Abstract
Chlorophenols are persistent environmental pollutants used in synthesizing dyes, drugs, pesticides, and other industrial products. The chlorophenols released from these processes seriously threaten the environment and human health. The present study describes 4-chlorophenol (4-CP) degradation activity and metagenome structure of a bacterial consortium enriched in a 4-CP-containing medium. The consortium utilized 4-CP as a single carbon source at a wide pH range, temperature, and in the presence of heavy metals. The immobilized consortium retained its degradation capacity for an extended period. The 4-aminoantipyrine colorimetric analysis revealed complete mineralization of 4-CP up to 200 mg/L concentration and followed the zero-order kinetics. The addition of glycerol and yeast extract enhanced the degradation efficiency. The consortium showed both ortho- and meta-cleavage activity of catechol dioxygenase. Whole genome sequence (WGS) analysis revealed the microbial compositions and functional genes related to xenobiotic degradation pathways. The identified genes were mapped on the KEGG database to construct the 4-CP degradation pathway. The results exhibited the high potential of the consortium for bioremediation of 4-CP contaminated sites. To our knowledge, this is the first report on WGS analysis of a 4-CP degrading bacterial consortium.
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
Introduction
Environmental pollution due to industrial development is a serious hazard to human health. Chlorophenols are chlorinated aromatic compounds that are widely employed in industry, agriculture, and medicine. They are found in wastewater, surface water, soil, and sediments, in bodies of aquatic organisms, animal milk, adipose tissue, and urine (Olaniran and Igbinosa 2011; Zada et al. 2021). In strongly contaminated groundwater the chlorophenol concentrations have been reported to range from 100 to 200 mg/L (Ettala et al. 1992). As per recommendations from the World Health Organisation (WHO) and United States Environmental Protection Agency (USEPA), the concentration of chlorophenol in drinking water and industrial effluent should not exceed 0.5 µg/L and 100 µg/L, respectively (Zada et al. 2021). More than 100,000 tonnes of chlorophenols are used worldwide each year (Nowak and Mrozik 2018). The extensive use of these compounds and their release into the environment, primarily through inappropriate disposal, pose a concern to humans, aquatic life, and wildlife due to their acute toxicity, carcinogenicity, and refractory nature (Nowak and Mrozik 2018; Zada et al. 2021). In particular, exposure to 4-chlorophenol (4-CP) causes a range of adverse health effects including diarrhoea, tremors, internal organs (kidney, lungs) dysfunction, mutations, abnormalities, muscular weakness, and coma in living organisms (Kwean et al. 2018; Swain et al. 2021). The 4-CP can be generated from paper industry during chlorine bleaching of pulp and wastewater treatment, as well as from the breakdown of more highly chlorinated phenols and phenoxy herbicides (Bjerketorp et al. 2018). The USEPA has classified 4-CP as a primary pollutant (Dan et al. 2021). Consequently, 4-CP pollutants ought to be managed carefully before being released into a public space
Various methods for the removal of 4-CP comprise adsorption, photocatalysis, electrochemical oxidation, etc. (Adhikari et al. 2017; Allaboun and Abu 2016; Duan et al. 2013; Saravanakumar et al. 2023). Although highly efficient, these techniques have limited use due to high cost and in some cases production of harmful intermediates. Bioremediation, on the other hand, is a cost-effective and environmentally friendly method for the reclamation of 4-CP contaminated environments (Patel and Kumar 2016; Arora and Bae 2014). It involves the use of microorganisms to degrade or transform pollutants into less toxic forms. Further, the bioremediation which can be conducted in situ reduces the need for costly excavation and transportation of contaminated material. Several bacterial species such as Bacillus sp., Pseudomonas sp., Rhodoccocus sp., Arthrobacter sp., and Alcaligens sp., have been reported to degrade 4-CP (Farrell and Quilty 2002; Konovalova et al. 2009; Nordin et al. 2005; Sandhibigraha et al. 2020; Swain et al. 2021). Swain et al. (2021) reported that Bacillus flexus was able to remove 78.12% of 4-CP at a concentration of 50 mg/L. According to Patel and Kumar (2016), a bacterial consortium consisting of Pseudomonas aeruginosa GF, Kocuria rhizophila 11Y, Bacillus endophyticus CP1R, and Bacillus cereus 3YS degraded 3-CP and 4-CP. However, alone none of these strains demonstrated chlorophenol degradation activity thus underscoring the importance of bacterial consortia for bioremediation of polluted environments. Environmental factors play a vital role in the biodegradation of pollutants in contaminated habitats which also require optimization to enhance the biodegradation efficiency (Fletcher et al. 2011; Swain et al. 2021; Wang and Sun 2021; Zhao et al. 2018).
The information on the degradation of chlorophenols using naturally developed bacterial consortia is limited. Therefore, the aim of present study was to characterize the 4-CP degradation by a laboratory developed bacterial consortium. The degradation activity was assessed at different 4-CP concentrations and in response to various environmental factors e.g., temperature, pH, heavy metals, and extra carbon sources. Further, whole genome sequencing (WGS) approach was used for the taxonomic and functional diversity analysis of the consortium.
