Abstract
Fipronil is a highly active, broad spectrum insecticide with increasing and wide use. The degradation of fipronil was studied in Brazilian soil under oxic and anoxic conditions. Under oxic conditions, the half-life of fipronil was 16.9 days, with fipronil sulfone as the main metabolite, and no further degradation during 30 days of incubation. This degradation was accompanied by an increase of bacterial and archaeal ammonia oxidizers, as well as denitrifiers, and microorganisms related to Sphingomonas. Under anoxic conditions, a half-life of 15.7 days for fipronil was obtained, with fipronil sulphide as the primary metabolite, and fipronil sulfone at lower concentrations, with no further degradation of these metabolites during 90 days of incubation. In these conditions, complete degradation of fipronil was accompanied by an increase of denitrifiers, iron-reducers and ammonia oxidizers and selection of microorganisms that are related to uncultured Clostridiales (family VIII). Sulphate reducers and methanogens and most of the microbial community were not affected by fipronil and its metabolites. Toxicity evaluation, using in vitro effect-based CALUX assays confirmed that the metabolites have a similar toxic potency as compared to the parent compound fipronil. Therefore, the potential (eco)toxicity of fipronil does not seem to decrease upon microbial degradation.
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
1 Introduction
Brazil is the world’s largest producer of sugar cane, and the increase in crop production is expected to reach 3.1% by 2019/2020 harvest, with 343 thousand tonnes of the product expected to be harvested in São Paulo state alone (UNICA, 2020). The São Paulo state is responsible for respectively 65% and 34% of ethanol and sugar production (UNICA, 2020). Productivity of sugarcane crops is maintained by soil management, such as addition of organic matter and nutrients, through the use of vinasse and filter cake, maintenance of sugarcane straw on the soil surface, crop rotation, development of new cultivars and use of pesticides (Bordonal et al., 2018). Due to restrictions of the use of organochlorine and organophosphate insecticides, the application of fipronil is increasing (Cappelini et al., 2018). This insecticide is applied in sugarcane plantation as Regent® 800 WG to control Lepidoptera and Orthoptera on crops and Coleoptera larvae in soils. As a result, the environmental fate of fipronil is becoming increasingly important.
Fipronil, 5-amino-1-[2,6-dichloro-4-(trifluoromethyl)phenyl]-4[(trifluoromethyl) sulfinyl]-1H-pyrazole-3-carbonitrile, is a phenyl pyrazole insecticide that reacts with the central nervous system of insects via blockage of chloride channels, regulated by gamma-aminobutyric acid (GABA) (Gunasekara et al., 2007). Although fipronil is harmful to biodiversity, a third of the world’s insecticide market is dominated by it due to its broad spectrum and high activity (Gunasekara et al., 2007; van der Sluijs et al., 2015).
Fipronil has four major degradation pathways, including oxidation, reduction, photolysis and hydrolysis, with formation of fipronil sulfone, fipronil sulphide, fipronil desulfinyl and fipronil amide, respectively (Gunasekara et al., 2007). Like fipronil, its metabolites are biologically active and may pose a risk to non-target organisms, such as pollinators (bees and bumblebees), butterflies and moths and earthworms (Bonmatin et al., 2015; Pisa et al., 2014). Although some metabolites are more toxic than fipronil itself (Schlenk et al., 2001), they can be further degraded by microorganisms (Masutti & Mermut, 2007; Tan et al., 2008), with reported half-life data of fipronil in soil ranging from 3 days to 7 months.
Many factors influence the degradation rate of fipronil, such as temperature, moisture, formulation and soil composition (Masutti & Mermut, 2007; Zhu et al., 2004). Furthermore, the availability of electron acceptors determines whether the insecticide is degraded under oxic and anoxic conditions. Moreover, the presence of microorganisms capable of biodegradation of the target compound fipronil and its metabolites is important. Under oxic conditions, microorganisms such as Burkholderia thailandensis, Paracoccus sp., Acinetobacter calcoaceticus, Acinetobacter oleivorans, Stenotrophomonas acidaminiphila (Proteobacteria phylum) and Bacillus thuringiensis (Firmicutes phylum) were found to degrade fipronil and metabolites (Abraham & Gajendiran, 2019; Cappelini et al., 2018; Kumar et al., 2012; Uniyal et al., 2016a, b). Under anoxic conditions, only a few reports describe the degradation of fipronil, and no information exist about the involved microorganisms (Brennan et al., 2009; Doran et al., 2009). Fertirrigation of sugarcane plantation with vinasse is a common practice, and literature shows that long-term application of raw sugarcane vinasse to soils can cause anoxic condition due to the excessive input of organic matter to soils (Fuess et al., 2017, 2018). For example, Fuess et al. (2018) showed that the applied organic load (AOL) into the soils could reach over 2.0 ton-BOD ha−1 what is equivalent a pollution load for population densities over 117 inhab ha−1. Hence, it is important to identify the anoxic degradability of fipronil and its metabolites.
In this research, oxic and anoxic biodegradation of fipronil and its metabolites were assessed in microcosms with soil from sugarcane plantation. Metabolites formed were identified, and fipronil degradation rate was determined. In order to assess the toxic potency of fipronil and its degradation products, several CALUX® reporter gene assays were performed. This CALUX® reporter gene assay panel consisted of seven human cell-based assays, each able to measure chemical interactions between a test compound and a specific nuclear receptor or cell signalling pathway (van der Burg et al., 2013). Finally, microorganisms were characterized in the microcosms by qPCR and 16S rDNA gene metagenomics.
2 Material and Methods
2.1 Chemicals
Fipronil (CAS 120068-37-3) and its metabolites (fipronil sulphide (CAS 120067-83-6), fipronil sulfone (CAS 120068-36-2), fipronil amide and fipronil desulfinyl (CAS 205650-65-3)) were purchased from AccuStandart® (100 μg mL−1 acetone). Stock solutions of individual compounds of 1 and 10 μg mL−1 were prepared in acetone (HPLC grade) and stored at −20°C.
