Abstract
The rhizosphere microbiome plays a significant role in the life of plants in promoting plant survival under adverse conditions. However, limited information is available about microbial diversity in saline environments. In the current study, we compared the composition of the rhizosphere microbiomes of the halophytes Urochloa, Kochia, Salsola, and Atriplex living in moderate and high salinity environments (Khewra salt mines; Pakistan) with that of the non-halophyte Triticum. Soil microbiomes analysis using pyrosequencing of 16S rRNA gene indicated that Actinobacteria were dominant in saline soil samples whereas Proteobacteria predominated in non-saline soil samples. Firmicutes, Acidobacteria, Bacteriodetes and Thaumarchaeota were predominant phyla in saline and non-saline soils, whereas Cyanobacteria, Verrucomicrobia, Gemmatimonadetes and the unclassified WPS-2 were less abundant. Sequences from Euryarchaeota, Ignavibacteriae, and Nanohaloarchaeota were identified only from the rhizosphere of halophytes. Dominant halophilic bacteria and archaea identified in this study included Agrococcus, Armatimonadetes gp4, Halalkalicoccus, Haloferula and Halobacterium. Our analysis showed that increases in soil salinity correlated with significant differences in the alpha and beta diversity of the microbial communities across saline and non-saline soil samples. Having a complete inventory of the soil bacteria from different saline environments in Pakistan will help in the discovery of potential inoculants for crops growing on salt-affected land.
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
Introduction
Salinization is one of the major problems that occur in more 100 countries of the world (Rengasamy 2006). Halophytes are plants that grow well in saline soil and water and compare well with forage crops like para grass, Salsola stocksii, Kochia indica and Atriplex amnicola (Ahmad et al. 2013). Therefore, these offer a better alternative where conventional crops cannot be raised and drainage is too expensive. They may contribute significantly to the developing world’s supply of food, fiber, fuel and fodder (Dagla and Shekhawat 2005). Salinity also affects microbial diversity, which plays a role in maintaining biogeochemical cycles (Tripathi et al. 2006). Rhizosphere microbial communities from extreme environments (saline, acidic, arid) are more complex than soils having a neutral pH and exhibiting moderate salinity. The rich microbial diversity of halophyte rhizospheres helps these plants cope with high salinity and also tolerate drought (Naz et al. 2009).
Biodiversity has been recognized as the primary factor that affects ecosystem functioning. For example, higher microbial diversity increases the resistance and stability of soil under stress conditions and influences nutrient cycling (Wagg et al. 2014; Zhang et al. 2015). The rhizosphere microbiome plays an important role in plant health because it serves as a “second genome” to the plant (Berendsen et al. 2012). Studies of rhizosphere microbiomes have revealed their influence on chemical exudates responsible for production and secretion of signaling molecules by both microbes and plants (Curlango-rivera et al. 2013). Rhizobacteria promote plant growth by increasing the availability and uptake of carbon, nitrogen, and minerals from soil (Dodd and Perez-Alfocea 2012; Delmont et al. 2014). Acting through a variety of mechanisms, certain microbes provide a source of fixed nitrogen and/or increase the bioavailability of soluble phosphorous for the plant. Many microbes also control plant pathogens by producing antimicrobial, antifungal, and insecticidal compounds as well as help plants withstand salt, drought, and heat (Craita and Tom 2013; Rolli et al. 2015).
Halophilic bacteria are classified according to the degree of their salt tolerance: slight halophiles (0.2–0.85 M NaCl), moderate halophiles (0.85–3.4 M NaCl), and extreme halophiles (3.4–5.1 M NaCl) (Ventosa et al. 2008). Halophiles use compatible solutes to balance their cytoplasmic osmotic potential and to protect their cells against drought, low oxygen, and high temperature (Yancey 2005). These microbes often have novel enzymes that function under salt stress conditions such as proteases, xylanases, cellulases, and amylases with polyextremophilic properties (Delgado-García et al. 2014). Certain enzymes that halophiles synthesize are useful for bioremediation of pollutants in saline habitats (Dastgheib et al. 2011) or are important biomolecules, e.g., exopolysaccharides and phytohormones (Liszka et al. 2012). Abiotic stresses, e.g., temperature, pH, salinity, and drought, affect the phytomicrobiome, directly or indirectly, via the host. However, the overall microbial composition in saline habitats is affected more by salinity than by other abiotic stresses (Ma and Gong 2013).
Scientists have developed a wide range of methods to study microbial diversity, community structure, and functions to understand plant-microbe interactions and soil biology (Rincon-Florez et al. 2013; Mukhtar et al. 2017). High throughput sequencing approaches have been developed to understand the complexity of microbial communities in a wide range of environments. Continuing advances in sequencing technology make it possible to study the dynamics of microbial population by using metagenomics and metatranscriptomics. Such approaches may uncover not only the composition of the microbiome community, but also the functional traits that the microbes use to survive under harsh conditions.
The microbial community associated with the Khewra salt mines, an extremely saline environment, have not been previously explored. Hence, the rhizosphere microbiomes of different halophytes from Khewra mines were thoroughly evaluated by high-throughput sequencing of the 16S rRNA gene, covering the V3–V4 regions. The main objective of this study was to compare and evaluate the differences in microbial communities from non-saline environment (rhizosphere of Triticum), moderately saline environment (rhizosphere of Urochloa and Kochia) and highly saline environment (rhizosphere of Salsola and Atriplex, non-rhizospheric and lake-bank soil samples) in Pakistan.
Materials and methods
Soil sampling
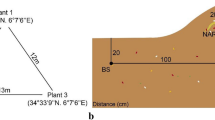
The Khewra salt mine, which is characterized by crystals of pure halite, is the second largest salt mine in the world (Ahmad et al. 2007). It is surrounded by a vast salt range that covers a 150-mile area from east to west. Rhizospheric soil samples were collected from three sites: highly saline soil from Khewra salt mines (73°08′N, 32°37′E); moderately saline soil from the Biosaline Research Station (BSRS) of the Nuclear Institute for Agriculture and Biology (NIAB) in Faisalabad (73°07′N, 31°42′E), and non-saline soil from the fields near Forman Christian College in Lahore (74°34′N, 31°54′E). Rhizospheric soil samples from the halophytes, S. stocksii and A. amnicola, non-rhizospheric soil, and lake-bank saline soil samples were collected from the Khewra salt mines (Fig. S1). In addition, soil samples from under the canopy of Urochloa mutica and K. indica were collected from the BSRS site in Faisalabad. Lastly, Triticum aestivum rhizospheric soil was collected from the Lahore site (Supplementary Table S1). No specific permissions were required to collect plant and soil samples from the Khewra salt mines because it is a public land and natural resource. We also collected plant (U. mutica and K. indica) and soil samples from the experimental fields of the NIAB, Faisalabad and from Forman Christian College, Lahore (wheat soil samples). An area approximately 1.1 km from the Khewra salt mines was surveyed. The sampling area was selected according to land use and vegetation cover. The vegetation of this area is classified as sub-tropical, dry evergreen forest. Suaeda, Salsola, Atriplex, Justica, Lantana, and Chrysopogon are the dominant plant genera in this region. Samples of rhizospheric soil from U. mutica, K. indica, and T. aestivum, grown under field conditions, were collected by gently removing the plants and carefully removing the soil attached to the roots. For non-rhizospheric saline soil samples, the upper 8–10 cm of mineral soil were collected. Lake-bank soil samples were collected from the bank of a salt lake (Fig. S2). At each site, soil samples of approximately 500 g each from four spatially separated plants from each species were collected at each site and placed in black sterile polythene bags. These samples were taken to the laboratory from the collection site in an ice box and stored at − 80 °C for pyrosequencing analysis.