Materials and methods
Chemicals and selection of bacterial consortium
All the chemicals and reagents were analytical grade. Catechol, 4-aminoantipyrine reagents, and 4-CP were obtained from Sigma-Aldrich (India). HPLC grade ethyl acetate, glycerol, and sucrose were purchased from SRL, India. Bushnell Haas medium (BHM), heavy metal salts, glucose, fructose, and yeast extract were purchased from Himedia, India.
BHM (Bushnell and Haas 1941) supplemented with 40 mg/L 4-CP was inoculated with 1.0 g soil sample collected from a pesticide contaminated rice field in Umshing area of Shillong, India (Latitude-25.61 N and Longitude-91.90 E). The inoculated flasks were incubated for two weeks in an incubator shaker at 110 rpm at 25±1 °C. The developed bacterial consortia were enriched by ten successive transfers every fifth day into fresh medium and by gradually increasing the 4-CP concentration from 40 to 100 mg/L. A total of five bacterial consortia with the capacity to utilize 4-CP as a carbon source were developed by enrichment method. A consortium showing the best degradation efficiency was selected and maintained in 100 mg/L 4-CP supplemented BHM for further study.
Microbial culture condition
All the experiments were carried out in a 250 mL Erlenmeyer flask containing 100 mL BHM supplemented with 4-CP. Acclimated microbial consortium were inoculated at initial OD600 = 0.04. The inoculated flasks were incubated at 25±1 °C in an incubator shaker (110 rpm). Uninoculated control flasks were incubated in parallel.
Determination of growth and 4-CP degradation
The bacterial consortium was inoculated in 4-CP amended BHM and the growth of the consortium was determined as an increase in cell density at λ600 using UV-VIS spectrophotometer (Chettri and Singh 2019). Measurement of residual 4-CP in the culture supernatant was determined by 4-aminoantipyrine colorimetric method (Farell and Quilty 1999). For gas chromatography mass spectrometry (GC-MS) analysis of residual 4-CP, a 5 mL culture supernatant was extracted twice in an equal volume of ethyl acetate. The ethyl acetate extract was evaporated to 1.0 mL final volume and passed through prewashed Isolute Solid Phase Extraction (SPE) column. The filtered extract was analyzed using a Thermo Scientific, Trace GC Ultra gas chromatograph coupled with a mass spectrometer (ITQ 1100 Ion Trap MS, Thermo Scientific) by injecting 1 µL of ethyl acetate extract (Chettri et al. 2016). The degradation percentage of 4-CP was calculated using Eq. (1):
The degradation of 4-CP was fitted into zero-order kinetics as described in Eq. (2)
where C0 is the initial concentration of 4-CP, Ct is the concentration of 4-CP at time t, t is the degradation period in hour and k is the biodegradation rate constant (Gaya et al. 2009). The half-life (t1/2) of biodegradation was calculated using the Eq. (3)
Effect of different process variables on biodegradation
The effect of various factors on the 4-CP degradation was evaluated in a bacterial consortium inoculated in 100 mL BHM containing 100 mg/L of 4-CP and incubated for 36 h under shaking condition (110 rpm); at six different temperatures (15, 20, 25, 30, 35 and 40 °C), five different pH (5, 6, 7, 8, and 9), in presence of 100 µM (Al3+, Cr3+, Fe2+, Mn2+, Pb2+, Sr2+, Zn2+) and 10 µM (Cu2+, Co2+, Ni2+) metal ions, and presence of 0.5 g/L (sucrose, glucose, fructose, yeast extract), and 0.05% glycerol as carbon sources. The concentration of glycerol is mentioned in percentage because it was available in liquid form.
Enzyme assay
The bacterial consortium was inoculated in 100 mg/L 4-CP supplemented BHM. After 24 h of incubation, the cells were collected by centrifuging at 13,000 rpm for 10 min at 4 °C. The pellet was washed twice and re-suspended in 33 mM Tris-HCl buffer (pH 7.6). The suspension was sonicated in an ice bath and centrifuged at 20,000 rpm at 4 °C for 20 min. The cell free extract was used for estimating catechol 1, 2-dioxygenase and catechol 2, 3-dioxygenase activities according to Farell and Quilty (1999). Total protein was assessed using bovine serum albumin (BSA) as the standard (Lowry et al. 1951).
Statistical analysis
Each experiment was repeated three times with three replicates each. The values from replicate experiments were used for calculating the mean ± standard deviation. One-way ANOVA was done in Microsoft Excel.
Whole genome sequencing (WGS)
Genomic DNA from 4-CP degrading bacterial consortium was extracted using c-TAB and Phenol: Chloroform extraction technique, followed by RNase A treatment. The extracted DNA's quality and quantity were assessed using NanoDrop, followed by 0.8% agarose gel electrophoresis. The paired-end sequencing library was created from QC-passed DNA samples using the illumina TruSeq Nano DNA Library Prep Kit. The library was sequenced on the Illumina NextSeq500 platform using 2 × 150 bp chemistry. Covaris M220 fragmented about 200 ng of DNA to produce a mean fragment distribution of 350 bp. Double stranded DNA fragments with 3’ or 5’ overhangs were generated by Covaris shearing and then subjected to end-repair. The End Repair Mix converts the 3’ or 5’ overhangs into blunt ends followed by adapter ligation to the fragments. AMPure XP beads were used to size select the ligated products. The ligated products were PCR amplified using the index primer according to the manufacturer's protocol. The PCR amplified library was analyzed on 4200 Tape Station system (Agilent Technologies) using high sensitivity D1000 Screen Tape.
