Abstract
The overexpression of CoA transferase (ctfAB), which catalyzes the reaction: acetate/butyrate + acetoacetyl-CoA → acetyl/butyryl-CoA + acetoacetate, was studied for its effects on acid reassimilation and butanol biosynthesis in Clostridium tyrobutyricum (Δack, adhE2). The plasmid pMTL007 was used to co-express adhE2 and ctfAB from Clostridium acetobutylicum ATCC 824. In addition, the sol operon containing ctfAB, adc (acetoacetate decarboxylase), and ald (aldehyde dehydrogenase) was also cloned from Clostridium beijerinckii NCIMB 8052 and expressed in C. tyrobutyricum (Δack, adhE2). Mutants expressing these genes were evaluated for their ability to produce butanol from glucose in batch fermentations at pH 5.0 and 6.0. Compared to C. tyrobutyricum (Δack, adhE2) without expressing ctfAB, all mutants with ctfAB overexpression produced more butanol, with butanol yield increased to 0.22 − 0.26 g/g (vs. 0.10 − 0.13 g/g) and productivity to 0.35 g/l h (vs. 0.13 g/l h) because of the reduced acetate and butyrate production. The expression of ctfAB also resulted in acetone production from acetoacetate through a non-enzymatic decarboxylation.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
With increasing concerns about environmental pollution and the diminishing oil supplies, increased attentions and efforts have focused on the development of next-generation or advanced biofuels (Demirbas 2009; Green 2011; Jiang et al. 2014). Biobutanol, which shares many similar fuel properties with gasoline and has a 30 % higher energy density than ethanol, is one of the most promising advanced biofuels with good prospect as a gasoline substitute (Dürre 2007; Xue et al. 2013). n-Butanol can be produced from biorenewable feedstocks in acetone-butanol-ethanol (ABE) fermentation, which was once the second largest industrial fermentation that can be traced back more than 100 years ago (Jones and Woods 1986). However, ABE fermentation processes are limited by low butanol yield, productivity, and titer and generally cannot compete with petroleum-based n-butanol that currently dominates in the market (Zhao et al. 2013). Metabolic engineering of solventogenic clostridia, mainly Clostridium acetobutylicum and Clostridium beijerinckii, have thus been intensively studied and used to manipulate the host strains to better understand their physiology and to develop robust strains for industrial application (Branduardi et al. 2014; Jang et al. 2012; Lee et al. 2008; Lütke-Eversloh 2014; Papoutsakis 2008; Wang et al. 2014).
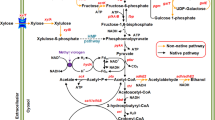
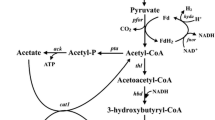
However, the progress to date has been limited because the biphasic nature of the ABE fermentation and the complex metabolic and regulatory pathways involved are difficult to manipulate and control (Zheng et al. 2009; Lehmann et al. 2012b). To overcome these problems, we have focused on the engineering of Clostridium tyrobutyricum, an acidogen which naturally can only produce butyrate and acetate (Liu et al. 2005), but not solvents, because of lacking some key enzymes (genes), including CoA transferase (ctfAB), acetoacetate decarboxylase (adc), and aldehyde dehydrogenase (ald), in the pathways leading to ABE production (see Fig. 1). In our recent study, we overexpressed a bifunctional aldehyde/alcohol dehydrogenase gene (adhE2) from C. acetobutylicum in C. tyrobutyricum and turned the mutant into an n-butanol producer (Yu et al. 2011). Compared to native solventogenic clostridia, the engineered C. tyrobutyricum has a much simpler butanol biosynthesis pathway and potentially can produce more butanol from glucose at a higher yield (>0.3 g/g). However, because large amounts of butyrate and acetate were also produced by this mutant in fermentation, the actual butanol yield was low, only ~0.1 g/g glucose consumed (Yu et al. 2011). Since ctfAB is usually overexpressed in solventogenic clostridia during the solventogenic phase to convert acetate and butyrate to acetyl-CoA and butyryl-CoA, respectively, it is desirable to also express ctfAB, together with adhE2, in C. tyrobutyricum to further increase butanol production.
Metabolic pathway in C. tyrobutyricum. The dotted lines show the pathways with missing genes in C. tyrobutyricum in comparison to other solvent producing Clostridium species. The boxes show the ABE products produced after introducing ald or adhE2 and ctfAB genes. The reversible reactions between butyryl-CoA and butyrate catalyzed by ptb and buk are hypothetical as these two genes have not been identified or annotated in the published draft genomic sequences of C. tyrobutyricum (ack acetate kinase, adc acetoacetate decarboxylase, adh alcohol dehydrogenase, adhE2 aldehyde/alcohol dehydrogenase, ald aldehyde dehydrogenase, bdh butanol dehydrogenase, buk butyrate kinase, ctfAB CoA-transferase, pta phosphotransacetylase, ptb phosphotransbutyrylase)
In the present study, adhE2 and ctfAB were co-expressed in C. tyrobutyricum strain Ct(Δack) with acetate kinase (ack) knockout, which could tolerate and produce butyrate at concentrations higher than 40 g/l (Liu et al. 2008). The ctfAB genes from C. acetobutylicum were expressed together with adhE2 in plasmid pMTL007 (Heap et al. 2007), and the effects of overexpressing ctfAB on fermentation kinetics were studied in stirred-tank bioreactors at pH 6.0 and 5.0. In addition, co-expressing ctfAB, ald, and adc genes obtained from the sol operon of C. beijerinckii with adhE2 in Ct(Δack) was also studied. The results showed that ctfAB expression not only significantly increased butanol and reduced acid production but also induced acetone production even in the absence of adc gene. This study demonstrated the beneficial effects of ctfAB on acid reassimilation and butanol biosynthesis in the non-native solventogenic C. tyrobutyricum with potential industrial application for n-butanol production. This study also provided new insights on the role of ctfAB in controlling acid reassimilation and its effects on solventogenesis.
Materials and methods
Bacterial strains, plasmids, and culture media
Table 1 shows the bacterial strains and recombinant plasmids developed and used in this study. C. tyrobutyricum Ct(Δack), a mutant strain of ATCC 25755 with ack knockout (Liu et al. 2008), was used as the host for all recombinant plasmids constructed in this work. The Clostridium cultures were grown in clostridial growth medium (CGM) with glucose as the carbon source at 37 °C under anaerobic conditions. The CGM contained (g/l): 4 tryptone, 2 yeast extract, 1.0 K2HPO4·3H2O, 0.5 KH2PO4, 2 (NH4)2SO4, 0.1 MgSO4·7H2O, and trace minerals (Zhu and Yang 2003). Escherichia coli strains used in the cloning were cultivated at 37 °C aerobically in liquid Luria-Bertani (LB) medium with agitation at 250 rpm and on LB agar plates. These media were sterilized by autoclaving at 121 °C for 30 min and after cooling supplemented with appropriate antibiotics: 25 μg/ml chloramphenicol, 45 μg/ml thiamphenicol, or 250 μg/ml cycloserine.
