyściak P, Turek K (2021) The effects of active double-layered furcellaran/gelatin hydrolysate film system with Ala-Tyr peptide on fresh Atlantic mackerel stored at −18°C. Food Chem 338:127867. https://doi.org/10.1016/j.foodchem.2020.127867
Jha B, Singh S (2023) Investigating antimicrobial peptide RI12 (K3W) as an effective bio-preservative against Listeria monocytogenes: a major foodborne pathogen. Arch Microbiol 205(12):367. https://doi.org/10.1007/s00203-023-03707-5
Jhong JH, Chi YH, Li WC (2019) AMP: an integrated resource for exploring antimicrobial peptides with functional activities and physicochemical properties on transcriptome and proteome data. Nucleic Acids Res 8(47):285–297. https://doi.org/10.1093/nar/gky1030
Jia L, Yarlagadda R, Reed CC (2015) Structure based thermostability prediction models for protein single point mutations with machine learning tools. PLoS ONE 10:e0138022. https://doi.org/10.1371/journal.pone.0138022
Jordan O, Gan BH, Alwan S, Perron K, Sublet E, Ducret V, Ye H, Borchard G, Reymond JL, Patrulea V (2024) Highly potent cationic chitosan derivatives coupled to antimicrobial peptide dendrimers to combat Pseudomonas aeruginosa. Adv Healthc Mater. https://doi.org/10.1002/adhm.202304118
Jordá-Vilaplana A, Fombuena V, García-García D, Samper MD, Sánchez-Nácher L (2014) Surface modification of polylactic acid (PLA) by air atmospheric plasma treatment. Eur Polym J 58:23–33. https://doi.org/10.1016/j.eurpolymj.2014.06.002
Joseph S, Karnik S, Nilawe P, Jayaraman VK, Idicula-Thomas S (2012) ClassAMP: a prediction tool for classification of antimicrobial peptides. IEEE/ACM Trans Comput Biol Bioinform 9(5):1535–1538. https://doi.org/10.1109/TCBB.2012.89
Kamech N, Vukicevic D, Ladram A, Piesse C, Vasseur J, Bojovic V, Simunic J, Juretic D (2012) Improving the selectivity of antimicrobial peptides from anuran skin. J Chem Inf Model 52(12):3341–3351. https://doi.org/10.1021/ci300328y
Kavousi K, Bagheri M, Behrouzi S, Vafadar S, Atanaki FF, Lotfabadi BT, Ariaeenejad S, Shockravi A, Moosavi-Movahedi AA (2020) IAMPE: NMR-assisted computational prediction of antimicrobial peptides. J Chem Inf Model 60(10):4691–4701. https://doi.org/10.1021/acs.jcim.0c00841
Khabbaz H, Karimi-Jafari MH, Saboury AA, BabaAli B (2021) Prediction of antimicrobial peptides toxicity based on their physico-chemical properties using machine learning techniques. BMC Bioinform 22:1–1. https://doi.org/10.1186/s12859-021-04468-y
Kumari A, Singh M, Sharma R, Kumar T, Jindal N, Maan S, Joshi VG (2023) Apoptin NLS2 homodimerization strategy for improved antibacterial activity and bio-stability. Amino Acids 55(10):1405–1416. https://doi.org/10.1007/s00726-023-03321-1
Lata S, Mishra NK, Raghava GPS (2009) AntiBP2: improved version of antibacterial peptide prediction. BMC Bioinform 11:1–7. https://doi.org/10.1186/1471-2105-11-S1-S19
Lawrence TJ, Carper DL, Spangler MK, Carrell AA, Rush TA, Minter SJ, Weston DJ, Labbé JL (2021) amPEPpy 1.0: a portable and accurate antimicrobial peptide prediction tool. Bioinformatics 37(14):2058–2060. https://doi.org/10.1093/bioinformatics/btaa917
Lee HT, Lee CC, Yang JR, Lai JZ, Chang KY (2015) A large-scale structural classification of antimicrobial peptides. Biomed Res Int 1:475062. https://doi.org/10.1155/2015/475062
Lee J, Ryu M, Bae D (2022) Development of DNA aptamers specific for small therapeutic peptides using a modified SELEX method. J Microbiol 60(7):659–667. https://doi.org/10.1007/s12275-022-2235-4
Lima KO, de Quadros CDC, da Rocha M, de Lacerda JTJG, Juliano MA, Dias M, Mendes MA, Prentice C (2019) Bioactivity and bioaccessibility of protein hydrolyzates from industrial byproducts of Stripped weakfish (Cynoscion guatucupa). LWT 111:408–413. https://doi.org/10.1016/j.lwt.2019.05.043
