Abstract
The salinization of grassland in arid and semi-arid areas is a serious environmental issue in China. Halophytes show extreme salt tolerance and are grown in saline-alkaline environments. Their rhizosphere microorganisms contribute significantly to plant stress tolerance. To study bacterial and fungal community structure changes in Chinese ryegrass (Leymus chinensis) rhizosphere soil under salt and alkali stress, pot experiments were conducted with different salt and alkali stress intensities. High-throughput sequencing was conducted, and the microbial diversity, community structure, and driving factors were analyzed in rhizosphere soil. The salinization of grassland in arid and semi-arid areas is a serious environmental issue in China. Halophytes show extreme salt tolerance and grow in saline-alkaline environments. A total of 549 species of bacteria from 28 phyla and 250 species from 11 phyla of fungi were detected in the rhizosphere soil of Leymus chinensis with different saline-alkali gradients. Alpha diversity analysis along saline-alkali gradients showed that bacterial community richness and diversity were the highest in the moderate saline-alkali group (pH = 8.28, EC = 160.4 μS·cm−1), while fungi had high richness and diversity in the control group (pH = 7.35, EC = 134.5 μS·cm−1). The bacteriophyta Proteobacteria, Acidobacteria, Plantomycetes, and the eumycota Ascomycota, Basidiomycota, and Glomeromycota were found with relative abundances of more than 10%. Saline-alkali gradients had significant effects on the abundance of the bacterial and fungal groups in the rhizosphere. The distribution of bacterial colony structure was not significant at the species level (P > 0.05). However, there were significant differences in the distribution of fungal structure and considerable differences in the composition of fungal species among the moderate saline-alkali group, severe saline-alkali group, and control group (P < 0.05). Correlation analysis showed that the bacterial phylum Gemmatimonadetes had a highly significant positive correlation with pH and EC (P < 0. 01). Saline-alkali stress significantly inhibited the abundance of the bacteria Latescibacteria, Cyanobacteria, and Bacteroides, and the fungi Zoopagomycota, Mortierllomycota, and Cryptomycota (P < 0. 05). Compared with fungi, bacterial community composition was most closely correlated with soil salinization. This report provided new insights into the responses of relationships between rhizosphere soil microorganisms and salt and alkali tolerance of plants.
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
Introduction
Saline-alkali soil occurs in hundreds of countries worldwide, with a total area of 1 × 109 hm2 (Gamalero et al. 2020). The total area of saline-alkali soil in China is about 3.6 × 107 hm2, ranking third globally (Lim et al. 2017). Grassland is the largest terrestrial ecosystem on earth. China is the second-largest grassland country globally, with a total grassland area of 3.31 × 108 hm2 (Wang and Wang 2019; Ondrasek and Rengel 2021). Land desertification, alkalization, and other grassland degradation caused by unreasonable human use have affected more than 80% of the total grassland area (Li and Fang 2017). The western Songnen Plain is one of the three major regions with saline-alkali soils in the world and currently has more than 2/3 of salt-affected soils (Wang and Wang 2019). Using efficient, low-cost, and adaptable methods to restore saline-alkali soil is a challenging goal.
The plant rhizosphere is the most active microhabitat in soils and the main area for plants to obtain nutrients. Colonial bacteria, fungi, and other microorganisms form a stable community structure through cooperation and competition, which is vital for plant growth and development, disease resistance, and stress resistance (Berendsen et al. 2012). Nutritional bacteria and phosphate-solubilizing bacteria can enhance soil nutrients and promote plant growth, thereby promoting the restoration of degraded grasslands (Pii et al. 2015; Ezawa and Saito 2018). Conversely, nitrifying bacteria and denitrifying bacteria increased soil nitrogen loss (Che et al. 2017). Abdel-Fattah and Abdul-Wasea (2012) showed that wheat plants displayed a high dependence on arbuscular mycorrhizal fungi in saline soils. Leymus chinensis has good salt and alkaline tolerance, can form a community on the alkali spots of saline-alkali land, and become a constructive species with good adaptability to the saline-alkali soil of the Songnen grassland (Jin-Huan et al. 2015; Wang et al. 2015). In the restoration of saline soil by planting halophytes, plants and microorganisms have their own characteristics and are interrelated in bioremediation and pollutant treatment. Hence, the digestion of organic pollutants by combined rhizosphere microorganisms is more effective than by a single microorganism (Umesh et al. 2020). The removal of pollutants by the symbiotic system of plants and rhizosphere microorganisms (Cui et al. 2018) is of great significance to understanding the physical and chemical properties and microbial diversity of saline-alkali tolerant plant soil as well as to isolate and screening saline-alkali tolerant bacteria to improve the physical and chemical properties of saline-alkali soil and promote plant life growth and development.