Plasmid construction
Plasmid pMTL007 was the basic vector from which the other constructs were derived. The DNA sequences for adhE2 (CA_P0035) and ctfAB (CA_P0163 and CA_P0164) genes were extracted and PCR-amplified from C. acetobutylicum ATCC 824 genomic DNA. The whole sol operon containing ald (Cbei_3832), ctfAB (Cbei_3833 and Cbei_3834), and adc (Cbei_3835) genes in pSOL and the truncated sol operon containing only ald and ctfAB genes in pSV6 were derived from C. beijerinckii NCIMB 8052 (ATCC 51743) genomic DNA. The thiolase (thl) promoter used to drive the constitutive expression of the genes mentioned above was from C. tyrobutyricum ATCC 25755 (Yu et al. 2012). The primers used in PCR amplification of these genes are listed in Table 1. The plasmid pMAD72 for the overexpression of adhE2 under the control of thl promoter has been described in details elsewhere (Yu et al. 2011). The plasmid pMAT was constructed from pMAD72 by inserting ctfAB after adhE2 at the SacII site using the Clontech infusion cloning kit. The plasmid pSOL was constructed by inserting the PCR-amplified sol operon from C. beijerinckii NCIMB 8052 together with thl promoter into pMTL007 between XhoI and SacII sites by infusion. The plasmid pSV6 contained hygromycin B resistance gene and the truncated sol operon (ald and ctfAB genes) and adhE2 under fac and thl promoters, respectively. The artificial promoter fac, which was constructed by combining the operator of the E. coli lacZ operon and the promoter of C. pasteurianum ferredoxin gene (Fox et al. 1996), was derived from the model Clostridium shuttle vector to direct the constitutive expression of heterologous genes in Clostridium (Heap et al. 2007). The hygromycin B resistance gene was cut from pGEMT-hygB vector with NcoI and ligated into pMAD72, and the PCR-amplified ald and ctfAB genes from C. beijerinckii genome were ligated into the plasmid with HindIII to generate plasmid pSV6. Figure 2 shows the schematic maps of these plasmids. These plasmids were amplified and stored in E. coli DH5α and transformed into C. tyrobutyricum Ct(Δack) via the donor cell, E. coli CA434, by conjugation described below.
Plasmid maps of pMAD72, pMAT, pSOL, and pSV6. The gray arrows show the promoters and genes (adc acetoacetate decarboxylase, adhE2 aldehyde/alcohol dehydrogenase, ald aldehyde dehydrogenase, ctfAB CoA-transferase, CatP chloramphenicol resistance gene, ColE1 gram-negative replicon, fac artificial Clostridium promoter, oriT origin of transfer, pCB102 gram-positive replicon, thl thiolase promoter, traJ TraJ protein for conjugation)
Transformation and mutant confirmation
All plasmids were transformed into Ct(Δack) by conjugation as previously described with some modifications (Yu et al. 2011). The plasmids were first transformed into E. coli CA434 using the heat shock method. Then, the transformants were cultivated in LB medium containing 25 μg/ml chloramphenicol at 37 °C overnight to reach optical density at 600 nm (OD600) of 1.5 − 2.0. The collected transformants were washed once using 1 ml sterile phosphate-buffered saline (PBS) and collected by centrifugation at 4000 × g for 2 min. The transformed donor cells were then mixed with 200 μl of C. tyrobutyricum cells precultured at 37 °C overnight, and the mixture was pipetted onto CGM agar plates in an anaerobic chamber and incubated at 37 °C for 8 − 24 h. Then, cells were recovered and resuspended in 1 ml of PBS and spread onto CGM plates containing 45 μg/ml thiamphenicol and 250 μg/ml cycloserine for 2 − 3 days to select for positive transformants, which were confirmed by PCR cloning and plasmid extraction. The transformants carrying the plasmids pMAD72, pMAT, pSOL, and pSV6 are designated as mutant strains Ct(Δack)-pMAD72, Ct(Δack)-pMAT, Ct(Δack)-pSOL, and Ct(Δack)-pSV6, respectively, and were obtained and stored at −80 °C.
Enzyme activity assay
The activity of ctfAB in cells was assayed under anaerobic conditions following the method previously described (Chen and Blaschek 1999). Each crude cell extract from 50 ml of cells present in an overnight culture was prepared in a buffer containing 50 mM 3-(N-morpholino) propanesulfonic acid (MOPS) (pH 7.0), 500 mM (NH4)2SO4, and 20 % (v/v) glycerol. The cell debris was removed by centrifugation at 13,000 rpm for 10 min at 4 °C. The assay mixture (1 ml) containing 110 mM Tris-HCl (pH 7.5), 5.5 % (v/v) glycerol, 20 mM MgCl2, 0.1 mM acetoacetyl-CoA, crude cell extract (20 to 100 μg), and 0.32 M potassium acetate (or butyrate) was purged with nitrogen to eliminate O2. The assay mixture without potassium acetate (or butyrate) was used as blank for negative control. The activity of CoA transferase was measured by monitoring the disappearance of acetoacetyl-CoA at 310 nm in a UV/Vis spectrophotometer (UV-1601, Shimadzu). One unit of enzyme activity is defined as the disappearance of 1 μmol of acetoacetyl-CoA per min. Protein concentration was measured by the Bradford dye-binding assay (Bio-Rad Laboratories, Hercules, CA) with bovine serum albumin as standard. The specific enzyme activity is reported as U/mg protein.
Fermentation kinetics
Batch fermentation kinetics were studied in a stirred-tank bioreactor containing 600 ml of CGM medium with glucose as the carbon source and 45 μg/ml thiamphenicol to prevent culture degeneration or plasmid loss. The bioreactor was sparged with nitrogen for ~30 min to reach anaerobic condition and then inoculated with an overnight culture at a volume ratio of 5 %. Unless otherwise noted, the bioreactor was maintained at 37 °C with the pH controlled at 5.0 or 6.0 by adding 40 % ammonium hydroxide. Samples were collected twice a day at regular intervals for analyses of cell density and concentrations of glucose, acetone, butanol, ethanol, acetate, and butyrate. Each fermentation condition was repeated at least once, and representative data with averages and standard deviations are reported.
