Abstract
Industrial units, manufacturing dyes, chemicals, solvents, and xenobiotic compounds, produce liquid and solid wastes, which upon conventional treatment are released in the nearby environment and thus are the major cause of pollution. Soil collected from contaminated Kharicut Canal bank (N 22°57.878′; E 072°38.478′), Ahmedabad, Gujarat, India was used for metagenomic DNA preparation to study the capabilities of intrinsic microbial community in dealing with xenobiotics. Sequencing of metagenomic DNA on the Genome Sequencer FLX System using titanium chemistry resulted in 409,782 reads accounting for 133,529,997 bases of sequence information. Taxonomic analyses and gene annotations were carried out using the bioinformatics platform Sequence Analysis and Management System for Metagenomic Datasets. Taxonomic profiling was carried out by three different complementary approaches: (a) 16S rDNA, (b) environmental gene tags, and (c) lowest common ancestor. The most abundant phylum and genus were found to be “Proteobacteria” and “Pseudomonas,” respectively. Metagenome reads were mapped on sequenced microbial genomes and the highest numbers of reads were allocated to Pseudomonas stutzeri A1501. Assignment of obtained metagenome reads to Gene Ontology terms, Clusters of Orthologous Groups of protein categories, protein family numbers, and Kyoto Encyclopedia of Genes and Genomes hits revealed genomic potential of indigenous microbial community. In total, 157,024 reads corresponded to 37,028 different KEGG hits, and amongst them, 11,574 reads corresponded to 131 different enzymes potentially involved in xenobiotic biodegradation. These enzymes were mapped on biodegradation pathways of xenobiotics to elucidate their roles in possible catalytic reactions. Consequently, information obtained from the present study will act as a baseline which, subsequently along with other “-omic” studies, will help in designing future bioremediation strategies in effluent treatment plants and environmental clean-up projects.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Many environmental habitats have lost their pristine characteristic since the beginning of industrialization on land as well as water bodies. In Gujarat (India), many industrial estates are situated within the “Golden Corridor” (a highly industrialized zone from Vapi to Mehsana). Industrial enterprises, manufacturing dyes, paints and pigments, pharmaceuticals, chemicals, solvents and textiles, release liquid wastes containing dyes, xenobiotic compounds and many other man-made products into the environment and thus are the major cause of ground and surface water pollution in these areas [1]. This has resulted in serious health problems of workers and people staying in slums surrounding these industrial estates. Hence, it becomes inevitable to develop novel bioremediation technologies for the treatment of industrial effluents, in order to reduce the impact of pollution on various sites in the vicinities of such industrial estates.
Implementation of efficacious bioremediation strategies relies on innate microbial community dynamics, structure, and function [2]. Depending on biotic and abiotic factors, microorganisms adapt to the environment and accordingly environmental conditions select for microorganisms featuring specific capabilities. Microbial communities are fundamental components of ecosystems playing a critical role in the catabolism and detoxification of anthropogenic/xenobiotic compounds [3, 4]. As microbial communities are involved in biogeochemical transformations within ecosystems, gaining insights into their metabolic dynamism of coping with environmental changes will help in responding to future environmental catastrophes. Analyses of soil microbial communities have provided the clue of the extent of damage in ecosystems. Many xenobiotic-contaminated areas have undergone a shift in microbial community composition [5]. Contaminated environments are enriched with pollutants and hence the bacteria capable of xenobiotic degradation are widely distributed in these environments. Adapted bacteria have evolved to utilize a variety of compounds that are present in the environment. Any individual microorganism is incapable of accomplishing all the metabolic reactions to degrade environmental pollutants. However, a sub-community comprising diverse organisms collectively interacts to perform all the metabolic reactions for bioremediation [3, 4]. More than 99 % of the microbes that exist in the environment cannot be cultivated easily [6–8], and consequently, most of the microbes in the environment have not been described and accessed for biotechnology or basic research [7]. Therefore, metagenomics-based analyses of entire microbial community become imperative to delineate the metabolic pathways responsible for biodegradation.
Metagenomics (also known as community genomics, ecogenomics, or environmental genomics), which aims to access the genomic potential of an environmental habitat either directly or after enrichment for specific capabilities, has had the greatest impact within the last few years [3, 6, 9, 10]. Two approaches, the function-driven analysis and the sequence-driven analysis, have been applied to obtain biological information from metagenomic libraries. The function-driven analysis is based on the identification of clones expressing a desired trait. The limitations of this approach are that it requires clustering of all genes required for expression of the function of interest in the host cell and the availability of an assay that can be performed efficiently on large libraries [11]. The sequence-driven analysis relies on the use of hybridization probes or PCR primers to screen metagenomic libraries for target genes or by large-scale random metagenome sequencing. There has been disagreement about the informative value of random metagenome sequencing as some scientists consider this approach as too undirected to yield biological understanding. Conversely, others stress that there is so little known about some divisions of Bacteria that any genomic sequence is helpful in guiding the design of experiments to reveal their biology and leading to significant discoveries [11]. Environmental sequencing was seen as a promising approach as early as in the 1990s. However, until a few years back, it was mostly used for sequencing of 16S rRNA genes to gain insights into the microbial composition of habitats. Since the development of next-generation sequencing technologies such as pyrosequencing [12], sequencing has become less expensive, faster, and less tedious. Consequently, more and more complete microbial genomes as well as environmental metagenomes are being sequenced to gain insights into functional aspects besides species composition [8]. Metagenomics is a burgeoning area that is generating enormous amounts of biological information. The development of new bioinformatics approaches and tools is allowing innovative mining of both existing and new data [9].
The environmental site analyzed in this study receives effluents from a variety of industries involved in manufacturing of various chemicals, dyes, solvents, paints, and many other xenobiotic compounds. Consequently, the intrinsic microbial community has to be capable of dealing with such a mixture of contaminants. Generally, a particular species or group of organisms may be tolerant to or might be able to degrade a particular class of compound(s). However, they are not able to cope with the variety of contaminants. It is very likely that different species degrade different toxic compounds and this concerted action may lead to environmental sites that are permissive for survival of microorganisms that do not feature specific degradative capabilities but live in syntrophic associations with other microorganisms. The study aims at the characterization of the microbial community inhabiting an industrially contaminated site for its taxonomic profile and catabolic gene potential by means of a sequence-driven metagenomic approach. Taxonomic profiling will provide insights into the composition of the microbial community capable of tolerating and/or degrading xenobiotic compounds. Functional characterization of metagenome sequence reads on the basis of Gene Ontology (GO) terms [13], Clusters of Orthologous Groups of proteins (COG) accessions [14], protein family (Pfam) numbers [15], and Kyoto Encyclopedia of Genes and Genomes (KEGG) database entries [16] will lead to elucidation of the catabolic potential of the indigenous microbial community. This approach will facilitate identification of genes essential for key catalytic steps in biodegradation pathways. Subsequently, complementary pathways and catalytic reactions, for biodegradation of specific xenobiotic compounds, missing in indigenous microbial population can be supplemented by an approach termed as “bioaugmentation.” Concisely, the aim of this study was to assess the genomic potential of the indigenous microbial community of the contaminated soil habitat.
Materials and Methods
Contaminated Site
The soil samples were collected from the contaminated banks of Kharicut Canal (N 22°57.878′; E 072°38.478′), flowing through Gujarat Industrial Development Corporation (GIDC) situated in Vatva (Ahmedabad, Gujarat, India) and into the Khari River. Soil, in incessant contact with the flowing contaminated river, along with little amount of contaminated canal water was collected in sterile containers. Soil samples were taken from the top layer (actually, sides of the bank are in contact with contaminated water) till 8 in. depth. The soil samples were stored at 4 °C.