To obtain high quality clean reads, the adapter sequences, ambiguous reads (reads with unknown nucleotides “N” >5%), and low-quality sequences (reads with>10% quality threshold (QV) < 20 phreds score) were removed using Trimmomatic v0.38. A minimum length of 100 nucleotides after trimming was used. The high quality paired-end reads (>20 phreds score) were used for de-novo assembly. These reads were assembled into scaffolds using CLC Workbench (ver.9), and the average size of scaffolds was ~677 bp. Prodigal 2.6.3 was used to predict genes from assembled scaffolds, and the total number of genes was estimated at 89,837, with a minimum gene length of 60 bp and a maximum gene length of 8,340 bp. The WGS sequenced raw data was deposited to the National Centre for Biotechnology Information (NCBI) under BioProject: PRJNA728811 and Sequence Read Archive (SRA): SRS18487309. The data summary can also be accessed at NCBI via https://www.ncbi.nlm.nih.gov/bioproject/PRJNA728811
Results
Effect of 4-CP concentration on growth and biodegradation activity
The bacterial consortium was inoculated in 4-CP (50–300 mg/L) amended BHM showed an increase in cell density with increasing 4-CP concentration up to 200 mg/L (Fig 1a). The maximum growth at 50 mg/L, 100 mg/L, 150 mg/L, 200 mg/L, and 300 mg/L was observed respectively, at 18 h, 24 h, 36 h, 48 h, and 60 h after incubation. However, growth was least at 300 mg/L concentration.
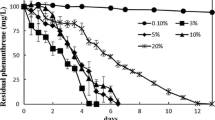
The amount of residual 4-CP was determined in the supernatant from the aforementioned cultures. The time needed for complete degradation increased with increasing 4-CP concentration (Fig 1b). The consortium required 18 h, 36 h, 60 h, and 120 h for complete degradation at 50 mg/L, 100 mg/L, 150 mg/L and 200 mg/L, respectively. At 300 mg/L concentration, the consortium could degrade only 28.8% of 4-CP after 144 h of incubation, resulting in incomplete elimination of the compound. A straight line with slope =−k and intercept = [4-CP]0 was obtained from a plot of [4-CP] against time, which conforms to the zero-order kinetics for 4-CP degradation. The values from Fig 1b were used to calculate the biodegradation rate constant (k) and half-life (t1/2) of degradation. The k and t1/2 values were found to be 2.89±0.0221 mg/L/h and 8.65 h at 50 mg/L; 2.83±0.006 mg/L/h and 17.64 h at 100 mg/L; 2.56±0.007 mg/L/h and 29.30 h at 150 mg/L; 1.70±0.0032 mg/L/h and 58.90 h at 200 mg/L; and 0.5632±0.019 mg/L/h and 266.34 h at 300 mg/L concentrations. The decline in degradation rate from 200 mg/L onward concentration reflects the upper limit of the bacterial consortium to degrade 4-CP. The degradation of 4-CP (100 mg/L) was assessed also by GC-MS based analysis. The absence of metabolite in GC-MS profile revealed complete degradation of 4-CP in 36 h. (Fig 1c and 1d). The consortium retained the 4-CP degradation activity 12 months after preservation in different matrices, e.g., Calcium alginate beads, sawdust, rice husk, charcoal, coconut husk, and firewood ash. The initial degradation activity of the preserved cells was highest for alginate beads. The preserved cells regained degradation ability comparable to freshly grown inoculum after two consecutive transfers.
Effect of temperature, pH, heavy metals, and carbon sources on biodegradation activity
A complete degradation of 4-CP was observed after 36 h incubation at temperatures ranging from 15 and 35 °C with a degradation rate of 2.7±0.02 mg/L/h. At 40 °C, the degradation rate decreased drastically to 0.87 mg/L/h, exhibiting just 19.9% and 31% degradation after 24 h and 36 h respectively (Fig 2a). As shown in Fig 2b, the ideal pH range for 4-CP degradation was between 7 and 9 as complete degradation was achieved within 36 h of incubation. At pH 6, about 82.5% of the total 4-CP was degraded in 36 h. However, when the initial pH was at 5, a drastic inhibition in 4-CP degradation was observed. The degradation rates were 0.03 mg/L/h, 2.11 mg/L/h, 2.66 mg/L/h, 2.75 mg/L/h, and 2.84 mg/L/h at pH 5, 6, 7, 8 and 9 respectively.
Similarly, a complete degradation occurred in Al3+, Cr3+, Fe2+, Mn2+, Pb2+, and Sr2+ supplemented conditions with a degradation rate of ~2.7±0.03 mg/L/h. Biodegradation of 4-CP was reduced to 64.96%, 57.47%, and 37.96% respectively in Ni2+, Zn2+, and Co2+ treated conditions (Fig 2c). The degradation rate was 1.74 mg/L/h (Ni2+), 1.53 mg/L/h (Zn2+), and 1.04 mg/L/h (Co2+). In contrast, Cu2+ treatment caused almost complete inhibition of 4-CP degradation. The addition of glycerol and yeast extract to the culture medium significantly increased the removal rate of 4-CP (Fig 2d). The degradation rates in sucrose, glycerol, glucose, fructose, and yeast extract amended medium were 2.71 mg/L/h, 2.89 mg/L/h, 2.38 mg/L/h, 2.63 mg/L/h, and 2.86 mg/L/h respectively. After 36 h incubation, complete removal of 4-CP was achieved in each carbon source amended media except for glucose.