Analytical methods
Cell growth was monitored by measuring the OD600 with a spectrophotometer (UV-16-1, Shimadzu, Columbia, MD). YSI 2700 Select Biochemistry Analyzer (Yellow Springs, OH) was used to assay the concentration of glucose in samples. Acetone, butanol, ethanol, acetate, and butyrate were analyzed with a gas chromatograph (GC, Shimadzu GC-2014) equipped with a flame ionization detector and a 30-m fused silica column (0.25-μm film thickness and 0.25-mm ID, Stabilwax-DA). The carrier gas was nitrogen at 1.47 ml/min (linear velocity: 35 cm/s). Samples were diluted 20 times with an internal standard buffer solution containing 0.5 g/l isobutanol, 0.1 g/l isobutyric acid, and 1 % phosphoric acid (for acidification) and injected (1 μl each) using an auto injector (AOC-20i Shimadzu). The column temperature was held at 80 °C for 3 min, raised to 150 °C at a rate of 30 °C/min, and held at 150 °C for 3.7 min. Both the injector and detector were set at 250 °C.
Statistical analysis
All batch fermentations were at least duplicated for each condition studied, and the means with standard errors for kinetic parameters such as product yields and productivities are reported. Student’s t test analysis with JMP software was performed to determine the significant difference (p < 0.05).
Results
Enzyme activity
To confirm the expression of ctfAB in the mutants, the CoA transferase activity was assayed with the parental strain Ct(Δack) as the negative control and C. acetobutylicum and C. beijerinckii as positive controls, and the results are shown in Table 2. As expected, the strains Ct(Δack) and Ct(Δack)-pMAD72 showed no or negligible CoA transferase activity while Ct(Δack)-pMAT, Ct(Δack)-pSOL, and Ct(Δack)-pSV6 all showed a high specific CoA transferase activity (0.11 to 0.26 U/mg protein) comparable to the positive controls, confirming the expression of ctfAB genes in these mutants. With CoA-transferase, these mutants can catalyze the transfer of CoA moiety from acetoacetyl-CoA to either butyrate or acetate, thus allowing the conversion of butyrate and acetate to butanol and ethanol, respectively.
Fermentation kinetics
Figure 3 shows the fermentation kinetics for Ct(Δack)-pMAD72, Ct(Δack)-pMAT, Ct(Δack)-pSOL, and Ct(Δack)-pSV6 at pH 6.0. All these mutants were able to produce butanol and ethanol because of the overexpression of adhE2 or ald gene. The former encodes a bifunctional aldehyde/alcohol dehydrogenase, which catalyzes the reaction from butyryl-CoA to butanol and acetyl-CoA to ethanol. It is noted that the genome of C. tyrobutyricum contains adh (alcohol dehydrogenase) and bdh (butanol dehydrogenase) genes (unpublished data). Therefore, overexpressing ald (aldehyde dehydrogenase) alone would be sufficient for C. tyrobutyricum to produce butanol and ethanol, as evidenced in the case with the mutant Ct(Δack)-pSOL (Fig. 3c).
For the mutant Ct(Δack)-pMAD72 overexpressing only adhE2, butanol production reached ~10 g/l, with large amounts of butyrate (~13.7 g/l) and acetate (~6.7 g/l) also produced (Fig. 3a). In contrast, for the mutants also expressing ctfAB, more butanol (12.3 to 13.4 g/l) and much less acids (3.1–4.5 g/l butyric acid, 1.6–2.6 g/l acetic acid) were produced (Fig. 3b–d). Clearly, with ctfAB genes, which are responsible for transferring the CoA moiety from acetoacetyl-CoA to butyrate and acetate, butyrate and acetate produced by the cells can be reassimilated back into butyryl-CoA and acetyl-CoA and reenter the main metabolic pathway (Wiesenborn et al. 1989a), leading to the production of butanol and ethanol, respectively. Therefore, much less acid accumulation and more butanol production were observed with Ct(Δack)-pMAT, Ct(Δack)-pSOL, and Ct(Δack)-pSV6. Compared to Ct(Δack)-pMAD72, the production of acetate and butyrate decreased 61–76 % and 67–77 %, respectively, while butanol production increased 20.6–31.4 %. While acetate production was significantly reduced, ethanol production did not increase but instead decreased in mutants overexpressing ctfAB. This could be due to that adhE2 overexpression shifted the metabolic flux from C2 (acetate) toward C4 (butyrate) biosynthesis (Yu et al. 2011), which is further discussed later in this paper.
It should be noted that Ct(ack)-pMAD72 consumed ~100 g/l of glucose during the fermentation, while only 60–70 g/l of glucose was consumed by the mutants with CoA transferase expression. The earlier and accelerated butanol production by these mutants led to an early threshold of butanol toxicity (Bowles and Ellefson 1985), which inhibited cell metabolism and resulted in incomplete glucose consumption. Nevertheless, the butanol titer produced by the mutants was higher than that by Ct(ack)-pMAD72 even though less glucose was consumed because of increased butanol yield.
Interestingly, the mutants overexpressing ctfAB also produced a significant amount of acetone (6.5–7.4 g/l) even in the absence of adc. In solventogenic clostridia, acetone is produced from acetoacetyl-CoA in two steps catalyzed by ctfAB and adc, respectively (Petersen and Bennett 1990). As expected, Ct(Δack)-pSOL overexpressing ctfAB and adc was able to produce 7.4 g/l acetone. However, Ct(Δack)-pMAT and Ct(Δack)-pSV6, which did not have the adc gene, also showed a comparable acetone production (6.5–7.0 g/l), suggesting a non-enzymatic decarboxylation of acetoacetate. This finding is consistent with a previous study by Han et al. (2011), who knocked out the adc gene in C. beijerinckii NCIMB 8052 and observed no obvious decrease in acetone production by the knockout mutant. Similarly, the downregulation of adc with antisense RNA resulted in 86 % decrease in the decarboxylase activity but only a 17 % reduction in acetone production (Tummala et al. 2003). Clearly, adc is not required for acetone production in C. tyrobutyricum although its presence appeared to give a slightly higher acetone production compared to the strains without the gene.
The fermentation kinetics for Ct(Δack)-pMAD72, Ct(Δack)-pMAT, Ct(Δack)-pSOL, and Ct(Δack)-pSV6 were also studied at pH 5.0 (see Figure S1 in Supplementary Materials). In general, similar kinetics was observed at both pH 5.0 and 6.0, although butanol and acetone production was lower at pH 5.0. Figures 4 and 5 illustrate the effects of ctfAB overexpression and pH on C. tyrobutyricum growth and fermentation kinetics, including specific growth rate, product titers, and butanol yield and productivity (also see Table S1 in Supplementary Materials). Compared to Ct(Δack)-pMAD72, the mutants overexpressing ctfAB had a much higher butanol yield (0.19–0.22 vs. 0.10 g/g glucose at pH 6.0, 0.18–0.26 vs. 0.14 g/g glucose at pH 5.0) and productivity (0.31–0.35 vs. 0.13 g/l h at pH 6.0, 0.23–0.24 vs. 0.13 g/l h at pH 5.0). These mutants had a comparable specific growth rate but a much lower final cell density compared to Ct(Δack)-pMAD72. The effects of ctfAB overexpression and pH on C. tyrobutyricum growth and fermentation kinetics are further discussed below.