Salt tolerant phytoremediation is an effective technique for improving saline soil, and microorganisms contribute significantly to plant stress resistance and soil fertility (Zhao et al. 2020a; Hou et al. 2021). Many scholars have isolated and screened excellent saline-alkali tolerant strains from Triticum aestivum, Zea mays, and other food crops, and used them for the development of subsequent inoculants (Wichern et al. 2006; Kamble et al. 2014; Li et al. 2020). However, there are relatively few studies on the community structure and dominant populations of salinity tolerant microbial communities in the rhizosphere of grassland forages (Yamamoto et al. 2018; Pankaj et al. 2020; Zhao et al. 2020b).
High throughput sequencing can quickly reveal the complexity and diversity of microbial communities in situ (Chu et al. 2017). In this study, the 16S rRNA of Leymus chinensis rhizosphere soil bacteria and fungi were sequenced by the MiSep high-throughput sequencing method to analyze the bacterial community structure of Leymus chinensis rhizosphere soils under different saline-alkali stress, to explore the relationship between bacterial community structure and environmental factors, to analyze and predict the functions of bacteria and fungi, and to link the halophytes to the host-specific microbial communities, which are of great importance for the development of phytoremediation techniques. This study provides a new theoretical basis for the growth of Leymus chinensis on the Songnen saline-alkali land and the restoration and reconstruction of degraded Leymus chinensis meadows.
Materials and methods
Experimental design
The soil used in the experiment was from the Songnen grassland, located in the saline-alkali eastern Eurasian grassland in Northeast China (44°45′ N, 123°45′ E). The main vegetation is Leymus chinensis. The soil was dug from the top 30 cm of the alkaline areas, whose pH was > 10, and the soluble salt content was higher than 0.2%. The soil was transported to a botanical garden on the campus of Jilin Jianzhu University in Changchun, Jilin Province, China. The saline-alkali soil and soil from the botanical garden were mixed in volume-based ratios of 2:1 (SS), 1:1 (SM), and one control (CK), each with three parallel samples (Table 1). Plant Leymus chinensis seeds in pots (April 20, 2020) and transplant them 2 weeks after germination. Leymus chinensis seeds were transplanted into each saline-alkali treated flowerpot with 15 plants per pot (35 cm high and 28 cm diameter at the top), and the final number of plants will be controlled at 10 plants per pot. Regular watering, remove weeds and let them grow naturally. Four months after transplanting, three plant samples were randomly collected from each flowerpot, the roots were shaken to remove the loose soil, and the residual soil was collected from the roots with a sterile brush as the rhizosphere soil (Hafees and Malik 2000). The soil samples were evenly mixed in equal quantities and divided into two parts. One part was loaded into a 15 mL centrifuge tube, put into a liquid nitrogen tank for quick freezing, and transferred to a − 80 ℃ refrigerator in the laboratory for high-throughput sequencing by Beijing Baimaike Biotechnology Co., Ltd. The other part was dried and screened naturally, sealed in a self-sealing bag, and used to determine the chemical properties of the soil.
Analytical methods
Soil environmental parameters
We determined soil physical and chemical indexes according to soil agrochemical analysis (Bao 2000). Soil pH was measured by A pH meter (PHSJ-5, Shanghai Precision & Scientific Instrument Co. Ltd., China. Electrical conductivity (EC) was measured in 5:1 water-soil extracts by a conductivity meter (DDS-307, Shanghai Precision & Scientific Instrument Co., Shanghai, China). Soil organic matter (SOM) was determined by the potassium dichromate bulk density method and total organic carbon (TOC) in soil by the potassium dichromate volumetric and external heating method.
DNA extraction and sequencing
To analyze the composition of bacterial and fungal communities in the rhizosphere samples with different saline-alkali stress treatments, bacterial and fungal profiling was carried out by Biomarker Technologies Corporation (Beijing, China) (http://www.biomarker.com.cn/). Microbial DNA from each sample was extracted by using the Soil DNA Isolation Kit (DP812, Beijing Tiangen Biochemical Technology Co., Ltd., China). A 16S full-length primers were 27F (5′-AGRGTTTGATYNTGGCTCAG-3′) and 1492R (5′-TASGGHTACCTTGTTASGACTT-3′ (Johnson & Spakowicz et al. 2019). ITS full-length primers were ITS1F (5′-CTTGGTCATTTAGAGGAAGTAA-3′) and ITS4 (5′-TCCTCCGCTTATTGATATGC-3′) (Sun et al. 2020; Tian et al. 2022). A genomic library was built by using the extracted DNA after PCR products were assessed. And the extracted DNA was sequenced on the single-molecule real-time (SMRT) sequencing platform of PacBio (Pacific Bioscience, MenloPark, CA, USA).
Data processing and analysis
The total circular consensus sequences (CCS) were generated before analysis. The procedures were as follows: CCS was exported by Smrt Link (v. 8.0), identified and filtering by using Lima (v1.7) (Martin 2011) and checked chimeras with Uchime (v. 4.2) (Edgar et al. 2011). The sequences were clustered at a 97% similarity level by Usearch (v. 10.0) (Edgar 2013), and an operational taxonomic unit (OTU) was obtained. QIIME2 was used for the classification of bacterial and fungal based on the Silva (v.132) and Unite (v.8.0) databases (Assainar et al. 2020).