2.2 Soil Samples
Soil samples were collected in APTA (Agência Paulista de Tecnologia Agropecuária – Região Centro Sul), in Piracicaba City, São Paulo, Brazil according to procedures previously described in Tomazini et al. (2020). The sandy-clay soil was used with the following granulometric characteristics: 55.74% of clay, 5.90% of silt and 38.36% of sand, as previously described (Tomazini et al., 2020).
2.3 Biodegradation of Fipronil
Biodegradation of fipronil was evaluated in batch bottles under oxic and anoxic conditions in Duran flasks of 250 mL containing 100 g of sieved and moisturized soil (set at 70% by adding sterile deionized water), to promote the bioavailability of fipronil in soil (Masutti & Mermut, 2007; Tan et al., 2008). Prior to filling the bottles, they were autoclaved at 121°C for 30 min. Filling of the bottles was performed under aseptic conditions. Before addition of fipronil, the soil was maintained for 30 min in ambient temperature for moisture stabilization. Fipronil was added to all flasks in a final measured concentration of 1.42 μg g−1 of wet soil or 1.08 μg g−1 of wet soil for respectively oxic and anoxic conditions and nitrate in concentration of 4 g NO3− g−1 of soil. The application of fipronil was chosen according to Regent® W800 label recommendation, 160–400 g.a.i ha−1, which corresponds to a final concentration of fipronil of 0.32–0.80 μg g−1 soil, just after application at 0–5 cm depth. The bottles were closed with air-permeable, sterile cellulose plugs for oxic condition. For anoxic condition, flasks were flushed with nitrogen for 5 min, sealed with rubber stoppers and closed with butyl covers under a N2 atmosphere. Abiotic control, for oxic and anoxic incubations, containing fipronil and autoclaved soil was taken along and for oxic and anoxic conditions. The soil was autoclaved for three consecutive days at 121°C under 15 psi chamber pressure for 30 min. All conditions, including the abiotic control, were performed in triplicate and incubated in a climate chamber at 37 ± 1°C in the dark. Samples for analyses were collected at day 0, 3, 6, 15, 20 and 30 days for the oxic incubations and at day 0, 10, 20, 30, 45, 60 and 90 for the anoxic incubations. In the abiotic control at oxic and anoxic conditions, fipronil at day 0 was measured in parallel series. Per sampling day, an amount of 2.7 g of soil was collected from the flasks and used for quantification of fipronil and metabolites, and 40 g was transferred to sterile Falcon flasks of 50 mL and stored at −20°C for molecular assays.
2.4 Extraction Procedure
Prior to quantification with gas chromatography, fipronil and metabolites were extracted from soil by 3 mL acetone and 3 mL hexane, following the procedures previously described in Tomazini et al. (2020). The recovery values of the method were between 81 and 108 %, with relative standard deviation (RSD) lower than 6%, with no effect of the used matrix. Limits of detection (LOD) from 0.002 to 0.006 μg g−1 and limits of quantification (LOQ) from 0.006 to 0.020 μg g−1 were reached for all analytes (Tomazini et al., 2020). In order to correct a matrix effect, the recovery test was done for each batch disassembled. Additionally, for each sampling round, a new calibration curve was done.
For the CALUX reporter gene bioassay analysis, the following extraction procedure was used. Soil samples were freeze-dried, and 10 g of each sample was extracted using accelerated solvent extraction (ASE) with acetone/hexane 1:1 (v/v). As a first clean-up step to remove any residual water, columns were filled with 15 g alumina with 0.5 cm of Na2SO4-anhydrous on top. The columns were equilibrated with 10 mL hexane. Then, the samples were loaded on the column and subsequently eluted with 3×50 mL hexane. The hexane was evaporated to dryness, and the extracts were dissolved in 600 μL dichloromethane. As a second clean-up step, gel permeation chromatography (GPC) was performed using two Styragel columns coupled in series (2× Waters Styragel Column, HR 0.5, 5 μm, 7.8 mm × 300 mm, THF), which were fitted with a guard column (Waters Styragel Guard Column, 20 μm, 4.6 mm × 30 mm, THF). The dicloromethane flow was set at 1mL/min. Samples were injected as 3 × 200 μL. The fraction used for CALUX analyses was collected in the 4.5–10 min time interval; this fraction contains components of MW 200–400. Dichloromethane was evaporated to dryness, and the extracts were dissolved in 150 μL DMSO.
2.5 Kinetics Analysis
First-order degradation rates for fipronil were obtained by fitting fipronil degradation data from each experiment to a first-order kinetic equation according to Martins and Mermoud (1998), and the corresponding half-lives (t1/2) were calculated.
2.6 Chemical Analyses
The GC-ECD used to detect fipronil and metabolites was a Shimadzu GC-2010 (Japan) chromatograph equipped with an autosampler. Chromatographic conditions are described in Tomazini et al. (2020).
2.7 Microbial
2.7.1 DNA Extraction
DNA from the soil that was used as inoculum and from biodegradation batches was extracted using PowerSoil® DNA Isolation Kit (Mobio) according to manufacturer’s instructions. Absorbance measurements (230, 260 and 280 nm) of the DNA solution were performed using ND-2000 spectrophotometer (Nanodrop Inc., Wilmington, DE) for quantification and determination of purity. DNA was extracted from the inoculum at t=0 day, from the biodegradation experiments at 6, 15 and 30 days of incubation under oxic condition and at 10, 20 and 60 days of incubation under anoxic condition.