Catechol dioxygenase activity
The bacterial consortium grown with 100 mg/L of 4-CP showed both catechol 1,2-dioxygenase and catechol 2,3-dioxygenase activities at 0.029±0.003 µmoles cis,cis-muconic acid/min/mg protein and 0.027±0.002 µmoles 2-hydroxymuconic semialdehyde/min/mg protein respectively.
Microbial community structure
The WGS statistics of the consortium generated a total of 89,837 high quality filtered reads and summary of metagenomics sequencing statistics were listed in Table 1. Taxonomic analysis of the predicted genes was carried out using Kaiju (kaiju.binf.ku.dk). The high throughput metagenomics sequencing data revealed the level of microbial diversity of the consortium. The microbial consortium was dominated by Bacteria (87.19 %), followed by unclassified sequences (12.73%), Eukaryotes (0.04%), Archaea (0.03%), and Viruses (0.01%).
The microbial community abundance of the consortium at phylum and genus level is presented in Fig 3. The bacterial domain was dominated mostly by the phyla Proteobacteria (73.27%) and Bacteroidota (10%), constituting more than 83% of the total bacterial community. At the class level, Alphaproteobacteria (42.13%), Betaproteobacteria (25.92%), and Gammaproteobacteria (4.41%) accounted for 72.46% of the total bacteria (Fig S1a). Burkholderiales (25.65%), Rhizobiales (20.22%), and Rhodobacterales (16.26%) were found to be the most common order (Fig S1b). The dominant bacterial families were Rhodobacteraceae (16.12%), Comamonadaceae (12.01%), Alcaligenaceae (11.43%), and Rhizobiaceae (10.66%) (Fig S1c). The bacterial genera at >1% abundance comprised Paracoccus (13.24%), Achromobacter (10.93%), Hydrogenophaga (6.74%), Rhizobium (5.27%), Leadbetterella (4.41%), Brevundimonas (4.16%), Acidovorax (3.76%), Mesorhizobium (2.79%), Shinella (2.58%), Aquamicrobium (1.34%), Sphingobacterium (1.32%) and Thermomonas (1.23%) (Fig 3b). Achromobacter denitrificans (8.72%), Leadbetterella bysophilia (4.41%) and Paracoccus denitrificans (4.22%) were the three most abundant species (Fig S1d).
Functional annotation
Functional annotation of genes was carried out using Cognizer (metagenomics.atc.tcs.com/cognizer), a comprehensive stand-alone framework that is capable of simultaneously providing clusters of orthologs genes (COG) and Kyoto encyclopedia of genes and genomes (KEGG) subsystem annotations to individual sequences constituting metagenomic datasets. Cognizer's innovative 'direct search' feature considerably reduces the total compute needs involved with functional analysis.
Classification of genes according to COG Categories
A total of 60,442 genes were assigned to COG categories. The functional annotation distribution of COG is presented in Fig 4. The COG categories at level 1 showed relative abundance of genes involved in Metabolism (46.32%), Poorly characterized (19.78), Cellular processes and signaling (18.55%), and Information storage and processing (15.36%). At level 2, General function prediction only [R] (12.41%) was the dominant function among the 24 categories, followed by Amino acid transport and metabolism [E] (12.26%), Inorganic ion transport and metabolism [P] (7.04%), Transcription [K] (6.90%), Energy transport and conversion [C] (6.80%) and Carbohydrate transport and metabolism [G] (6.10%) (Thresholds level 6%). The lowest number of genes were assigned to RNA processing and modification [A] (0.01%), Extracellular structures [W] (0.01%), and Chromatin structures and dynamics [B] (0.04%).
Classification of KEGG functional categories and 4-CP degradation pathway
A total of 87,981 genes were assigned to KEGG orthology (KO). The distribution of KEGG categories at level 1 revealed unclassified genes (40%), metabolism (34%), environmental information processing (16%), genetic information processing (5%), and cellular processes (4%). The five most abundant categories at level 2 of KEGG metabolic pathways were carbohydrate metabolism (15.5%), amino acid metabolism (14.53%), membrane transport (13.54 %), energy metabolism (7.68%), and xenobiotics biodegradation and metabolism (7.66%) (Fig 5). The mapping of obtained KO ids in xenobiotic biodegradation and metabolism pathway revealed 4780 genes corresponding to 308 different enzymes. The number of possible reactions of these enzymes in xenobiotic degradation is presented in Table 2. Majority of these genes were mapped in benzoate degradation (ko-00362) and chlorocyclohexane and chlorobenzene degradation (ko-00361), drug metabolism-other enzymes (ko-00983), xylene degradation (ko-00622), and metabolism of xenobiotics by cytochrome-P450 (ko-00980). Based on the gene mappings in KEGG database, the 4-CP degradation pathway which is a component of chlorocyclohexane and chlorobenzene degradation (Fig S2), was constructed (Fig 6). The list of enzymes of 4-CP degradation pathway is presented in Table 3. The pathway shows 4-chlorocatechol as the first intermediate in degradation process which is subsequently cleaved through both ortho- and meta-ring cleavage by catechol 1,2-dioxygenase and catechol 2,3-dioxygenase. This result was confirmed by the presence of catechol 1,2-dioxygenase and catechol 2,3-dioxygenase activities. The 4-CP degradation by microbial consortium followed the chlorocyclohexane and chlorobenzene degradation pathways, and then the benzoate degradation pathways (Fig S3). Finally, the metabolites are metabolized through citric acid cycle.
