Abstract
Biomass production is currently explored in microalgae, macroalgae and land plants. Microalgal biofuel development has been performed mostly in green algae. In the Japanese tradition, macrophytic red algae such as Pyropia yezoensis and Gelidium crinale have been utilized as food and industrial materials. Researches on the utilization of unicellular red microalgae such as Cyanidioschyzon merolae and Porphyridium purpureum started only quite recently. Red algae have relatively large plastid genomes harboring more than 200 protein-coding genes that support the biosynthetic capacity of the plastid. Engineering the plastid genome is a unique potential of red microalgae. In addition, large-scale growth facilities of P. purpureum have been developed for industrial production of biofuels. C. merolae has been studied as a model alga for cell and molecular biological analyses with its completely determined genomes and transformation techniques. Its acidic and warm habitat makes it easy to grow this alga axenically in large scales. Its potential as a biofuel producer is recently documented under nitrogen-limited conditions. Metabolic pathways of the accumulation of starch and triacylglycerol and the enzymes involved therein are being elucidated. Engineering these regulatory mechanisms will open a possibility of exploiting the full capability of production of biofuel and high added-value oil. In the present review, we will describe the characteristics and potential of these algae as biotechnological seeds.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Biomass production has been important resources for human activity both as nutrition, materials and energy source. Traditionally, crop plants served as nutritional sources, whereas woody plants or trees were used as construction materials and fuels. Algae have rarely been regarded as probable materials for either nutrition or construction, while no one had ever imagined using algae as fuels before the break of current trends in biofuel production (Menetrez 2012; De Bhowmick et al. 2015). Currently, extensive efforts have been made on the biotechnology of microalgae, notably, green algae such as Chlamydomonas reinhardtii (Merchant et al. 2012; Li-Beisson et al. 2015), or marine algae such as Nannochloropsis (Pal et al. 2011; Iwai et al. 2015).
In the Japanese tradition, however, people have long been exploiting various macrophyte marine algae from the beginning of the ancient imperial era (more than 1300 years ago as documented in the oldest chronicle Kojiki published in 712): the red alga Pyropia yezoensis known as “(Asakusa) Nori” (“Asakusa” refers to the algae grown in Tokyo Bay in Edo Era), the brown algae in Laminariaceae, notably Saccharina japonica known as “Konbu”, the brown algae Sargassum fusiforme (formerly Hizikia) known as “Hiziki” and Undaria pinnatifida known as “Wakame” are still common food stuff in daily Japanese life (Fig. 1). The red algae in Gelidiaceae, typically Gelidium crinale known as “Tengusa”, are used to produce agar or “Kanten”, which is used as gelling additives in food production or industry (as well as in microbiology laboratories!). Cyanobacterial mat of Aphanothece sacrum known as “Suizenji nori” is at the crisis of extinction in the natural habitats, but has been used as delicacy in the local cuisine.
Marine macro algae used in daily Japanese meal. a Nori (Pyropia yezoensis) as dried sheets (left) and Nori rolls (right) with rice and Kanpyo (a cucurbit Lagenaria siceraria var. hispida) inside, a kind of Sushi; b Konbu (Saccharina japonica) as dried thalli (left) and Kobumaki (right), cooked in soy sauce and sugar and served with Kanpyo (a dish for New Year); c dried Hiziki (Sargassum fusiforme), commonly cooked with soy beans in soy sauce and sugar; d dried Wakame (Undaria pinnatifida), commonly used in soy soup for Japanese breakfast
In spite of this tradition of seaweed or algal consumption, Japanese people did not try to use seaweeds or algae as fuels, not only because marine products are wet, but also due to their low content of oil. They were mainly used as food and mineral resources. This does not mean that the algae do not produce oil. In the present review, we will present recent understandings on the lipid biosynthesis and the production of triacylglycerol (TAG) in red algae, or specifically in red microalgae in the hope of using this knowledge in biotechnological developments. Oils can be used as either high added-value products or biofuel depending on its composition. Readers are referred to some reviews on algal lipids in general (see e.g., Guschina and Harwood 2006; Qin et al. 2012; Zienkiewiz et al. 2016).
Two types of red algal oils
Red algae are taxonomically classified into two classes, Cyanidiales that include unicellular species that are blue-green in appearance and found in hot springs (such as Cyanidioschyzon merolae) and Rhodophytina that include marine species of both unicellular (such as Porphyridium) and macrophyte (such as Pyropia) forms. Apart from taxonomic classification, there are two types of red algae that produce different kinds of oil. Most marine red algae (and some Cyanidiales species) produce oil containing high levels of polyunsaturated fatty acids (PUFA), whereas the oil of C. merolae contains no PUFA and suited for biofuel. PUFA such as arachidonic (20:4 or ARA) and eicosapentaenoic (20:5 or EPA) acids are more suited for nutrients or healthy products rather than biofuel. We first explain general characteristics of lipid biosynthesis in red algae, and then present individual red algae and their potential biotechnological use.