2.7.2 qPCR Analyses
Samples were analysed by qPCR using specific primers for 16S rDNA gene (total bacteria, total archaea), nirK gene (denitrification), Geobacter 16S rDNA gene (iron-reducers), drsA gene (sulphate reducers) mcrA gene (methanogens), bacterial Amoa and archaeal Amoa genes. Thus, total bacteria, total archaea, denitrifying bacteria, iron-reducing bacteria, sulphate-reducing bacteria and the methanogenic archaea communities were quantified. All qPCR analyses were performed on an IQ5 real-time PCR system (Bio-Rad) using different temperature programmes dependent on the used protocol. The temperature programme for denitrifying bacteria was 95°C for 3 min, 46 cycles 95°C for 15 s, 63°C for 30 s, 72°C for 30 s, 72°C for 5 min, 95°C for 1 min, 56°C for 1 min and 75 cycles 58°C for 10 s. The total bacteria, total archaea, methanogens, sulphate-reducing bacteria and iron-reducing bacteria were measured using the following temperature programme: 94°C for 3 min, 35 cycles 94°C for 30 s, 58°C for 30 s, 72°C for 30 s, 72°C for 5 min, 95°C for 1 min, 58°C for 1 min, 75 cycles 58°C for 10 s and a final hold of 12°C. For the detection of the microbial groups, different forward and reverse primers in a final concentration of 0.6 pmol μL−1 were used (Table 1).
The protocols used for qPCR in this research experiment contained the fluorescent dye SYBR® Green I. For every reaction 22 μL of SYBR®, Green I mix was used, and 3 μL of DNA was added on the plate.
2.7.3 Microbial Metagenomic and Sequence Data Analyses
For metagenomic analysis, the 16S rDNA genes were amplified using the primer set described in Klindworth et al. (2013) containing adapter overhangs, according to manufacturer’s recommendations (Illumina manual). Amplicon sequencing was performed using an Illumina MiSeq platform, following the manufacturer’s guidelines, for paired end samples, using the MiSeq v3 kits.
Raw reads were subjected to quality control using SolexaQA++ dynamic trim (Cox et al., 2010) and a homemade Perl script (available upon request) to remove short reads (less than 50bp). Paired ends needed reorganization using PAIRFQ (open license) and were joined with PANDASEQ (Masella et al., 2012), using standard configuration. Singlets which passed quality control were added to the joined pairs and processed alongside. Chimeric sequences were checked with Uchime (Edgar et al., 2011) in the de novo mode within CATCh (Mysara et al., 2015). To ease computing, exact replicates were joined with grep in command line, and operational taxonomic units (OTUs) were clustered with SWARM (Mahé et al., 2014). Finally, OTUs were annotated in Mothur 1.36 (Schloss et al., 2009) with classifiy.seqs, against Silva SSU database (v1.32) (Quast et al., 2013).
2.7.4 CALUX Reporter Gene Assays
ERα CALUX, AR CALUX (Sonneveld et al., 2005) and the PXR CALUX (van der Burg et al., 2013) were used, which are all human U2-OS cell lines stably transfected with an expression construct for the human oestrogen (ER), androgen (AR) or pregnane X receptor (PXR) and a reporter construct consisting of multimerized responsive elements for the cognate receptor coupled to a minimal promoter element (TATA) and a luciferase gene. Furthermore, three U2-OS-based singly transfected lines expressing only a reporter gene were used. They were designed to selectively measure activation of the endoplasmic reticulum stress pathway (ESRE), the tumour suppressor protein p53 pathway or the oxidative stress Nrf2 pathway, respectively (van der Burg et al., 2013; van der Linden et al., 2014). The Cytotox CALUX was used as a control line for non-specific effects and consists of human U2-OS cells stably transfected with an expression construct constitutively expressing the luciferase gene (van der Linden et al., 2014).
Cells were maintained as described previously (Sonneveld et al., 2005). The automated CALUX assays were carried out on a Hamilton STARlet as described earlier (van der Burg et al., 2015).
GraphPad Prism was used to fit a sigmoidal curve through the data (sigmoidal dose response). The EC10 concentration was defined as the concentration where the response elicited by the test compound equals 10% of the maximum response of the reference compound. For antagonist experiments, EC20 values were determined instead, which was defined as the concentration where the test compound causes 20% inhibition of the basal signal elicited by the receptor agonist which was set to 100% (van Vugt-Lussenburg et al., 2018).
Results are expressed in PC10 (Log[(M)]). PC10 being the concentration where the signal elicited by the compound equals 10% of the maximum signal elicited by the reference compound.
3 Results and Discussion
3.1 Fipronil Biodegradation
Fipronil concentrations decreased under both oxic and anoxic conditions (Fig. 1, Supplement Material Tables S3 and S4), whereas our abiotic controls showed only a limited decrease (< 19 %) in fipronil concentration (Supplement Material; Tables S1, S2). These data show that microbial biodegradation was the main removal mechanism of fipronil.
At oxic conditions, fipronil was degraded with a lag phase of 6 days, and 0.83 μg g−1 was degraded during the first 15 days, corresponding to 58% fipronil removal. The removal followed first-order kinetics with t1/2 of 16.9 days (Table 2). After 15 days, fipronil showed only a small further decrease (Fig. 1a). The main metabolite was fipronil sulfone, indicating the oxidative degradation pathway. Fipronil sulfone was produced in the batches, and its net production was more than its degradation. Furthermore, fipronil sulphide was detected at 0.12 μg g−1 at day 30. Another known intermediate, fipronil desulfinyl, was not detected in these batches. Similar results were observed by Tan et al. (2008) where both fipronil sulfone and fipronil sulphide were detected as intermediates. In contrast, only 16% removal of 1 μg g−1 fipronil in soil after 30 days was shown in another study. Fipronil sulfone was detected as main metabolite, followed by fipronil sulphide, and both were further degraded (Masutti & Mermut, 2007).
Under anoxic conditions, 1.08 μg g−1 of fipronil was added to soil samples and completely degraded within 60 days, without a lag phase, with 70% of removal during the first 20 days (Fig. 1b). The removal followed first-order kinetics with a half-life of 15.7 days (Table 2). Fipronil sulphide was the main metabolite, followed by fipronil sulfone, which was detected at lower concentrations (Fig.1b), indicating reductive degradation of fipronil. These metabolites were produced during the first 20 days and not further degraded during the 90 days incubation, as observed by others (Doran et al., 2009; Tan et al., 2008). For example, Tan et al. (2008) observed degradation of 1.8 μg g−1 of fipronil within 60 days via a reductive pathway with formation of fipronil sulphide that was not further degraded.