Effects of ctfAB
CoA transferase encoded by ctfAB plays an important role in the reassimilation and conversion of acetate and butyrate, produced in the acidogenesis phase, to solvents (acetone, butanol, and ethanol) in the solventogenic phase in solventogenic Clostridium. The overexpression of ctfAB in C. tyrobutyricum Ct(Δack) thus had pronounced effects on cell growth, acid production, and butanol production. Without ctfAB, the strain Ct(Δack)-pMAD72 produced much more butyrate (Fig. 4a, b) and acetate (Fig. 4c, d), and less butanol (Fig. 4e, f), compared to the mutants overexpressing CoA-transferase. Ct(Δack)-pMAD72 also grew slower initially with a longer lag phase of ~30 h but reached a higher final cell density (Fig. 4g, h). On the other hand, the overexpression of CoA transferase caused an earlier production of butanol, which was toxic to cells and thus resulted in lower cell density and earlier termination of the fermentation with less glucose consumption and total metabolites (solvents and acids) produced (Fig. 5a, b). Nevertheless, overexpressing ctfAB resulted in over 100 % increase in butanol yield (Fig. 5c, d) and productivity (Fig. 5e) but negligible effect on the specific growth rate (Fig. 5f).
Clearly, ctfAB expression improved butanol production by reassimilating and converting butyrate and acetate to their corresponding alcohols, resulting in 21–31 % higher butanol titer and over 100 % increase in butanol yield (from 0.10 to 0.22 g/g glucose) and productivity (from 0.13 to 0.35 g/l h) at pH 6.0. The mutants with CoA transferase expression also had a much shorter lag phase, although a similar specific growth rate, indicating that CoA transferase expression allowed cells to grow sooner by limiting the accumulation of butyrate, which is an inhibitor to cell growth (Zhu and Yang 2003). However, the accelerated production of butanol by these mutants led to an early threshold of butanol toxicity (Bowles and Ellefson 1985), lower cell density reached in the stationary phase, and earlier termination of the fermentation with less glucose consumption.
Although ctfAB had pronounced effects on decreasing butyrate and acetate production and increasing butanol production, it showed negligible effect on ethanol production (Fig. 5a, b). This can be attributed to the fact that C. tyrobutyricum, as a native high butyrate-tolerant and producing strain, has a high metabolic flux from acetyl-CoA to butyryl-CoA, which favors butanol production over ethanol production. Therefore, the expression of CoA transferase in C. tyrobutyricum increased its butanol production but had little effect on ethanol production.
It is noted that the reduction in butyrate production was much more than the reduction in acetate production in the presence of ctfAB. For example, on average, butyrate production decreased 73 and 85 %, while acetate production decreased 67 and 59 % at pH 6.0 and pH 5.0, respectively (see Table S1 in Supplementary Materials). Apparently, more butyrate has been converted by CoA transferase than acetate, although the in vitro enzyme activity assay showed a similar rate for CoA transfer to butyrate or acetate (see Table 2). This finding is consistent with a previous study showing that in vivo CoA transferase had a much higher activity toward butyrate than acetate (Wiesenborn et al. 1989a).
Effects of pH
In general, more butanol was produced at a higher rate at pH 6.0 than at pH 5.0 because the optimal pH for aldehyde/alcohol dehydrogenase (adhE2) is around 6.5 (Fontaine et al. 2002). In addition, at pH 5.0, most acids would be present in the undissociated form, which is toxic to cells (Maddox et al. 2000). On the other hand, the CoA transferase did not seem to be much affected by the pH between 5.0 and 6.0, as their effects on decreasing acid production and increasing butanol yield and productivity were similar at both pHs. The lack of effect on increasing butanol titer at pH 5.0 by the mutants Ct(Δack)-pMAT and Ct(Δack)-pSOL as compared to Ct(Δack)-pMAD72 could be attributed to the low aldehyde/alcohol dehydrogenase activities at the acidic pH. This problem was alleviated by co-expressing ald and adhE2 in Ct(Δack)-pSV6, which produced significantly more butanol (11 vs. <9 g/l) compared to the other mutants.
For all mutants expressing ctfAB, more acetone was also produced at pH 6.0 than at pH 5.0 (see Table S1 in Supplementary Materials), because of higher cell activity at pH 6.0. Moreover, the butanol/acetone ratio was lower at pH 6.0 (1.7–1.9 g/g) than at pH 5.0 (2.1–2.6 g/g), suggesting that pH 6.0 was more favorable for acetone production, probably because CoA transferase activity was higher at pH 6.0 than at 5.0 (Wiesenborn et al. 1989a).
Effects of different ctfAB and ald genes
No significant difference in the fermentation kinetics were found for Ct(Δack)-pMAT and Ct(Δack)-pSOL. The former expressed adhE2 and ctfAB from C. acetobutylicum ATCC 824, while the latter expressed ald, ctfAB, and adc from C. beijerinckii. Different from C. acetobutylicum, C. beijerinckii does not bear any mega-plasmid and ald, ctfAB, and adc are located on its chromosome. Also, the ald gene in C. beijerinckii, unlike adhE gene from C. acetobutylicum, only has aldehyde dehydrogenase activity. Nevertheless, butanol and acid production levels in both mutant strains were similar, suggesting that the native bdh and adh genes in C. tyrobutyricum genome are functional. In addition, both strains produced acetone at a similar level, indicating that the adc gene encoding an acetoacetate decarboxylase is not required for acetone production from acetoacetate, as also found for C. beijerinckii by Han et al. (2011). Overexpressing both adhE2 and ald in Ct(Δack)-pSV6 gave the best butanol production among the mutants studied, probably because of the increased aldehyde dehydrogenase activity. The effect was more pronounced at pH 5.0, at which the activities of aldehyde/alcohol dehydrogenase might be limited because the optimal pH for the enzyme activity is around neutral (Fontaine et al. 2002).