Genomics-based elucidation of lipid biosynthesis in red algae
Composition and biosynthesis of lipids have been intensively studied in model algae such as Chlamydomonas reinhardtii (Merchant et al. 2012; Sakurai et al. 2014; Li-Beisson et al. 2015; Zienkiewicz et al. 2016) or related species such as C. debaryana (Toyoshima and Sato 2015), but understanding of entire pathways of biosynthesis of lipids, including TAG, was quite limited in red algae until recently. Simple lists of genes involved in lipid synthesis have been repeatedly presented: such as Riekof et al. (2005) and Li-Beisson et al. (2015) for C. reinhardtii, Sato and Moriyama (2007) for C. merolae, and Misra et al. (2012) for several algal species including these algae and two species of Ostreococcus.
Genomic data are the key information for the estimation of metabolic pathways and hence metabolic engineering. All the 121 genes involved in lipid metabolism in C. merolae were estimated (Mori et al. 2016) by comparative genomics based on homolog clustering (Sato 2009). Intracellular localization of 113 enzymes involved in the metabolism of fatty acids and lipids in C. merolae (excepting those encoded by the plastid genome) was studied by Mori et al. (2016) using transient expression of GFP fusion proteins. The results confirmed the previous results of Sato and Moriyama (2007), and enabled construction of the complete metabolic map of lipid metabolism in C. merolae, which is essentially applicable to other red algae in Cyanidiales, as well as to red algae in general with minor modifications (Fig. 2). We believe that this model on red algae can even be extended, with minor modifications, to Chromophyta (brown algae, diatoms, Eustigmatophyceae etc) that originated by secondary endosymbiosis of red algal cell (Cavalier-Smith 2003). In this model, some interesting characteristics emerge: lack of stearate desaturation in the plastid and phycobilisomes as a source of TAG accumulation.
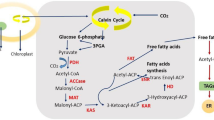
Generalized pathway of lipid biosynthesis in red algae. a Generalized lipid metabolism in red algae. The model was based on Mori et al. (2016), but modified and simplified according to the comparative genomic data (Table S1). b Comparison of fatty acid elongation (FAE) and desaturation (Des) pathways in five red algae. Note that the pathway starts from 18:0, but not 18:1, because stearoyl-ACP desaturase is not present in the plastid in red algae. Δ + number indicates the position of desaturation. Abbreviations that are not defined in the text (see also legends for Tables 2 and 3): G3P glycerol 3-phosphate, CDP cytidine diphosphate, DAG diacylglycerol, PGP phosphatidylglycerol phosphate, LPA lysophosphatidic acid, LPC lysophosphatidylcholine, SCD stearoyl-CoA Δ9-desaturase
Lack of stearate desaturation in the red algal plastid
The most important trait of red algae in lipid biosynthesis is the lack of stearoyl acyl-carrier-protein (ACP) desaturase (Fig. 2a, Table S1), which is ubiquitous in green algae and land plants, as well as some actinobacteria (see Cluster 1324 of Dataset Gclust2012_42 in the Gclust comparative genomic database at http://gclust.c.u-tokyo.ac.jp/, Sato 2009). Since this soluble enzyme is localized in the plastid stroma, the primary products of plastid fatty acid synthesis (FAS) are palmitic acid (16:0) and oleic acid (18:1) in green algae and plants (See the legends for Tables 2, 3 for the names of lipids and fatty acids). The plastid FAS in red algae produces saturated acids, such as 16:0 and stearic acid (18:0), which are then transported to cytosol, and activated to become acyl CoAs, which are further used in acyl lipid synthesis, desaturation or elongation in ER. The products of elongation and desaturation are dependent on algal species (Fig. 2b): In C. merolae, the end product of elongation and desaturation is 20:2, whereas trienoic acids are also produced in G. sulphuraria. In marine red algae, the products are predominantly C20 PUFA, such as ARA and EPA (Guschina and Harwood 2006), which are then transported back to plastids for the synthesis of galactolipids, which retain most of ARA and EPA within the cell.
Many previous labeling studies used labeled 18:1 or other unsaturated fatty acids as precursors (Nichols and Appleby 1969; Shiran et al. 1996; Khozin et al. 1997) to show the desaturation and elongation pathways, which are now turned out to be localized in the ER, or outside plastids. That is why a flow of fatty acids from TAG to monogalactosyl diacylglycerol (MGDG) was detected in P. cruentum (Khozin-Goldberg et al. 2000). In contrast, labeling with acetate resulted in rapid labeling of 16:0 in MGDG in C. merolae (Sato and Moriyama 2007). Linoleic acid (18:2) is, however, provided for MGDG synthesis by the rapid turnover of phosphatidylcholine (PC) in the ER (Sato and Moriyama 2007; Sato et al. 2016; Toyoshima et al. 2016). The collaboration of plastid and ER for the synthesis of plastid MGDG is a unique characteristic of this alga.
The lack of stearoyl ACP desaturase is universal in red algae and Chromophyta. The abundant ARA and EPA, or docosahexaenoic acid (22:6 or DHA) present in marine red algae or chromophyte algae are all produced by the elongation and desaturation pathway starting from 18:0 (but not from 18:1, as usually mentioned) in the ER. The only desaturase present in the red algal plastids is FAD4 (Cluster 7570 in Gclust database; Gao et al. 2009), which introduces a trans double bond in 16:0 to produce ∆3-trans-hexadecenoic acid (16:1). This acid is specifically present at the sn-2 position of plastid phosphatidylglycerol (PG) in all photosynthetic eukaryotes, and the desaturation is believed to occur within PG molecule.