Discussion
Degradation of 4-CP has been studied using both bacterial isolates and defined bacterial consortium. Swain et al. (2021) reported 78% and 38% degradation by free cells and 90.34% and 49.44% by immobilized cells respectively at 50 mg/L and 200 mg/L of 4-CP by Bacillus flexus in 7 days. Patel et al. (2016) using a defined bacterial consortium reported 63% removal of 200 mg/L 4-CP in 168 h. Past studies have focused on using either pure isolates or defined consortium for chlorophenol biodegradation. In our study, the bacterial consortium enriched from pesticide-contaminated soil showed complete biodegradation of 4-CP up to 200 mg/L within 120 h of incubation. Moreover, the retention of 4-CP degradation activity after immobilization for 12 months reflects our consortium's high usefulness for its application under field conditions.
The enriched consortium’s ability to degrade 4-CP across a wide temperature range illustrates its resilience to fluctuating temperatures or seasonal variations. Monsalvo et al. (2009) have reported the complete removal of 4-CP (105 mg/L) in acclimated sequencing batch reactors (SBR) industrial sludge at 20 °C–35 °C. At temperatures over 35 °C, the consortium's capacity to breakdown 4-CP decreased significantly, which is most likely caused by inhibition of the cell's multienzyme complex system (Bandyopadhyay et al. 1998).
In soil ecosystems, pH values can be highly variable ranging from 2.5 to 11 (Bartha and Bossert 1984). The consortium's capacity to operate from pH 6-9 enables it to target and remediate 4-CP contaminated sites at wide pH profiles. Since chlorophenols exist in the unionized hydrophobic form at acidic pH, their toxicity to microorganisms is usually high as they may readily infiltrate the lipid membranes of the microorganisms (Penttinen 1995). This phenomenon explains the low activity of our consortium at pH-5. The result suggests that alkaline pH is more conducive in the removal of 4-CP. Similar observations have been reported for degradation of various chlorophenols by bacterial isolates and defined bacterial consortium (Farag et al. 2021; Gallego et al. 2009).
There are reports on the simultaneous presence of heavy metals in pesticide contaminated soil (Tariq et al. 2016). Heavy metals in contaminated soil affect microbial nutrient cycling activity. Therefore, it is desirable to apply heavy metal tolerant bacteria in bioremediation of contaminated soils. Heavy metals at low concentrations stimulate microbial growth and become toxic at high concentrations (Giller et al. 1998). Our consortium showed 4-CP degradation activity in presence of various metals except Cu2+. Silva et al. (2012) reported stimulation of catechol 1,2-dioxygenase and catechol 2,3-dioxygenase activities, the key enzymes responsible for 4-CP degradation, in presence of Mn2+ and Fe3+. The inhibition of microbial activities in Cu2+ treated conditions might be due to an increase in the production of reactive oxygen species, a reduction in the level of enzyme activity, and delay carbohydrate metabolism (Kuo and Genthner 1996; Ma et al. 2022).
Microorganisms display extraordinary diversity with respect to the utilization of carbon sources. Decreased degradation activity in glucose amended medium could be due to microorganism’s strong preference for simple carbon sources, resulting in a decrease in pH of the culture medium (Fakhruddin and Quilty 2005) and depletion in dissolved oxygen concentration (Wang and Sun 2020a). The high efficiency of carbon sources to stimulate microbial growth and biodegradation of toxic compounds at lower concentrations has been reported (Wang and Sun 2020b; Ziagova et al. 2009).
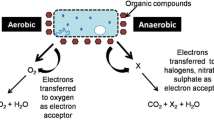
Chlorocatechols form a key intermediate in aerobic degradation of chlorinated aromatic compounds. The cleavage of the catechol ring occurs via two pathways (1) ortho-cleavage (intradiol) catalyzed by catechol 1,2-dioxygenase and (2) meta-cleavage (extradiol) catalyzed by catechol 2,3-dioxygenase (Farell and Quilty 1999). The bacterium Rhodococcus opacus 6a was found to have ortho-cleavage pathway (Konovalova et al. 2009). A microbial consortium comprising of Pseudomonas and Actinomycetes and a fungus Trichoderma harzamanium showed 4-CP degradation via a meta-cleavage pathway (Farrell and Quilty 1999). Presence of enzyme activity of both ortho- and meta-cleavage pathways for 4-CP degradation underscores the high utility of consortium for bioremediation of organic pollutants.