3.2 Microbial Quantitation by qPCR
qPCR analyses were performed on the DNA extracted from the inoculum (time 0) and oxic and anoxic soil samples to gain insight in the microbial community present, functional genes and their development in time. Total archaea increased linearly throughout the incubation period, while total bacteria numbers increased mainly during the first 15 and 20 days for oxic and anoxic incubations, respectively (Fig. 2).
Our data were analysed for the presence of specific functional groups, such as denitrifiers, iron reducers, sulphate reducers, methanogens and ammonia oxidizers. Bacterial and archaeal ammonia oxidizers were identified in oxic incubations, as well as denitrifiers (Fig. 2a). This shows that neither fipronil nor fipronil sulfone affected the presence of these functional guilds. In the anoxic incubations, a sequential follow-up of denitrifiers, iron-reducers and ammonia oxidizer was detected at the beginning of the incubation period until the 20th day of incubation. This coincides with the degradation of fipronil, followed by an increase of sulphate reducers and methanogens at the end of incubation period (Fig. 2b). These results indicate that denitrifiers, iron-reducers and/or ammonia oxidizers could have been involved in fipronil degradation. Furthermore, the formed metabolite fipronil sulphide did not affect sulphate reducers and methanogens that were selected by the anoxic environment and organic matter present in soil.
3.3 Characterization of the Microbial Community
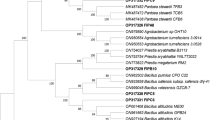
To get insight into the taxonomic groups that were present in the inoculum and the incubations with fipronil, 16S rDNA genes were accessed through metagenomic analysis. Amplicon sequencing provided 5,164,454 raw reads, counting both ends sequenced. A few short sequences were removed in the reverse end in each sample. We formed 2,531,226 joined reads after quality control and kept 42,321 high-quality singlets to the analysis in total. Altogether 2,125,485 reads were annotated as OTUs, although part of it is unknown. The inoculum comprised 246,641 reads, the oxic condition added up to 986,369 reads (average 328,790±42,607) and the anoxic condition summed 892,475 reads (average 297,492±48,901). The Bacteria domain was predominant in the inoculum (97.51%) and oxic (97.44±0.28%) and anoxic (96.63±0.56%) incubations.
At higher taxonomic levels (phylum/class), a diverse composition was shown in the soil used as inoculum for the biodegradation of fipronil, as shown in Fig. 3a. The major phyla Proteobacteria, Actinobacteria and Chloroflexi observed in our study were reported to comprise 38%, 30% and 4.3% of global soil bacteria (Delgado-Baquerizo et al., 2018). Proteobacteria and Actinobacteria were also reported in contaminated soils with representatives capable of degrading other pesticides (Alvarez et al., 2017; Kaur et al., 2015).
Taxonomic affiliation at level of phyla (a) and genera (b) of the sequences from the inoculum (soil) and from the flasks with fipronil at 6, 15 and 30 days of incubation time for oxic condition and at 10, 20 and 6 days of incubation time for anoxic condition. Note the break between 40 and 50%, to improve visualization of the less frequent groups. Cut-off value of 1.5%
The overall structure of the inoculum microbes was maintained throughout the incubation period in the soil spiked with fipronil and incubated under oxic condition (Fig. 3a). However, the presence of fipronil seemingly selected some of these groups, such as Alpha- and Gammaproteobacteria. In our study, the most abundant alphaproteobacterial genus after fipronil exposure was Sphingomonas (2.73±0.57%, Fig. 3b), with the highest relative abundance (3.35%) detected after 15 days of incubation, corresponding to the complete degradation of fipronil (Fig. 1a). Sphingomonas strains have the capability to degrade herbicides (Adkins, 1999), insecticides (Yu et al., 2013) and heavy metals (Chien et al., 2008). Our results suggest that this genus is also involved in the degradation of fipronil.
For the anoxic incubation, a selection of Firmicutes (from 2.17 to 39.6±6.42%) was observed, mainly Clostridia (from 0.34 to 30.48±6.77%; Fig. 2a). Uncultured Clostridiales (family XVIII) which averaged 11.54±7.15% peaked at the 20th day of incubation (18.07%; Fig. 3b), corresponding to the largest reduction of fipronil (Fig. 1b). These results suggest that microorganisms of this genus could be involved in fipronil degradation anaerobically. Although there is no report about the capability of Clostridiales to degrade fipronil, different Clostridium sp. was reported in hydrocarbon contaminated soil (Bastida et al., 2010). Strains of Clostridium are also capable of degrading 2,4,6-trinitrotoluene under anaerobic condition (Ederer et al., 1997); Lewis et al., 1997) and both Fenpropathrin (Zhang et al., 2011) and Dinoseb via co-metabolism (Hammill & Crawford, 1996). Ktedonobacteria, Alphaproteobacteria and Actinobacteria were the classes most impacted by the presence of fipronil in the anoxic condition (Fig. 3a). Ktedonobacteria comprises aerobic microorganisms (Cavaletti et al., 2006) that were slightly affected by fipronil under aerobic condition (Fig. 3a); therefore, their reduction was most likely related to anoxic condition or to the presence of metabolites (i.e. fipronil sulphide) instead of fipronil itself. The same can be inferred about the decay of both Alphaproteobacteria and Actinobacteria, despite the existence of aerobic and anaerobic representatives of these classes (Alvarez et al., 2017; Kaur et al., 2015).
3.4 Toxicity of Fipronil and the Spiked Soil Extracts
In order to assess the biological activity of fipronil and its transformation products, an effect profile was generated using a panel of seven CALUX reporter gene assays, covering different toxicological endpoints, each in the absence and presence of a metabolic fraction mimicking hepatic metabolism. The panel was also used to analyse extracts of the spiked soil samples.