Liu B, Zhang W, Gou S, Huang H, Yao J, Yang Z, Liu H, Zhong C, Liu B, Ni J, Wang R (2017) Intramolecular cyclization of the antimicrobial peptide Polybia-MPI with triazole stapling: influence on stability and bioactivity. J Pept Sci 23(11):824–832. https://doi.org/10.1002/psc.3031
Lombardi L, Maisetta G, Batoni G, Tavanti A (2015) Insights into the antimicrobial properties of hepcidins: advantages and drawbacks as potential therapeutic agents. Molecules 20(4):6319–41. https://doi.org/10.3390/molecules20046319
Lu D, Chen Y, Xie Q, Qiu Z, Zhang H, Sun P, Pan J, Wang Y (2022) Preparation of bioactive peptides from marine industrial waste for moon cake preservation by coating. J Food Process Preserv 46(12):e17221. https://doi.org/10.1111/jfpp.17221
Lund MN, Ray CA (2017) Control of Maillard reactions in foods: strategies and chemical mechanisms. J Agric Food Chem 65(23):4537–4552. https://doi.org/10.1021/acs.jafc.7b00882
Luo X, Chen H, Song Y, Qin Z, Xu L, He N, Tan Y, Dessie W (2023) Advancements, challenges and future perspectives on peptide-based drugs: focus on antimicrobial peptides. Eur J Pharm Sci 181:106363. https://doi.org/10.1016/j.ejps.2022.106363
Luz C, Calpe J, Saladino F, Luciano FB, Fernandez-Franzón M, Mañes J, Meca G (2018) Antimicrobial packaging based on ϵ-polylysine bioactive film for the control of mycotoxigenic fungi in vitro and in bread. J Food Process Preserv 42:e13370. https://doi.org/10.1111/jfpp.13370
Malheiros PdaS, Sant’Anna V, de Souza BM, Brandelli A, de Melo Franco BDG (2012) Effect of liposome-encapsulated nisin and bacteriocin-like substance P34 on Listeria monocytogenes growth in Minas frescal cheese. Int J Food Microbiol 156(3):272–277. https://doi.org/10.1016/j.ijfoodmicro.2012.04.004
Meena M, Prajapati P, Ravichandran C, Sehrawat R (2021) Natamycin: a natural preservative for food applications-a review. Food Sci Biotechnol 30(12):1481–1496. https://doi.org/10.1007/s10068-021-00981-1
Meher PK, Sahu TK, Saini V, Rao AR (2017) Predicting antimicrobial peptides with improved accuracy by incorporating the compositional, physico-chemical and structural features into Chou’s general PseAAC. Sci Rep 7(1):1–12. https://doi.org/10.1038/srep42362
Meira SMM, Zehetmeyer G, Werner JO, Brandelli A (2017) A novel active packaging material based on starch-halloysite nanocomposites incorporating antimicrobial peptides. Food Hydrocoll 63:561–570. https://doi.org/10.1016/j.foodhyd.2016.10.013
Melo MC, Maasch JR, de la Fuente-Nunez C (2021) Accelerating antibiotic discovery through artificial intelligence. Commun Biol 4(1):1050. https://doi.org/10.1038/s42003-021-02586-0
Mironov PA, Paramonov AS, Reznikova OV, Safronova VN, Panteleev PV, Bolosov IA, Ovchinnikova TV, Shenkarev ZO (2024) Dimerization of the β-hairpin membrane-active cationic antimicrobial peptide capitellacin from marine polychaeta: an NMR structural and thermodynamic study. Biomolecules 14(3):332. https://doi.org/10.3390/biom14030332
Mitchell JB (2014) Machine learning methods in chemoinformatics. Wiley Interdiscipl Rev 4:468–481. https://doi.org/10.1002/wcms.1183
Mohanty DP, Mohapatra S, Misra S, Sahu DP (2016) Milk derived bioactive peptides and their impact on human health–a review. Saudi J Biol Sci 23(5):577–583. https://doi.org/10.1016/j.sjbs.2015.06.005
Molinos AC, Abriouel H, Lopez RL, Valdivia E, Omar NB, Galvez A (2008) Combined physico-chemical treatments based on enterocin AS-48 for inactivation of Gram-negative bacteria in soybean sprouts. Food Chem Toxicol 46:2912–2921. https://doi.org/10.1016/j.fct.2008.05.035
Mozafari MR, Khosravi-Darani K, Borazan GG, Cui J, Pardakhty A, Yurdugul S (2008) Encapsulation of food ingredients using nanoliposome technology. Int J Food Prop 11(4):833–844. https://doi.org/10.1080/10942910701648115