Soil physicochemical characteristics
The physical and chemical properties of each soil sample were determined based on 400 g of dried and sieved soil. Electrical conductivity (dS/m) was measured by 1:1 (w/v) soil to water mixtures at 25 °C (Adviento-Borbe et al. 2006). The pH was measured by making a 1:2.5 (w/v) soil to water suspension whereas moisture (%), temperature (°C) and texture class were determined according to Anderson and Ingram (1993). Organic matter (Corg) was calculated by Walkley and Black method (1934), phosphorous (P) was estimated by extraction with sodium bicarbonate (Olsen et al. 1954), and calcium and magnesium were detected by atomic absorption spectrometry. Nitrate ions were measured by the Kjeldahl method and potential acidity (H + Al) was determined using an equation based on the pH in SMP buffer solution (pH SMP). Cation exchange capacity (CEC) is defined as the capacity to retain and release cations (Ca2+, Mg2+, K+ and Na+), and sodium adsorption ratio (SAR) is the measure of the sodicity of soil, which is calculated as the ratio of sodium to magnesium and calcium.
DNA extraction and pyrosequencing
Metagenomic DNA was extracted from 0.5 g of soil using a FastPrep® instrument (MP Biomedicals, USA) according to the manufacturer’s instructions. The concentration of metagenomic DNA was qualitatively determined on 0.8% (w/v) agarose gel and quantified using Nanodrop technology (NanoDrop 200c Thermo Scientific, USA). In total, 28 DNA samples seven rhizospheric and non-rhizospheric saline soil samples (with four replicates each) were used for amplification of 16S rRNA gene and pyrosequencing.
Amplification of 16S rRNA gene and pyrosequencing
The V3–V4 region of the 16S rRNA gene was amplified using primers F515 (5′-G TGCCAGCMGCCGCGG-3′) and R907 (5′-CCGTCAATTCMTTTRAG TTT-3′), which were linked with unique identifier and adapter sequences (Supplementary Table S2). The detailed PCR conditions for amplicon sequencing were the same as described previously (Mirza et al. 2014). Briefly, a 50 µl PCR amplification reaction contained 1X buffer, 0.2 µM of each primer, 1.8 mM MgCl2, 200 µM deoxynucleoside triphosphates (dNTPs), 20 ng of template, and 1 µl FastStart high-fidelity PCR system enzyme (Roche Applied Sciences). The PCR conditions were 3 min at 95 °C, followed by 30 cycles of denaturation at 94 °C for 45 s, primer annealing at 54 °C for 45 s, extension at 72 °C for 1 min, and a final extension for 7 min. Amplified PCR products were purified with Agencourt AMPure beads (Beckman Coulter, Brea, CA). Purified PCR products from different samples were pooled in equimolar concentrations. Pyrosequencing was performed on the mixture with the 454 GS FLX sequencer (454 Life Sciences) at the Utah State University Center for Integrated Biosystems.
Bioinformatics and statistical analyses
Sequences were processed and sorted using the default parameters in QIIME 1.3 (Caporaso et al. 2010). An offset of 10 nucleotides was set in order to remove the first 10 bases of each sequence, and high quality sequences with an average length of 375 bases were selected. A total of 63,450 high quality sequences were clustered into operational taxonomic units (OTUs) with 3% difference using UCLUST for 28 samples with sequence numbers ranging from 872 to 5457. For the identification of chimeric sequences, Chimera Slayer software was used (DeSantis et al. 2006). The cleaned sequences were analyzed using RDP classifier naïve Bayesian algorithm (Wang et al. 2007) with an 80% confidence threshold. For each taxonomic level (phylum, class, order, family and genus) Good’s coverage was calculated by using 97% similarity cutoff (Good 1953).
Alpha and beta diversity were determined using QIIME alpha_rerefaction.py and beta_diversity_through_plots.py commands respectively. The rarefaction curves for observed OTUs and Shannon index were presented in Fig. S3. Alpha diversity was calculated at sequence depth of 2152 reads per soil sample, as alpha diversity indices are correlated with the number of sequences. The selected maximum sampling depth corresponded to minimum number of reads obtained from any of the remaining sequenced samples. Beta diversity was analyzed by using Principal component analysis (PCoA). A matrix was calculated using the weighted and unweighted UniFrac distances among samples at a sequence depth of 872 reads per soil sample. The robustness of the UPGMA tree was calculated with Jackknife test, based on 1000 replicates and the beta_diversity.py, UPGMA_cluster.py and tree_compare.py commands. The similarity percentages (SIMPER) was performed pairwise comparisons of groups of non-saline, moderately and highly saline soil samples and found the average contributions of each OUT to the average overall Bray-Curtis dissimilarity between soil samples through the decomposition of Bray-Curtis dissimilarity (Henderson and Seaby 2014). The decomposition of dissimilarity was calculated using Community analysis package 5 (CAP) for the difference of abundance of each OTU in each soil sample (Henderson and Seaby 2014). To detect the taxonomic classifications that were significantly abundant in non-saline, moderately and highly soils, Wilkcoxon’s non-parametric rank-sum test and LDA using the LEfSe program was used (Segata et al. 2011). One-way ANOVA was applied to analyze differences among saline and non-saline soil samples. Detrended correspondence analysis (DCA) was used to measure the correlation between soil physicochemical properties and community structure. DCA was performed using PAST (PAleontological STatistics) 3.12 (Hammer et al. 2001). Spearman’s rank correlation test explains the degree of dependence among the variables, but the coefficient does not suppose that the relationship among the variables is linear (Gautier 2001). Canonical corresponding analysis (CCA) was used to show overall patterns of microbial diversity in different soil samples by using PAST 3.12 (Hammer et al. 2001). To explain the differences in the composition of taxa inside the data matrix community, a heatmap (relative abundance matrix) was generated at class level using XLSTAT 7.0 software (Fahmy 2003). Variation partitioning was performed on the basis of canonical correspondence analysis and then applied on the community data to disentangle the specific effects of soil salinity on the microbial community structure. The variation in microbial diversity explained by the soil salinity was calculated using partitioning analysis (Borcard et al. 1992; Buttigieg and Ramette 2014) as implemented in the vegan R package with some modifications according to our data. To calculate the variation explained by one group (highly saline soils) within our dataset, we calculated the variation explained on the complete community matrix and compared this to a matrix from which all OTUs from highly saline soils had been removed. The difference in these variation partitionings was taken as the variation explained by highly saline soil group in the context of other groups, e.g., non-saline soil group. Sequence data analyzed through pyrosequencing was submitted to the NCBI Sequence Read Archive (SRA) under ID project PRJNA309754.