Metagenomic DNA Preparation
Twenty grams of contaminated river bank soil sample was used for total community DNA preparation using the Zhou et al. [17] protocol with some modifications. The method was based on lysis with a high salt extraction buffer [10 % (w/v) sucrose, 1 % (w/v) CTAB, 1.5 M NaCl, 100 mM Tris–Cl (pH 8.0), 100 mM EDTA (pH 8.0), 25 mM sodium phosphate buffer (pH 8.0)] and extended heating (1–2 h) in the presence of sodium dodecyl sulfate, lysozyme, and proteinase K along with mechanical shearing. Ribolyzer (FastPrepTM FP120, Thermo Savant, USA) with ribolyzer tubes (Lysing Matrix B, 2 ml tubes, MP Biomedicals, USA) was used for mechanical cell lysis. An additional step of powdered activated charcoal treatment was given before chloroform washes [18]. The precipitation was done by adding polyethylene glycol (PEG) 10,000 at a final concentration of 5 % and incubating at 4 °C overnight. DNA concentration was measured by means of a NanoDrop spectrophotometer (NanoDrop Technologies Inc., Delaware, USA) and analyzed by gel electrophoresis. Further qualitative and quantitative confirmation was done using Quant-iT PicoGreen dsDNA kit (Invitrogen) and the Tecan Infinite 200 Microplate Reader (Tecan Group Ltd., Switzerland).
Sequencing of the Metagenomic DNA on the GS FLX Titanium Platform
Sequencing of the metagenomic DNA was done by applying the whole genome shotgun sequencing approach on the Genome Sequencer FLX system (Roche Applied Science, Manheim, Germany) by applying titanium chemistry. Five micrograms of DNA was used to generate a whole genome shotgun library according to the protocol given by the manufacturer. After titration, 6.5 DNA copies per bead were used for the main sequencing run. After emulsion PCR and subsequent bead recovery, 790,000 DNA beads were loaded on quarter of the PicoTiter Plate and subjected to sequencing. The reads were assembled into contigs by means of the Genome Sequencer De Novo Assembler Software (Roche Applied Science, Mannheim, Germany). The subsequent taxonomic and functional analyses of metagenome data were carried out using Sequence Analysis and Management System for Metagenomic Datasets (MetaSAMS) [19].
Taxonomic Analysis
The taxonomic interpretation of the metagenome sequences was accomplished by applying three different approaches, namely, 16S rDNA, environmental gene tags (EGTs), and lowest common ancestor (LCA).
Taxonomic Profiling Based on 16S rDNA Sequences Using RDP Classifier
The microbial composition of the contaminated site was characterized by using fragments of 16S rDNA as phylogenetic anchors. 16S rDNAs were detected in a BLAST search of all metagenome reads vs. the 16S rRNA database [20]. All sub-regions of reads having a BLAST hit with an E value <1 × 10−10 were phylogenetically classified using the Ribosomal Database Project (RDP) Classifier [21]. The RDP classifier predicted the taxonomic origin of 16S rDNA up to the rank of genus.
Taxonomic Profiling Based on EGTs Using CARMA
Phylogenetic algorithm CARMA [22] was used with standard parameters. Phylogenetic trees were constructed for matching Pfam accessions and found EGTs were classified into a higher order taxonomy based on their phylogenetic relationships to family members with known taxonomic affiliations. In the following, regions of reads matching a Pfam are called EGTs.
Taxonomic Profiling Based on LCA of Multiple Blast Hits
The reads were compared against known species in a given taxonomy obtained from the NCBI database using BLAST. The fragments that could not be unequivocally assigned to a specific taxon were assigned to an inner node of the taxonomy using the LCA of all sequences to which the read might be assigned [21, 23–26].
Allocation and Mapping of Metagenome Single Reads on Microbial Genomes
Metagenome single reads were mapped on available microbial genomes by aligning to the sequenced genome(s) based on a BLASTN search of reads vs. the genome sequence(s) available at the NCBI database. An E value cutoff of 1 × 10−50 was set. The coverage of reference genome sequence by reads was visualized using the MetaSAMS [19].
Functional Characterization
To assess the contaminated soil microbial community and their genetic potential for biodegradation, the metagenome data were functionally annotated. The analysis pipeline included gene predictions and different BLAST tools: BLAST2x vs. the SWISSPROT protein database (E value cutoff of 1 × 10−10) and BLAST2x vs. the COG protein database (E value cutoff of 1 × 10−10). Furthermore, Hidden Markov model-based search vs. Pfam and Tigrfam was applied. For the prediction of coding sequences, the Glimmer 3 was used.
Characterization of Genes in Metagenome Reads According to GO Terms
Metagenome reads were annotated for their GO terminologies [13] in MetaSAMS [19], which provided gene and gene product details.
Characterization and Classification of Genes in Contigs According to COG Categories
Using MetaSAMS [19], the predicted genes in large contigs were classified according to COG [14]. Alignment to COG was done using BLASTX with an E value cutoff of 1 × 10−10. Subsequently, they were functionally annotated according to their best BLAST hit.
Characterization of Genes in Metagenome Reads According to Pfams
Metagenome reads were annotated for their Pfams [15] in MetaSAMS [19]. Pfam protein family members found in the contaminated site metagenome were predicted using the algorithm CARMA [22].
Characterization and Gene Annotation According to KEGG Database
Using MetaSAMS [19], the predicted genes in large contigs were further characterized for their Enzyme Commission (EC) numbers by a BLAST search against the KEGG database [16]. However, the large contigs contain only 3.54 % of total bases sequenced. Consequently, the EC numbers were detected in all metagenome single reads by their best BLAST hit against the KEGG database. The EC numbers detected in large contigs were mapped on KEGG pathways involved in xenobiotic compound biodegradation. The EC numbers detected in all metagenome single reads were compared to a list of enzymes involved in biodegradation of xenobiotic compounds available in the University of Minnesota Biocatalysis/Biodegradation Database (UM-BBD; http://umbbd.msi.umn.edu/) [27]. UM-BBD was developed in 1995 and is regularly updated. On the day of comparison, it contained information on 1,246 compounds, 878 enzymes, 1,345 reactions, 275 biotransformation rules, and had 510 microorganism entries.
Data Availability
Sequence data of Genome Sequencer (GS) FLX titanium has been deposited at NCBI and the accession number is SRX209034.
Results
Metagenome Analysis
Contaminated canal bank soil collected from the industrial area was used for metagenomic DNA preparation using the protocol of Zhou et al. [17] with some modifications. By using PEG 10,000, we could remove humic acids and other impurities. Consequently, there was no need for further purification steps (such as gel permeation chromatography and/or other chromatographic techniques). The obtained DNA was of high molecular weight and was pure enough for PCR, restriction digestion, ligation, and other further studies. Details about purity ratio (260/280:1.93) and quality of metagenomic DNA with respect to humic acids (260/230:2.13) have been described in Table 1. A quarter of a run on the GS FLX platform using titanium chemistry generated 409,782 sequence reads amounting to a total of 133,529,997 bases of sequence information. The average read length was 325.9 bases and the GC content varied from 5.83 to 88.46 %. In total, 87,331 reads (21.31 % of total reads) comprising 6,550,595 bases (4.91 % of total bases) were assembled into 11,038 contigs. After removing small contigs (<500 bp), 5,592 contigs comprising of 4,727,081 bases (3.54 % of total bases) were obtained. The largest contig had a size of 6,617 bases and the average contig length was around 845.3 bases. Statistical data summarizing the sequencing details are given in Table 2. Assembly statistics indicates that the sequencing approach is far from saturation.