The alpha diversity index (including ACE, Chao1, Simpson, and Shannon) was calculated with the Mothur software (v.1.30) (Schloss et al. 2011), and the sample dilution curve and grade relative abundance curve were drawn. Further analyses were conducted on the platform of BMKCloud (www.biocloud.net). Nonmetric multidimensional scaling (NMDS) was analyzed by using Bray–Curtis similarity (Wu et al. 2019). Network analysis of microbial communities was conducted based on Sparcc (Friedman and Alm 2012), and the top 50 genera with the highest correlation were given. Spearman correlation analysis was used to analyze the relationship between environmental factors (for example, pH, SOM, and TOC) and microorganisms (Friedman and Alm 2012).
The soil environmental parameters were expressed as mean value ± standard error (n = 3), determined by one-way analysis of variance at a 95% confidence interval. Duncan's method was used for multiple comparisons, and independent samples T-student test was performed. Statistical analysis was performed with Excel (v 2007) and SPSS (v 19.0). Significant differences between the treatments of various indexes were tested (P < 0.05), and the Origin (v 8.5) software was used for graphing.
Results
Microbial community diversity in different saline-alkali soil treatments
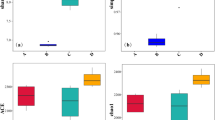
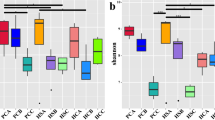
Chao1 and ACE indexes were used to measure species abundance, and Shannon and Simpson’s indexes were used to measure species diversity (Grice et al. 2009). According to Table 2, the Shannon index of the bacterial community varied between groups in the order SS > SM > CK and the Simpson index SM > SS > CK. According to the Shannon and Simpson indexes, the diversity of the bacterial community in saline-alkali soil was higher than that in the control group. The Chao index varied in the order SM > SS > CK. The order of the OTU number was the same as that of the Chao index, and the order of the ACE index was SM > CK > SS. Based on the Chao and ACE indexes of the bacterial community, it could be inferred that the rhizosphere soil in moderately saline-alkali land had the largest bacterial community richness.
The Shannon index order of the fungal community was opposite to that of bacteria. The Simpson index order of fungi was the same as that of bacteria, and the Chao index order was CK > SM > SS. The order of the fungi OTU number was the same as that of the Chao index, and the ACE index order was CK > SM > SS. The fungal alpha diversity results showed that the richness and diversity of the fungal community were the highest in the CK group and the lowest in the SS group.
Species composition of the microbial community in different saline-alkali soil treatments
According to the species distribution analysis at the phylum level, the relative abundance of bacteria in ten phyla in the rhizosphere soil of the three treatment groups was more than 1%. In terms of relative abundances, Proteobacteria, Acidobacteria, and Planctomycetes were dominant in the three treatments, with more than 10% relative abundance. Among the three groups of samples, there were differences in the abundance of Proteobacteria, Acidobacteria, Plantomycetes, Gemmatimonadetes, Actinobacteria, Bacteroidetes, Verrucomicrobia, Rokubacteria, and Chloroflex, but the abundance differences were not significant (P > 0.05) (Fig. 1). At the species level, there were ten species in the three groups, and their relative abundances were higher than 1%. The results showed that the distribution of bacterial colony structure was not significant (P > 0.05).
The relative abundance of soil fungi in the three treatment groups was more than 1%, of which Ascomycota accounted for the largest proportion, i.e., 57.37% in the SS group, 49.46% in the SM group, and 35.92% in the CK group. The relative abundance of Basidiomycota and Glomeromycota exceeded 10%. There were significant differences in the distribution of fungal colony structure (P < 0.05). In addition, there were great differences in fungal species composition between the three groups. The results of species distribution analysis showed significant differences in the distribution of fungal colony structure (P < 0.05). Moreover, there were considerable differences in fungal species composition between the three groups (Fig. 2).
Differences between microbial communities in different saline-alkali soil treatments
NMDS divided the bacteria and fungi in different degrees of saline-alkali soil into three apparent communities. In the samples of the SS, SM, and CK groups, the bacteria and fungi in the same saline-alkali group were close (Fig. 3), and the bacteria of each saline-alkali group could be clearly distinguished. There were significant differences in the composition of bacterial and fungal communities between different saline-alkali groups.