Nie T, Meng F, Zhou L, Lu F, Bie X, Lu Z, Lu Y (2021) In silico development of novel chimeric lysins with highly specific inhibition against Salmonella by computer-aided design. J Agric Food Chem 69(12):3751–3760. https://doi.org/10.1021/acs.jafc.0c07450
Ning HQ, Wang ZS, Li YQ, Tian WL, Sun GJ, Mo HZ (2019) Effects of glycinin basic polypeptide on the textural and physicochemical properties of Scomberomorus niphonius surimi. LWT 114:108328. https://doi.org/10.1016/j.lwt.2019.10832
Noonan J, Williams WP, Shan X (2017) Investigation of antimicrobial peptide genes associated with fungus and insect resistance in maize. Int J Mol Sci 18(9):1938. https://doi.org/10.3390/ijms18091938
Okella H, Okello E, Mtewa AG, Ikiriza H, Kaggwa B, Aber J, Ndekezi C, Nkamwesiga J, Ajayi CO, Mugeni IM, Ssentamu G, Ochwo S, Odongo S, Tolo CU, Kato CD, Engeu PO (2022) ADMET profiling and molecular docking of potential antimicrobial peptides previously isolated from African catfish, Clarias gariepinus. Front Mol Biosci 9:1039286. https://doi.org/10.3389/fmolb.2022.1039286
Oshiro KG, Candido ES, Chan LY, Torres MD, Monges BE, Rodrigues SG, Porto WF, Ribeiro SM, Henriques ST, Lu TK, de la Fuente-Núñez C (2019) Computer-aided design of mastoparan-like peptides enables the generation of nontoxic variants with extended antibacterial properties. J Med Chem 62(17):8140–8151. https://doi.org/10.1021/acs.jmedchem.9b00915
Peng J, Zheng F, Wei L, Lin H, Jiang J, Hui G (2018) Jumbo squid (Dosidicus gigas) quality enhancement using complex bio-preservative during cold storage. J Food Meas Character 12(1):78–86. https://doi.org/10.1007/s11694-017-9618-y
Porto WF, Silva ON, Franco OL (2012) Prediction and rational design of antimicrobial peptides. In: Faraggi E (ed) Protein structure. InTech, London, pp 377–396
Porto WF, Fensterseifer IC, Ribeiro SM, Franco OL (2018) Joker: an algorithm to insert patterns into sequences for designing antimicrobial peptides. Biochim Biophys Acta Gen Subj 1862:2043–2052. https://doi.org/10.1016/j.bbagen.2018.06.011
Przybylski R, Firdaous L, Châtaigné G, Dhulster P, Nedjar N (2016) Production of an antimicrobial peptide derived from slaughterhouse by-product and its potential application on meat as preservative. Food Chem 211:306–313. https://doi.org/10.1016/j.foodchem.2016.05.074
Rai M, Pandit R, Gaikwad S, Kövics G (2016) Antimicrobial peptides as natural bio-preservative to enhance the shelf-life of food. J Food Sci Technol 53:3381–3394. https://doi.org/10.1007/s13197-016-2318-5
Rouhi A, Yousefi Y, Falah F, Azghandi M, Behbahani BA, Tabatabaei-Yazdi F, Vasiee A (2024) Exploring the potential of melittin peptide: expression, purification, anti-pathogenic properties, and promising applications as a bio-preservative for beef slices. LWT 199:116083. https://doi.org/10.1016/j.lwt.2024.116083
Rounds T, Straus SK (2020) Lipidation of antimicrobial peptides as a design strategy for future alternatives to antibiotics. Int J Mol Sci 21(24):9692. https://doi.org/10.3390/ijms21249692
Santos JC, Sousa RC, Otoni CG, Moraes AR, Souza VG, Medeiros EA, Espitia PJ, Pires AC, Coimbra JS, Soares NF (2018) Nisin and other antimicrobial peptides: production, mechanisms of action, and application in active food packaging. Innov Food Sci Emerg Technol 48:179–194. https://doi.org/10.1016/j.ifset.2018.06.008
Santos-Filho NA, Righetto GM, Pereira MR, Piccoli JP, Almeida LMT, Leal TC, Camargo ILBC, Cilli EM (2021) Effect of C-terminal and N-terminal dimerization and alanine scanning on antibacterial activity of the analogs of the peptide p-BthTX-I. Pept Sci. https://doi.org/10.1002/pep2.24243
Selvarajan V, Tram NDT, Xu J, Ngen STY, Koh JJ, Teo JWP, Yuen TY, Ee PLR (2023) Stapled β-hairpin antimicrobial peptides with improved stability and activity against drug-resistant Gram-negative bacteria. J Med Chem 66(13):8498–8509. https://doi.org/10.1021/acs.jmedchem.3c00140