Taxonomic Profiling of the Microbial Community by Using Three Different Complementary Approaches
To deduce the taxonomic composition of the underlying microbial community, three different complementary approaches were carried out: (1) classification based on 16S rRNA gene sequences by means of the RDP Classifier, (2) classification based on EGTs applying the CARMA software, and (3) the LCA classification based on BLAST results. Taxonomic classification was carried out at all taxonomic ranks.
Out of a total of 409,782 reads, 16S rRNA gene-specific sequences were identified in 1,510 reads (∼0.4 % of all reads). According to the RDP Classifier, Bacteria is the dominant domain (99.9 %; 1,508 reads) and only two reads were classified as Archaea. Proteobacteria (47 %) is the most abundant phylum followed by Firmicutes (28 %), Bacteroidetes (9 %), and others (16 %). At rank class, Gammaproteobacteria (22 %) is the most abundant followed by Clostridia (21 %), Anaerolineae (8 %), and others (49 %). At rank order, Clostridiales (22 %) is the most abundant followed by Pseudomonadales (16 %), Bacteroidales (7 %), and others (55 %). However, at rank family Pseudomonadaceae (19 %) is the most abundant followed by Clostridiaceae (10 %), Caldilineaceae (9 %), and others (62 %). At rank genus, Pseudomonas (21 %) is the most abundant followed by Clostridium (7 %), Shewanella (5 %), and others (67 %). The taxonomic composition of the community as deduced from 16S rDNA sequence classification is depicted in Fig. 1.
View of taxa among Bacteria at all ranks classified according to 16S rDNA (RDP Classifier). The taxa abundant and playing a role in xenobiotic biodegradation are displayed. Below the names at each rank, numbers are represented wherein the first number specifies the total number of reads classified to that taxa and the second number written in brackets specifies the reads that can be accurately classified only till this rank and cannot be classified at lower ranks
Microbial taxonomic classification based on EGTs was carried out in parallel to the 16S rRNA gene fragment analysis. Out of a total of 409,782 reads, EGTs were identified in 90,158 reads (∼22 % of all reads) by applying the CARMA software. Based on EGTs, Bacteria (92.5 %) is the dominant domain followed by Eukaryota (5 %) and Archaea (2 %), respectively. Proteobacteria (54 %) is the most abundant phylum followed by Firmicutes (17 %), Bacteroidetes (9 %), and others (20 %). At rank class, Gammaproteobacteria (23 %) is the most abundant followed by Clostridia (11 %), Alphaproteobacteria (11 %), and others (55 %). At rank order, Pseudomonadales (12 %) is the most abundant followed by Clostridiales (10 %), Bacteroidales (7 %), and others (71 %). At rank family, Pseudomonadaceae (14 %) is the most abundant followed by Clostridiaceae (8 %), Bacteroidaceae (5 %), and others (73 %). At rank genus, Pseudomonas (15 %) is the most abundant taxon followed by Clostridium (7 %), Shewanella (6 %), and others (72 %). Taxonomic classification based on CARMA results is shown in Table 3.
Thirdly, microbial classification was carried out based on the LCA approach analyzing all hits obtained in a BLAST search. Out of a total of 409,782 reads, 66,682 reads (∼16 % of all reads) were classified according to the LCA method. Based on LCA classification, Bacteria (98.8 %, 65,982 reads) is the dominant domain followed by Archaea (1 %, 677 reads) and Eukaryota (0.1 %, 11 reads), respectively. Proteobacteria (77 %) is the most abundant phylum followed by Actinobacteria (9 %), Firmicutes (8 %), and others (6 %). At rank class, Gammaproteobacteria (45 %) is the most abundant followed by Betaprotobacteria (13 %), Alphaproteobacteria (11 %), and others (31 %). At rank order, Pseudomonadales (30 %) is the most abundant followed by Alteromonadales (13 %), Burkholderiales (7 %), and others (50 %). At rank family, Pseudomonadaceae (32 %) is the most abundant followed by Shewanellaceae (14 %), Bifidobacteriaceae (4 %), and others (50 %). At rank genus, Pseudomonas (32 %) is the most abundant taxon followed by Shewanella (14 %), Bifidobacterium (5 %), and others (49 %). These results are summarized in Table 4.
The most abundant phyla (Fig. 2a) and genera (Fig. 2b) are depicted for all the three different complementary classification approaches. Out of a total of 1,508 16S rDNA reads, 574 reads (38 %) and 290 reads (19 %) could be assigned to taxa at ranks phylum and genus, respectively. Comparatively, out of a total of 83,649 EGTs identified, 71,705 EGTs (86 %) and 39,476 EGTs (47 %) could be assigned to taxa at ranks phylum and genus, respectively. According to the classification based on the lowest common ancestor approach, out of a total of 65,982 reads, 65,378 reads (99 %) and 53,676 reads (81 %) could be assigned to taxa at ranks phylum and genus, respectively. Rarefaction curves, of all the three microbial classifications (RDP, CARMA, and LCA) accomplished in this study, are shown for rank phylum (Supplement Fig. S1a) and rank genus (Supplement Fig. S1b). At rank phylum, 13, 36, and 27 taxa, and at rank genus, 67, 545, and 340 taxa were identified according to 16S rDNA, EGT, and LCA assignments, respectively.
Allocation and Mapping of Metagenome Single Reads to Microbial Genomes
Metagenome reads were mapped on genomes of microorganisms and the results are shown in Fig. 3. The highest number of reads was allocated to the Pseudomonas stutzeri A1501 (9,511 reads, 2.3 %) genome followed by Shewanella baltica OS223 (4,747 reads, 1.2 %), S. baltica OS185 (4,219 reads, 1 %), S. baltica OS195 (4,153 reads, 1 %), S. baltica OS155 (3,961 reads, 0.97 %), and P seudomonas flourescens PfO-1 (3,226 reads, 0.8 %).
Analysis of Genetic Potential of Contaminated Soil Indigenous Microbial Community
Functional annotation and characterization of genes allowed deeper insights into genetic potential, possessed by indigenous microbial community, for the biodegradation of xenobiotic compounds. Metagenome dataset analysis, with respect to GO terms, COG categories, Pfams, and KEGG hits, is described in the following subsections.
Characterization of Genes in Metagenome Reads According to GO Terms
Metagenome reads were characterized for their GO terms [13]. One hundred forty-three thousand six hundred forty-four reads (35.05 % of all reads) were classified according to GO terms. GO terms for carbohydrate metabolism (0005975), glycolysis (0006096), TCA cycle (0006099), electron carrier activity (0009055), hydrolase activity (0016787; 0016811), transferase (0016740), transporter (0005215), aromatic compound metabolism (0006725; 0019439), chromate transporter (0015109; 0015703), nitrate reductase (0009325; 0008940), monooxygenase (0004497), arsenic response (0046685; 0015105), peroxidase (0004601), organomercury catabolic process (0046413), catechol (0018576), cytochrome c-oxidase (0004129), sulfur metabolism (0006790; 0008146), nitrile hydratase (0018822), anaerobic electron transport chain (0019645), response to oxidative stress (0006979), oxidoreductase activity (0016705; 0016616; 0016651; 0016712; 0016625; 0016620; 001655; 0016669; 0016614; 0016730), and others that are associated with biodegradation pathways and are represented in the metagenome dataset analyzed are described in Supplement Table S2.