Relative abundance of the top 20 bacteria (Fig. 4) showed that uncultured_bacterium_o_RCP2-54, uncultured_bacterium_o_IMCC26256, uncultured_bacterium_o_Gaiellales, uncultured_bacterium_f_Burkholderiaceae, uncultured_bacterium_c_MB-A2-108, and uncultured_bacterium_c_BD2-11_terrestrial_group had higher relative abundance in the SS group than in other saline-alkali treatments. However, only three bacteria had higher relative abundance in the SM group, that is, uncultured_bacterium_p_FBP, uncultured_bacterium_f_SM2D12 and uncultured_bacterium_f_SM2D12. While in CK group, higher relative abundances bacteria were uncultured_bacterium_p_Latescibacteria, uncultured_bacterium_o_PLTA13, uncultured_bacterium_o_NB1-j, uncultured_bacterium_f_bacteriap25, uncultured_bacterium_c_vadinHA49 and uncultured_bacterium_f_Ilumatobacteraceae. In all groups, uncultured_bacterium_f_bacteriap25, uncultured_bacterium_f_Saprospiraceae and uncultured_bacterium_c_MB-A2-108 had highest abundance than other bacterial communities.
Relative abundance of analysis of the top 20 fungi (Fig. 4) showed that Wardomyces, Tomentella, and Preussia had higher relative abundances in the SS group than in the other saline-alkali treatments. Talaromyces, Neosetophoma, Neocosmospara, and Conocybe had higher relative abundances in the SM group than the other groups. Solicoccozyma and Podospora had higher relative abundances in the CK group than in the other groups. Preussia, Podospora, Neosetophoma, neocosmospara, and Conocybe had more abundance in all groups.
Microbial network in different saline-alkali soil treatments
The bacteria in different saline-alkali soil treatments were analyzed on the gene level (Fig. 5). The results showed that Candidatus_Udaeobacter and Ferraginibacter, beta_proteobacterium_LWH83, and Candidatus_Udaeobacter, beta proteobacterium_ LWH83 and Ferraginibacter, uncultured_bacterium_c_MB-A2-108, and uncultured_bacterium_f_ Geminicoccaceae, whose correlation coefficient of the analysis was greater than 0.9, and there was a positive correlation.
Network analysis bacteria (upper panel) and fungi species (lower panel) in different saline-alkali soil treatments. The circle represents the species, the size of the circle represents the average abundance of the species; the line represents the correlation between two species, the thickness of the line represents the strength of the correlation, the color of the line: orange represents positive correlation, green represents negative correlation
Coefficients for the correlations between Cladosporium and Fusarium, Glomus, and Acremonium, as well as Paraphoma and Discoconium, were greater than 0.9, and these correlations were negative. The correlation coefficients between Lobulomyces and Scytalidium, Spitellomyces and Syncephalis, Clonostachys and Cercophora, Tausonia, and Melanocarpus, as well as Dioszegia and Hannella, were greater than 0.9, and correlations were positive.
Correlation of environmental factors with microorganisms in different saline-alkali soil treatments
According to correlation analysis between bacteria on the phylum level and soil environmental factors (Fig. 6), Germmatimonadetes had a highly significant, positive correlation with pH and EC. Latescibacteria, Cyanobacteria, and Bacteroidetes showed a significant, negative correlation with pH and EC (P < 0.05). Total organic carbon (TOC) was positively correlated with Nitrospirae, Latencibacteria, Cyanobacteria, and Bacteroides and negatively correlated with Actinobacteria and Gemmatimonades (P < 0.05). Soil organic matter (SOM) was positively correlated with Latescibacteria and Cyanobacteria and negatively correlated with Gemmatimonadetes (P < 0.05). Zoopagomycota, Mortierellomycota, and Cryptomycota were negatively correlated with pH and EC (P < 0.05). Mortierellomycota, Cryptomycota, and Entomophthoromycota were positively correlated with TOC (P < 0.05). Mortierellomycota and Cryptomycota were positively correlated with SOM (P < 0.05).