Phycobilisomes as a source of TAG accumulation
TAG is accumulated under nitrogen deprivation in C. merolae, as in many other algae (Fig. 3f, g; Toyoshima et al. 2016; Takusagawa et al. 2016). Both oil bodies and starch granules are accumulated in the cytosol, because starch (or glycogen) synthesis takes place in the cytosol in red algae (Ball et al. 2011). Comparison of fatty acid composition suggested that the acyl groups for the synthesis of TAG comes from PC, or acyl CoA pool which is in rapid equilibrium with PC. Labeling studies on C. merolae using [32P]phosphate indicated slow labeling of PC (Sato et al. 2016), whereas the acyl groups are rapidly turned over (Sato and Moriyama 2007). This indicates important metabolic roles of PC in desaturation and lipid remodeling to provide fatty acids for TAG synthesis (Sato et al. 2016; Toyoshima et al. 2016). Similar active turnover of PC is known in plants (Bates 2016). In contrast with C. reinhardtii, nitrogen deprivation did not result in large-scale degradation of plastid lipids or global degeneration of plastids in C. merolae at least in the initial phase (Toyoshima et al. 2016). Phycobilisomes (light-harvesting pigment-protein complexes on the thylakoid membranes) are, however, degraded to provide nitrogen source for the cellular metabolism and carbon source for the synthesis of lipids and starch. The conversion of phycobilisomes to storage materials could be a good strategy of efficient oil production in red algae.
Unicellular model red algae. Upper panels (a–g), Cyanidioschyzon merolae; lower panels (h, i), Porphyridium purpureum a, b, h are fluorescence micrographs of 4′,6-diamidino-2-phenylindole (DAPI)-stained cells; c, d, i are corresponding differential interference images. Panels b, d show a dividing cell. Panels e–g are electron micrographs of C. merolae cells in stationary culture (e), and after nitrogen deprivation for 2 days (f, g). Cp chloroplast (plastid), Mt mitochondrion, N nucleus, O oil body, S starch granule
Comparative genomics of lipid biosynthesis
Enzymes involved in major lipid biosynthesis in C. merolae were estimated from the genome sequences (Sato and Moriyama 2007; Mori et al. 2016). As a result, enzymes for the fatty acid and lipid biosynthesis were identified. The list of all genes involved in lipid metabolism was extended to include five red algae, and the results are provided as Table S1. In the current map, we do not include betaine lipid biosynthesis, because this pathway was not detected in the complete genome sequences (Table S1). The comparative genomic database for red algae is available as Dataset Gclust2016R in the Gclust server.
Various red algae and their biotechnological potentials
Table 1 presents comparison of model red algae with model green algae, Chlamydomonas reinhardtii and their relatives. Roughly speaking, the two types of algae are comparable in the capacity of production of oil and carbohydrate. But the actual growth conditions are quite different, and this difference can be exploited for better cultivation. We briefly explain characteristics of representative red algae.
Cyanidioschyzon merolae
Cyanidioschyzon merolae is a small unicellular red alga, having a very simple cell structure, comprising one each of mitochondrion, plastid, and microbody per cell (Fig. 3a–e). It lives in acidic hot springs containing sulfuric acid (at about 40–50 °C, at pH 1.5–2.5). This growth condition allows culturing under open air without special cares such as autoclaving of the medium. Tolerance to high concentration (up to 100%) of CO2 as well as nitrate and sulfate/sulfite allows use of exhaust gas of industry for the culture. Temperatures higher than 40 °C favorable for its growth can be obtained also by exhaust heat from the industry, but this alga can also grow at ambient temperatures. All these properties are suited for its use in biotechnology. The nuclear (16,546,747 bp), mitochondrial (32,211 bp) and plastid (149,987 bp) genomes have been completely sequenced (Matsuzaki et al. 2004; Nozaki et al. 2007). Molecular genetic tools for genetic manipulation are now available for transient expression of fluorescent proteins (Ohnuma et al. 2014; Moriyama et al. 2014a, b) as well as targeted gene disruption (Fujiwara et al. 2013).
Lipid profile of C. merolae is quite simple (Table 2). Unlike many other eukaryotes, phosphatidylserine (PS) and cardiolipin (CL) are not detected (Sato and Moriyama 2007). TAG accumulates under nitrogen depletion (Table 2; Toyoshima et al. 2016; Takusagawa et al. 2016). Major fatty acids are 16:0, 18:1 and 18:2. No PUFAs are formed even at low temperatures (Sato and Moriyama 2007; Toyoshima et al. 2016: Table 3). The TAG accumulated under nitrogen deprivation is also rich in the three fatty acids, namely, 16:0, 18:1 and 18:2, and is suitable for biodiesel after conversion to fatty acid methyl esters. Currently, this is the only red alga that can produce TAG for biofuel. Because of its simple pathway of lipid synthesis, this alga could be a good seed for further manipulation in biotechnology.