Results
Physicochemical characteristics of soil and microbial community structure
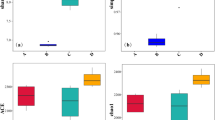
Soil samples from the seven different sites were characterized by a great spatial variability and covered a significant variation in soil salinity, pH, organic matter, vegetation type, texture class, CEC, and SAR. EC (EC1:1) ranged from 1.11 ± 0.23 to 6.24 ± 1.21 dS/m, with the highest values in lake-bank soil samples and the lowest values in Triticum (non-halophyte) soil samples; pH values ranged from 7.65 ± 0.45 to 8.54 ± 0.57, temperature from 22 ± 2 to 32 ± 3 °C, and moisture from 20 ± 3 to 35 ± 4%. Total organic matter ranged from 29.29 ± 2.56 to 36.25 ± 4.87 g/Kg. The available P, K, Ca, and Mg contents were different in Triticum (non-halophyte) rhizospheric soil than in the other soil samples (Supplementary Table S3). CEC values ranged from 56.46 ± 4.51 to 81.15 ± 7.45 mg/dm3 and SAR values from 6.68 ± 2.56 to 13.45 ± 4.29, with the highest values in non-rhizospheric saline soil samples and the lowest in Triticum soil samples. Differences in community structure among different soil samples were explained by DCA (Fig. 1). Soil sample replicates of Urochloa and Kochia and lake-bank showed greater variation in microbial community composition from site to site than other saline soil samples. At each site, certain bacterial and archaeal species prevailed better than others. The microbial communities expressed differently from point to point because of variations in environmental factors like salinity and pH. DCA also revealed that Triticum (non-halophyte) soil samples had significant differences in physicochemical characteristics compared to saline soil samples.
General analysis of pyrosequencing at phylum, class and genus level
Almost all reads (99%) were assigned at the domain level, 70–85% reads were classified at the phylum level and only 30–49% reads were classified to the genus level (Fig. S3). The maximum number of phyla (20) was identified from the rhizospheric soil sample of Urochloa and the minimum number of phyla (17) was from the rhizospheric soil samples of Triticum. Sequences assigned by the RDP classifier at the class level suggested an uneven distribution in soil samples from saline and non-saline environments (Fig. 2). A total of 56 different bacterial and archaeal classes were identified in this study, with the maximum number of classes (42) in non-rhizospheric saline soil samples and the minimum (30) in the rhizospheric soil samples of Kochia. Sequence analysis at the genus level showed that Urochloa soil samples had the maximum identifiable bacterial and archaeal genera (170) and the Triticum (non-halophyte) soil had the minimum identifiable bacterial genera (99) (Fig. 2).
Alpha and beta diversity patterns
Number of observed OTUs, alpha diversity (Shannon index), beta diversity, and phylogenetic diversity were calculated at 97% DNA similarity. The alpha diversity patterns were highly variable across the saline and non-saline soil samples (Fig. 3). All Chao1 values were greater than the observed OTUs, indicating that more OTUs could be retrieved by additional sequencing (Table S5). Rarefaction curves were obtained using only those sequences assigned to a genus level. The rarefaction curves did not reach saturation in all the soil samples from non-saline, moderately and highly saline environments (Fig. S4). Results of observed OTUs and alpha diversity showed that Urochloa rhizosphere had more microbial diversity as compared to other halophytes and non-halophyte soil samples (Fig. 3). PCoA using unweighted Unifrac distances showed that non-saline soils were clustered separately from moderately and highly saline soils (Fig. 4a). Similar results were observed in the UPGMA tree of unweighted UniFrac distances (Fig. 4b). Non-saline soils (Triticum rhizosphere) were grouped in one cluster, moderately saline soils (Urochloa and Kochia) and highly saline soils (Salsola and Atriplex, non-rhizospheric and lake-bank) formed other two clusters (Fig. 4b). All the soil samples were analyzed in UPGMA tree of unweighted UniFrac at 100% jackknife support (JS). These results showed that microbial communities from moderately saline soils (Urochloa and Kochia) had more diversity as compared to other highly saline (Salsola and Atriplex, non-rhizospheric and lake-bank) and non-saline (Triticum) soil samples.
Microbial diversity comparisons between saline and non-saline soils
Rhizospheric soil microbial communities in moderately saline, highly saline and non-saline environments showed similar patterns of relative abundance in the major groups at the phylum level. A total of 24 bacterial and archaeal phyla were identified from saline and non-saline soil samples. The dominant phyla in all soil samples were Actinobacteria (16.14–36.34%), Proteobacteria (20.51–35.27%), Firmicutes (3.11–15.58%), Acidobacteria (1.11–5.51%), and Bacteroidetes (1.11–3.92%) (Fig. S5). Saline soils (Urochloa, Kochia, Salsola and Atriplex) showed greater abundance of Actinobacteria (7.23%), Firmicutes (4.53%), Chloroflexi (2.21%), Thaumarchaeota (1.26%), and Euryarchaeota (0.42%) as compare to non-saline soils (Triticum). Sequences of the 16S rRNA gene from Proteobacteria (5.22%), Acidobacteria (2.41%), and Planctomycetes (0.31%) were more abundant in the rhizosphere of Triticum. Sequences related to phylum Chlamydiae were found in the rhizosphere of Urochloa (0.14%), Salsola (0.084%), and lake-bank (0.11%) soil samples. Sequences belonging to the bacterial phylum Hydrogenedentes were detected only from the rhizosphere of Urochloa (0.14%) and Salsola (0.12%). Members of the phylum Ignavibacteriae were identified from the rhizosphere of Salsola (0.047%) and non-rhizospheric (0.11%) saline soil samples. Sequences from the phylum Nanohaloarchaeota were detected only from the rhizosphere of Salsola (0.16%). Canonical correspondence analysis showed that microbial communities in the rhizosphere of Urochloa and Kochia (moderately saline environment) were similar. From highly saline environment, microbial communities were similar in the rhizosphere of Salsola and Atriplex, but lake-bank soil samples showed totally different microbial diversity as compared to other samples. Microbial community present in the rhizosphere of Triticum (non-halophyte) showed a great variation from the saline samples. For example, bacterial phyla BRC1 and Fusobacteria were only present in the rhizosphere of wheat. This could be due to the differences in soil salinity at different sites (Fig. 5). Pearson correlation analysis was performed between different soil samples (halophyte and non-halophyte) and microbial diversity at the phylum level (Table 1). Our results showed that Actinobacteria significantly correlated with the rhizosphere of halophytes and Proteobacteria with the non-halophyte, whereas Planctomycetes correlated least with the rhizosphere of halophytes and Bacteroidetes with the non-halophyte (Triticum).