Acid reassimilation
To further illustrate the effects of CoA-transferase on acid reassimilation, batch fermentations of Ct(Δack)-pMAD72 and Ct(Δack)-pMAT were studied at pH 6.0 in media initially also containing ~20 mM acetate or butyrate. As expected, for the strain Ct(Δack)-pMAT expressing ctfAB, both acetate and butyrate were kept at a relatively low level (less than 3–5 g/l) compared to the control strain Ct(Δack)-pMAD72 without ctfAB, which produced large amounts of acetate and butyrate (Fig. 6). In fact, a notable decrease in the acetate level after peaking at ~24 h was observed for Ct(Δack)-pMAT (Fig. 6a, b) but not for Ct(Δack)-pMAD72 (Fig. 6c, d). These results clearly demonstrated that ctfAB played an important role in acid reassimilation. It is noted that without ctfAB, there was a long lag phase of ~24 h, especially when ~20 mM butyrate was added in the medium. Interestingly, the butyrate concentration decreased from 2.4 to 0.4 g/l during the lag phase (Fig. 6d). The apparent butyrate uptake by Ct(Δack)-pMAD72 suggested the existence of a reverse reaction from butyrate to butyryl-CoA, possibly catalyzed by phosphotransbutyrylase (Ptb) and butyrate kinase (Buk), which has also been proposed for C. acetobutylicum (Jiang et al. 2009; Lehmann et al. 2012a; Jang et al. 2012; Millat et al. 2014). Nevertheless, this butyrate uptake pathway seemed to work only in the lag phase, not during the exponential growth phase, and required energy (ATP). Once the butyrate level was reduced to a non-inhibiting level, normal cell growth started and butyrate (and acetate) was produced, which generated more ATP to support fast cell growth. No acetate uptake by Ct(Δack)-pMAD72 was observed (Fig. 6c), again confirming that acetate reassimilation required the CoA transferase (ctfAB).
Discussion
The sol operon containing adhE or ald and ctfA and ctfB (encoding two protein subunits for the CoA-transferase) is responsible for the production of ABE in solventogenic clostridia (Cornillot et al. 1997; Nair et al. 1999; Nair and Papoutsakis 1994). The ability to reassimilate acetate and butyrate is critical to the biphasic ABE fermentation. Failure to do so by solventogenic clostridia can cause acid crash, a phenomenon often observed in industrial ABE fermentation (Wang et al. 2011; Maddox et al. 2000). As evidenced in this study and many other studies, the CoA transferase encoded by ctfAB is responsible for transferring CoA from acetoacetyl-CoA to butyrate and acetate, forming acetoacetate, butyryl-CoA, and acetyl-CoA, which are then converted to ABE in the reactions catalyzed by the enzymes encoded by adc and adhE, respectively (Lee et al. 2008). Also, acetone production is usually coupled with the reassimilation of acids, as mutants with disrupted acetone-producing pathway also showed a significantly increased acid production (Sillers et al. 2008; Lee et al. 2008, 2009; Jang et al. 2012). It is thus generally believed that the reassimilation of acids in C. acetobutylicum is controlled by the expression of ctfAB during the metabolic shift from acidogenesis to solventogenesis (Lehmann et al. 2012b).
However, studies with ctfAB-disrupted mutants also suggested the existence of a CoA-transferase-independent butyrate uptake pathway involving Ptb and Buk, which normally catalyze the reactions from butyryl-CoA to butyryl phosphate and then to butyrate, respectively (Jiang et al. 2009; Lehmann et al. 2012a; Jang et al. 2012). Butyrate uptake through the reverse Ptb-Buk pathway was demonstrated with a mutant of C. acetobutylicum overexpressing ptb and buk (Walter et al. 1994), as well as by using a mathematical model simulating the metabolic pathways in ABE-producing network (Millat et al. 2014). In addition, purified Ptb from C. acetobutylicum ATCC 824 also showed an increased catalytic activity for the reverse reaction of butyryl phosphate to butyryl-CoA as the pH decreased below 6.0 (Wiesenborn et al. 1989b). For the first time, our study also showed the possible existence of the reverse Ptb-Buk pathway for butyrate uptake by a native butyrate-producing C. tyrobutyricum, although ptb and buk genes have not been found or annotated in the recently published draft genome of C. tyrobutyricum ATCC 25755 (Bassi et al. 2013; Jiang et al. 2013), probably because of the incomplete annotation (only ~50 %). The existence of Ptb and Buk in C. tyrobutyricum was partially proved by testing their enzyme activities in a previous study (Zhang et al. 2012); however, further verification would be necessary.
All previous studies on ctfAB and acid reassimilation were conducted with type strains of solventogenic clostridia, such as C. acetobutylicum ATCC 824 and C. beijerinckii NCIMB 8052, which have complex biphasic physiology involving highly regulated metabolic and transcriptional networks (Alsaker et al. 2010; Dürre et al. 2002; Girbal et al. 1995; Janssen et al. 2012; Rydzak et al. 2011; Thormann et al. 2002; Schwarz et al. 2012; Wang et al. 2013). Several genes located on two operons (sol and adc) are involved in the biphasic ABE fermentation (Fischer et al. 1993; Gerischer and Dürre 1990; Petersen and Bennett 1990), and they are tightly regulated by several transcription factors, including spo0A and solR (Alsaker et al. 2004; Thormann et al. 2002; Nair et al. 1999; Ravagnani et al. 2000; Steiner et al. 2011; Tomas et al. 2004). However, the regulatory mechanism is highly complicated, involving many additional genes and transcription factors controlling not only acidogenesis and solventogenesis but also sporulation and clostridia life cycle and remains unclear (Nicolaou et al. 2010; Xu et al. 2015). In contrast, C. tyrobutyricum, as a native ctfAB and adc deficient strain without the complex biphasic physiology, provides a novel (simpler) system to study acid reassimilation by ctfAB and its effects on cell growth and solvent production. This cannot be easily done with C. acetobutylicum as its ctfAB disruption would also influence the expression of adhE located within the same cistronic operon, compromising alcohol production by the mutant (Tummala et al. 2003; Sillers et al. 2009).
The metabolically engineered C. tyrobutyricum can also be used as a novel host for n-butanol production with several advantages over conventional solventogenic clostridia (Ma et al. 2015). Its high tolerance to butyrate, as well as butanol, and strong carbon flux toward C4 products would favor the production of n-butanol, instead of ethanol, when adhE2 is overexpressed (Yu et al. 2011). Butanol is the desirable product as it has superior biofuel properties compared to ethanol. While overexpressing adhE in C. acetobutylicum ATCC 824 increased both ethanol and butanol production, increasing the flux from acetyl-CoA to acetoacetyl-CoA by also overexpressing thl (encoding thiolase) decreased C2 metabolites (acetate and ethanol) and increased acetone and butyrate production (Sillers et al. 2009). Clearly, an enhanced intracellular butyryl-CoA pool could improve butanol production and selectivity. On the other hand, a butyrate-negative mutant strain of C. acetobutylicum ATCC 824 showed elevated ethanol titer with depressed butanol production (Lehmann et al. 2012b). Constitutively expressing adhE2 in C. tyrobutyricum also enables the mutant strain to continuously produce n-butanol throughout the fermentation without subjecting to life cycle regulation and acid crash as often encountered by C. acetobutylicum. Further expression of CoA transferase in C. tyrobutyricum not only increased its butanol production, with more than 100 % increase in butanol yield and productivity, but also facilitated the production of acetone. A previous study also showed that overexpressing adc and ctfAB in C. acetobutylicum led to earlier induction of acetone formation, with enhanced acetone (95 %), butanol (37 %), and ethanol (90 %) production (Mermelstein et al. 1993).