Porphyridium purpureum
Porphyridium purpureum (previously cruentum) is a marine unicellular red alga, which has also been studied for a long time in basic studies (Fig. 3h, i). The nuclear genome (19.7 Mbp; Bhattacharya et al. 2013) and the plastid genome (217,694 bp; Tajima et al. 2014) have been sequenced. Genetic manipulation in P. purpureum was reported (Lapidot et al. 1999, 2002), but no further attempts have been made. The plastid genome of the red algae in general has a large capacity of protein-coding genes (more than 200, see Tajima et al. 2014). Red algal plastids retain many prokaryotic enzymes involved in the gene expression, and in this respect, they are distinct from green algae and plants (Sato 2001). This could be a target for engineering not only in P. purpureum (Lapidot et al. 2002) but also in C. merolae.
As this alga excretes highly viscous materials, characterization and synthesis of polysaccharides have been studied (Merchuk et al. 1998; Geresh et al. 2009). Porphyridium was found to contain semi-amylopectin rather than glycogen that was found in Cyanidium and Galdieria (Shimonaga et al. 2008). Lipid profile of P. purpureum (Khozin-Goldbert et al. 2000) is qualitatively similar to that of C. merolae (Table 2). In contrast, fatty acid profile is quite different: P. purpureum contains high levels of ARA and EPA (Table 3; Nichols and Appleby 1969; Ohta et al. 1992; Oh et al. 2009). Putative genes for the desaturases producing ARA and EPA were identified (Table S1). Detailed structural analysis of SQDG by mass spectrometry was reported (Naumann et al. 2007).
Process engineering of P. purpureum was studied in Posten laboratory in Germany (Fleck-Schneider et al. 2007; Sastre 2010). They developed a large plant for mass cultivation of this alga using sunlight, with least possible energy input for stirring and aeration. A problem of P. purpureum could be the production of viscous mucilage, which prevents harvesting and processing. We found that the cells can be grown at a reasonable rate at a temperature as high as 35 °C (Tajima et al. 2014), while the production of mucilage was reduced, although the color of the cells turns orange, in contrast with dark red at lower temperatures.
Other red algae
Galdieria sulphuraria is another thermophilic red alga belonging to Cyanidiales. Massive horizontal gene transfer (HGT) was found to contribute to the extremophilic properties of this alga (Schönknecht et al. 2013). HGT is not common in other thermophilic algae in Cyanidiales, such as C. merolae and Cyanidium caldarium (Ciniglia et al. 2004). This could be the reason for the facultative heterotrophy of G. sulphuraria (Sakurai et al. 2016). Detailed lipidomic analysis (Vítová et al. 2016) indicated that this alga also contains uncommon lipids, such as diacylglyceryl-N,N,N-trimethylhomoserine (DGTS), diacylglyceryl-O-2′-(hydroxymethyl)-(N,N,N-trimethyl)-β-alanine (DGTA), and phosphosulfocholine (PSC) (Table 2). DGTS and DGTA are known as betaine lipids, which were initially not detected in red algae (Sato 1992), but later reported to be present in several species of red algae (Dembitsky 1996). Curiously, however, the gene BTA1 encoding the enzyme catalyzing the biosynthesis of DGTS (Riekhof et al. 2005), or its bacterial homologs, btaA and btaB (the corresponding two domains are fused in BTA1), was not detected in the genome sequences of G. sulphuraria or Chondrus crispus (Table S1). Another point on the lipid composition of G. sulphuraria as reported by Vítová et al. (2016) is the unusual paucity of common plastid lipids, MGDG and DGDG, in autotrophic culture, in which large plastids must be present (Fig. 1 in Schönknecht et al. 2013). Further studies will be needed to elucidate these paradoxes. Differences in lipid and fatty acid composition in various growth conditions (Sakurai et al. 2016) were reported. Notably, TAG level markedly increased (up to 11 µg mL−1) by addition of glucose in the heterotrophic culture.
We have to mention two marine macrophytic red algae that are well characterized. Lipid analysis of C. crispus was reported (Pettitt et al. 1989), but detailed structural studies were recently published (Melo et al. 2015). Based on the genomic data now available (Collén et al. 2013), we can identify enzymes involved in lipid metabolism (Table S1). Another well-characterized red alga is Pyropia (formerly Porphyra) yezoensis, which is an important food in Japan (Fig. 1a). Commercially available sheets of this alga (laver of “Nori”) are dry, but all lipid components are preserved without notable oxidation for a long time (Kayama et al. 1983). Lipid composition of Pyropia yezoensis was studied in detail (Araki et al. 1986, 1987, 1989). Lipid and fatty acid compositions in various species of Gracilaria were also reported from the same group (Araki et al. 1990). These macrophytic red algae contain high levels of ARA and EPA, especially in MGDG (Table 3). The contents in TAG are not, however, very high.