Heatmap of environmental factors and soil microorganisms in different saline-alkali soil treatments. The saline-alkali soil and soil from the botanical garden were mixed in volume-based ratios of 2:1 (SS), 1:1 (SM), and control soil (CK). EC, electrical conductance; TOC, total organic carbon; SOM, soil organic matter. *Indicates significant difference (p < 0.05); **Indicates extremely significant difference (p < 0.01)
Discussion
Saline-alkali soil treatment effects on microbial community structure
Rhizosphere microorganisms are regulators of nutrient transformations and transport between soil and roots. Under saline-alkali stress, the growth of rhizosphere microorganisms was inhibited, and the richness of the community decreased significantly (Baumann et al. 2013). In the present study, the Shannon index, Simpson index, and OTU size showed that the diversity of the bacterial community of saline-alkali treatments was higher than that in the control group. However, the richness and diversity of the fungal community were the highest in the CK group. Compared with fungi, most bacteria were, therefore, saline-alkali tolerant. Under high saline-alkali stress, the relative abundance of the rhizosphere bacteria Proteobacteria, Acidobacteria, and Plantomycetes exceeded 10%, among which Proteobacteria had the largest relative abundance and therefore an absolute advantage. The diversity of fungi in rhizosphere soil of halophytes was not rich at the phylum level, the Ascomycota were dominant, and there were significant differences between saline-alkali groups. Zhu et al. (2020) showed that the relative abundance of Acidobacteria was highest in an original, undisturbed Leymus chinensis meadow. Zhang et al. (2017a) reported that the abundance of bacteria in the peanut rhizosphere of saline-alkali soils was higher at the phylum level, with average abundances of Proteobacteria and Actinobacteria of about 35 and 25%, respectively. Li et al. (2018) studied rhizosphere soil microorganisms of salt-tolerant plants and found that the three most abundant microorganisms of salt claw and black fruit medlar plants were Proteobacteria, Firmicutes, and Bacteroidetes. Under saline-alkali stress, the relative abundance of Ascomycota in the rhizosphere of Karelinia caspia was the highest, accounting for 94.8% (Li et al. 2021). Accordingly, we can infer that saline-alkali stress significantly inhibits the diversity of rhizosphere microorganisms. Three bacterial phyla of the Leymus chinensis rhizosphere, i.e., Proteobacteria, Acidobacteria, and Plantomycetes, as well as Ascomycota fungi, possess strong adaptability to saline-alkali soil environments, and their relative abundance increased in saline-alkali stress environments. Proteobacteria use ammonia, methane, and other nutrients produced by organic decomposition for their growth and metabolism. Increases in its relative abundance are conducive to microbial nitrogen fixation. There was no significant difference in the relative abundance of Acidobacteria with increasing saline-alkali conditions (P > 0.05), which was consistent with the results of Liu et al. (2015) on the fungal community in the black soil area of Northeast China. However, Jones et al. (2009) analyzed soil samples from North and South America and found that the relative abundance of Acidobacteria was negatively correlated with pH. Different subpopulations of Acidobacteria may, therefore, respond differently to changes in soil environmental factors.
In our study, the relative abundance of Ascomycete was higher than that of Basidiomycota. With increasing saline-alkali conditions, the abundance of Ascomycete increased significantly in our study. There were significant differences in fungal species composition between different saline-alkali treatments. These results are consistent with a previous study (Li et al. 2021), which studied the rhizosphere soil of Kalidium foliatum, Lycium ruthenicum, Karelinia Caspia and found Phragmites australis Ascomycetes were dominant. This phylum is the largest group of fungi, according to the literature. It is mainly found in saline-alkali deserts and saprophytic arid areas, and its habitats are similar to barren meadows and deserts (Li et al. 2021; HaShem et al. 2019). Glomus can adapt to a wide range of soil salinity but prefers alkaline and neutral soils (Gai et al. 2006).
Relationships between environmental factors and the soil microbial
Soil microbial community structure is affected by vegetation type, soil physical and chemical properties, latitude, soil temperature, and climate change (Zhang et al. 2017b). Plants actively select the structure of the rhizosphere soil microbial community. Plants change the rhizosphere environment through root activities to selectively enhance or reduce the abundance or diversity of some rhizosphere soil microbial groups (Ziegler et al. 2013). Soil properties are important drivers of soil bacterial community structure, but soil pH appears to be a major factor influencing community composition (Wakelin et al. 2008; Lauber et al. 2009; Ondrasek et al. 2012). Insufficient carbon sources limit the number of microorganisms in saline-alkali soil, thereby affecting the decomposition of organic matter (Kuzyakov et al. 2000). The decomposition of organic matter will provide a variety of macro and micronutrients for plant growth (Yang et al. 2009; Ondrasek et al. 2019). In this study, the correlation between rhizosphere soil microorganisms and environmental factors was assessed by correlation analysis. Different responses of bacteria and fungi to environmental factors are caused by direct competition for survival (de Boer et al. 2005). The results showed that measured pH, EC, TOC, and SOM were the main environmental factors causing differences in rhizosphere soil microbial community structure of Leymus chinensis saline-alkali land. Gemmatimonadetes was positively correlated with pH and EC and negatively correlated with TOC and SOM. The composition of bacterial communities is closely correlated with soil pH compared to fungi due to fungi requiring the use of specific compounds as substrates and having greater energy requirements (Bahram et al. 2018). Studies have shown that the abundance of Gemmatimonadetes is mainly related to soil type and environmental factors. Besides water content, soil acidity may play a secondary role in restricting Blastomonas in these soils, which is consistent with the results of the present study (Jennifer et al. 2011; Ren et al. 2020). The abundance of the phylum Gemmatimonadetes is correlated to soil drought stress (DeBruyn et al. 2011). Gemmatimonadetes clone sequences are well adapted to not only arid but also oligotrophic conditions. They can survive the saline-alkali stress and starvation, possibly forming a resting stage (Zhou et al. 2007; Pasic et al. 2010). Research on grassland soil by Nacke et al. (2011) showed that relative abundances of Bacteroidetes and Actinobacteria in the analyzed soils significantly increased with higher pH values. Saline-alkali stress significantly inhibited the abundance of the bacterial phyla Latescibacteria, Cyanobacteria, and Bacteroides, and the fungal phyla Zoopagomycota, Mortierllomycota, and Cryptomycota. Through saline-alkali research on non-halophytes of wild oats, the abundance of Mortierellomycota and Cryptomycota in non-rhizosphere soils increased (Nuccio et al. 2016). Most species of Zoopagomycota and Mortierellomycota are saprophytic and the main decomposers of soil organic carbon (Khomich et al. 2017). Microorganisms participate in the decomposition of soil organic matter, nutrient mineralization, and the formation of soil aggregates, which can indicate the quality of the soil environment. Its diversity is easily affected by pH and EC. Our results showed that different salinity was one of the reasons for differences in dominant microbial groups. The pH value and EC synergistically inhibit the growth of bacteria, such as Latescibacteria, Cyanobacteria, Bacteroides, and fungi, such as Zoopagomycota, Mortierllomycota, and Cryptomycota, resulting in the decline of community diversity. These functional microorganisms enriched in plant rhizosphere can directly characterize soil nutrient transformation and the environmental adaptability of microorganisms.