Fipronil and the transformation products showed similar activities; they were cytotoxic at concentrations above 10–30 μM, and they were all active as oestrogen receptor agonists, androgen receptor antagonists and agonists on the xenobiotic sensing receptor PXR in the lower μM range (Table 3). For fipronil and fipronil sulfone, oxidative stress (Nrf2 CALUX) was observed at 30 μM. Interestingly, the compounds were only active on the oestrogen receptor assay ERα CALUX in the presence of S9; this indicates that the compounds require bioactivation before they can activate the receptor. For the other assays, the addition of rat liver S9 did not have a significant effect on the activity, occasionally even abolished the activity instead. For fipronil, activation of the PXR receptor (Lemaire et al., 2006) as well as androgen antagonist activity (Ait-Aissa et al., 2010) has been described before. Its activity as an oestrogen receptor agonist and androgen receptor antagonist could be linked to previously observed reproductive toxicity in vivo (Ohi et al., 2004). Also oxidative stress has been observed before, in vitro as well as in vivo (Mossa et al., 2015; Slotkin & Seidler, 2010).
The spiked soil extracts (oxic incubation only) were also analysed using the CALUX assays. This time, only the ERα CALUX and p53 GENTOX CALUX were performed in the presence of metabolic enzymes. No significant difference was observed between the different time points, which was expected since the fipronil transformation products had shown a similar activity on the bioassays as the parent compound fipronil (Table 3).
While the pure compounds were exposed on the bioassays with concentrations up to 1E-4M, the extracts could be exposed only to a concentration of 1E-6M (Cytotox, ESRE, Nrf2) or 1E-5M (ERα, AR-anti, PXR, p53 GENTOX) for technical reasons. As a consequence, the cytotoxicity and ESRE- and Nrf2-activation observed at higher concentrations for the pure compounds (Table 3) could not be observed for the extracts (Table 4). ERα CALUX activation was observed in the absence of metabolic enzymes at two time points. Furthermore, the oestrogenic activity of all four samples was greatly enhanced in the presence of metabolic enzymes as was also observed for the pure compounds. Anti-androgenic activity could not be detected for the extracts, but PXR activation was observed at all four time points. This shows that the activity profile of the spiked soil sample extracts matches that of the pure compounds.
4 Conclusions
To conclude, we have shown that fipronil degradation in the soil takes place under both oxic and anoxic conditions. Different metabolites were produced in each condition, since fipronil sulfone was the main metabolite under oxic condition, whereas fipronil sulphide was formed under anoxic condition. Neither intermediates affected functional microbial groups present in soil though, such as nitrate or sulphate reducers. Nevertheless, fipronil and its metabolites produced under these conditions presented similar toxic potency. Therefore, the potency of fipronil does not necessarily decrease upon microbial degradation.
Finally, since Sphingomonas was seemingly selected in aerobic condition, whereas Clostridiales was selected in anoxic condition, the degradation of fipronil in contaminated soil should be stimulated by bioaugmentation with either type of bacteria, depending on the aeration conditions of the soil.
References
Abraham, J., & Gajendiran, A. (2019). Biodegradation of fipronil and its metabolite fipronil sulfone by Streptomyces rochei strain AJAG7 and its use in bioremediation of contaminated soil. Pesticide Biochemistry and Physiology, 155, 90–100. https://doi.org/10.1016/j.pestbp.2019.01.011.
Adkins, A. (1999). Degradation of the phenoxy acid herbicide diclofop-methyl by Sphingomonas paucimobilis isolated from a Canadian prairie soil. Journal of Industrial Microbiology & Biotechnology, 23, 332–335. https://doi.org/10.1038/sj.jim.2900744.
Ait-Aissa, S., Laskowski, S., Laville, N., Porcher, J. M., & Brion, F. (2010). Anti-androgenic activities of environmental pesticides in the MDA-kb2 reporter cell line. Toxicology In Vitro, 24, 1979–1985.
Alvarez, A., Saez, J. M., Davila Costa, J. S., Colin, V. L., Fuentes, M. S., Cuozzo, S. A., Benimeli, C. S., Polti, M. A., & Amoroso, M. J. (2017). Actinobacteria: Current research and perspectives for bioremediation of pesticides and heavy metals. Chemosphere, 166, 41–62.
Bastida, F., Nicolás, C., Moreno, J. L., Hernández, T., & García, C. (2010). Tracing changes in the microbial community of a hydrocarbon-polluted soil by culture-dependent proteomics. Pedosphere, 20, 479–485. https://doi.org/10.1016/S1002-0160(10)60037-9.
Bonmatin, J. M., Giorio, C., Girolami, V., Goulson, D., Kreutzweiser, D. P., Krupke, C., Liess, M., Long, E., Marzaro, M., Mitchell, E. A., Noome, D. A., Simon-Delso, N., & Tapparo, A. (2015). Environmental fate and exposure; neonicotinoids and fipronil. Environmental Science and Pollution Research, 22, 35–67. https://doi.org/10.1007/s11356-014-3332-7.
Bordonal, R.d. O., Carvalho, J. L. N., Lal, R., Figueiredo, E. B., Oliveira, B. G., & La Scala Jr., N. (2018). Sustainability of sugarcane production in Brazil. A review. Agronomy for Sustainable Development, 38, 1–23. https://doi.org/10.1007/s13593-018-0490-x.
Brennan, A. A., Harwood, A. D., You, J., Landrum, P. F., & Lydy, M. J. (2009). Degradation of fipronil in anaerobic sediments and the effect on porewater concentrations. Chemosphere, 77, 22–28. https://doi.org/10.1016/j.chemosphere.2009.06.019.
Cappelini, L. T. D., Alberice, J. V., Eugênio, P. F. M., Pozzi, E., Urbaczek, A. C., Diniz, L. G. R., Carrilho, E. N. V. M., Carrilho, E., & Vieira, E. M. (2018). Burkholderia thailandensis: The main bacteria biodegrading fipronil in fertilized soil with assessment by a QuEChERS/GC-MS Method. Journal of the Brazilian Chemical Society, 29, 1934–1943. https://doi.org/10.21577/0103-5053.20180069.
Cavaletti, L., Monciardini, P., Bamonte, R., Schumann, P., Ronde, M., Sosio, M., & Donadio, S. (2006). New lineage of filamentous, spore-forming, gram-positive bacteria from soil. Applied and Environmental Microbiology, 72, 4360–4369. https://doi.org/10.1128/AEM.00132-06.