Although overexpressing ctfAB increased butanol yield and productivity by more than 100 %, the final butanol titer in the fermentation only increased 20 to 30 %. This is because butanol production is also limited by butanol toxicity and the availability of NADH (see Fig. 1). Butanol toxicity can be alleviated by removing butanol in situ during fermentation (Xue et al. 2012, 2014), increasing butanol tolerance via adaptation (Yang and Zhao 2013) and metabolic engineering (Lütke-Eversloh and Bahl 2011; Tomas et al. 2003), whereas NADH availability can be increased by inhibiting hydrogen production (Datta and Zeikus 1985), redox engineering (Ventura et al. 2013; Wang et al. 2012), and using artificial electron carriers such as methyl viologen (Du et al. 2015) and more reduced substrates such as mannitol (Yu et al. 2012). With further metabolic and process engineering, it is possible to produce butanol at a higher titer of ~20 g/l using C. tyrobutyricum Ct(Δack) overexpressing ctfAB and adhE2.
In conclusion, overexpressing ctfAB facilitated the reassimilation of butyrate and significantly increased butanol production from glucose by C. tyrobutyricum Ct(Δack) overexpressing adhE2, resulting in over 100 % increase in butanol yield and productivity. Co-expressing ctfAB with adhE2 also led to the production of acetone to a high level of ~50 % of that for butanol, turning the native acidogenic C. tyrobutyricum into an ABE producer with high yields. Further improvement in butanol production can be achieved by engineering the cells for higher butanol tolerance and increasing the NADH level available for butanol biosynthesis during the fermentation. This study demonstrated the essential role of CoA-transferase in acetate and butyrate reassimilation and also suggested possible existence of an exclusive Pta-Buk reverse pathway for butyrate uptake by C. tyrobutyricum.
References
Alsaker KV, Spitzer TR, Papoutsakis ET (2004) Transcriptional analysis of spo0A overexpression in Clostridium acetobutylicum and its effect on the cell's response to butanol stress. J Bacteriol 186:1959–1971
Alsaker KV, Paredes C, Papoutsakis ET (2010) Metabolite stress and tolerance in the production of biofuels and chemicals: gene-expression-based systems analysis of butanol, butyrate, and acetate stresses in the anaerobe Clostridium acetobutylicum. Biotechnol Bioeng 105:1131–1147
Bassi D, Fontana C, Gazzola S, Pietta E, Puglisi E, Cappa F, Cocconcelli PS (2013) Draft genome sequence of Clostridium tyrobutyricum strain UC7086, isolated from grana padano cheese with late-blowing defect. Genome Announc 1:e00614–13
Bowles LK, Ellefson WL (1985) Effects of butanol on Clostridium acetobutylicum. Appl Environ Microbiol 50:1165–1170
Branduardi P, De Ferra F, Longo V, Porro D (2014) Microbial n-butanol production from clostridia to non-clostridial hosts. Eng Life Sci 14:16–26
Chen C-K, Blaschek HP (1999) Effect of acetate on molecular and physiological aspects of Clostridium beijerinckii NCIMB 8052 solvent production and strain degeneration. Appl Environ Microbiol 65:499–505
Cornillot E, Nair RV, Papoutsakis ET, Soucaille P (1997) The genes for butanol and acetone formation in Clostridium acetobutylicum ATCC 824 reside on a large plasmid whose loss leads to degeneration of the strain. J Bacteriol 179:5442–5447
Datta R, Zeikus JG (1985) Modulation of acetone butanol ethanol fermentation: utilization of carbon monoxide to inhibit hydrogen production and to enhance butanol yields. Appl Environ Microbiol 48:764–770
Demirbas A (2009) Political, economic and environmental impacts of biofuels: a review. Appl Energy 86:s108–s117
Du Y, Jiang W, Yu M, Tang IC, Yang ST (2015) Metabolic process engineering of Clostridium tyrobutyricum Δack-adhE2 for enhanced n-butanol production from glucose: effects of methyl viologen on NADH availability, flux distribution and fermentation kinetics. Biotechnol Bioeng 112:705–715
Dürre P (2007) Biobutanol: an attractive biofuel. Biotechnol J 2:1525–1534
Dürre P, Böhringer M, Nakotte S, Schaffer S, Thormann K, Zickner B (2002) Transcriptional regulation of solventogenesis in Clostridium acetobutylicum. J Mol Microbiol Biotechnol 4:295–300
Fischer RJ, Helms J, Dürre P (1993) Cloning, sequencing, and molecular analysis of the sol operon of Clostridium acetobutylicum, a chromosomal locus involved in solventogenesis. J Bacteriol 175:6959–6969
Fontaine L, Meynial-Salles I, Girbal L, Yang X, Croux C, Soucaille P (2002) Molecular characterization and transcriptional analysis of adhE2, the gene encoding the NADH-dependent aldehyde/alcohol dehydrogenase responsible for butanol production in alcohologenic cultures of Clostridium acetobutylicum ATCC 824. J Bacteriol 184:821–830
Fox ME, Lemmon MJ, Mauchline ML, Davis TO, Giaccia AJ, Minton NP, Brown JM (1996) Anaerobic bacteria as a delivery system for cancer gene therapy: in vitro activation of 5-fluorocytosine by genetically engineered clostridia. Gene Ther 3:173–178
Gerischer U, Dürre P (1990) Cloning, sequencing, and molecular analysis of the acetoacetate decarboxylase gene region from Clostridium acetobutylicum. J Bacteriol 172:6907–6918
Girbal L, Croux C, Vasconcelos I, Soucaille P (1995) Regulation of metabolic shifts in Clostridium acetobutylicum ATCC 824. FEMS Microbiol Rev 17:287–298
Green EM (2011) Fermentative production of butanol–the industrial perspective. Curr Opin Biotechnol 22:337–343
Han B, Gopalan V, Ezeji C (2011) Acetone production in solventogenic Clostridium species. Appl Microbiol Biotechnol 91:565–576
Heap JT, Pennington OJ, Cartman ST, Carter GP, Minton NP (2007) The Clostron: a universal gene knock-out system for the genus Clostridium. J Microbiol Methods 70:452–464