The bacterial consortium developed in presence of xenobiotic exhibits high xenobiotic metabolizing capacity. Metagenomic analysis of such consortium provides direct and in-depth knowledge of relative abundance and metabolic roles of bacterial groups in the metabolism of the xenobiotic. The WGS analysis of our consortium revealed a high abundance of Proteobacteria (73.27%) and Bacteroidota (10%). Several studies using 16S rRNA metagenomic approach reported a similar composition of 4-CP degrading bacterial community at the phylum level (Zhao et al. 2018). Gómez-Acata et al. (2018) reported the dominance of Proteobacteria followed by Actinomycetota and Bacillota in 4-CP treated SBR inoculated with wastewater sludge from an aerobic wastewater plant. Moreno-Andrade et al. (2020) observed increases in the abundance of Proteobacteria and Bacteroidota in 4-CP degrading acclimated granules of SBR with a tremendous increase in Bacteroidota members from <1% to 40.6%. The dominant genera Paracoccus, Achromobacter, Hydrogenophage, Rhizobium, Leadbetterella, Brevundimonas, Acidovorax, Mesorhizobium, Shinella, Aquamicrobium, Sphingobacterium and Thermomonas have been found coexisting in acclimated granules of SBR degrading 4-CP (Moreno-Andrade et al. 2020). The high abundance of the genus Paracoccus in 4-CP fed SBR (Gómez-Acata et al. 2018), and synergistic contribution of Paracoccus and Achromobacter in degradation of xenobiotics has been reported (Gao et al. 2021). An increase in relative abundance of genus Leadbetterella in presence of phenolic compounds was observed by Gómez-Acata et al. (2017). The bacterium Achromatobacter denitrificans has been reported in degradation of various xenobiotics (Benjamin et al. 2016; Mawad et al. 2016), thus highlighting the potential of our consortium in bioremediation.
WGS of industrial discharges have provided an in depth understanding of the microbial xenobiotic degradation activity (Pandit et al. 2021; Shah et al. 2013). However, the number of mapped reactions of xenobiotic degradation pathways can vary depending on the sources of microbial samples. In our study, the total number of mapped reactions in degradation of xenobiotics e.g., benzoate, chlorocyclohexane and chlorobenzene, styrene, metabolism of xenobiotics by cytochrome P450, and atrazine respectively were 50, 35, 12, 25, and 8. However, in a study reported by Shah et al. (2013), these values were 24, 5, 1, 16, and 2 respectively. It is important to note that our consortium was raised in 4-CP amended BHM whereas Shah et al. (2013) used a contaminated environmental soil sample. The presence of a large number of enzymes of xenobiotic degradation (Table 2) mapped in KEGG database reflects the high potential of our consortium in bioremediation of a wide range of xenobiotics including 4-CP.
Conclusion
The enriched bacterial consortium efficiently degraded 4-CP in the presence of heavy metals, carbon sources, at pH 7-9, and temperatures from 15 °C-35 °C. WGS analysis revealed the dominance of phyla Proteobacteria and Bacteroidota, the bacterial genera Paracoccus, Achromobacter, Hydrogenophage, Rhizobium, Leadbetterella, Brevundimonas, Acidovorax, Mesorhizobium, Shinella, Aquamicrobium, and the consortium’s capacity to devour xenobiotic compounds.
Data availability
All data generated or analyzed during this study are included in the article.
References
Adhikari S, Sarkar D, Madras G (2017) Hierarchical design of CuS architectures for visible light photocatalysis of 4-chlorophenol. ACS Omega 2:4009–4021
Allaboun H, Abu Al-Rub FA (2016) Removal of 4-chlorophenol from contaminated water using activated carbon from dried date pits: equilibrium, kinetics, and thermodynamics analyses. Materials 9:251
Arora PK, Bae H (2014) Bacterial degradation of chlorophenols and their derivatives. Microb Cell Factories 13:1–17
Bandyopadhyay K, Das D, Maiti BR (1998) Kinetics of phenol degradation using Pseudomonas putida MTCC 1194. Bioprocess Eng 18:373–377
Bartha R, Bossert I (1984) The fate of petroleum in the soil ecosystems. Petroleum Microbiology. Macmillan, New York, pp 435–473
Benjamin S, Kamimura N, Takahashi K, Masai E (2016) Achromobacter denitrificans SP1 efficiently utilizes 16 phthalate diesters and their downstream products through protocatechuate 3,4-cleavage pathway. Ecotoxicol Environ Saf 134:172–178
Bjerketorp J, Röling WF, Feng XM, Garcia AH, Heipieper HJ, Håkansson S (2018) Formulation and stabilization of an Arthrobacter strain with good storage stability and 4-chlorophenol-degradation activity for bioremediation. Appl Microbiol Biotechnol 102:2031–2040
Bushnell LD, Haas HF (1941) The utilization of certain hydrocarbons by microorganisms. J Bacteriol 41:653–673