Potentials of red algae as biofuel feedstocks and high-added value products
Biofuel production in microalgae is now highlighted (De Bhowmick et al. 2015; Zienkiewicz et al. 2016), but conventional algal biotechnology focused on green algae, such as those in the genus Chlamydomonas. Table 1 compares red algae and Chlamydomonas for their potentials in biotechnology. Obviously, red algae have potentials as both biofuel feedstocks and producers of high-added value compounds. Representative values of lipid content and productivity of lipids and carbohydrates are comparable in red and green algae. Growth properties are markedly different: Cyanidiales such as C. merolae grows in acidic hot springs, and this property makes these groups of red algae unique organisms suitable for biotechnology. They are easy to grow in an acidic medium without sterilization, and exhaust gas and heat of industry can be used for their growth (Table 1). Sea water can be tolerated to some extent as stressful conditions to increase the content of oil. Marine red algae such as P. purpureum also provide good resource for biotechnology, because sea water is available everywhere in a country such as Japan. This microalga is usually grown in sterilized artificial sea water, but growth at high temperature (up to 35 °C) could be favorable for its maintenance in axenic state. In contrast, standard green algae such as Chlamydomonas reinhardtii grows in fresh water at neutral pH at ambient temperature. The growth medium must be sterilized for their growth. However, acid-tolerant strains of Chlamydomonas exist. These can be used more easily in large-scale cultivation. Growth rate of both red algae and green algae depends on light intensity. The microalgae can grow faster under the light with higher intensity, if enough CO2 is provided. In this respect, very rapid growth is achieved in C. merolae under high light with high concentration of CO2. This is realizable in open air with industrial exhaust gas and heat. In contrast, rapid growth and high yield of oil are difficult to achieve in Chlamydomonas even at high light and higher concentration of CO2.
Genetic manipulation is now possible in both red and green algae. But homologous recombination is only possible in C. merolae. A promising example of genetic modification of C. merolae is the introduction of cyanobacterial acyl-ACP reductase, which increased the accumulation of TAG (Sumiya et al. 2015). Genetic engineering in completely sequenced, unicellular microalgae has a great advantage in efficient development. The genome sizes of red algae are, in general, small (10–30 Mbp), whereas the genomes of Chlamydomonas are fairly larger, namely, about 70–140 Mbp (Hirashima et al. 2016). Although there are other microalgae having small genomes, red algae are definitely the organisms of choice for genetic modification, because paucity of introns also characterizes red algae. C. merolae has only 27 introns in the entire 4775 protein-coding nuclear genes! We do not need to isolate cDNA, and we can manipulate the genome just like prokaryotic genomes.
Red algae provide two types of organisms, one suitable for biofuels and the other suitable for high added-value products. C. merolae does not contain PUFA, and this property is good as biofuel production. The simple metabolic pathway as well as ease in genetic manipulation makes this alga a versatile photosynthetic organism to use in bioengineering. Metabolic maps of lipid metabolism (Mori et al. 2016) and carbon metabolism (Moriyama et al. 2014a) are ready to use in C. merolae. ARA and EPA are representative high added-value compounds produced by many red algae. If the flow of these acids into plastid MGDG could be switched to TAG synthesis, we would expect production of TAG enriched in ARA and EPA. The use of the desaturase genes in different organisms might also be promising.
Moreover, C. merolae serves as a model for engineering other red algae, both microalgae and macrophytes, in planning the strategy of genetic modification. Lipid biotechnology in microalgae has been mostly studied in green algae, but it now jumped up into a new era of red algal technology.
References
Araki S, Sakurai T, Omata T, Kawaguchi A, Murata N (1986) Lipid and fatty acid composition in the red alga Porphyra yezoensis. Jap. J Phycol 34:94–100
Araki S, Sakurai T, Kawaguchi A, Murata N (1987) Positional distribution of fatty acids in glycerolipids of the marine red alga, Porphyra yezoensis. Plant Cell Physiol 28:761–766
Araki S, Sakurai T, Oohusa T, Kayama M, Sato N (1989) Characterization of sulfoquinovosyl diacylglycerol from marine red algae. Plant Cell Physiol 30:775–781