Jang YS, Lee JY, Lee J, Park JH, Im JA, Eom MH, Lee J, Lee SH, Song H, Cho JH, Seung DY, Lee SY (2012) Enhanced butanol production obtained by reinforcing the direct butanol-forming route in Clostridium acetobutylicum. MBio 3:e00314–12
Janssen H, Grimmler C, Ehrenreich A, Bahl H, Fischer RJ (2012) A transcriptional study of acidogenic chemostat cells of Clostridium acetobutylicum—solvent stress caused by a transient n-butanol pulse. J Biotechnol 161:354–365
Jiang Y, Xu C, Dong F, Yang Y, Jiang W, Yang S (2009) Disruption of the acetoacetate decarboxylase gene in solvent-producing Clostridium acetobutylicum increases the butanol ratio. Metab Eng 11:284–291
Jiang L, Zhu L, Xu X, Li Y, Li S, Huang H (2013) Genome sequence of Clostridium tyrobutyricum ATCC 25755, a Butyric acid-overproducing strain. Genome Announc. 30:1(3)
Jiang W, Zhao J, Wang Z, Yang ST (2014) Stable high-titer n-butanol production from sucrose and sugarcane juice by Clostridium acetobutylicum JB200 in repeated batch fermentations. Bioresour Technol 163:172–179
Jones DT, Woods DR (1986) Acetone-butanol fermentation revisited. Microbiol Rev 50:484–524
Lee SY, Park JH, Jang SH, Nielsen LK, Kim J, Jung KS (2008) Fermentative butanol production by Clostridia. Biotechnol Bioeng 101:209–228
Lee JY, Jang YS, Lee J, Papoutsakis ET, Lee SY (2009) Metabolic engineering of Clostridium acetobutylicum M5 for highly selective butanol production. Biotechnol J 4:1432–1440
Lehmann D, Hönicke D, Ehrenreich A, Schmidt M, Weuster-Botz D, Bahl H, Lütke-Eversloh T (2012a) Modifying the product pattern of Clostridium acetobutylicum: physiological effects of disrupting the acetate and acetone formation pathways. Appl Microbiol Biotechnol 94:743–754
Lehmann D, Radomski N, Lütke-Eversloh T (2012b) New insights into the butyric acid metabolism of Clostridium acetobutylicum. Appl Microbiol Biotechnol 96:1325–1339
Liu X, Zhu Y, Yang ST (2005) Butyric acid and hydrogen production by Clostridium tyrobutyricum ATCC 25755 and mutants. Enzym Microb Technol 38:521–528
Liu X, Zhu Y, Yang ST (2008) Construction and characterization of ack deleted mutant of Clostridium tyrobutyricum for enhanced butyric acid and hydrogen production. Biotechnol Prog 22:1265–1275
Lütke-Eversloh T (2014) Application of new metabolic engineering tools for Clostridium acetobutylicum. Appl Microbiol Biotechnol 98:5823–5837
Lütke-Eversloh T, Bahl H (2011) Metabolic engineering of Clostridium acetobutylicum: recent advances to improve butanol production. Curr Opin Biotechnol 22:634–647
Ma C, Kojima K, Xu N, Mobley J, Zhou L, Yang ST, Liu X (2015) Comparative proteomics analysis of high n-butanol producing metabolically engineered Clostridium tyrobutyricum. J Biotechnol 193:108–119
Maddox IS, Steiner E, Hirsch S, Wessner S, Gutierrez NA, Gapes JR, Schuster KC (2000) The cause of “acid crash” and “acidogenic fermentations” during the batch acetone-butanol-ethanol (ABE-) fermentation process. J Mol Microbiol Biotechnol 2:95–100
Mermelstein LD, Papoutsakis ET, Petersen DJ, Bennett GN (1993) Metabolic engineering of Clostridium acetobutylicum ATCC 824 for increased solvent production by enhancement of acetone formation enzyme activities using a synthetic acetone operon. Biotechnol Bioeng 42:1053–1060
Millat T, Voig C, Janssen H, Cooksley CM, Winzer K, Minton NP, Bah H, Fischer RJ, Wolkenhauer O (2014) Coenzyme A-transferase-independent butyrate re-assimilation in Clostridium acetobutylicum-evidence from a mathematical model. Appl Microbiol Biotechnol 98:9059–9072
Nair RV, Papoutsakis ET (1994) Expression of plasmid-encoded aad in Clostridium acetobutylicum M5 restores vigorous butanol production. J Bacteriol 176:5843–5846
Nair RV, Green EM, Watson DE, Bennett GN, Papoutsakis ET (1999) Regulation of the sol locus genes for butanol and acetone formation in Clostridium acetobutylicum ATCC 824 by a putative transcriptional repressor. J Bacteriol 181:319–330
Nicolaou SA, Gaida SM, Papoutsakis ET (2010) A comparative view of metabolite and substrate stress and tolerance in microbial bioprocessing: from biofuels and chemicals, to biocatalysis and bioremediation. Metab Eng 12:307–331
Papoutsakis ET (2008) Engineering solventogenic clostridia. Curr Opin Biotechnol 19:420–429
Petersen DJ, Bennett GN (1990) Purification of acetoacetate decarboxylase from Clostridium acetobutylicum ATCC 824 and cloning of the acetoacetate decarboxylase gene in Escherichia coli. Appl Environ Microbiol 56:3491–3498
Ravagnani A, Jennert KC, Steiner E, Grünberg R, Jefferies JR, Wilkinson SR, Young DI, Tidswell EC, Brown DP, Youngman P, Morris JG, Young M (2000) Spo0A directly controls the switch from acid to solvent production in solvent-forming clostridia. Mol Microbiol 37:1172–1185
Rydzak T, Levin DB, Cicek N, Sparling R (2011) End-product induced metabolic shifts in Clostridium thermocellum ATCC 27405. Appl Microbiol Biotechnol 92:199–209
Schwarz KM, Kuit W, Grimmler C, Ehrenreich A, Kengen SWM (2012) A transcriptional study of acidogenic chemostat cells of Clostridium acetobutylicum—cellular behavior in adaptation to n-butanol. J Biotechnol 161:366–377
Sillers R, Chow A, Tracy B, Papoutsakis ET (2008) Metabolic engineering of the non-sporulating, non-solventogenic Clostridium acetobutylicum strain M5 to produce butanol without acetone demonstrate the robustness of the acid-formation pathways and the importance of the electron balance. Metab Eng 10:321–332
Sillers R, Al-Hinai MA, Papoutsakis ET (2009) Aldehyde-alcohol dehydrogenase and/or thiolase overexpression coupled with CoA transferase downregulation lead to higher alcohol titers and selectivity in Clostridium acetobutylicum fermentations. Biotechnol Bioeng 102:38–49