CCA of the microbial communities from the rhizosphere of halophytes (Urochloa, Kochia, Salsola, and Atriplex) from moderately and highly saline and non-saline environments (Triticum) at the phylum level. Among all the variables tested, these were significantly associated with the three different sites
The relative distribution of bacterial groups at the class level showed significant differences among saline and non-saline soils. Different bacterial and archaeal classes were clustered and plotted with respect to the different soil sample values (saline and non-saline) in the heatmap (Fig. 6). The cluster structure showed a different distribution pattern in non-saline soils (Triticum) as compared to saline soils. The main group of bacterial and archaeal classes (Actinobacteria, Alphaproteobacteria, Gammaproteobacteria, Deltaproteobacteria, Bacilli, Sphingobacteriia and Nitrososphaeria) shared similar composition and abundance among the different saline soils.
The heatmap reports the normalized values of the taxonomic assignments at class level. Each value has been normalized following this criterion: \(\mathop {\text{X}}\nolimits_{{{\text{ij}}}}^{{{\text{norm}}}} = {{{\text{X}}_{{{\text{ij}}}} } \mathord{\left/ {\vphantom {{{\text{X}}_{{{\text{ij}}}} } {\sum\nolimits_{{{\text{k = 1}}}}^{{\text{N}}} {{\text{X}}_{{{\text{ik}}}} } }}} \right. \kern-\nulldelimiterspace} {\sum\nolimits_{{{\text{k = 1}}}}^{{\text{N}}} {{\text{X}}_{{{\text{ik}}}} } }},\) where Xij is the occurrence of the phyla ‘j’ in the soil sample ‘i’ and N is the number of saline and non-saline soil samples (in this study 7). Using this transformation, sequences assigned to each phylum can be compared in all soil samples independently from its order of magnitude
Soil salinity influences the halophyte microbiome (genus level)
Of 232 different phylotypes identified, 46 were commonly detected in all saline and non-saline soil samples. Our results also showed that 32 phylotypes were detected only from Urochloa, 18 from Kochia, 22 from Salsola, 30 from Atriplex, 30 from non-rhizospheric saline soil samples, 27 from lake-bank soil samples, and 27 from the rhizosphere of Triticum (Fig. S6). Total dissimilarity between pairs of non-saline, moderately and highly saline soils and the relative distribution of each bacterial or archaeal family to the observed dissimilarity was analyzed by using SIMPER test. Results of SIMPER analysis showed the difference in microbial diversity across non-saline and saline soil samples. Non-saline soils (T. rhizosphere) indicated the maximum dissimilarity, approximately 32% with moderately saline and 45% with highly saline soils (Fig. 7a). SIMPER analysis also showed that the overall differences between non-saline and saline soil samples were due to the detection of a wide range of taxa, each contributing a relatively small percentage of the differences. For identification of specific bacterial genera that were significantly abundant in one site (non-saline soils) than the others (saline soils), LEfSe (LDA effective size) analysis was performed. The results of LEfSe analysis showed that 7 bacterial genera (Bacillus, Azospirillum, Burkholderia, Enterobacter, Microbacterium, Aeromonas, and Mesorhizobium) were more abundant in non-saline soils (T. rhizosphere) as compared to moderately and highly saline soils (Fig. 7b). Similarly, 13 bacterial genera were found to be abundant in moderately saline soils (Urochloa and Kochia rhizospheres) as compared to non-saline and highly saline soils. The top 5 bacterial genera based on LDA scores were Virgibacillus, Arthrobacter, Klebsiella, Pseudomonas, and Azotobacter from moderately saline soils (Fig. 7b). From highly saline soils (Salsola and Atriplex, non-rhizospheric and lake-bank soils), 7 bacterial genera (Halobacillus, Aciditerrimonas, Methylobacillus, Dechloromonas, Patulibacter, Oceanobacillus, and Micrococcus) were more abundant as compared to non-saline and moderately saline soils (Fig. 7b). These results also confirmed the abundance of bacterial phyla Proteobacteria, Actinobacteria, Firmicutes, and Bacteroidetes, as the top bacterial genera from all the soil samples were belonging to these phyla (Fig. 7b). From the rhizosphere of Urochloa, Arthrobacter (17.52%), Virgibacillus (12.34%), and Pseudomonas (9.57%) were found to be abundant bacterial genera while from Kochia rhizosphere, Rubrobacter (12.95%), Asanoa (10.67%), and Planococcus (7.85%) were abundant bacterial genera (Table 2). From highly saline environment, Aciditerrimonas (14.62%), Terrimonas (8.54%), and Citrobacter (6.32%) were identified from the rhizosphere of Salsola, Halobacillus (17.52%), Staphylococcus (11.23%), and Rhizobium (4.14%) were abundant in the rhizosphere of Atriplex, Oceanobacillus (14.23%), Patulibacter (9.89%), and Kocuria (7.98%) and Xanthomonas (10.29%), Streptococcus (8.38%), and Clostridium (6.93%) were identified as dominant genera from lake-bank soil samples (Table 2).
Variations in the bacterial genera detected across non-saline, moderately, and highly saline environments. a Comparison of community structure between non-saline, moderately and highly saline soils by SIMPER analysis. Bars represent the standard error. b The results of LEfSe analysis, which identified bacterial genera that were significantly abundant in one site as compared to other sites. Bacterial genera with the LDA score of more than 3 are shown
Bacterial and archaeal genera with specific functions (PGPR activities) were also characterized from all saline and non-saline soil samples. N2-fixing bacteria were found to be abundant in the rhizosphere of halophytes (Kochia, Salsola) as well as in Triticum (non-halophyte). P-solubilizers and IAA producers were more abundant in the rhizosphere of halophytes as compared to the non-halophyte. Bacterial genera involved in biocontrol and bioremediation of toxic compounds were abundant in the non-rhizospheric saline soil samples. Rhizospheric and non-rhizospheric saline soil samples had high numbers of halophilic bacterial and archaeal genera (Table 3).
Strains belonging to 14 PGPR genera (Bacillus, Arthrobacter, Ensifer, Burkholderia, Nocardioides, Gemmatimonas, and Mesorhizobium) were identified from all soil samples (Fig. 8). Sequences from 14 PGPR genera Aeromonas, Bradyrhizobium, Citrobacter, Delftia, Flavobacterium, Kocuria, and Skermanella were detected only from the saline environments (Urochloa, Kochia, Salsola, and Atriplex) while Achromobacter, Azotobacter, Pseudomonas, Klebsiella, Lysobacter, Micrococcus, and Pantoea were detected only from the non-saline environment (Triticum).