Shabir U, Ali S, Magray AR, Ganai BA, Firdous P, Hassan T, Nazir R (2018) Fish antimicrobial peptides (AMPs) as essential and promising molecular therapeutic agents: a review. Microb Pathog 114:50–56. https://doi.org/10.1016/j.micpath.2017.11.039
Sharma A, Singla D, Rashid M, Raghava GPS (2014) Designing of peptides with desired half-life in intestine-like environment. BMC Bioinform 15:1–8. https://doi.org/10.1186/1471-2105-15-282
Shwaiki LN, Arendt EK, Lynch KM (2020) Study on the characterization and application of synthetic peptide Snakin-1 derived from potato tubers—action against food spoilage yeast. Food Control 118:107362. https://doi.org/10.1016/j.foodcont.2020.107362
Singh RP, Heldman DR (2001) Introduction to food engineering. Gulf Professional Publishing, San Diego
Singh SS, Akhtar MN, Sharma D, Mandal SM, Korpole S (2021) Characterization of iturin V, a novel antimicrobial lipopeptide from a potential probiotic strain Lactobacillus sp. M31. Probiotics Antimicrob Proteins 13(6):1766–1779. https://doi.org/10.1007/s12602-021-09796-2
Singh A, Duche RT, Wandhare AG, Sian JK, Singh BP, Sihag MK, Singh KS, Sangwan V, Talan S, Panwar H (2023) Milk-derived antimicrobial peptides: overview, applications, and future perspectives. Probiotics Antimicrob Proteins 15:44–62. https://doi.org/10.1007/s12602-022-10004-y
Spaller BL, Trieu JM, Almeida PF (2013) Hemolytic activity of membrane-active peptides correlates with the thermodynamics of binding to 1-palmitoyl-2-oleoyl-sn-glycero-3-phosphocholine bilayers. J Membr Biol 246:257–262. https://doi.org/10.1007/s00232-013-9525-z
Sultana A, Luo H, Ramakrishna S (2021) Harvesting of antimicrobial peptides from insect (Hermetia illucens) and its applications in food packaging. Appl Sci 11(15):6991. https://doi.org/10.3390/app11156991
Tajer L, Paillart JC, Dib H, Sabatier JM, Fajloun Z, Abi Khattar Z (2024) Molecular mechanisms of bacterial resistance to antimicrobial peptides in the modern era: an updated review. Microorganisms 12(7):1259. https://doi.org/10.3390/microorganisms12071259
Tang R, Tan H, Dai Y, Li L, Huang Y, Yao H, Cai Y, Yu G (2023) Application of antimicrobial peptides in plant protection: making use of the overlooked merits. Front Plant Sci 14:1139539. https://doi.org/10.3389/fpls.2023.1139539
Tian L, Zhang D, Su P, Wei Y, Wang Z, Wang PX, Dai CJ, Gong GL (2019) Design, recombinant expression, and antibacterial activity of a novel hybrid magainin–thanatin antimicrobial peptide. Prep Biochem Biotechnol 49(5):427–434. https://doi.org/10.1080/10826068.2018.1476875
Timmons PB, Hewage CM (2020) HAPPENN is a novel tool for hemolytic activity prediction for therapeutic peptides which employs neural networks. Sci Rep 10:10869. https://doi.org/10.1038/s41598-020-67701-3
Torres MDT, de la Fuente-Núñez C (2019) Toward computer-made artificial antibiotics. Curr Opin Microbiol 51:30–38. https://doi.org/10.1016/j.mib.2019.03.004
Udenigwe CC, Fogliano V (2017) Food matrix interaction and bioavailability of bioactive peptides: two faces of the same coin? J Funct Foods 35:9–12. https://doi.org/10.1016/j.jff.2017.05.029
Veltri D, Kamath U, Shehu A (2018) Deep learning improves antimicrobial peptide recognition. Bioinformatics 34(16):2740–2747. https://doi.org/10.1093/bioinformatics/bty179
Vishnepolsky B, Grigolava M, Managadze G, Gabrielian A, Rosenthal A, Hurt DE, Tartakovsky M, Pirtskhalava M (2022) Comparative analysis of machine learning algorithms on the microbial strain-specific AMP prediction. Brief Bioinform. https://doi.org/10.1093/bib/bbac233
Waghu FH, Idicula-Thomas S (2020) Collection of antimicrobial peptides database and its derivatives: applications and beyond. Protein Sci 29(1):36–42. https://doi.org/10.1002/pro.3714