Characterization and Classification of Genes According to COG Categories
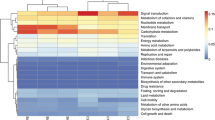
Assembled contigs were analyzed by assigning predicted functions to genes based on COGs [14]. In total, 5,822 hits corresponding to 1,650 different COG accessions were identified and subsequently classified into 22 classes based on functional categories (Fig. 4). Amongst all functional COG categories, the class “energy production and conversion (C)” was characterized for various kinds of oxidoreductases, which have been reported for dye and other xenobiotic compound degradation under stress conditions. The other classes such as “inorganic ion transport and metabolism (P)” and “coenzyme metabolism (H),” “secondary metabolites biosynthesis, transport, and catabolism (Q),” and “signal transduction mechanisms (T)” are associated with transport of ions/compounds and other metabolic processes. COG categories/accessions important in xenobiotic biodegradation are described in Supplement Table S3.
Categorization of assembled reads according to Clusters of Orthologous Groups of proteins. Categories are abbreviated as follows: B, chromatin structure and dynamics; L, replication, recombination and repair; K, transcription; J, translation, ribosomal structure and biogenesis; D, cell cycle control, cell division, chromosome partitioning; M, cell wall/membrane/envelope biogenesis; N, cell motility; O, posttranslational modification, protein turnover, chaperones; T, signal transduction mechanisms; U, intracellular trafficking, secretion and vesicular transport; V, defense mechanisms; C, energy production and conversion; E, amino acid transport and metabolism; F, nucleotide transport and metabolism; G, carbohydrate transport and metabolism; H, coenzyme transport and metabolism; I, lipid transport and metabolism; P, inorganic ion transport and metabolism; Q, secondary metabolite biosynthesis, transport and catabolism; Z, cytoskeleton; R, general function prediction only; and S, function unknown
Characterization of EGTs According to Pfams
Metagenome single reads were characterized according to Pfam categories [15]. In total, 96,125 reads (23.5 % of all reads) were assigned to Pfam entries. In particular, Pfams representing enzymes with a predicted role in xenobiotic tolerance and biodegradation were analyzed. The Pfam entries compiled include key enzymes involved in biodegradation such as the dyp-type peroxidase family (PF04261), catechol dioxygenase (PF04444), 2-nitropropane dioxygenase (PF03060), dioxygenase (PF00775), phenol hydroxylase (PF07976; PF06099), NADH ubiquinone oxidoreductase (PF01058), copper amine oxidase (PF07833; PF01179; PF02727), aromatic ring-opening dioxygenase (PF07746), cytochrome oxidase (PF02322), cytochrome ubiquinol oxidase (PF01654), nitrate reductase (PF02613; PF03892; PF02665), and others. Many transporter proteins such as chromate transporter (PF02417), benzoate membrane transport protein (PFO3594), mercuric transport protein (PF02411), ABC nitrate/sulfonate/biocarbonate family transporter (PF09821), BCCT family transporter (PF02028), arsenical pump membrane transport protein (PF02040), ferrous ion transport protein B (PF07664; PF02421), and others were identified in the metagenome sequenced. Moreover, many genes for proteins essential for resistance to metals or for tolerance to xenobiotics such as organic solvent tolerance protein (PF04453), toxic anion resistance protein (PF05816), copper resistance protein (PF04234), arsenical resistance operon trans-acting repressor ArsD (PF06953), tellurium resistance protein (PF10138), chromate resistance exported protein (PF09828), tellurite resistance protein (PF05099), cadmium resistance transporter (PF03596), toluene tolerance (PF05494), and others were also identified. Besides these, many other Pfam members predicted to play a general role in biodegradation such as bacterial stress protein (PF02342), electron transfer flavoprotein-ubiquinone oxidoreductase (PF05187), sulfotransferase (PF00685), sulfur oxidation protein (PF08770), stringent starvation protein (PF04386), and many other proteins involved in energy transfer essential for catabolism were also identified. Identified Pfam members possibly involved in xenobiotic biodegradation are listed and described in Supplement Table S4.
Characterization of Genes According to KEGG Database
In total, 2,772 KEGG hits corresponding to 670 different EC numbers were identified and characterized in large contigs. The EC numbers were mapped on 21 different KEGG pathways (Table 5). Eighteen, 16, and 11 enzymatic functions were mapped on the benzoate degradation via CoA ligation, metabolism of xenobiotics by cytochrome P450, and 1,2-methylnaphthalene degradation pathways, respectively. A schematic figure showing mapped enzymes on the benzoate degradation via CoA ligation pathway is presented in Fig. 5.
Moreover, EC numbers were searched and characterized in all metagenome reads as well. In total, 157,024 reads (corresponding to 37,028 different EC numbers) were analyzed for producing hits to the KEGG database. Subsequently, these EC numbers were mapped on a selected list of enzymes involved in biodegradation of xenobiotic compounds available in UM-BBD. Eleven thousand five hundred seventy-four reads corresponding to 131 different enzymes, such as benzyl alcohol dehydrogenase, azoreductase, lignin peroxidase, catechol 1,2 dioxygenase, catechol 2,3 dioxygenase, acetoacetyl CoA reductase, protocatechuate 3,4 dioxygenase, protocatechuate 4,5 dioxygenase, formate dehydrogenase, citronellal dehydrogenase, carbon monoxide dehydrogenase, 2,5-dichloro-2,5-cyclohexadiene-1,4-diol dehydrogenase, enoyl CoA reductase, trimethylamine dehydrogenase, butyryl CoA dehydrogenase, benzoyl-CoA reductase, glutaryl CoA dehydrogenase, NAD(P)H nitroreductase, and others, involved in biodegradation were identified. These enzymes could be mapped on biodegradation pathways of complex compounds such as benzoate, toluene, dinitro/trinitrotoluene, xylene, cyclohexane, dyes, 1,2-dichloroethane, citronellol, nitroglycerin, styrene, trichloroethane, 2-nitropropane, dimethylether, phthalate, biphenyl, nitrilotriacetate, 2-aminobenzenesulfonate, 2,4-dichlorophenoxyacetic acid, 3-fluorobenzoate, 4-fluorobenzoate, chromium, anthracene, octane, propylene, pentaerythritol tetranitrate, gallate, octane, thiocyanate, hexahydro-1,3,5-trinitro-1,3,5-triazine, and many others. Eighty-eight enzymes and their possible reactions and their probable participation in biodegradation pathways are described in Supplement Table S5.
Enzymes in Xenobiotic Biodegradation
In the present study, many sequence reads corresponded to azoreductases, chromate reductase, aresenite oxidase, arsenite reductase, benzoate 1,2-dioxygenase, phenol 2-monooxygenase, catechol 1,2-dioxygenase, catechol 2,3-dioxygenase, NAD(P)H reductase, benzoyl-CoA reductase, 4-hydroxy benzoyl-CoA reductase, 2,5-dichloro-2,5-cyclohexadiene-1,4-diol dehydrogenase, and other enzymes playing a direct role in biodegradation processes. Selected xenobiotic compound-degrading enzymes and their probable source organisms identified by means of BLAST searches of metagenome sequences are described in Table 6.