Steiner E, Dago AE, Young DI, Heap JT, Minton NP, Hoch JA, Young M (2011) Multiple orphan histidine kinases interact directly with Spo0A to control the initiation of endospore formation in Clostridium acetobutylicum. Mol Microbiol 80:641–654
Thormann K, Feustel L, Lorenz K, Nakotte S, Dürre P (2002) Control of butanol formation in Clostridium acetobutylicum by transcriptional activation. J Bacteriol 184:1966–1973
Tomas CA, Welker NE, Papoutsakis ET (2003) Overexpression of groESL in Clostridium acetobutylicum results in increased solvent production and tolerance, prolonged metabolism, and changes in the cell's transcriptional program. Appl Environ Microbiol 69:4951–4965
Tomas CA, Beamish J, Papoutsakis ET (2004) Transcriptional analysis of butanol stress and tolerance in Clostridium acetobutylicum. J Bacteriol 186:2006–2018
Tummala SB, Junne SG, Papoutsakis ET (2003) Antisense RNA downregulation of coenzyme A transferase combined with alcohol-aldehyde dehydrogenase overexpression leads to predominantly alcohologenic Clostridium acetobutylicum fermentations. J Bacteriol 185:3644–3653
Ventura S, Hu H, Jahng D (2013) Enhanced butanol production in Clostridium acetobutylicum ATCC 824 by double overexpression of 6-phosphofructokinase and pyruvate kinase genes. Appl Microbiol Biotechnol 97:7505–7516
Walter KA, Mermelstein LD, Papoutsakis ET (1994) Studies of recombinant Clostridium acetobutylicum with increased dosages of butyrate formation genes. Ann N Y Acad Sci 721:69–72
Wang S, Zhang Y, Dong H, Mao S, Zhu Y, Wang R, Luan G, Li Y (2011) Formic acid triggers the "Acid Crash" of acetone-butanol-ethanol fermentation by Clostridium acetobutylicum. Appl Environ Microbiol 77:1674–1680
Wang S, Zhu Y, Zhang Y, Li Y (2012) Controlling the oxidoreduction potential of the culture of Clostridium acetobutylicum leads to an earlier initiation of solventogenesis, thus increasing solvent productivity. Appl Microbiol Biotechnol 93:1021–1030
Wang Q, Venkataramanan KP, Huang H, Papoutsakis ET, Wu CH (2013) Transcription factors and genetic circuits orchestrating the complex, multilayered response of Clostridium acetobutylicum to butanol and butyrate stress. BMC Syst Biol 7:120
Wang J, Yang X, Chen CC, Yang ST (2014) Engineering clostridia for butanol production from biorenewable resources: from cells to process integration. Curr Opin Chem Eng 6:43–54
Wiesenborn DP, Rudolph FB, Papoutsakis ET (1989a) Coenzyme A transferase from Clostridium acetobutylicum ATCC 824 and its role in the uptake of acids. Appl Environ Microbiol 55:323–329
Wiesenborn DP, Rudolph FB, Papoutsakis ET (1989b) Phosphotransbutyrylase from Clostridium acetobutylicum ATCC 824 and its role in acidogenesis. Appl Environ Microbiol 55:317–322
Williams DR, Young DI, Young M (1990) Conjugative plasmid transfer from Escherichia coli to Clostridium acetobutylicum. Microbiology 136:819–826
Xu M, Zhao J, Yu L, Tang IC, Xue C, Yang ST (2015) Engineering Clostridium acetobutylicum with a histidine kinase knockout for enhanced n-butanol tolerance and production. Appl Microbiol Biotechnol 99:1011–1022
Xue C, Zhao J, Lu C, Yang ST, Bai F, Tang IC (2012) High-titer n-butanol production by Clostridium acetobutylicum JB200 in fed-batch fermentation with intermittent gas stripping. Biotechnol Bioeng 109:2746–2756
Xue C, Zhao XQ, Liu CG, Chen LJ, Bai FW (2013) Prospective and development of butanol as an advanced biofuel. Biotechnol Adv 31:1575–1584
Xue C, Zhao J, Sun JX, Jie CL, Bai FW, Yang ST (2014) Integrated butanol recovery for an advanced biofuel: current state and prospects. Appl Microbiol Biotechnol 98:3463–3474
Yang ST, Zhao JB (2013) Adaptive engineering of Clostridium for increased butanol production. US Patent 8450093
Yu M, Zhang Y, Tang IC, Yang ST (2011) Metabolic engineering of Clostridium tyrobutyricum for n-butanol production. Metab Eng 13:373–382
Yu M, Du Y, Jiang W, Chang WL, Yang ST, Tang IC (2012) Effects of different replicons in conjugative plasmids on transformation efficiency, plasmid stability, gene expression and n-butanol biosynthesis in Clostridium tyrobutyricum. Appl Microbiol Biotechnol 93:881–889
Zhang Y, Yu M, Yang ST (2012) Effects of ptb knockout on butyric acid fermentation by Clostridium tyrobutyricum. Biotechnol Prog 28:52–59
Zhao J, Lu C, Chen CC, Yang ST (2013) Biological production of butanol and higher alcohols. In: Yang ST, El-Enshasy HA, Thongchul N (eds) Bioprocessing technologies in biorefinery for sustainable production of fuels, chemicals, and polymers. John Wiley & Sons, New York, pp 235–261
Zheng Y, Li L, Xian M, Ma Y, Yang J, Xu X, He D (2009) Problems with the microbial production of butanol. J Ind Microbiol Biotechnol 36:1127–1138
Zhu Y, Yang ST (2003) Adaptation of Clostridium tyrobutyricum for enhanced tolerance to butyric acid in a fibrous-bed bioreactor. Biotechnol Prog 19:365–372
Acknowledgments
This work was supported in part by the National Science Foundation STTR program (IIP-1026648). We are grateful to Prof. N. P. Minton, University of Nottingham, UK for providing the donor E. coli CA434 and plasmid pMTL007 used in this study.
Compliance with ethical standards
This research does not involve human participants or animals.
Conflict of interest
The authors declare no conflict of interests.
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
ESM 1
(PDF 190 kb)
Rights and permissions
About this article
Cite this article
Yu, L., Zhao, J., Xu, M. et al. Metabolic engineering of Clostridium tyrobutyricum for n-butanol production: effects of CoA transferase. Appl Microbiol Biotechnol 99, 4917–4930 (2015). https://doi.org/10.1007/s00253-015-6566-5
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00253-015-6566-5