Araki S, Sakurai T, Oohusa T, Kayama M, Nisizawa K (1990) Content of arachidonic and eicosapentaenoic acids in polar lipids from Gracilaria (Gracilariales, Rhodophyta). Hydrobiologia 204/205:513–519
Ball S, Colleoni C, Cenci U, Raj JN, Tirtiaux C (2011) The evolution of glycogen and starch metabolism in eukaryotes gives molecular clues to understand the establishment of plastid endosymbiosis. J Exp Bot 62:1775–1801
Bates PD (2016) Understanding the control of the acyl flux through the lipid metabolic network of plant oil biosynthesis. Biochim Biophys Acta 1861:1214–1225
Bhattacharya D, Price DC, Chan CX, Qiu H, Rose N, Ball S, Weber AP, Arias MC, Henrissat B, Coutinho PM, Krishnan A, Zäuner S, Morath S, Hilliou F, Egizi A, Perrineau MM, Yoon HS (2013) Genome of the red alga Porphyridium purpureum. Nat Commun 4:1941
Cavalier-Smith T (2003) Genomic reduction and evolution of novel genetic membranes and protein-targeting machinery in eukaryote–eukaryote chimaeras (meta-algae). Phil Trans R Soc Lond B 358:109–134
Ciniglia C, Yoon HS, Pollio A, Pinto G, Bhattacharya D (2004) Hidden biodiversity of the extremophilic Cyanidiales red algae. Mol Ecol 13:1827–1837
Collén J, Porcel B, Carré W, Ball SG, Chaparro C, Tonon T, Barbeyron T, Michel G, Noel B, Valentin K, Elias M, Artiguenave F, Arun A, Aury JM, Barbosa-Neto JF, Bothwell JH, Bouget FY, Brillet L, Cabello-Hurtado F, Capella-Gutiérrez S, Charrier B, Cladière L, Cock JM, Coelho SM, Colleoni C, Czjzek M, Da Silva C, Delage L, Denoeud F, Deschamps P, Dittami SM, Gabaldón T, Gachon CM, Groisillier A, Hervé C, Jabbari K, Katinka M, Kloareg B, Kowalczyk N, Labadie K, Leblanc C, Lopez PJ, McLachlan DH, Meslet-Cladiere L, Moustafa A, Nehr Z, Nyvall Collén P, Panaud O, Partensky F, Poulain J, Rensing SA, Rousvoal S, Samson G, Symeonidi A, Weissenbach J, Zambounis A, Wincker P, Boyen C (2013) Genome structure and metabolic features in the red seaweed Chondrus crispus shed light on evolution of the Archaeplastida. Proc Natl Acad Sci USA 110:5247–5252
De Bhowmick G, Koduru L, Sen R (2015) Metabolic pathway engineering towards enhancing microalgal lipid biosynthesis for biofuel application—A review. Renew Sustain Energy Rev 50:1239–1253
Dembitsky VM (1996) Betaine ether-linked glycerolipids: chemistry and biology. Prog Lipid Res 35:1–51
Fleck-Schneider P, Lehr F, Posten C (2007) Modelling of growth and product formation of Porphyridium purupureum. J Biotechnol 132:134–141
Fujiwara T, Ohnuma M, Yoshida M, Kuroiwa T (2013) Gene targeting in the red alga Cyanidioschyzon merolae: Single- and multi-copy insertion using authentic and chimeric selection markers. PLOS ONE 8:e73608
Gao J, Ajjawi I, Manoli A, Sawin A, Xu C, Froehlich JE, Last RL, Benning C (2009) FATTY ACID DESATURASE 4 of Arabidopsis encodes a protein distinct from characterized fatty acid desaturases. Plant J 60:832–839
Geresh S, Arad S, Levy-Ontman O, Zhang W, Tekoah Y, Glaser R (2009) Isolation and characterization of poly- and oligosaccharides from the red microalga Porphyridium sp. Carbohydrate Res 344:343–349
Guschina IA, Harwood JL (2006) Lipids and lipid metabolism in eukaryotic algae. Prog Lipid Res 45:160–186
Hirashima T, Tajima N, Sato N (2016) Draft genome sequences of four species of Chlamydomonas containing phosphatidylcholine. Genome Announc 4:e01070–16
Iwai M, Hori K, Sasaki-Sekimoto Y, Shimojima M, Ohta H (2015) Manipulation of oil synthesis in Nannochloropsis strain NIES-2145 with a phosphorus starvation-inducible promoter from Chlamydomonas reinhardtii. Front Microbiol 6:912
Karpagam R, Preeti R, Ashokkumar B, Varalakshmi P (2015) Enhancement of lipid production and fatty acid profiling in Chlamydomonas reinhardtii, CC1010 for biodiesel production. Ecotoxicol Environ Saf 121:253–257
Kayama M, Imayoshi J, Araki S, Ogawa H, Oohusa T, Ueno T, Saito M (1983) Changes in the lipids of dried laver “Nori” at different water activities. Bull Japan Soc Sci. Fisheries 49:787–793 (AbstractEnglish)
Khozin I, Adlerstein D, Bigongo C, Heimer YM, Cohen Z (1997) Elucidation of the biosynthesis of eicosapentaenoic acid in the microalga Porphyridium cruentum II. Studies with radiolabeled precursors. Plant Physiol 114:223–230
Khozin-Goldberg I, Yu HZ, Adlerstein D, Didi-Cohen S, Heimer YM, Cohen Z (2000) Triacylglycerols of the red microalga Porphyridium cruentum can contribute to the biosynthesis of eukaryotic galactolipids. Lipids 35:881–889
Lapidot M, Raveh D, Sivan A, Arad S, Shapira M (1999) Molecular analysis of the AhaS gene of Porphyridium sp. (Rhodophyta) and of a mutant resistant to sulfometuron methyl. J Phycol 35:1233–1236