Ecological interpretation of overall microbial diversity patterns
A multivariate variation partitioning approach was used to study effects of soil salinity on the overall variation of rhizosphere microbiome based on the most abundant OTUs. The results showed that significant environmental variables (moderately saline, highly saline, and non-saline soil samples) included in the most parsimonious multivariate models were qualitatively different for the resident OTU, the phylum level OTU, OTUall (all sequences) and SSOrel (reduced sequence) (Fig. 9a). This study also included rare types, the resident single-sequence OTU relative (SSOrel) data sets or sequence related to highly salt tolerant bacteria and archaea (Halomonas, Halobacillus, Halococcus, Halobacterium and Halalkalicoccus) to better understand the patterns of rare fraction of microbial community. A combination of environmental parameters (moderately saline, highly saline, and non-saline soil samples) could explain 59% of their overall biological variation while 41% variations were unexplained (Fig. 9b).
Partitioning between the biological variations in the bacterial community structure based on soil salinity. Three explanatory matrices were used here, containing variables pertaining to moderately saline, highly saline and non-saline soil samples. Variation explained on basis of a the resident OTU, the phylum level OTU, OTUall and SSOrel b highly salt tolerant bacteria and archaea (Halomonas, Halobacillus, Halococcus, Halobacterium and Halalkalicoccus)
Discussion
In this study, we determined that the microbial community structure and the abiotic factors like salinity affect the microbial community. The rhizospheric soil samples were taken from moderately saline, highly saline, and non-saline environments. Using metagenomic analysis, we provided information about different bacterial and archaeal genera with potential functions in the rhizosphere microbiomes of halophytes and non-halophytes. Increases in salinity cause changes in soil organic matter, leaching and erosion, loss of nutrients, and changes in both quantity and composition of soil microbial communities (Langenheder et al. 2010; Yousuf et al. 2012). We determined that the differences in microbial communities were strongly correlated with specific soil properties such as SAR and CEC, both of which can be correlated with salinity (Fig. 1). The link among plants, rhizosphere microbial communities and soil properties are often described as complex derivatives of the ecosystem and any modification in this relationship might affect the microbial structure and the ecological processes (Fierer et al. 2012; Pan et al. 2014).
In general, microbial diversity from non-saline soils (T. rhizosphere) was found to be less as compared to moderately and highly saline soils. The number of observed OTUs and Shannon index were maximum from the rhizosphere of Urochloa and Kochia (moderately saline soils). These plants grow under moderate saline environments (4.14 EC1:1 dS/m) and the physiological conditions of these plants influence the development of rhizosphere microbial communities. Some OTUs were commonly identified from all the saline soil samples and some OTUs were identified only from the rhizosphere of Urochloa and Kochia (Fig. 3). The number of observed OTUs and Shannon index for halophytes Salsola and Atriplex which are growing under highly saline environments (5.19 EC1:1 dS/m) was found to be less as compared to moderately saline soils and more as compare to non-saline soils, but the abundance of certain bacterial groups especially the Firmicutes were more in these soils as compared to non-saline and moderately saline soils (Urochloa and Kochia rhizosphere). Beta diversity analysis by PCoA using unweighted Unifrac distances and UPGMA tree also showed the differences in microbial diversity in three sites: non-saline, moderately and highly saline soils (Fig. 4).
Sequence analysis through the RDP classifier showed that the rhizosphere microbiomes of halophytes and the non-halophyte Triticum had different abundances of bacterial and archaeal phyla. Microbial communities from moderately saline environments exhibited more diversity compared to highly saline and non-saline environments at the phylum level (Fig. S5). The decrease in microbial diversity with increased salinity might be due to decline in hydrolytic and oxidative enzyme activities and carbon cycling (Ghollarata and Raiesi 2007). Some of the bacterial and archaeal phyla, namely Actinobacteria, Proteobacteria, Firmicutes, Acidobacteria, Planctomycetes, Bacteroidetes, Chloroflexi, Gemmatimonadetes, and Cyanobacteria were found to be abundant in salt-affected soils as compared to non-saline soils. Sequences from Euryarchaeota were identified only from the rhizosphere of halophytes. Overall microbial diversity composition was significantly different among the three sites; non-saline, moderately and highly saline soils.
Our results showed that members of Actinobacteria, Arthrobacter, Solirubrobacter, Patulibacter, Kocuria, and Actinomycetospora were identified from moderate and high salinity environments. These strains were found to promote plant growth and to degrade a variety of hazardous agricultural and industrial pollutants including atrazine, butane, p-nitrophenol, trinitrophenol, and vinyl chloride (Deng et al. 2015). The main difference in previous studies and our current findings was that Actinobacteria were more abundant in moderately saline environments especially in the rhizosphere of Kochia at genus level. Overall, bacterial genera Arthrobacter and Kocuria were found to be abundant in all the saline soils. These bacteria have ability to survive in hypersaline environments. They contain glutamic acid, glycine, alanine, and meso-diaminopimelic acid as cell-wall-bound amino acids (Bodenhausen et al. 2013; Niemho et al. 2013). Bacterial genera Euzebya, Marmoricola, Aciditerrimonas, and Aeromicrobium, which were detected only in highly saline environments (rhizospheres of Salsola and Atriplex and non-rhizospheric soil samples), are mostly saccharolytic bacteria (they obtain carbon and energy by carbohydrate hydrolysis) and contribute to carbon cycling in soil ecosystems (Krivushin et al. 2015). They also show the ability to degrade various hazardous compounds and can be used as biocatalysts in the production of fossil fuels and bioactive steroids (Deng et al. 2015).
Among Proteobacteria the Gammaproteobacteria were the most abundant. Bacterial genera Devosia, Halomonas, Burkholderia, Rubrobacter, Geobacter, Azospirillum, and Aeromonas were more abundant in the rhizospheres of halophytes as compared to non-halophyte (T. rhizosphere). These bacteria are involved in plant growth promotion, antibacterial and antifungal activities, and bioremediation of various toxic compounds and are also a source of naturally occurring bioactive products (Bodenhausen et al. 2013). Bacterial genera Azoarcus, Kosakonia, Dongia, and Sorangium identified only from the rhizosphere of Urochloa and Kochia (moderate salinity) were previously reported from the rhizosphere of grasses (para grass and kallar grass) as having good plant growth-promoting abilities (Malik et al. 1997; Mukhtar et al. 2016). These results indicated that microbial communities in the rhizosphere of each plant vary to some extent even if these plants were collected from the same environments. For example, microbial diversity from the rhizosphere of Urochloa showed different community composition when compared to Salsola.
Members of the Firmicutes Bacillus, Staphylococcus, Planococcus, Sporosarcina were dominant in saline soils as compared to non-saline soils. Bacillus-like organisms are dominant in saline environments due to their ability of spore-forming and Gram-positive cell walls (Schimel et al. 2007). It is well known that Bacillus-like organisms play an important ecological role in biogeochemical cycles in different ecosystems such as marine waters and saline soils (Taprig et al. 2013; Mukhtar et al. 2017). Halophilic Bacillus strains promote plant growth, produce industrially important enzymes (proteases, amylases, cellulases and lipases) and involved in bioremediation of different toxic chemicals and pollutants from saline environments (López-López et al. 2010; Mukhtar et al. 2018). Planococcus, and Planomicrobium were dominant genera identified from moderately saline environments (rhizosphere of Urochloa and Kochia). They are involved in calcite precipitation and biological cementation (Ma and Gong 2013; Tsuda et al. 2015).