Conclusions
Saline-alkali stress enhanced soil ion concentrations and reduced available soil nutrients, resulting in changes in the diversity of the microbial community in the rhizosphere of Leymus chinensis. Fungi were more susceptible to saline-alkali stress than the bacteria based on diversity indexes ACE and Chao1. The bacterial colony structure distribution at the species level did not differ among the saline-alkali treatment groups (P > 0.05). However, the composition of fungal species was quite different between the three soil groups (P < 0. 05). Saline-alkali stress was positively correlated with Gemmatimonadetes. Therefore, it was suggested to screen the halo-tolerant plant Leymus chinensis growth-promoting rhizobacteria from the perspective of soil bacteria. Here, we provided new knowledge on the mechanisms involved in the effects of different levels of saline-alkali conditions on the soil microbial diversity and community of the saline-alkali soil rhizosphere of Leymus chinensis.
Data availability
The datasets used and/or analyzed during the current study are available from the corresponding author on reasonable request.
References
Abdel-Fattah GM, Abdul-Wasea AA (2012) Arbuscular mycorrhizal fungal application to improve growth and tolerance of wheat Triticum aestivum L plants grown in saline soil. Acta Physiol. Plant. 34(1):267–277
Assainar SK, Abbott LK, Mickan BS et al (2020) Polymer-coated rock mineral fertilizer has potential to substitute soluble fertilizer for increasing growth, nutrient uptake, and yield of wheat. Biol Fertil Soils 56(3):381–394
Bahram M, Hildebrand F, Forslund SK et al (2018) Structure and function of the global topsoil microbiome. Nature 560(7717):233–237
Bao SD (2020) Soil agrochemical analysis. China Agriculture Press, Beijing, China (In Chinese)
Baumann K, Dignac MF, Rumpel C et al (2013) Soil microbial diversity affects soil organic matter decomposition in a silty grass land soil. Biogeochemistry 114:201–212
Berendsen RL, Pieterse C, Bakker P (2012) The rhizosphere microbiome and plant health. Trends Plant Sci 17(8):478–486
Che R, Wang F, Wang W et al (2017) Increase in ammonia-oxidizing microbe abundance during degradation of alpine meadows may lead to greater soil nitrogen loss. Biogeochemistry 136(3):341–352
Chu DM, Ma J, Prince AL et al (2017) Maturation of the infant microbiome community structure and function across multiple body sites and in relation to mode of delivery. Nat Med 23(3):314–326
Cui JT, Li YN, Wang CY et al (2018) Characteristics of the rhizosphere bacterial community across different cultivation years in saline-alkaline paddy soils of Songnen Plain of China. Can J Microbiol 64:925–936
de Boer W, Folman LB, Summerbell RC et al (2005) Living in a fungal world: impact of fungi on soil bacterial niche development. FEMS Microbiol 29:795–811
DeBruyn JM, Nixon LT, Fawaz MN et al (2011) Global biogeography and quantitative seasonal dynamics of Gemmatimonadetes in soil. Appl Environ Microbiol 77:6295–6300
Ezawa T, Saito K (2018) How do arbuscular mycorrhizal fungi handle phosphate? new insight into fine-tuning of phosphate metabolism. New Phytol 220(4):1116–1121
Edgar RC, Haas BJ, Clemente JC et al (2011) UCHIME improves sensitivity and speed of chimera detection. Bioinformatics 27:2194–2200
Edgar RC (2013) UPARSE: highly accurate OTU sequences from microbial amplicon reads. Nat Methods 10:996–998
Friedman J, Alm EJ (2012) Inferring correlation networks from genomic survey data. PLoS Comput Biol. 8(9):e1002687
Gai JP, Christie P, Feng G et al (2006) Twenty years of research on community composition and species distribution of arbuscular mycorrhizal fungi in China: a review. Mycorrhiza 16(4):229–239
Gamalero E, Bona E, Todeschini V et al (2020) Saline and arid soils: impact on bacteria, plants, and their interaction. Biology 9(6):116–142
Grice EA, Kong HH, Conlan S et al (2009) Topographical and temporal diversity of the human skin microbiome. Science 324(5931):1190–1192
Hafees FY & Malik KA (2000) Manual on biofertilizer technology. Pakistan: Nibge.