Lapidot M, Raveh D, Sivan A, Arad S, Shapira M (2002) Stable chloroplast transformation of the unicellular red alga Porphyridium species. Plant Physiol 129:7–12
Li-Beisson Y, Beisson F, Riekhof W (2015) Metabolism of acyl-lipids in Chlamydomonas reinhardtii. Plant J 82:504–522
Matsuzaki M, Misumi O, Shin-i T, Maruyama S, Takahara M, Miyagishima S, Mori T, Nishida K, Yagisawa F, Nishida K, Yoshida Y, Nishimura Y, Nakao S, Kobayashi T, Momoyama Y, Higashiyama T, Minoda A, Sano M, Nomoto H, Oishi K, Hayashi H, Ohta F, Nishizaka S, Haga S, Miura S, Morishita T, Kabeya Y, Terasawa K, Suzuki Y, Ishii Y, Asakawa S, Takano H, Ohta N, Kuroiwa H, Tanaka K, Shimizu N, Sugano S, Sato N, Nozaki H, Ogasawara N, Kohara Y, Kuroiwa T (2004) Genome sequence of the ultrasmall unicellular red alga Cyanidioschyzon merolae 10D. Nature 428:653–657
Melo T, Alves E, Azevedo V, Martins AS, Neves B, Domingues P, Calado R, Abreu MH, Domingues MR (2015) Lipidomics as a new approach for the bioprospecting of marine macroalgae—unraveling the polar lipid and fatty acid composition of Chondrus crispus. Algal Res 8:181–191
Menetrez MY (2012) An overview of algae biofuel production and potential environmental impact. Environ Sci Technol 46:7073–7085
Merchant SS, Kropat J, Liu B, Shaw J, Warakanont J (2012) TAG, You’re it! Chlamydomonas as a reference organism for understanding algal triacylglycerol accumulation. Curr Opin Biotechnol 23:352–363
Merchuk JC, Ronen M, Giris S, Arad S (1998) Light/dark cycles in the growth of the red microalga Porphyridium sp. Biotechnol Bioeng 59:705–713
Misra N, Panda PK, Parida BK, Mishra BK (2012) Phylogenomic study of lipid genes involved in microalgal biofuel production—candidate gene mining and metabolic pathway analyses. Evol Bioinformatics 8:545–564
Mori N, Moriyama T, Toyoshima M, Sato N (2016) Construction of global acyl lipid metabolic map by comparative genomics and subcellular localization analysis in the red alga Cyanidioschyzon merolae. Front Plant Sci 7:958
Moriyama T, Sakurai K, Sekine K, Sato N (2014a) Subcellular distribution of central carbohydrate metabolism pathways in the red alga Cyanidioschyzon merolae. Planta 240:585–598
Moriyama T, Tajima N, Sekine K, Sato N (2014b) Localization and phylogenetic analysis of enzymes related to organellar genome replication in the unicellular rhodophyte Cyanidioschyzon merolae. Genome Biol Evol 6:228–237
Naumann I, Darsow KH, Walter C, Lange HA, Buchholz R (2007) Identification of sulfoglycolipids from the alga Porphyridium purpureum by matrix-assisted laser desorption/ionisation quadrupole ion trap time-of-flight mass spectrometry. Rapid Commun Mass Spectrom 21:3185–3192
Nichols BW, Appleby RS (1969) The distribution and biosynthesis of arachidonic acid in algae. Phytochemistry 8:1907–1915
Nozaki H, Takano H, Misumi O, Terasawa K, Matsuzaki M, Maruyama S, Nishida K, Yagisawa F, Yoshida Y, Fujiwara T, Takio S, Tamura K, Chung SJ, Nakamura S, Kuroiwa H, Tanaka K, Sato N, Kuroiwa T (2007) A 100%-complete sequence reveals unusually simple genomic features in the hot spring red alga Cyanidioschyzon merolae. BMC Biol 5:28
Oh SH, Han JG, Kim Y, Ha JH, Kim SS, Jeong MH, Jeong HS, Kim NY, Cho JS, Yoon WB, Lee SY, Kang DH, Lee HY (2009) Lipid production in Porphyridium cruentum grown under different culture conditions. J Biosci Bioeng 108:429–434
Ohnuma M, Yokoyama T, Inouye T, Sekine Y, Kuroiwa T, Tanaka K (2014) Optimization of polyethylene glycol (PEG)-mediated DNA introduction conditions for transient gene expression in the unicellular red alga Cyanidioschyzon merolae. J Gen Appl Microbiol 60:156–159
Ohta S, Chang T, Aozasa O, Kondo M, Miyata H (1992) Sustained production of arachidonic and eicosapentaenoic acids by the red alga Porphyridium purpureum cultured in a light/dark cycle. J Ferment Bioeng 74:398–402
Pal D, Khozin-Goldberg I, Cohen Z (2011) The effect of light, salinity, and nitrogen availability on lipid production by Nannochloropsis sp. Appl Microbiol Biotechnol 90:1429–1441
Pettitt TR, Jones AL, Harwood JL (1989) Lipids of the marine red algae, Chondrus crispus and Polysiphonia lanosa. Phytochemistry 28:399–405
Qin S, Lin H, Jiang P (2012) Advances in genetic engineering of marine algae. Biotechnol Adv 30:1602–1613
Riekhof WR, Sears B, Benning C (2005) Annotation of genes involved in glycerolipid biosynthesis in Chlamydomonas reinhardtii: discovery of the betaine lipid synthase BTA1Cr. Eukaryot Cell 4:242–252
Rodolfi L, Chini ZG, Bassi N, Padovani G, Biondi N, Bonini G, Tredici MR (2009) Microalgae for oil: strain selection, induction of lipid synthesis and outdoor mass cultivation in a low-cost photobioreactor. Biotechnol Bioeng 102:100–112