Chien, C., Kuo, Y., Chen, C., Hung, C., Yeh, C., & Yeh, W. (2008). Microbial diversity of soil bacteria in agricultural field contaminated with heavy metals. Journal of Environmental Sciences, 20, 359–363. https://doi.org/10.1016/S1001-0742(08)60056-X.
Cox, M. P., Peterson, D. A., & Biggs, P. J. (2010). SolexaQA: At-a-glance quality assessment of Illumina second-generation sequencing data. BMC Bioinformatics, 11, 485. https://doi.org/10.1186/1471-2105-11-485.
Delgado-Baquerizo, M., Oliverio, A. M., Brewer, T. E., Benavent-González, A., Eldridge, D. J., Bardgett, R. D., Maestre, F. T., Singh, B. K., & Fierer, N. (2018). A global atlas of the dominant bacteria found in soil. Science, 359, 320–325. https://doi.org/10.1126/science.aap9516.
Dhillon, A., Teske, A., Dillon, J., Stahl, D. A., & Sogin, M. L. (2003). Molecular characterization of sulfate-reducing bacteria in the Guaymas basin. Applied and Environmental Microbiology, 69, 2765–2772. https://doi.org/10.1128/AEM.69.5.2765-2772.2003.
Doran, G., Eberbach, P., & Helliwell, S. (2009). Sorption and degradation of fipronil in flooded anaerobic rice soils. Journal of Agricultural and Food Chemistry, 57, 10296–10301. https://doi.org/10.1021/jf902644x.
Ederer, M. M., Lewis, T. A., & Crawford, R. L. (1997). 2,4,6-Trinitrotoluene (TNT) transformation by clostridia isolated from a munition-fed bioreactor: comparison with non-adapted bacteria. Journal of Industrial Microbiology & Biotechnology, 18, 82–88. https://doi.org/10.1038/sj.jim.2900257.
Edgar, R. C., Haas, B. J., Clemente, J. C., Quince, C., & Knight, R. (2011). UCHIME improves sensitivity and speed of chimera detection. Bioinformatics, 27, 2194–2200. https://doi.org/10.1093/bioinformatics/btr381.
Francis, C. A., Roberts, K. J., Beman, J. M., Santoro, A. E., & Oakley, B. B. (2005). Ubiquity and diversity of ammonia-oxidizing archaea in water columns and sediments of the ocean. Proceedings of the National Academy of Sciences of the United States of America, 102, 14683–14688. https://doi.org/10.1073/pnas.0506625102.
Fuess, L. T., Rodrigues, I. J., & Garcia, M. L. (2017). Fertirrigation with sugarcane vinasse: Foreseeing potential impacts on soil and water resources through vinasse characterization. Journal of Environmental Science and Health Part A, 52(11), 1063–1072. https://doi.org/10.1080/10934529.2017.1338892.
Fuess, L. T., Garcia, M. L., & Zaiat, M. (2018). Seasonal characterization of sugarcane vinasse: assessing environmental impacts from fertirrigation and the bioenergy recovery potential through biodigestion. Science of the Total Environment, 634, 29–40. https://doi.org/10.1016/j.scitotenv.2018.03.326.
Gunasekara, A. S., Truong, T., Goh, K. S., Spurlock, F., & Tjeerdema, R. S. (2007). Environmental fate and toxicology of fipronil. Journal of Pesticide Science, 32, 189–199. https://doi.org/10.1584/jpestics.R07-02.
Hales, B. A., Edwards, C., Ritchie, D. A., Hall, G., Pickup, R. W., & Saunders, J. R. (1996). Isolation and identification of methanogen-specific DNA from blanket bog peat by PCR amplification and sequence analysis. Applied and Environmental Microbiology, 62, 668–675. https://doi.org/10.1128/aem.62.2.668-675.1996.
Hammill, T. B., & Crawford, R. L. (1996). Degradation of 2-sec-butyl-4,6-dinitrophenol (dinoseb) by Clostridium bifermentans KMR-1. Applied and Environmental Microbiology, 62, 1842–1846.
Henry, S., Baudoin, E., López-Gutiérrez, J. C., Martin-Laurent, F., Brauman, A., & Philippot, L. (2004). Quantification of denitrifying bacteria in soils by nirK gene targeted real-time PCR. Journal of Microbiological Methods, 59, 327–335. https://doi.org/10.1016/j.mimet.2004.07.002.
Holmes, D. E., Finneran, K. T., O’Neil, R. A., & Lovley, D. R. (2002). Enrichment of members of the family Geobacteraceae associated with stimulation of dissimilatory metal reduction in uranium-contaminated aquifer sediments. Applied and Environmental Microbiology, 68, 2300–2306. https://doi.org/10.1128/AEM.68.5.2300-2306.2002.
Kaur, G., Sharma, R., Singh, K., & Sharma, P. K. (2015). Delineating bacterial community structure of polluted soil samples collected from cancer prone belt of Punjab, India. 3 Biotech, 5, 727–734. https://doi.org/10.1007/s13205-014-0270-5.
Klindworth, A., Pruesse, E., Schweer, T., Peplies, J., Quast, C., Horn, M., & Glöckner, F. O. (2013). Evaluation of general 16S ribosomal RNA gene PCR primers for classical and next-generation sequencing-based diversity studies. Nucleic Acids Research, 41, e1. https://doi.org/10.1093/nar/gks808.
Kumar, R., Singh, B., & Gupta, V. K. (2012). Biodegradation of fipronil by Paracoccus sp. in different types of soil. Bulletin of Environmental Contamination and Toxicology, 88, 781–787. https://doi.org/10.1007/s00128-012-0578-y.
Lane, D. (1991). 16S/23S sequencing. In E. Stackebrandt & M. Goodfellow (Eds.), Nucleic acids techniques in bacterial systematics (pp. 205–248). Chichester: Wiley.