Chettri B, Mukherjee A, Langpoklakpam JS, Chattopadhyay D, Singh AK (2016) Kinetics of nutrient enhanced crude oil degradation by Pseudomonas aeruginosa AKS1 and Bacillus sp. AKS2 isolated from Guwahati refinery. India Environ Pollut 216:548–558
Chettri B, Singh AK (2019) Kinetics of hydrocarbon degradation by a newly isolated heavy metal tolerant bacterium Novosphingobium panipatense P5. ABC Bioresour Technol 294:122–190
Dan X, Luo Z, Dai M, Zhang M, Yue X, Xie S (2021) Oxidative degradation of p-chlorophenol by ferrate (VI): kinetics, intermediates and pathways. J Environ Chem Eng 9:105810
Duan X, Tian L, Liu W, Chang L (2013) Study on electrochemical oxidation of 4-chlorophenol on a vitreous carbon electrode using cyclic voltammetry. Electrochim Acta 94:192–197
Ettala M, Koskela J, Kiesilä A (1992) Removal of chlorophenols in a municipal sewage treatment plant using activated sludge. Water Res 26:797–804
Fakhruddin ANM, Quilty B (2005) The influence of glucose and fructose on the degradation of 2-chlorophenol by Pseudomonas putida CP1. World J Microbiol Biotechnol 21:1541–1548
Farag AM, Fawzy A, El-Naggar MY, Ghanem KM (2021) Biodegradation and enhancement of 2,4-dichlorophenol by marine halophilic Bacillus subtilis AAK. Egypt J Aquat Res 47:117–123
Farrell A, Quilty B (1999) Degradation of mono-chlorophenols by a mixed microbial community via a meta-cleavage pathway. Biodegradation 10:353–362
Farrell A, Quilty B (2002) Substrate-dependent autoaggregation of Pseudomonas putida CP1 during the degradation of mono-chlorophenols and phenol. J Ind Microbiol Biotechnol 28:316–324
Fletcher KE, Costanza J, Pennell KD, Löffler FE (2011) Electron donor availability for microbial reductive processes following thermal treatment. Water Res 45:6625–6636
Gallego A, Gemini V, Rossi S, Fortunato MS, Planes E, Gómez CE, Korol SE (2009) Detoxification of 2,4,6-trichlorophenol by an indigenous bacterial community. Int Biodeterior Biodegrad 63:1073–1078
Gao Y, Liu M, Zhao X, Zhang X, Zhou F (2021) Paracoccus and Achromobacter bacteria contribute to rapid biodegradation of imidacloprid in soils. Ecotoxicol Environ Saf 225:112785
Gaya UI, Abdullah AH, Zainal Z, Hussein MZ (2009) Photocatalytic treatment of 4-chlorophenol in aqueous ZnO suspensions: Intermediates, influence of dosage and inorganic anions. J Hazard Mater 168:57–63
Ge T, Han J, Qi Y, Gu X, Ma L, Zhang C, Huang D (2017) The toxic effects of chlorophenols and associated mechanisms in fish. Aquat Toxicol 184:78–93
Giller KE, Witter E, Mcgrath SP (1998) Toxicity of heavy metals to microorganisms and microbial processes in agricultural soils: a review. Soil Biol Biochem 30:1389–1414
Gómez-Acata S, Esquivel-Ríos I, Pérez-Sandoval MV, Navarro-Noya Y, Rojas-Valdez A, Thalasso F, Dendooven L (2017) Bacterial community structure within an activated sludge reactor added with phenolic compounds. Appl Microbiol Biotechnol 101:3405–3414
Gómez-Acata S, Vital-Jácome M, Pérez-Sandoval MV, Navarro-Noya YE, Thalasso F, Luna-Guido M, Dendooven L (2018) Microbial community structure in aerobic and fluffy granules formed in a sequencing batch reactor supplied with 4-chlorophenol at different settling times. J Hazard Mater 342:606–616
Konovalova EI, Solyanikova IP, Golovleva LA (2009) Degradation of 4-chlorophenol by the strain Rhodococcus opacus 6a. Microbiology 78:805–807
Kuo CW, Genthner B (1996) Effect of added heavy metal ions on biotransformation and biodegradation of 2-chlorophenol and 3-chlorobenzoate in anaerobic bacterial consortia. Appl Environ Microbiol 62:2317–2323
Kwean OS, Cho SY, Yang JW, Cho W, Park S, Lim Y, Kim HS (2018) 4-Chlorophenol biodegradation facilitator composed of recombinant multi-biocatalysts immobilized onto montmorillonite. Bioresour Technol 259:268–275
Lowry OH, Rosebrough NJ, Farr AL, Randall RJ (1951) Protein measurement with the Folin phenol reagent. J Biol Chem 193:265–275
Ma WJ, Cheng YF, Jin RC (2022) Comprehensive evaluation of the long-term effect of Cu2+ on denitrifying granular sludge and feasibility of in situ recovery by phosphate. J Hazard Mater 422:126901
Mawad AM, Hesham AEL, Mostafa YM, Shoriet A (2016) Pyrene degrading Achromobacter denitrificans ASU-035: growth rate, enzymes activity, and cell surface properties. Rend Lincei 27:557–563
Monsalvo VM, Mohedano AF, Casas JA, Rodríguez JJ (2009) Cometabolic biodegradation of 4-chlorophenol by sequencing batch reactors at different temperatures. Bioresour Technol 100:4572–4578
Moreno-Andrade I, Valdez-Vazquez I, López-Rodríguez A (2020) Effect of transient pH variation on microbial activity and physical characteristics of aerobic granules treating 4-chlorophenol. J Environ Sci Health Part A 55:878–885
Nordin K, Unell M, Jansson JK (2005) Novel 4-chlorophenol degradation gene cluster and degradation route via hydroxyquinol in Arthrobacter chlorophenolicus A6. Appl Environ Microbiol 71:6538–6544