Discussion
Industrial estates [situated in GIDC, Vatva, Ahmedabad], manufacturing dyes, chemicals, solvents and other xenobiotic compounds, produce liquid and solid wastes which upon conventional treatment are released in the nearby environment. Due to persistent release of a variety of toxic wastes, the surrounding water and soil bodies have become highly contaminated with many different xenobiotic compounds [1]. Consequently, to analyze the microbial community inhabiting an industrially contaminated site in terms of its composition, diversity, gene content, metabolic capabilities, and role of specific organisms in xenobiotic biodegradation, a metagenomic approach was pursued.
The first step in any metagenomic analysis consists of isolating high-quality DNA from environmental samples in an unbiased manner. Methods for nucleic acid extraction from soil may be limited and biased by incomplete cell lysis, DNA adsorption to soil surfaces, and co-extraction of enzymatic inhibitors from soil, and loss, degradation, or damage of DNA [28]. Organic matter is the major source of inhibitors that are co-extracted from soil along with metagenomic DNA. In particular, humic acids interfere with enzymatic manipulations of DNA and thus pose the major problem [29, 30]. The presence of organic contaminants and heavy metals can greatly influence the recovery of total community DNA [31]. Moreover, it is more tedious to obtain high-quality DNA from soil samples contaminated by discharges from industrial estates as they consist of dyes, aromatic compounds, amines, paints, chemicals, solvents, and other xenobiotic pollutants besides regular contaminants like humic acids.
On taxonomic analyses, the most abundant phylum and genus were found to be “Proteobacteria” and “Pseudomonas,” respectively. Many studies such as identification of nitrogen-incorporating bacteria in petroleum-contaminated Arctic soils [32], identification of bacteria utilizing biphenyl, benzoate, and naphthalene in long-term contaminated soil [33], assessing the suitability of bioremediation for the treatment of hydrocarbon-impacted soil [34], diversity, and structure of soil microbial communities on exposure to chromium and arsenic [35], bacterial community analyses in Permafrost soils along the China–Russia crude oil pipeline by pyrosequencing [36], dynamics of bacterial communities in unpolluted soils after spiking with phenanthrene [37], and others have analyzed the bacterial communities in different kinds of contaminated soil and have reported the abundance and/or selection of Proteobacteria. 16S RNA-based pyrosequencing data have shown that although Proteobacteria are present in normal soil, however, their phylogenetic diversity increases in hydrocarbon-contaminated soils [32, 34].
In the present study, taxonomic profiling was carried out by three different complementary approaches to obtain a complete picture. Each of the methods has certain advantages and limitations, and consequently, some variations were observed. The classification based on 16S rRNA sequences (RDP Classifier) is the most widely used approach. However, only very few reads representing 16S rRNA gene fragments are present in metagenome dataset. Consequently, two other approaches were also used for taxonomic classification of the community. The discrepancies found between them were that the phylum Proteobacteria was predicted to be 47 and 54 % by the 16S rDNA and EGT classification, respectively. However, this phylum was predicted to be 77 % when the classification was done by the LCA analysis. As a result of higher representation of Proteobacteria, other phyla, namely Firmicutes and Bacteroidetes, were underrepresented when classified based on LCA in comparison to the other two approaches. This can be explained by the fact that as the lowest common ancestor is selected and that many genes important for survival in contaminated sites and as well as xenobiotic biodegradation might have been transferred horizontally from Proteobacteria to other phyla. The fact that phylum Proteobacteria is present in higher proportion and plays a very active role in biodegradation is supported by the observations in the studies related to heavy metal contamination, waste-water treatments and other contaminated sites all over the world. Besides this major difference, other small variations are that the phylum Cyanobacteria was classified to be 1 % according to EGT classification, whereas it was not detected based on 16S rDNA and only 0.04 % reads were classified to this phylum based on LCA. At the genus level, Bacteroides could not be classified based on RDP and was underrepresented based on LCA, whereas the genus Pseudomonas and Shewanella were overrepresented by classification based on LCA. All other predictions corroborated among all three approaches.
Rarefaction analyses were carried out to determine whether the metagenome sequencing approach was carried out to saturation which is a prerequisite to deduce the complete taxonomic profile of the community. The rarefaction curves are nearly reaching the plateau phase (saturation) at rank genus for classifications based on EGTs and the LCA. However, since only very few 16S rDNA sequences were identified in the metagenome dataset, the corresponding rarefaction curve did not reach the plateau phase.
Mapping of metagenome reads onto bacterial genomes suggested that organisms related to the identified species were enriched at the site contaminated with xenobiotics and presumably play an active role in biodegradation. Many reports have described the roles and mechanisms of reference strains in biodegradation and bioremediation. P. stutzeri has been reported to degrade carbon tetrachloride [38], phenanthrene [39], o-xylene [40], and dye effluents [41, 42]. Moreover, analysis of the genome sequence of many different Pseudomonas species has revealed details about oxygenases, oxidoreductases, ferredoxins and cytochromes, dehydrogenases, sulfur metabolism proteins, and others [43]. Moreover, this genus also possesses many operons coding for the catabolism of a large number of aromatic compounds and gene clusters encoding enzymes that are predicted to be involved in the metabolism of non-natural substrates [44]. S. baltica plays an important role in the bioremediation of sites contaminated with organic pollutants (such as naphthalene, styrene, and others), radionuclides, and heavy metals [45, 46]. The genus Rhodobacter has been reported for nitrophenol [47], chlorobenzenes, and azo dye decolorization [48].
In the last decade, enzymes involved in environmental bioremediation gained considerable importance, and thus, various new approaches have been applied for detailed studies on some classes of relevant enzymes. Many of the enzymes active in degradation pathways are linked from their protein phylogeny and not strictly linked to the taxonomical affiliation of their host bacteria [49], indicating that the genes encoding those catabolic enzymes are involved in very dynamic events. Even observations in other studies, such as the presence of tmo-like genes in phylogenetically distant strains of Pseudomonas, Mycobacterium, and Bradyrhizobium [50], and others suggest towards horizontal gene transfer. To characterize the catabolic potential for biodegradation, it is necessary to take into consideration the broad diversity of catabolic routes evolved by microorganisms and also the diversity of enzymes of a given gene family or even between gene families [51]. Therefore, the catabolic gene potential has to be analyzed independently as any presumption that is based on taxonomic profiles only allows for very vague statements regarding functional assignments and will result in circumstantial and unresolved associations [51].
Microbial activities of importance in biodegradation, such as oxidation, reduction, binding, immobilization, volatilization, or transformation, are carried out by enzymes such as oxidases, reductases, oxygenases, and many others. However, only few specific enzymes are involved in biodegradation. Conversely, there are many enzymes which by their specific role are involved in cellular metabolic functions but under stress conditions, induced by pollutants such as hydrocarbons, dyes, and aromatic and xenobiotic compounds, they perform alternate functions in metabolic pathways involved in biodegradation. Several enzymes are endowed with promiscuous activities [4]. The term “catalytic promiscuity” describes the capability of an enzyme to catalyze different reactions, called secondary activities, at its active site. Furthermore, from a basic point of view, studies of catalytic promiscuity offer clues to understand the natural evolution of enzymes and to translate this into in vitro adaptation of enzymes to specific human needs [52].