Lemaire, G., Mnif, W., Pascussi, J. M., Pillon, A., Rabenoelina, F., Fenet, H., Gomez, E., Casellas, C., Nicolas, J. C., Cavailles, V., Duchesne, M. J., & Balaguer, P. (2006). Identification of new human pregnane X receptor ligands among pesticides using a stable reporter cell system. Toxicological Sciences, 91, 501–509.
Lewis, T. A., Ederer, M. M., Crawford, R. L., & Crawford, D. L. (1997). Microbial transformation of 2,4,6-trinitrotoluene. Journal of Industrial Microbiology & Biotechnology, 18, 89–96.
Mahé, F., Rognes, T., Quince, C., de Vargas, C., & Dunthorn, M. (2014). Swarm: Robust and fast clustering method for amplicon-based studies. PeerJ, 2014, e593. https://doi.org/10.7717/peerj.593.
Martins, J. M., & Mermoud, A. (1998). Sorption and degradation of four nitroaromatic herbicides in mono and multi-solute saturated/unsaturated soil batch systems. Journal of Contaminant Hydrology, 33, 187–210. https://doi.org/10.1016/S0169-7722(98)00070-9.
Masella, A. P., Bartram, A. K., Truszkowski, J. M., Brown, D. G., & Neufeld, J. D. (2012). PANDAseq: Paired-end assembler for illumina sequences. BMC Bioinformatics, 13, 31. https://doi.org/10.1186/1471-2105-13-31.
Masutti, C. S. M., & Mermut, A. R. (2007). Degradation of fipronil under laboratory conditions in a tropical soil from Sirinhaém Pernambuco. Journal of Environmental Science and Health Part B, 42, 33–43. https://doi.org/10.1080/03601230601017981.
Mossa, A. H., Swelam, E. S., & Mohafrash, S. S. M. (2015). Sub-chronic exposure to fipronil induced oxidative stress, biochemical and histopathological changes in the liver and kidney of male albino rats. Toxicology Reports, 2, 775–784.
Muyzer, G., & Ramsing, N. B. (1995). Molecular methods to study the organization of microbial communities. Water Science and Technology, 32, 1–9. https://doi.org/10.1016/0273-1223(96)00001-7.
Mysara, M., Saeys, Y., Leys, N., Raes, J., & Monsieurs, P. (2015). CATCh, an ensemble classifier for chimera detection in 16s rRNA sequencing studies. Applied and Environmental Microbiology, 81, 1573–1584. https://doi.org/10.1128/AEM.02896-14.
Ohi, M., Dalsenter, P. R., Andrade, A. J., & Nascimento, A. J. (2004). Reproductive adverse effects of fipronil in Wistar rats. Toxicology Letters, 146, 121–127.
Okano, Y., Hristova, K. R., Leutenegger, C. M., Jackson, L. E., Denison, R. F., Gebreyesus, B., Lebauer, D., & Scow, K. M. (2004). Application of real-time PCR to study effects of ammonium on population size of ammonia-oxidizing bacteria in soil. Applied and Environmental Microbiology, 70, 1008–1016. https://doi.org/10.1128/AEM.70.2.1008-1016.2004.
Pisa, L. W., Amaral-Rogers, V., Belzunces, L. P., Bonmatin, J. M., Downs, C. A., Goulson, D., Kreutzweiser, D. P., Krupke, C., Liess, M., Mcfield, M., Morrissey, C. A., Noome, D. A., Settele, J., Simon-Delso, N., Stark, J. D., Van Der Sluijs, J. P., Van Dyck, H., & Wiemers, M. (2014). Effects of neonicotinoids and fipronil on non-target invertebrates. Environmental Science and Pollution Research, 22, 68–102. https://doi.org/10.1007/s11356-014-3471-x.
Quast, C., Pruesse, E., Yilmaz, P., Gerken, J., Schweer, T., Yarza, P., Peplies, J., & Glöckner, F. O. (2013). The SILVA ribosomal RNA gene database project: Improved data processing and web-based tools. Nucleic Acids Research, 41, D590–D596. https://doi.org/10.1093/nar/gks1219.
Schlenk, D., Huggett, D. B., Allgood, J., Bennett, E., Rimoldi, J., Beeler, A. B., Block, D., Holder, A. W., Hovinga, R., & Bedient, P. (2001). Toxicity of fipronil and its degradation products to Procambarus sp.: Field and laboratory studies. Archives of Environmental Contamination and Toxicology, 41, 325–332. https://doi.org/10.1007/s002440010255.
Schloss, P. D., Westcott, S. L., Ryabin, T., Hall, J. R., Hartmann, M., Hollister, E. B., Lesniewski, R. A., Oakley, B. B., Parks, D. H., Robinson, C. J., Sahl, J. W., Stres, B., Thallinger, G. G., Van Horn, D. J., & Weber, C. F. (2009). Introducing mothur: Open-source, platform-independent, community-supported software for describing and comparing microbial communities. Applied and Environmental Microbiology, 75, 7537–7541. https://doi.org/10.1128/AEM.01541-09.
Slotkin, T. A., & Seidler, F. J. (2010). Oxidative stress from diverse developmental neurotoxicants: Antioxidants protect against lipid peroxidation without preventing cell loss. Neurotoxicology and Teratology, 32(2), 124–131.
Sonneveld, E., Jansen, H. J., Riteco, J. A. C., Brouwer, A., & van der Burg, B. (2005). Development of androgen-and estrogen-responsive bioassays, members of a panel of human cell line-based highly selective steroid-responsive bioassays. Toxicological Sciences, 83, 136–148. https://doi.org/10.1093/toxsci/kfi005.
Tan, H., Cao, Y., Tang, T., Qian, K., Chen, W. L., & Li, J. (2008). Biodegradation and chiral stability of fipronil in aerobic and flooded paddy soils. Science of the Total Environment, 407, 428–437. https://doi.org/10.1016/j.scitotenv.2008.08.007.
Tomazini, R., Grosseli, G. M., de Sousa, D. N. R., Fadini, P. S., Saia, F. T., Langenhoff, A., van der Zaan, B., & Mozeto, A. A. (2020). Development of a simple method to quantify fipronil and its intermediates in soil. Analytical Methods, 12, 3242. https://doi.org/10.1039/d0ay00924e.