Wang W, Feng G, Li X, Ruan C, Ming J, Zeng K (2021) Inhibition of three citrus pathogenic fungi by peptide PAF56 involves cell membrane damage. Foods 10(9):2031. https://doi.org/10.3390/foods10092031
Wang R, Wang T, Zhuo L, Wei J, Fu X, Zou Q, Yao X (2024) Diff-AMP: tailored designed antimicrobial peptide framework with all-in-one generation, identification, prediction and optimization. Brief Bioinform 25(2):pbbae078. https://doi.org/10.1093/bib/bbae078
Wei D, Zhang X (2022) Biosynthesis, bioactivity, biotoxicity and applications of antimicrobial peptides for human health. Biosaf Health 25:118–134. https://doi.org/10.1016/j.bsheal.2022.02.003
Win TS, Malik AA, Prachayasittikul V, Wikberg JEE, Nantasenamat C, Shoombuatong W (2017) HemoPred: a web server for predicting the hemolytic activity of peptides. Fut Med Chem 9(3):275–291. https://doi.org/10.4155/fmc-2016-0188
Xiao J, Niu L (2015) Antilisterial peptides released by enzymatic hydrolysis from grass carp proteins and activity on controlling L. monocytogenes inoculated in surimi noodle. J Food Sci 80(11):M2564-9. https://doi.org/10.1111/1750-3841.13104
Xiao X, Wang P, Lin W-Z, Jia J-H, Chou K-C (2013) iAMP-2L: a two-level multi-label classifier for identifying antimicrobial peptides and their functional types. Anal Biochem 436:168–177. https://doi.org/10.1016/j.ab.2013.01.019
Yang S, Li J, Aweya JJ, Yuan Z, Weng W, Zhang Y, Liu GM (2020) Antimicrobial mechanism of Larimichthys crocea whey acidic protein-derived peptide (LCWAP) against Staphylococcus aureus and its application in milk. Int J Food Microbiol 335:108891. https://doi.org/10.1016/j.ijfoodmicro.2020.108891
Yang X, Wang Y, Jiang H, Song R, Liu Y, Guo H, Meng D (2023) Antimicrobial peptide CB-M exhibits direct antifungal activity against Botrytis cinerea and induces disease resistance to gray mold in cherry tomato fruit. Postharvest Biol Technol 196:112184. https://doi.org/10.1016/j.postharvbio.2022.112184
Yonezawa A, Kuwahara J, Fujii N, Sugiura Y (1992) Binding of Tachyplesin I to DNA revealed by footprinting analysis: significant contribution of secondary structure to DNA binding and implication for biological action. Biochemistry 31:2998–3004. https://doi.org/10.1021/bi00126a022
Zasloff M (2002) Antimicrobial peptides of multicellular origin. Nature 415:389–395. https://doi.org/10.1038/415389a
Zhang S, Luo L, Sun X, Ma A (2021) Bioactive peptides: a promising alternative to chemical preservatives for food preservation. J Agric Food Chem 69(42):12369–12384. https://doi.org/10.1021/acs.jafc.1c04020
Zhao Y, Zhang M, Qiu S, Wang J, Peng J, Zhao P, Zhu R, Wang H, Li Y, Wang K, Yan W (2016) Antimicrobial activity and stability of the D-amino acid substituted derivatives of antimicrobial peptide polybia-MPI. AMB Express 6:1–11. https://doi.org/10.1186/s13568-016-0295-8
Zhong C, Liu T, Gou S, He Y, Zhu N, Zhu Y, Wang L, Liu H, Zhang Y, Yao J, Ni J (2019) Design and synthesis of new N-terminal fatty acid modified-antimicrobial peptide analogues with potent in vitro biological activity. Eur J Med Chem 182:111636. https://doi.org/10.1016/j.ejmech.2019.111636
Funding
The authors declare that no funds, grants, or other support were received during the preparation of this manuscript.
Author information
Authors and Affiliations
Contributions
All authors contributed to writing of the manuscript. SS and VGJ reviewed and edited the manuscript. All authors finally read and approved the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Singh, S., Jha, B., Tiwari, P. et al. Recent approaches in the application of antimicrobial peptides in food preservation. World J Microbiol Biotechnol 40, 315 (2024). https://doi.org/10.1007/s11274-024-04126-4
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s11274-024-04126-4