Consequently, to deduce the genetic potential of the microbial community of the contaminated soil habitat and its adaptive features regarding biodegradation of xenobiotic compounds, functional annotation and characterization of genes were carried out in many different ways. Metagenome single reads and contigs were annotated according to GO terms, COG accessions, Pfams, and KEGG hits thus assigning predicted functions to coding sequences. The predicted enzymes were mapped on biodegradation pathways to elucidate the probable catabolism pathways of inhabiting microbial community.
Many other studies have also been carried out to understand the microbial metabolism involved in xenobiotic biodegradation. Kim et al. [53] have described a stepwise overview of degradation pathway in which high molecular weight polycyclic aromatic hydrocarbons are degraded into the β-ketoadipate pathway through protocatechuate and then mineralized to carbon dioxide via the TCA cycle. Many enzymes characterized by them have been identified in our metagenome reads. Perez-Pantoja et al. [54] in their review have described the catabolic potential of Cupriavidus necator JMP134 in aromatic compound degradation. Of the 140 aromatic compounds tested, 60 served as a sole carbon and energy source for this strain, strongly correlating with those catabolic abilities predicted from genomic data. Almost all the main ring-cleavage pathways for aromatic compounds are found in C. necator: the β-ketoadipate pathway, with its catechol, chlorocatechol, methylcatechol, and protocatechuate ortho ring-cleavage branches; the (methyl)catechol meta ring-cleavage pathway; the gentisate pathway; the homogentisate pathway; the 2,3-dihydroxyphenylpropionate pathway; the (chloro)hydroxyquinol pathway; the (amino)hydroquinone pathway; the phenylacetyl-CoA pathway; the 2-aminobenzoyl-CoA pathway; the benzoyl-CoA pathway; and the 3-hydroxyanthranilate pathway. C. necator has been identified in our metagenome reads and correspondingly the possibilities of above described capabilities. Pieper and Seeger [55] have described the microbial metabolism for degradation of polychlorinated biphenyls in their review. They have described the biphenyl upper pathway and the enzymes involved in it followed by lower pathways for the degradation of products (2-hydroxypenta-2,4-dienoates and benzoates) of upper pathway. Suenaga et al. [56] from their studies have concluded that the complete pathways generally reported as “upper” and/or “lower” pathway modules are extremely rare. Instead, they identified various types of gene subsets, suggesting that aromatic compounds in the natural environment are degraded through the concerted actions of various fragmental pathways. Denef et al. [57, 58], using a combined approach of genetics, transcriptomics, and proteomics, showed that all three presumed benzoate pathways act in a coordinated manner in Burkholderia xenovorans LB400. Consequently, these studies also provide further proof of the complex interconnected regulatory network controlling aromatic catabolic pathway. Till few years back, these kinds of studies were carried out mainly on understanding an individual microorganism. However, in recent years since the developments in next-generation sequencing, the focus has shifted towards understanding the microbial metabolism in metagenome or enriched metagenome (community genomics).
In the present study, many enzymes (class of enzymes) playing a role in xenobiotic biodegradation have been identified in metagenomic data set. This provides us with information about the capabilities of indigenous microbial population. Many studies have reported the roles of oxygenase systems (dioxygenases and monooxygenases) in biodegradation of xenobiotic compounds. Brennerova et al. [59] described the exceptionally high extradiol dioxygenase diversity at a site highly contaminated with aliphatic and aromatic hydrocarbons which indicated that this function confers a positive biological fitness to the indigenous microbial community members. Witzig et al. [60] assessed the toluene/biphenyl dioxygenase gene diversity in benzene, toluene, ethylbenzene, and xylene (BTEX)-polluted soils and also concluded that indigenous bacteria seem to possess a genotypic flexibility, which is important for their adaptation and evolution while facing challenging and continuously changing conditions in these ecosystems. Cavalca et al. [50] studied gene fragments corresponding to toluene monooxygenase, catechol 1,2-dioxygenase, catechol 2,3-dioxygenase, and toluene dioxygenase in bacterial communities inhabiting a BTEX-polluted groundwater. Iwai et al. [61] have developed a microarray to detect di- and monooxygenases involved in benzene degradation and for the rapid profiling of benzene oxygenase gene diversity in contaminated soils. Suenaga et al. [62] studied the molecular basis for adaptive evolution in novel extradiol dioxygenases retrieved from the metagenome. This kind of studies will be of immense help in understanding the adapted novel/modified/improved role of enzymes. van Hellemond et al. [63] discovered a new enzyme belonging to the family of styrene monooxygenases from a metagenome analysis. Gene-targeted metagenomics approach of Iwai et al. [64] revealed extensive diversity of aromatic dioxygenase genes. Sipila et al. [65] have studied the phylogenetic diversity of extradiol dioxygenases in both polluted and pristine soil. Yagi and Madsen [66] studied the diversity, abundance, and consistency of dioxygenase gene expression and biodegradation in a shallow contaminated aquifer over a short-term as well as long-term period.
The detection of genes corresponding to enzymes involved in a wide variety of reactions and operating in many unrelated biodegradation pathways corroborates well with the fact that the site of study receives effluents from a variety of industries involved in manufacturing of various chemicals, dyes, solvents, paints, and other xenobiotic compounds. The microbial community analysis at the metagenome level gives an insight into the repertoire of catabolic genes available “in situ” to deal with environmental pollution. Subsequent analysis of the microbial community at the transcriptome, proteome, and metabolome level will endow with the information about which of the genes are active and which are not leading to the accumulation of recalcitrant xenobiotics. In this regard, obtained knowledge will be useful in designing bioremediation strategies to clean up the contaminated environmental sites.