UNICA. (2020). UNICA - UNIÃO DA INDÚSTRIA DE CANA-DE-AÇÚCAR. https://www.unicadata.com.br/. Accessed 22 Jun 2020
Uniyal, S., Paliwal, R., Sharma, R. K., & Rai, J. P. N. (2016a). Degradation of fipronil by Stenotrophomonas acidaminiphila isolated from rhizospheric soil of Zea mays. Biotech, 6, 1–10. https://doi.org/10.1007/s13205-015-0354-x.
Uniyal, S., Paliwal, R., Verma, M., Sharma, R. K., & Rai, J. P. N. (2016b). Isolation and characterization of fipronil degrading Acinetobacter calcoaceticus and Acinetobacter oleivorans from rhizospheric zone of Zea mays. Bulletin of Environmental Contamination and Toxicology, 96, 833–838. https://doi.org/10.1007/s00128-016-1795-6.
van der Burg, B., van der Linden, S., Man, H., Winter, R., Jonker, L., van Vugt-Lussenburg, B., & Brouwer, A. (2013). A panel of quantitative calux® reporter gene assays for reliable high-throughput toxicity screening of chemicals and complex mixtures. In J. Wiley & S. Inc (Eds.), High-Throughput Screening Methods in Toxicity Testing (pp. 519–532).
van der Burg, B., Pieterse, B., Buist, H., Lewin, G., van der Linden, S. C., Man, H. Y., Rorije, E., Piersma, A. H., Mangelsdorf, I., Wolterbeek, A. P. M., Kroese, E. D., & van Vugt-Lussenburg, B. (2015). A high throughput screening system for predicting chemically-induced reproductive organ deformities. Reproductive Toxicology, 55, 95–103. https://doi.org/10.1016/j.reprotox.2014.11.011.
van der Linden, S. C., von Bergh, A. R. M., van Vught-Lussenburg, B. M. A., Jonker, L. R. A., Teunis, M., Krul, C. A. M., & van der Burg, B. (2014). Development of a panel of high-throughput reporter-gene assays to detect genotoxicity and oxidative stress. Mutation Research, Genetic Toxicology and Environmental Mutagenesis, 760, 23–32. https://doi.org/10.1016/j.mrgentox.2013.09.009.
van der Sluijs, J. P., Amaral-Rogers, V., Belzunces, L. P., Bijleveld Van Lexmond, M. F., Bonmatin, J. M., Chagnon, M., Downs, C. A., Furlan, L., Gibbons, D. W., Giorio, C., Girolami, V., Goulson, D., Kreutzweiser, D. P., Krupke, C., Liess, M., Long, E., Mcfield, M., Mineau, P., Mitchell, E. A., Morrissey, C. A., Noome, D. A., Pisa, L., Settele, J., Simon-Delso, N., Stark, J. D., Tapparo, A., Van Dyck, H., Van Praagh, J., Whitehorn, P. R., & Wiemers, M. (2015). Conclusions of the worldwide integrated assessment on the risks of neonicotinoids and fipronil to biodiversity and ecosystem functioning. Environmental Science and Pollution Research, 22, 148–154.
van Vugt-Lussenburg, B. M. A., van der Lee, R. B., Man, H. Y., Middelhof, I., Brouwer, A., Besselink, H., & van der Burg, B. (2018). Incorporation of metabolic enzymes to improve predictivity of reporter gene assay results for estrogenic and anti-androgenic activity. Reproductive Toxicology, 75, 40–48. https://doi.org/10.1016/j.reprotox.2017.11.005.
Vetriani, C., Jannasch, H. W., Macgregor, B. J., Stahl, D. A., & Reysenbach, A. L. (1999). Population structure and phylogenetic characterization of marine benthic Archaea in deep-sea sediments. Applied and Environmental Microbiology, 65, 4375–4384. https://doi.org/10.1128/aem.65.10.4375-4384.1999.
Wagner, M., Roger, A. J., Flax, J. L., Brusseau, G. A., & Stahl, D. A. (1998). Phylogeny of dissimilatory sulfite reductases supports an early origin of sulfate respiration. Journal of Bacteriology, 180, 2975–2982. https://doi.org/10.1128/jb.180.11.2975-2982.1998.
Yu, F. B., Shan, S. D., Luo, L. P., Guan, L. B., & Qin, H. (2013). Isolation and characterization of a Sphingomonas sp. strain F-7 degrading fenvalerate and its use in bioremediation of contaminated soil. Journal of Environmental Science and Health Part B Pesticides Food Contaminants and Agricultural Wastes, 48, 198–207. https://doi.org/10.1080/03601234.2013.730299.
Zhang, S., Yin, L., Liu, Y., Zhang, D., Luo, X., Cheng, J., Cheng, F., & Dai, J. (2011). Cometabolic biotransformation of fenpropathrin by Clostridium species strain ZP3. Biodegradation, 22, 869–875. https://doi.org/10.1007/s10532-010-9444-y.
Zhu, G., Wu, H., Guo, J., & Kimaro, F. M. E. (2004). Microbial degradation of fipronil in clay loam soil. Water, Air, and Soil Pollution, 153, 35–44. https://doi.org/10.1023/B:WATE.0000019928.67686.b1.
Funding
This work was supported by a BE-Basic Foundation grant from the Dutch Ministry of Economic Affairs and a grant from the Brazilian FAPESP organization (2012/51496-3). The scholarship of Rafaella Tomazini was supported by Coordination for the Improvement of Higher Education Personnel – CAPES.
Author information
Authors and Affiliations
Corresponding author
Additional information
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
ESM 1
(DOCX 22 kb)
Rights and permissions
About this article
Cite this article
Tomazini, R., Saia, F.T., van der Zaan, B. et al. Biodegradation of Fipronil: Transformation Products, Microbial Characterisation and Toxicity Assessment. Water Air Soil Pollut 232, 123 (2021). https://doi.org/10.1007/s11270-021-05071-w
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s11270-021-05071-w