Hashem A, Tabassum B, Abd_Allah EF, (2019) Bacillus subtilis: a plant-growth promoting rhizobacterium that also impacts biotic stress. Saudi J Biol Sci 26(6):1291–1297
Hou YL, Zeng WZ, Hou ML et al (2021) Responses of the soil microbial community to salinity stress in maize fields. Biology 10:1114
Jennifer MD, Lauren TN, Mariam NF et al (2011) Global biogeography and quantitative seasonal dynamics of gemmatimonadetes in soil. Appl Environ Microbiol 77(17):6295–6300
Jin-Huan L, Anjum SA, Mei-Ru L et al (2015) Modulation of morpho-physiological traits of Leymus Chinensis (Trin) through exogenous application of brassinolide under salt stress. J Anim Plant Sci 25(4):1055–1062
Jones RT, Robeson MS, Lauber CL et al (2009) A comprehensive survey of soil acidobacterial diversity using pyrosequencing and clone library analyses. ISME J 3(4):442–453
Johnson JS, Spakowicz DJ, Hong BY et al (2019) Evaluation of 16S rRNA gene sequencing for species and strain-level microbiome analysis. Nat Commun 10(1):1–11
Kamble PN, Gaikwad VB, Kuchekar SR et al (2014) Microbial growth, biomass, community structure and nutrient limitation in high pH and salinity soils from Pravaranagar (India). Eur J Soil Biol 65:87–95
Khomich M, Davey L, Kauserud H et al (2017) Fungal communities in Scandinavian lakes along a longitudinal gradient. Fungal Ecol 27:36–46
Kuzyakov Y, Friedel JK, Stahr K (2000) Review of mechanisms and quantification of priming effects. Soil Biol Biochem 32:1485–1498
Lauber CL, Hamady M, Knight R et al (2009) Pyrosequencing-based assessment of soil pH as a predictor of soil bacterial community structure at the continental scale. Appl Environ Microbiol 75:5111–5120
Lim SJ, Shin MN, Son JK et al (2017) Evaluation of soil pore-water salinity using a Decagon GS3 sensor in saline-alkali reclaimed tidal lands. Comput Electron Agric 132:49–55
Li MY, Wang JL, Zhou Q et al (2021) Analysis on the rhizosphere fungal community structure of four halophytes in Southern Xinjiang. Acta Ecol Sin 41(21):1–12 ((In Chinese))
Li JD, Fang JY (2017) Study on the ecological function of Chinese grassland. Science Press, Beijing, China (In Chinese)
Liu JJ, Sui YY, Yu ZH et al (2015) Soil carbon content drives the biogeographical distribution of fungal communities in the black soil zone of northeast China. Soil Bio Biochem 83:29–39
Li XZ, Sun P, Zhang YN et al (2020) A novel pgpr strain kocuria rhizophila y1 enhances salt stress tolerance in maize by regulating phytohormone levels, nutrient acquisition, redox potential, ion homeostasis, photosynthetic capacity and stress-responsive genes expression. Environ. Exp. Bot. 174:104023
Li Y, Yang XD, Qin L et al (2018) The bacterial diversity and community structures in rhizosphere soil of two halophytes, Lycium ruthenicum and Kalidium capsicum. Acta Ecol Sin 38(9):3118–3131 ((In Chinese))
Martin M (2011) Cutadapt removes adapter sequences from high-throughput sequencing reads. Embnet 17:10–12
Nacke H, Thürmer A, Wollherr A et al (2011) Pyrosequencing-based assessment of bacterial community structure along different management types in German forest and grassland soils. PLoS One 6(2):e17000
Nuccio EE, Anderson-Furgeson J, Estera KY et al (2016) Climate and edaphic controllers influence rhizosphere community assembly for a wild annual grass. Ecology 97(5):1307–1318
Ondrasek G, Begić HB, Zovko M et al (2019) Biogeochemistry of soil organic matter in agroecosystems & environmental implications. Sci Total Environ 658:1559–1573
Ondrasek G, Rengel Z (2021) Environmental salinization processes: detection, implications & solutions. Sci. Total Environ. 754(39):142432
Ondrasek G, Rengel Z, Romic D et al (2012) Salinity decreases dissolved organic carbon in the rhizosphere and increases trace element phyto-accumulation. Eur J Soil Sci 63(5):685–693
Pankaj U, Singh DN, Mishra P et al (2020) Autochthonous halotolerant plant growth-promoting rhizobacteria promote bacoside a yield of Bacopa monnieri (L) Nash and phytoextraction of salt-affected soil. Pedosphere 30(5):671–683
Pasic L, Kovce B, Sket B et al (2010) Diversity of microbial communities colonizing the walls of a Karstic cave in Slovenia. FEMS Microbiol Ecol 71:50–60
Pii Y, Mimmo T, Tomasi N et al (2015) Microbial interactions in the rhizosphere beneficial influences of plant growth-promoting rhizobacteria on nutrient acquisition process. a review Biol Fertil Soil 51(4):403–415