Abbreviations
- MetaSAMS:
-
Sequence Analysis and Management System for Metagenomic Datasets
- EGTs:
-
Environmental gene tags
- LCA:
-
Lowest common ancestor
- GO:
-
Gene Ontology
- COG:
-
Clusters of Orthologous Groups of proteins
- Pfam:
-
Protein family
- KEGG:
-
Kyoto Encyclopedia of Genes and Genomes
References
Moosvi S, Keharia H, Madamwar D (2005) Decolourization of textile dye Reactive Violet 5 by a newly isolated bacterial consortium RVM 11.1. World J Microbiol Biotechnol 21:667–672
Desai C, Pathak H, Madamwar D (2010) Advances in molecular and "-omics" technologies to gauge microbial communities and bioremediation at xenobiotic/anthropogen contaminated sites. Bioresour Technol 101:1558–1569
Shah V, Jain K, Desai C, Madamwar D (2011) Metagenomics and integrative ‘-omics’ technologies in microbial bioremediation: current trends and potential applications. In: Marco D (ed) Metagenomics: current innovations and future trends. Caister Academic Press, Norfolk, pp 211–240
Shah V, Jain K, Desai C, Madamwar D (2012) Molecular analyses of microbial communities involved in bioremediation. In: Satyanarayana T, Johri BN (eds) Microbes in environmental management and biotechnology. Springer, Amsterdam, pp 221–247
Desai C, Parikh RY, Vaishnav T, Shouche YS, Madamwar D (2009) Tracking the influence of long-term chromium pollution on soil bacterial community structures by comparative analyses of 16S rRNA gene phylotypes. Res Microbiol 160:1–9
Schloss PD, Handelsman J (2005) Metagenomics for studying unculturable microorganisms: cutting the Gordian knot. Genome Biol 6:229
Kimura N (2006) Metagenomics: access to unculturable microbes in the environment. Microb Environ 21:201–215
Handelsman J (2004) Metagenomics: application of genomics to uncultured microorganisms. Microbiol Mol Biol Rev 68:669–685
Ward N (2006) New directions and interactions in metagenomics research. FEMS Microbiol Ecol 55:331–338
Handelsman J, Rondon MR, Brady SF, Clardy J, Goodman RM (1998) Molecular biological access to the chemistry of unknown soil microbes: a new frontier for natural products. Chem Biol 5:R245–R249
Schloss PD, Handelsman J (2003) Biotechnological prospects from metagenomics. Curr Opin Biotechnol 14:303–310
Margulies M, Egholm M, Altman WE, Attiya S, Bader JS, Bemben LA, Berka J, Braverman MS, Chen YJ, Chen Z et al (2005) Genome sequencing in microfabricated high-density picolitre reactors. Nature 437:376–380
Ashburner L, Ball CA, Blake JA, Botstein D, Butler H, Cherry JM, Davis AP, Dolinski K, Dwight S, Eppig JT et al (2000) Gene ontology: tool for the unification of biology. The Gene Ontology Consortium. Nat Genet 25:25–29
Tatusov RL, Natale DA, Garkavtsev IV, Tatusova TA, Shankavaram UT, Rao BS, Kiryutin B, Galperin MY, Fedorova ND, Koonin EV (2001) The COG database: new developments in phylogenetic classification of proteins from complete genomes. Nucleic Acids Res 29:22–28
Finn RD, Mistry J, Tate J, Coggill P, Heger A, Pollington JE, Gavin OL, Gunasekaran P, Ceric G, Forslund K, Holm L, Sonnhammer EL, Eddy SR, Bateman A (2010) The Pfam protein families database. Nucleic Acids Res 38:D211–D222
Ogata H, Goto S, Sato K, Fujibuchi W, Bono H, Kanehisa M (1999) KEGG: Kyoto Encyclopedia of Genes and Genomes. Nucleic Acids Res 27:29–34
Zhou J, Bruns MA, Tiedje JM (1996) DNA recovery from soils of diverse composition. Appl Environ Microbiol 62:316–322
Desai C, Madamwar D (2007) Extraction of inhibitor-free metagenomic DNA from polluted sediments, compatible with molecular diversity analysis using adsorption and ion-exchange treatments. Bioresour Technol 98:761–768
Zakrzewski M, Bekel T, Ander C, Pühler A, Rupp O, Stoye J, Schlüter A, Goesmann A (2012) MetaSAMS—a novel software platform for taxonomic classification, functional annotation and comparative analysis of metagenome datasets. J Biotechnol. doi:10.1016/j.jbiotec.2012.09.013
Ludwig W, Strunk O, Westram R, Richter L, Meier H, Yadhukumar BA, Lai T, Steppi S, Jobb G, Förster W et al (2004) ARB: a software environment for sequence data. Nucleic Acids Res 32:1363–1371
Wang Q, Garrity GM, Tiedje JM, Cole JR (2007) Naïve Bayesian classifier for rapid assignment of rRNA sequences into the new bacterial taxonomy. Appl Environ Microbiol 73:5261–5267
Krause L, Daiz NN, Goesmann A, Kelley S, Nattkemper TW, Rohwer F, Edwards RA, Stoye J (2008) Phylogenetic classification of short environmental DNA fragments. Nucleic Acids Res 36:2230–2239
Clemente JC, Jansson J, Valiente G (2010) Accurate taxonomic assignment of short pyrosequencing reads. Pac Symp Biocomput p 3–9
Cole JR, Wang Q, Cardenas E, Fish J, Chai B, Farris RJ, Kulam-Syed-Mohideen AS, McGarrell DM, Marsh T, Garrity GM, Tiedje JM (2009) The Ribosomal Database Project: improved alignments and new tools for rRNA analysis. Nucleic Acids Res 37:D141–D145
Liu Z, DeSantis TZ, Andersen GL, Knight R (2008) Accurate taxonomy assignments from 16S rRNA sequences produced by highly parallel pyrosequencers. Nucleic Acids Res 36:e120
Huson DH, Auch AF, Qi J, Schuster SC (2007) Megan analysis of metagenomic data. Genome Res 17:377–386
Gao J, Ellis LB, Wackett LP (2010) The University of Minnesota Biocatalysis/Biodegradation Database: improving public access. Nucleic Acids Res 38:D488–D491
Miller DN, Bryant JE, Madsen EL, Ghiorse WC (1999) Evaluation and optimization of DNA extraction and purification procedures for soil and sediment samples. Appl Environ Microbiol 65:4715–4724
Yeates C, Gillings MR, Davison AD, Altavilla N, Veal DA (1998) Methods for microbial DNA extraction from soil for PCR amplification. Biol Proced Online 14:40–47
Holben WE, Jansson JK, Chelm BK, Tiedje JM (1988) DNA probe method for the detection of specific microorganisms in the soil bacterial community. Appl Environ Microbiol 54:703–711
Fortin N, Beaumier D, Lee K, Greer CW (2004) Soil washing improves the recovery of total community DNA from polluted and high organic content sediments. J Microbiol Methods 56:181–191
Bell TH, Yergeau E, Martineau C, Juck D, Whyte LG, Greer CW (2011) Identification of nitrogen-incorporating bacteria in petroleum-contaminated arctic soils by using [15N] DNA-based stable isotope probing and pyrosequencing. Appl Environ Microbiol 77:4163–4171
Uhlik O, Wald J, Strejcek M, Musilova L, Ridl J, Hroudova M, Vlcek C, Cardenas E, Mackova M, Macek T (2012) Identification of bacteria utilizing biphenyl, benzoate, and naphthalene in long-term contaminated soil. PLoS One 7:e40653
Adetutu EM, Smith RJ, Weber J, Aleer S, Mitchell JG, Ball AS, Juhasz AL (2013) A polyphasic approach for assessing the suitability of bioremediation for the treatment of hydrocarbon-impacted soil. Sci Total Environ 450-451:51–58
Sheik CS, Mitchell TW, Rizvi FZ, Rehman Y, Faisal M, Hasnain S, McInerney MJ, Krumholz LR (2012) Exposure of soil microbial communities to chromium and arsenic alters their diversity and structure. PLoS One 7:e40059
Yang S, Wen X, Jin H, Wu Q (2012) Pyrosequencing investigation into the bacterial community in permafrost soils along the China-Russia Crude Oil Pipeline (CRCOP). PLoS One 7:e52730
Ding GC, Heuer H, Smalla K (2012) Dynamics of bacterial communities in two unpolluted soils after spiking with phenanthrene: soil type specific and common responders. Front Microbiol 3:290