Bacterial genera related to Bacteroidetes (Flavobacterium, Salinibacter, Cytophaga, Adhaeribacter, and Pontibacter) were identified from the rhizospheres of Urochloa, Salsola and Atriplex and are important in the bioremediation of complex organic compounds. They have also been reported in previous studies from marine environments (Vaisman and Oren 2009; Wang et al. 2012). All members of the phylum Thaumarchaeota identified so far are ammonia oxidizers and have important roles in biogeochemical cycles such as carbon and nitrogen cycles (Yanagawa et al. 2014; Doxey et al. 2015). Sequences attributed to the genus Nitrososphaera were frequently detected in the rhizosphere of Salsola and Atriplex (highly saline soils) compared to moderately saline and non-saline soils.
Bacterial genera belonging to Acidobacteria (Gp2, Gp4, Gp6, Gp21, and Gp25) were abundant in the rhizosphere of halophytes (Salsola and Atriplex) from highly saline soils. Members of Acidobacteria were a dominant part of microbial communities from saline soils and marine sediments (Ghosh et al. 2010; Mukhtar et al. 2016). Bacterial genera Sphaerobacter and Litorilinea (phylum Chloroflexi) were identified from Salsola and Atriplex plants. They have been previously reported from hypersaline wastewater (Valenzuela-Encinas et al. 2008; Rincon-Florez et al. 2013). Bacterial genera Armatimonas, Fimbriimonas and Chthonomonas (phylum Armatimonadetes) were detected from the rhizosphere of Urochloa, Salsola and Atriplex. These genera have been studied from various habitats including hypersaline, marine and hot spring water, and anaerobic sludge (DeBruyn et al. 2011; Tamaki et al. 2011).
Archaeal sequences belonging to Euryarchaeota were abundant in the rhizospheres of the halophytes and non-rhizospheric saline soil samples. Members of Euryarchaeota from marine environments are reported to be involved in bioremediation, and they can break down complex lipoprotein molecules to survive in extreme conditions (Garrity and Holt 2001; Iverson et al. 2012). Sequences assigned to Nanohaloarchaeota were identified only in the rhizosphere of Urochloa and Salsola. Previous studies reported that they are abundant in surface waters of hypersaline lakes worldwide (Narasingarao et al. 2012; Pan et al. 2014). However, from the rhizosphere of halophytes, identification of Nanohaloarchaeota was reported for the first time in this study. From the rhizosphere of Urochloa, unclassified bacteria were more prevalent as compared to unclassified bacteria from other soil samples. Previous studies on metagenomics have also reported a higher proportion of unclassified bacteria from different environments; however, these studies need thorough excellent sequencing procedures (Chakraborty et al. 2015).
Sequences related to plant growth-promoting rhizobacteria (PGPR) such as Arthrobacter, Bacillus, Burkholderia, Kocuria, Nocardioides, and Planococcus were relatively more abundant in the rhizospheres of the halophytes compared to that of the non-halophyte (Fig. 8). PGPR genera Achromobacter, Klebsiella, and Lysobacter were identified only from the rhizosphere of Triticum (Fig. 8). Application of these PGPR strains as a biofertilizer has resulted in improved growth and grain yield of various crops including maize, wheat, rice, and sugarcane (Gupta et al. 2015). Overall, our pyrosequencing results analysis revealed significant differences in the microbial community composition from saline and non-saline environments.
Conclusion
To our knowledge, this is the first study of moderate and high salinity soils of the Khewra salt mines of Pakistan that is based on a metagenomic analysis. The soil microbiomes of the halophytes Urochloa and Kochia from a moderately saline environment, the high saline-adapted halophytes Salsola and Atriplex, the non-rhizospheric and lake-bank samples of a high salinity environment, and a non-halophyte (Triticum) exhibited significant differences in microbial populations especially at the genus level. At each site, specific bacterial and archaeal species dominated. The Salsola and lake-bank soil samples showed great variation even within biological replicates and from site to site. In the soils studied here, we observed significant differences in alpha and beta diversity in microbial communities from saline (both moderate and high) and non-saline soils. The rhizosphere microbiomes of the halophytes (Urochloa and Kochia) from the moderately saline soils showed more alpha diversity compared to highly saline (Salsola and Atriplex) and non-saline (Triticum) soils. By contrast, soil microbiomes of Salsola and Atriplex and non-rhizopspheric soil samples from highly saline environments showed more beta diversity compared to the other soil samples. A novel representative of Nanohaloarchaeota (Candidatus Nanosalina) was detected only in moderately saline soils. More than 230 bacterial and archaeal genera with potential PGPR functions (N2 fixation, P solubilization, IAA, exopolysaccharide production, and biocontrol and bioremediation activity) were identified from both moderately and highly saline soils. With deeper study, we may find that many of inhabitants of the microbial communities from naturally salt-affected soils will prove to have significant biotechnological potential for promoting crop growth in harsh environments.