Nowak A, Mrozik A (2018) Degradation of 4-chlorophenol and microbial diversity in soil inoculated with single Pseudomonas sp. CF600 and Stenotrophomonas maltophilia KB2. J Environ Manage 215:216–229
Olaniran AO, Igbinosa EO (2011) Chlorophenols and other related derivatives of environmental concern: properties, distribution and microbial degradation pro-cesses. Chemosphere 83:1297–1306
Pandit PR, Kumar R, Kumar D, Patel Z, Pandya L, Kumar M, Joshi C (2021) Deciphering the black box of microbial community of common effluent treatment plant through integrated metagenomics: tackling industrial effluent. J Environ Manage 289:112448
Patel BP, Kumar A (2016) Multi-substrate biodegradation of chlorophenols by defined microbial consortium. 3 Biotech 6:1–10
Penttinen OP (1995) Chlorophenols in aquatic environments: structure-activity correlations. Finnish Zoological and Botanical Publishing Board, In Annales Zoologici Fennici, pp 287–294
Roque F, Diaz K, Ancco M, Delgado D, Tejada K (2018) Biodepuration of domestic sewage, textile effluents and acid mine drainage using starch-based xerogel from recycled potato peels. Water Sci Technol 77:1250–1261
Sandhibigraha S, Chakraborty S, Bandyopadhyay T, Bhunia B (2020) A kinetic study of 4-chlorophenol biodegradation by the novel isolated Bacillus subtilis in batch shake flask. Environ Eng Res 25:62–70
Saravanakumar K, Yun K, Maheskumar V, Yea Y, Jagan G, Park CM (2023) Construction of novel In 2S3/Ti3C2 MXene quantum dots/SmFeO3 Z-scheme heterojunctions for efficient photocatalytic removal of sulfamethoxazole and 4-chlorophenol: Degradation pathways and mechanism insights. Chem Eng J 451:138933
Shah V, Zakrzewski M, Wibberg D, Eikmeyer F, Schlüter A, Madamwar D (2013) Taxonomic profiling and metagenome analysis of a microbial community from a habitat contaminated with industrial discharges. Microb Ecol 66:533–550
Silva AS, Camargo FADO, Andreazza R, Jacques RJS, Baldoni DB, Bento FM (2012) Enzymatic activity of catechol 1, 2-dioxygenase and catechol 2,3-dioxygenase produced by Gordonia polyisoprenivorans. Quim Nova 35:1587–1592
Swain G, Sonwani RK, Singh RS, Jaiswal RP, Rai BN (2021) Removal of 4-chlorophenol by Bacillus flexus as free and immobilized system: effect of process variables and kinetic study. Environ Technol Innov 21:101356
Tariq SR, Shafiq M, Chotana GA (2016) Distribution of heavy metals in the soils associated with the commonly used pesticides in cotton fields. Scientifica 2016:1–11
Wang J, Sun Z (2020a) Exploring the effects of carbon source level on the degradation of 2,4,6-trichlorophenol in the co-metabolism process. J Hazard Mater 392:122293
Wang J, Sun Z (2020b) Effects of different carbon sources on 2,4,6-trichlorophenol degradation in the activated sludge process. Bioprocess Biosyst Eng 43:2143–2152
Wang J, Sun Z (2021) Successful application of municipal domestic wastewater as a co-substrate in 2,4,6-trichlorophenol degradation. Chemosphere 280:130707
Zada A, Khan M, Khan MA, Khan Q, Habibi-Yangjeh A, Dang A, Maqbool M (2021) Review on the hazardous applications and photodegradation mechanisms of chlorophenols over different photocatalysts. Environ Res 195:110742
Zhao J, Li Y, Chen X, Li Y (2018) Effects of carbon sources on sludge performance and microbial community for 4-chlorophenol wastewater treatment in sequencing batch reactors. Bioresour Technol 255:22–28
Ziagova M, Kyriakou G, Liakopoulou-Kyriakides M (2009) Co-metabolism of 2,4-dichlorophenol and 4-Cl-m-cresol in the presence of glucose as an easily assimilated carbon source by Staphylococcus xylosus. J Hazard Mater 163:383–439
Acknowledgements
Authors are thankful to the Biochemistry Department of North Eastern Hill University, India for providing an infrastructural facility for completing this study. Fellowship grant from Council of Scientific and Industrial Research (CSIR), Govt. of India to LK (19/06/2016(i)EU-V) is gratefully acknowledged.
Funding
Not applicable.
Author information
Authors and Affiliations
Contributions
AKS, LK, and NAS designed the study. LK performed the detailed experiments with help from JN. LKand WJL performed bioinformatic analysis. AKS and LK prepared the manuscript. All authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare that they have no conflicting interests.
Ethical approval
Not applicable.
Consent to participate
Not applicable.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Kipgen, L., Singha, N.A., Lyngdoh, W.J. et al. Degradation and metagenomic analysis of 4-chlorophenol utilizing multiple metal tolerant bacterial consortium. World J Microbiol Biotechnol 40, 56 (2024). https://doi.org/10.1007/s11274-023-03855-2
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s11274-023-03855-2