Sepulveda-Torres LC, Rajendran N, Dybas MJ, Criddle CS (1999) Generation and initial characterization of Pseudomonas stutzeri KC mutants with impaired ability to degrade carbon tetrachloride. Arch Microbiol 171:424–429
Grimberg SJ, Stringfellow WT, Aitken MD (1996) Quantifying the biodegradation of phenanthrene by Pseudomonas stutzeri P16 in the presence of a nonionic surfactant. Appl Environ Microbiol 62:2387–2392
Arenghi FLG, Barbieri P, Bertoni G, de Lorenzo V (2001) New insights into the activation of o-xylene biodegradation in Pseudomonas stutzeri OX1 by pathway substrates. EMBO Rep 5:409–414
Bafana A, Devi SS, Krishnamurthi K, Chakrabarti T (2007) Kinetics of decolourisation and biotransformation of direct black 38 by C. hominis and P. stutzeri. Appl Microbiol Biotechnol 74:1145–1152
Itoh K, Kitade Y, Nakanishi M, Yatome C (2002) Decolorization of methyl red by a mixed culture of Bacillus sp. and Pseudomonas stutzeri. J Environ Sci Health A Tox Hazard Subst Environ Eng 37:415–421
Wu X, Monchy S, Taghavi S, Zhu W, Ramos J, van der Lelie D (2011) Comparative genomics and functional analysis of niche-specific adaptation in Pseudomonas putida. FEMS Microbiol Rev 35:299–323
Nelson KE, Weinel C, Paulsen IT, Dodson RJ, Hilbert H, Martins dos Santos P, Fouts DE, Gill SR, Pop M, Holmes M et al (2002) Complete genome sequence and comparative analysis of metabolically versatile Pseudomonas putida KT2440. Environ Microbiol 4:799–808
Venkateswaran K, Moser DP, Dollhopf ME, Lies DP, Saffarini DA, Mac-Gregor BJ, Ringelberg DB, White DC, Nishijima M, Sano H, Burghardt J, Stackenbrandt E, Nealson KH (1999) Polyphasic taxonomy of the genus Shewanella and description of Shewanella oneidensis sp. Int J Syst Bacteriol 49:705–724
Tiedje JM (2002) Shewanella—the environmentally versatile genome. Nat Biotechnol 20:1093–1094
Roldan MD, Blasco R, Caballero FJ, Castillo F (1998) Degradation of p-nitrophenol by the phototrophic bacterium Rhodobacter capsulatus. Arch Microbiol 169:36–42
Song ZY, Zhou JT, Wang J, Yan B, Du CH (2003) Decolorization of azo dyes by Rhodobacter sphaeroides. Biotechol Lett 25:1815–1818
Perez-Pantoja D, Donoso R, Junca H, Gonzalez B, Pieper DH (2009) Phylogenomics of aerobic bacterial degradation of aromatics. In: Timmis KN (ed) Handbook of hydrocarbon and lipid microbiology. Springer, Berlin, pp 1356–1397
Cavalca L, Dell’Amico E, Andreoni V (2004) Intrinsic bioremediability of an aromatic hydrocarbon-polluted groundwater: diversity of bacterial population and toluene monoxygenase genes. Appl Microbiol Biotechnol 64:576–587
Vilchez-Vargas R, Junca H, Pieper DH (2010) Metabolic networks, microbial ecology and ‘omics’ technologies: towards understanding in situ biodegradation processes. Environ Microbiol 12:3089–3104
Mandrich L, Merone L, Manco G (2010) Hyperthermophilic phosphotriesterases/lactonases for the environment and human health. Environ Technol 31:1115–1127
Kim SJ, Kweon O, Jones RC, Edmondson RD, Cerniglia CE (2008) Genomic analysis of polycyclic aromatic hydrocarbon degradation in Mycobacterium vanbaalenii PYR-1. Biodegradation 19:859–881
Perez-Pantoja D, De la Iglesia R, Pieper DH, Gonzalez B (2008) Metabolic reconstruction of aromatic compounds degradation from the genome of the amazing pollutant-degrading bacterium Cupriavidus necator JMP134. FEMS Microbiol Rev 32:736–794
Pieper DH, Seeger M (2008) Bacterial metabolism of polychlorinated biphenyls. J Mol Microbiol Biotechnol 15:121–138
Suenaga H, Koyama Y, Miyakoshi M, Miyazaki R, Yano H, Sota M, Ohtsubo Y, Tsuda M, Miyazaki K (2009) Novel organization of aromatic degradation pathway genes in a microbial community as revealed by metagenomic analysis. ISME J 3:1335–1348
Denef VJ, Patrauchan MA, Florizone C, Park J, Tsoi TV, Verstraete W, Tiedje JM, Eltis LD (2005) Growth substrate- and phase-specific expression of biphenyl, benzoate, and C1 metabolic pathways in Burkholderia xenovorans LB400. J Bacteriol 187:7996–8005
Denef VJ, Klappenbach JA, Patrauchan MA, Florizone C, Rodrigues JL, Tsoi TV, Verstraete W, Eltis LD, Tiedje JM (2006) Genetic and genomic insights into the role of benzoate-catabolic pathway redundancy in Burkholderia xenovorans LB400. Appl Environ Microbiol 72:585–595
Brennerova MV, Josefiova J, Brenner V, Pieper DH, Junca H (2009) Metagenomics reveals diversity and abundance of meta-cleavage pathways in microbial communities from soil highly contaminated with jet fuel under air-sparging bioremediation. Environ Microbiol 11:2216–2227
Witzig R, Junca H, Hecht HJ, Pieper DH (2006) Assessment of toluene/biphenyl dioxygenase gene diversity in benzene-polluted soils: links between benzene biodegradation and genes similar to those encoding isopropylbenzene dioxygenases. Appl Environ Microbiol 72:3504–3514
Iwai S, Kurisu F, Urakawa H, Yagi O, Kasuga I, Furumai H (2008) Development of an oligonucleotide microarray to detect di- and monooxygenase genes for benzene degradation in soil. FEMS Microbiol Lett 285:111–121
Suenaga H, Mizuta S, Miyazaki K (2009) The molecular basis for adaptive evolution in novel extradiol dioxygenases retrieved from the metagenome. FEMS Microbiol Ecol 69:472–480
van Hellemond EW, Janssen DB, Fraaije MW (2007) Discovery of a novel styrene monooxygenase originating from the metagenome. Appl Environ Microbiol 73:5832–5839
Iwai S, Chai B, Sul WJ, Cole JR, Hashsham SA, Tiedje JM (2010) Gene-targeted-metagenomics reveals extensive diversity of aromatic dioxygenase genes in the environment. ISME J 4:278–285
Sipila TP, Keskinen AK, Akerman ML, Fortelius C, Haahtela K, Yrjala K (2008) High aromatic ring cleavage diversity in birch rhizosphere: PAH treatment specific changes of I.E.3 group extradiol dioxygenases and 16S rRNA bacterial communities in soil. ISME J 2:968–981
Yagi JM, Madsen EL (2009) Diversity, abundance, and consistency of microbial oxygenase expression and biodegradation in a shallow contaminated aquifer. Appl Environ Microbiol 75:6478–6487
Acknowledgments
V.S. and D.M. acknowledge the financial support by the German Academic Exchange Service (DAAD) and Department of Biotechnology, New Delhi. M.Z and F.G.E. acknowledge the receipt of a scholarship from the CLIB Graduate Cluster “Industrial Biotechnology” co-financed by the Ministry of Innovation of North Rhine-Westphalia (Germany). The authors thank Dr. Asha Parmar, BRD School of Biosciences, Sardar Patel University, Gujarat, India for discussions and critical suggestions during work plan and manuscript preparation. The bioinformatics and technological support of the Bioinformatics and Genomics Platforms at the Center for Biotechnology (CeBiTec, Bielefeld University, Germany) is gratefully acknowledged.
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
ESM 1
(DOC 467 kb)
Rights and permissions
About this article
Cite this article
Shah, V., Zakrzewski, M., Wibberg, D. et al. Taxonomic Profiling and Metagenome Analysis of a Microbial Community from a Habitat Contaminated with Industrial Discharges. Microb Ecol 66, 533–550 (2013). https://doi.org/10.1007/s00248-013-0253-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00248-013-0253-9