References
Adviento-Borbe MA, Doran JW, Drijber RA, Dobermann A (2006) Soil electrical conductivity and water content affect nitrous oxide and carbon dioxide emissions in intensively managed soils. J Environ Qual 35:1999–2010
Ahmad K, Hussain M, Ashraf M, Luqman M, Ashraf MY, Khan ZI (2007) Indigenous vegetation of Soon valley at the risk of extinction. Pak J 39(3):679–690
Ahmad MJ, Arshad M, Iqbal A, Khalid M, Akhtar N (2013) Rice production in salt-affected soils of Pakistan using different reclamation techniques. In: Shahid SA, Abdelfattah MA, Taha FK (eds), Developments in soil salinity assessment and reclamation: innovative thinking and use of marginal soil and water resources in irrigated agriculture Springer, Dordrecht, pp 283–293
Anderson JM, Ingram JS (1993) Tropical soil biology and fertility: a handbook of methods, 2nd edn. CAB International, Wallingford, pp 93–94
Berendsen RL, Pieterse CMJ, Bakker AHM (2012) The rhizosphere microbiome and plant health. Trends in Plant Sci 17:478–486
Bodenhausen N, Horton MW, Bergelson J (2013) Bacterial communities associated with the leaves and the roots of Arabidopsis thaliana. PLoS ONE 8:e56329
Borcard D, Legendre P, Drapeau P (1992) Partialling out the spatial component of ecological variation. Ecology 14:1045
Buttigieg PL, Ramette A (2014) A Guide to statistical analysis in microbial ecology: a community-focused, living review of multivariate data analyses. FEMS Microbiol Ecol 90:543–550
Caporaso JG, Kuczynski J, Stombaugh J, Bittinger K et al (2010) QIIME allows analysis of high throughput community sequencing data. Nat Methods 7:335–336
Chakraborty A, Bera A, Mukherjee A et al (2015) Changing bacterial profile of Sundarbans, the world heritage mangrove: impact of anthropogenic interventions. World J Microbiol Biotechnol 31(4):593–610
Craita EB, Tom G (2013) Plant tolerance to high temperature in a changing environment: scientific fundamentals and production of heat stress-tolerant crops. Front Plant Sci 4:1–18
Curlango-rivera G, Huskey DA, Mostafa A, Kessler JO, Xiong Z, Hawes MC (2013) Intraspecies variation in cotton border cell production: rhizosphere microbiome implications. Am J Bot 100:1706–1712
Dagla HR, Shekhawat NS (2005) In vitro multiplication of Haloxylon recurvum (Moq.)—a plant for saline soil reclamation. J Plant Biol 7:155–160
Dastgheib SM, Amoozegar MA, Khajeh K, Ventosa A (2011) A halotolerant Alcanivorax sp. strain with potential application in saline soil remediation. Appl Microbiol Biotech 90:305–312
DeBruyn J, Nixon L, Fawaz M, Johnson M, Radosevich M (2011) Global biogeography and quantitative season dynamics of Gemmatimonadetes in Soil. Appl Environ Microbiol 77(17):6295–6300
Delgado-García M, Aguilar CN, Contreras-Esquivel JC, Rodríguez-Herrera R (2014) Screening for extracellular hydrolytic enzymes production by different halophilic bacteria. Mycopathy 12(1):17–23
Delmont TO, Francioli D, Jacquesson S, Laoudi S, Mathieu A, Nesme J et al (2014) Microbial community development and unseen diversity recovery in inoculated sterile soil. Biol Fert Soil 50:1–8
Deng S, Chang X, Zhang Y, Ren L, Jiang F, Qu Z, Peng F (2015) Nocardioides antarcticus sp. nov., isolated from marine sediment of Ardley cove. Int J Syst Evol Microbiol 65(8):2615–2621
DeSantis TZ, Hugenholtz P, Larsen N, Rojas M, Brodie EL, Keller K et al (2006) Green genes, a chimera checked 16S rRNA gene database and workbench compatible with ARB. Appl Environ Microbiol 72:5069–5072
Dodd IC, Perez-Alfocea F (2012) Microbial amelioration of crop salinity stress. J Exp Bot 63(9):3415–3428
Doxey AC, Kurtz DA, Lynch MDJ, Sauder LA, Neufeld JD (2015) Aquatic metagenomes implicate Thaumarchaeota in global cobalamin production. ISME J 9:461–471
Fahmy T (2003) XLSTAT-Pro 7.0 (XLSTAT). Addinsoft, Paris
Fierer N, Leff JW, Adams BJ et al (2012) Cross-biome metagenomic analyses of soilmicrobial communities and their functional attributes. Proc Natl Acad Sci USA 109:21390–21395
Garrity GM, Holt JG (2001) Phylum AII. Euryarchaeota phy. nov. In: Boone DR, Castenholz RW (eds) Bergey’s manual of systematic bacteriology volume 1: the archaea and the deeply branching and phototrophic bacteria, 2nd edn. Springer-Verlag, New York, pp 169–192
Gautier TD (2001) Detecting trends using Spearman’s rank correlation coefficient. Environ Forens 2:359–362
Ghollarata M, Raiesi F (2007) The adverse effects of soil salinization on the growth of Trifolium alexandrinum L. and associated microbial and biochemical properties in a soil from Iran. Soil Biol Biochem 39:1699–1702
Ghosh A, Dey N, Bera A, Tiwari A, Sathyaniranjan K, Chakrabarti K, Chattopadhyay D (2010) Culture independent molecular analysis of bacterial communities in the mangrove sediment of Sundarban, India. Saline Syst 6:1–5
Good IJ (1953) The population frequencies of species and the estimation of population parameters. Biometrica 40:237–264
Gupta G, Parihar SS, Ahirwar NK, Snehi SK, Singh V (2015) Plant growth promoting rhizobacteria (PGPR): current and future prospects for development of sustainable agriculture. J Microb Biochem Technol 7:96–102
Hammer Ø, Harper DAT, Ryan PD (2001) PAST: paleontological statistics software package for education and data analysis. Palaeont Electron 4(1):9–13
Henderson PA, Seaby RMH (2014) Community analysis package version 5. Pisces Conservation Ltd, Lymington
Iverson V, Morris RM, Frazar CD, Berthiaume CT, Morales RL, Armbrust EV (2012) Untangling genomes from metagenomes: revealing an uncultured class of marine Euryarchaeota. Science 335:587–590
Krivushin K, Kondrashov F, Shmakova L, Tutukina M, Petrovskaya L, Rivkinaa E (2015) Two metagenomes from late Pleistocene northeast Siberian permafrost. Genome Announc 3(1):1380–1400
Langenheder S, Bulling MT, Solan M, Prosser JI (2010) Bacterial biodiversity–ecosystem functioning relations are modified by environmental complexity. PLoS ONE 5:e1083
Liszka M, Clark M, Schneider E, Clark DS (2012) Nature versus nurture: developing enzymes that function under extreme conditions. Ann Rev Chem Biomol Eng 3:77–102
López-López A, Yarza P, Richter M, Suárez-Suárez A et al (2010) Extremely halophilic microbial communities in anaerobic sediments from a solar saltern. Environ Microbiol Rep 2:258–271
Ma B, Gong J (2013) A meta-analysis of the publicly available bacterial and archaeal sequence diversity in saline soils. World J Microbiol Biotechnol 29:2325–2334
Malik KA, Bilal R, Mehnaz S, Rasool G, Mirza MS, Ali S (1997) Association of nitrogen-fixing, plant growth promoting rhizobacteria (PGPR) with kallar grass and rice. Plant Soil 194:37–44
Mirza BS, Muruganandam S, Meng X, Sorensen DL, Dupont RR, McLean JE (2014) Arsenic(V) reduction in relation to Iron(III) transformation and molecular characterization of the structural and functional microbial community in sediments of a basin-fill aquifer in Northern Utah. Appl Environ Microbiol 80:3198–3208