Ren H, Huang B, Fernández-García V et al (2020) Biochar and rhizobacteria amendments improve several soil properties and bacterial diversity. Microorganisms 8(4):502–519
Schloss PD, Gevers D, Westcott SL (2011) Reducing the effects of PCR amplification and sequencing artifacts on 16S rRNA-based studies. PLoS One. 6
Sun T, Miao J, Saleem M et al (2020) Bacterial compatibility and immobilization with biochar improved tebuconazole degradation, soil microbiome composition and functioning. J Hazard Mater 398:122941
Tian Y, Liu Y, Yue L et al (2022) Bacterial inoculant and sucrose amendments improve the growth of rheum palmatum L. by reprograming its metabolite composition and altering its soil microbial community. Int J Mol Sci 23(3):1694
Umesh PD, Narain SP, Mishra P et al (2020) Autochthonous halotolerant plant growth-promoting rhizobacteria promote bacoside a yield of bacopa monnieri (l.) nash and phytoextraction of salt-affected soil. Pedosphere 30(5):97–109
Wakelin SA, Macdonald LM, Rogers SL et al (2008) Habitat selective factors influencing the structural composition and functional capacity of microbial communities in agricultural soils. Soil Biol Biochem 40:803–813
Wang DL, Wang L (2019) A new perspective on the concept of grassland management. Chin Sci Bull 64(11):1106–1113
Wang LX, Fang C, Wang K (2015) Physiological responses of LeymusChinensis to long-term salt, alkali and mixed salt-alkali stresses. J Plant Nutr 38(4):526–540
Wichern J, Wichern F, Joergensen RG (2006) Impact of salinity on soil microbial communities and the decomposition of maize in acidic soils. Geoderma 137:100–108
Wu PF, Li DX, Kong LF et al (2019) The diversity and biogeography of microeukaryotes in the euphotic zone of the northwestern Pacific Ocean. Sci Total Environ 698:134289
Yamamoto K, Shiwa Y, Ishige T et al (2018) Bacterial diversity associated with the rhizosphere and endosphere of two halophytes: Glaux maritima and Salicornia europaea. Front Microbiol 9:2878
Yang J, Kloepper JW, Ryu CM (2009) Rhizosphere bacteria help plants tolerate abiotic stress. Trends Plant Sci 14(1):1–4
Zhang Y, Liu X, Cong J et al (2017) The microbially mediated soil organic carbon loss under degenerative succession in an alpine meadow. Mol Ecol 26(14):3676–3686
Zhang ZM, Ci DW, Zhang GC et al (2017) Diversity of microbial community structure in the spermosphere of saline-alkali soil in Shandong area. Acta Microbiol Sin 57(4):582–596
Zhao S, Liu JJ, Banerjee S et al (2020) Biogeographical distribution of bacterial communities in saline agricultural soil. Geoderma 361:114095
Zhao Y, Liu Z, Wu J (2020) Grassland ecosystem services: a systematic review of research advances and future directions. Landsc Ecol 35:1–22
Zhou J, Gu Y, Zou C et al (2007) Phylogenetic diversity of bacteria in an earth-cave in Guizhou province, southwest of China. J Microbiol 45:105–111
Zhu RF, Liu JL, Wang JL et al (2020) Study on bacterial community diversity and environmental factors in rhizosphere soil of Leymuschinensis. Acta Agrestia Sinica 28(3):652–660
Ziegler M, Engel M, Welzl G et al (2013) Development of a simple root model to study the effects of single exudates on the development of bacterial community structure. J Microbiol Methods 94(1):30–36
Funding
This work was supported by the Department of Science and Technology of Jilin Province [grant number 20200403004SF]; the Jilin Province Development and Reform Commission [grant number 2019C056-6]; the Education Department of Jilin Province [JJKH20210283KJ].
Author information
Authors and Affiliations
Contributions
Binshuo Liu: writing–original draft. Yunhang Hu: investigation. Ying Wang: data analysis. Honghai Xue: writing–review and editing. Zhonghe Li and Ming Li: conceptualization and methodology.
Corresponding author
Ethics declarations
Ethics approval
Not applicable.
Consent to participate
Not applicable.
Consent for publication
Not applicable.
Competing of interest
The authors declare no competing interests.
Additional information
Responsible Editor: Robert Duran
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Liu, B., Hu, Y., Wang, Y. et al. Effects of saline-alkali stress on bacterial and fungal community diversity in Leymus chinensis rhizosphere soil. Environ Sci Pollut Res 29, 70000–70013 (2022). https://doi.org/10.1007/s11356-022-20270-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11356-022-20270-6