Sakurai K, Moriyama T, Sato N (2014) Detailed identification of fatty acid isomers sheds light on the probable precursors of triacylglycerol accumulation in photoautotrophically grown Chlamydomonas reinhardtii. Eukaryot Cell 13:256–266
Sakurai T, Aoki M, Ju X, Ueda T, Nakamura Y, Fujiwara S, Umemura T, Tsuzuki M, Minoda A (2016) Profiling of lipid and glycogen accumulations under different growth conditions in the sulfothermophilic red alga Galdieria sulphuraria. Bioresour Technol 200:861–866
Sastre RMR (2010) Kopplung physiologischer und verfahrenstechnischer Parameter beim Wachstum and bei der Produktbildung der Rotalge Porphyridium purpureum. Dissertation, Universität Karlsruhe 2009
Sato N (1992) Betaine lipids. Bot Mag 105:185–197
Sato N (2001) Was the evolution of plastid genetic machinery discontinuous? Trends Plant Sci 6:151–156
Sato N (2009) Gclust: trans-kingdom classification of proteins using automatic individual threshold setting. Bioinformatics 25:599–605
Sato N, Moriyama T (2007) Genomic and biochemical analysis of lipid biosynthesis in the unicellular rhodophyte Cyanidioschyzon merolae: lack of plastidic desaturation pathway results in mixed pathway of galactolipid synthesis. Eukaryot Cell 6:1006–1017
Sato N, Mori N, Hirashima T, Moriyama T (2016) Diverse pathways of biosynthesis of phosphatidylcholine in algae as estimated by labeling studies and genomic sequence analysis. Plant J 87:281–292
Satyanarayana KG, Mariano AB, Vargas JVC (2011) A review on microalgae, a versatile source for sustainable energy and materials. Int J Energy Res 35:291–311
Schönknecht G, Chen WH, Ternes CM, Barbier GG, Shrestha RP, Stanke M, Bräutigam A, Baker BJ, Banfield JF, Garavito RM, Carr K, Wilkerson C, Rensing SA, Gagneul D, Dickenson NE, Oesterhelt C, Lercher MJ, Weber AP (2013) Gene transfer from bacteria and archaea facilitated evolution of an extremophilic eukaryote. Science 339:1207–1210
Shimonaga T, Konishi M, Oyama Y, Fujiwara S, Satoh A, Fujita N, Colleoni C, Buléon A, Putaux JL, Ball SG, Yokoyama A, Hara Y, Nakamura Y, Tsuzuki M (2008) Variation in storage α-glucans of the Porphyridiales (Rhodophyta). Plant Cell Physiol 49:103–116
Shiran D, Khozin I, Heimer YM, Cohen Z (1996) Biosynthesis of eicosapentaenoic acid in the microalga Porphyridium cruentum. I: the use of externally supplied fatty acids. Lipids 31:1277–1282
Siaut M, Cuiné S, Cagnon C, Fessler B, Nguyen M, Carrier P, Beyly A, Beisson F, Triantaphylidès C, Li-Beisson Y, Peltier G (2011) Oil accumulation in the model green alga Chlamydomonas reinhardtii: characterization, variability between common laboratory strains and relationship with starch reserves. BMC Biotechnol 11:7. doi:10.1186/1472-6750-11-7
Sumiya N, Kawase Y, Hayakawa J, Matsuda M, Nakamura M, Era A, Tanaka K, Kondo A, Hasunuma T, Imamura S, Miyagishima S (2015) Expression of cyanobacterial acyl-ACP reductase elevates the triacylglycerol level in the red alga Cyanidioschyzon merolae. Plant Cell Physiol 56:1962–1980
Tajima T, Sato S, Maruyama F, Kurokawa K, Ohta H, Tabata S, Sekine K, Moriyama T, Sato N (2014) Analysis of the complete plastid genome of the unicellular red alga Porphyridium purpureum. J Plant Res 127:389–397
Takusagawa M, Nakajima Y, Saito T, Misumi O (2016) Primitive red alga Cyanidioschyzon merolae accumulates storage glucan and triacylglycerol under nitrogen depletion. J Gen Appl Microbiol 62:111–117
Toyoshima M, Sato N (2015) High-level accumulation of triacylglycerol and starch in photoautotrophically grown Chlamydomonas debaryana NIES-2212. Plant Cell Physiol 56:2447–2456
Toyoshima M, Mori N, Moriyama T, Misumi O, Sato N (2016) Analysis of triacylglycerol accumulation under nitrogen deprivation in the red alga Cyanidioschyzon merolae. Microbiology 162:803–812
Vítová M, Goecke F, Sigler K, Řezank T (2016) Lipidomic analysis of the extremophilic red alga Galdieria sulphuraria in response to changes in pH. Algal Res 13:218–226
Zienkiewicz K, Du ZY, Ma W, Vollheyde K, Benning C (2016) Stress-induced neutral lipid biosynthesis in microalgae—molecular, cellular and physiological insights. Biochim Biophys Acta 1861:1269–1281
Acknowledgements
The studies in the authors’ laboratory have been supported in part by the CREST project on algal biotechnology from Japan Science and Technology Agency.
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Sato, N., Moriyama, T., Mori, N. et al. Lipid metabolism and potentials of biofuel and high added-value oil production in red algae. World J Microbiol Biotechnol 33, 74 (2017). https://doi.org/10.1007/s11274-017-2236-3
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s11274-017-2236-3