Mukhtar S, Mirza MS, Awan HA, Maqbool A, Mehnaz S, Malik KA (2016) Microbial diversity and metagenomic analysis of the rhizosphere of para grass (Urochloa mutica) growing under saline conditions. Pak J Bot 48:779–791
Mukhtar S, Ishaq A, Hassan S, Mehnaz S, Mirza MS, Malik KA (2017) Comparison of microbial communities associated with halophyte (Salsola stocksii) and non-halophyte (Triticum aestivum) using culture-independent approaches. Pol J Microbiol 66:375–386
Mukhtar S, Mirza MS, Mehnaz S, Mirza BS, Malik KA (2018) Diversity of Bacillus-like bacterial community in the rhizospheric and non-rhizospheric soil of halophytes (Salsola stocksii and Atriplex amnicola) and characterization of osmoregulatory genes in halophilic Bacilli. Can J Microbiol 64:567–579
Narasingarao P, Podell S, Ugalde JA, Brochier-Armanet C, Emerson JB, Brocks JJ, Heidelbert KB, Banfield JF, Allen EE (2012) De novo metagenomic assembly reveals abundant novel major lineage of Archaea in hypersaline microbial communities. ISME J 6:81–93
Naz I, Bano A, Hassan T (2009) Morphological, biochemical and molecular characterization of rhizobia from halophytes of Khewra salt range and Attock. Pak J Bot 41(6):3159–3168
Niemho N, Suriyachadkun C, Tamura T, Thawai C (2013) Asanoa siamensis sp. nov., isolated from soil from a temperate peat swamp forest. Int J Syst Evol Microbiol 63(1):66–71
Olsen SR, Cole CV, Watanabe FS, Dean LA (1954) Estimation of available phosphorus in soils by extraction with sodium bicarbonate. USDA Circular, 939. USDA, Washington, DC, pp 1–19
Pan Y, Cassman N, Hollander M, Mendes LW, Korevaar H, Geerts RHEM, van Veen JA, Kuramae EE (2014) Impact of long-term N, P, K, and NPK fertilization on the composition and potential functions of the bacterial community in grassland soil. FEMS Microbiol Ecol 90(1):195–205
Rengasamy R (2006) World salinization with emphasis on Australia. J Exp Bot 57:1017–1023
Rincon-Florez VA, Carvalhais LC, Schenck PM (2013) Culture-independent molecular tools for soil and rhizosphere microbiology. Diversity 5:581–612
Rolli E, Marasco R, Vigani G, Ettoumi B, Mapelli F et al (2015) Improved plant resistance to drought is promoted by the root-associated microbiome as a water stress-dependent trait. Environ Microbiol 17:316–331
Schimel J, Balser TC, Wallenstein M (2007) Microbial stress-response physiology and its implications for ecosystem function. Ecology 88:1386–1394
Segata N, Izard J, Waldron L, Gevers D, Miropolsky L, Garrett W, Huttenhower C (2011) Metagenomic biomarker discovery and explanation. Gen Biol 12(6):R60
Tamaki H, Tanaka Y, Matsuzawa H, Muramatsu M, Meng XY et al (2011) Armatimonas rosea gen. nov., sp. nov., of a novel bacterial phylum, Armatimonadetes phyl. nov., formally called the candidate phylum OP10. Int J Syst Evol Microbiol 61:1442–1447
Taprig T, Akaracharanya A, Sitdhipol J, Visessanguan W, Tanasupawat S (2013) Screening and characterization of protease-producing Virgibacillus, Halobacillus and Oceanobacillus strains from Thai fermented fish. J Appl Pharm Sci 3:025–030
Tripathi S, Kumari S, Chakraborty A, Gupta A, Chakrabarti K et al (2006) Microbial biomass and its activities in salt-affected coastal soils. Biol Fertil Soil 42:273–277
Tsuda K, Nagano H, Ando A, Shima J, Ogawa J (2015) Isolation and characterization of psychrotolerant endospore-forming Sporosarcina species associated with minced fish meat (surimi). Int J Food Microbiol 199:15–22
Vaisman N, Oren A (2009) Salisaeta longa gen. nov., sp. nov., a red, halophilic member of the Bacteroidetes. Int J Syst Evol Microbiol 59:2571–2574
Valenzuela-Encinas C, Neria-Gonzalez I, Alcantara-Hernandez RJ, Enriquez-Aragon JA, Estrada-Alvarado I, Hernandez-Rodriguez C, Dendooven L, Marsch R (2008) Phylogenetic analysis of the archaeal community in an alkaline-saline soil of the former lake Texcoco (Mexico). Extremophiles 12:247–254
Ventosa A, Mellado E, Sanchez-Porro C, Marquez MC (2008) Halophile and halotolerant microorganisms from soils. In: ds Dion P, Nautiyal CS (eds) Microbiology of extreme soils. Springer-Verlag, Berlin, pp 87–115
Wagg C, Bender SF, Widmer F, van der Heijden MG (2014) Soil biodiversity and soil community composition determine ecosystem multifunctionality. Proc Natl Acad Sci USA 111(14):5266–5270
Walkley A, Black IA (1934) An examination of Degtjareff method for determining soil organic matter and a proposed modification of the chromic acid titration method. Soil Sci 37:29–37
Wang Q, Garrity GM, Tiedje JM, Cole JR (2007) Naive Bayesian classifier for rapid assignment of rRNA sequences into the new bacterial taxonomy. Appl Environ Microbiol 73:5261–5267
Wang Y, Sheng H, He Y, Wu J, Jiang Y, Tam N, Zhou H (2012) Comparison of the levels of bacterial diversity in freshwater, intertidal wetland, and marine sediments by using millions of illumina tags. Appl Environ Microbiol 78:8264–8271
Yanagawa K, Breuker A, Schippers A, Nishizawa M, Ijiri A, Hirai M, Takaki Y (2014) Microbial community stratification controlled by the subsea floor fluid flow and geothermal gradient at the Iheya north hydrothermal field in the Mid-Okinawa trough (integrated ocean drilling program expedition 331). Appl Environ Microbiol 80(19):6126–6135
Yancey PH (2005) Organic osmolytes as compatible, metabolic and counteracting cytoprotectants in high osmolarity and other stresses. J Exp Biol 208:2819–2830
Yousuf B, Sanadhya P, Keshri J. Jha B (2012) Comparative molecular analysis of chemolithoautitrophic bacterial diversity and community structure from coastal saline soils, Gujarat, India. BMC Microbiol 12:150
Zhang Y, Cao C, Guo L, Wu Q, Cui Z (2015) Soil properties, bacterial community composition, and metabolic diversity responses to soil salinization of a semiarid grassland in northeast China. J Soil Water Conserv 70(2):110–120
Acknowledgements
We are highly thankful to Higher Education Commission [Project # HEC (FD/2012/1843)] and Pakistan Academy of Sciences [Project # 5-9/PAS/2012/969] for research grants. We would like to express our gratitude to Mr. Mukhtar Ahmad (Assistant Professor), Dyal Singh College, Lahore, for assistance in statistical analyses. We are grateful to Prof. Ann M. Hirsch (UCLA) for the comments on the manuscript.
Author information
Authors and Affiliations
Contributions
SM: Conducted experiment and prepared manuscript; BSM: pyrosequencing and data analysis; SM: manuscript preparation; MSM: supervised research and manuscript preparation; JM: pyrosequencing and data analysis; KAM: guided in experiment plan and edited manuscript.
Corresponding author
Ethics declarations
Conflict of interest
The authors declared that they have no conflict of interest in the publication.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Mukhtar, S., Mirza, B.S., Mehnaz, S. et al. Impact of soil salinity on the microbial structure of halophyte rhizosphere microbiome. World J Microbiol Biotechnol 34, 136 (2018). https://doi.org/10.1007/s11274-018-2509-5
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s11274-018-2509-5