Abstract
Paratuberculosis (PTB) is a chronic infectious enteritis of ruminants, caused by Mycobacterium avium subspecies paratuberculosis (MAP) that brings huge economic loss to the dairy farmers. The study was conducted to explore the association of selected SNPs in IFNG, SLC11A1, ANKRA2 and PGLYRP1 genes with resistance to PTB disease in Indian cattle population. A case-control resource population was established based on the results of diagnostic tests used for detection of MAP infection status viz. ELISA, Johnin PPD test, faecal microscopy and IS900 blood PCR. The PCR-RFLP method was used for genotyping of SNPs. SNPs rs109453173 in SLC11A1, rs110853455 in IFNG and rs41933863 in ANKRA2 genes were significantly (P<0.05) associated with resistance to MAP infection. For SNP rs109453173, GG genotype and G allele was found to be associated with resistance against MAP infection than CC and CG genotypes and C allele, respectively. For SNP rs110853455, AG genotype was found to be associated with susceptibility to MAP infection than AA and GG genotype. For SNP rs41933863, the AG genotype provided three and six times more resistance against MAP infection than GG and AA genotype. The results of this study are suggestive of SNPs rs109453173, rs110853455 and rs41933863 as potential markers for screening MAP resistant cattle and a breeding programme favouring GG genotype and G allele for rs109453173, AG genotype for rs41933863 and against AG genotype for rs110853455 might confer resistance against MAP infection in Indian cattle. However, investigation of these SNPs in an independent and larger population will warrant the strength of association for resistance against MAP infection in cattle.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Mycobacterium avium subspecies paratuberculosis (MAP), the causative agent of bovine paratuberculosis (PTB) or Johne’s disease in ruminants, is also reported in pigs, horses, camels, alpaca, deer, llama, rabbits, weasel, fox and stoat (Larsen et al. 1971; Beard et al. 2001a, b; Pavlik et al. 2004; Miller et al. 2017; Juste et al. 2018; Fox et al. 2020). It is a World Organisation for Animal Health (OIE) listed disease, recognized as a chronic debilitating intestinal illness of cattle as early as 1826 (OIE 2020) and till today it continues to be a fatal, untreatable, chronic, granulomatous infection of the intestine (Whittington et al. 2004). PTB is characterised by slowly progressive wasting, severe diarrhoea and animals generally die from cachexia (Kumar et al. 2019a). PTB is a disease of economic importance for the dairy sector, since it causes major loss due to reduced production, high replacement cost, premature culling, infertility, low feed conversion efficiency and predisposes the herd to other diseases (Garcia and Shalloo 2015). Hence, PTB is one of the costliest diseases of cattle globally, which cause annual loss of $200 to $1,500 million in the US (Losinger 2005). In India, the reduction in milk yield due to PTB causes economic loss of Rs 54,442.5 /cow/lactation (Rawat et al. 2014). Whittington and coworker reported the prevalence of PTB in 20 % of the herds worldwide, which, may reach up to 40 % in some developed countries (Whittington et al. 2019). In India, the sero-prevalence of PTB was approximately 29.0 % in north India, 27.58 % in Uttar Pradesh (Narnaware and Tripathi 2017), 23.3 % Punjab (Singh et al. 2008), 15.14 % in Bengal (Gupta et al. 2012), 13.39 % in Gujarat, 16.26 % in Andhra Pradesh and 37.7 % in West Bengal (Bhutediya et al. 2017).
The zoonotic role of MAP has not yet been proved but MAP has been occasionally found in Crohn’s disease (CD) patients and symptoms of CD in humans resemble PTB in ruminants. Hence, it leads to the proposition that MAP is a potential food safety concern. Milk obtained from MAP infected animals is most likely source of infection and MAP has also been reported in both pasteurized and unpasteurized dairy products (Shankar et al. 2010). The MAP DNA was also found in some commercial baby formulas, using both IS900 and F57 qPCR (Hruska et al. 2005, 2011) but the presence of MAP DNA in milk and milk formulas indicates that MAP is there, dead or alive. Being able to culture MAP from milk can be more of a problem. Study found that in an infected herd, more than 81 % of the MAP-positive colostrum or MAP-positive teat swabs came from potential environmental seeding of MAP rather than direct shedding (Pithua et al. 2011). MAP organism had been isolated from the humans with type I diabetes, Blau syndrome (Sechi and Dow 2015) and multiple sclerosis (Cossu et al. 2013). Taking into consideration, the zoonotic potential, animal welfare concerns and economic loss associated with PTB, animal health authorities aim to decrease the prevalence of PTB disease.
Management practices like maintaining farm hygiene, vaccination of animals and test and cull programmes, which are primarily used to control the incidence of PTB in dairy herd, have failed to eliminate the MAP infection in cattle. The test and cull programme involves huge cost, time and manpower; moreover, this approach is less efficient due to the varied sensitivity of available diagnostic tests (Nielsen and Toft 2008). In the present scenario, vaccination is the most effective mean to control PTB (Juste and Perez 2011) since it prevents the clinical outcome of MAP infection; however, animals continue to excrete MAP bacilli and serve as a source of infection. Also, vaccinated and infected animals are difficult to differentiate due to the lack of DIVA (Differentiating Infected from Vaccinated Animals) based diagnostic tests (Bastida and Juste 2011). Due to the long incubation period (3-14 years in cattle and 1-7 years in sheep), absence of pathognomonic symptoms, variable sensitivity and specificity of diagnostic methods, PTB disease is difficult to diagnose and control (Pritchard et al. 2017). Hence, MAP infection continues to spread and requires new approaches to control PTB in animals. Since, differences in the host susceptibility to PTB have been associated with genetic factors (Bishop and MacKenzie 2003) and this difference is heritable which is evidenced by low to medium estimates of heritability to MAP infection (Gonda et al. 2006). The possibility of using marker assisted selection (MAS) have been elucidated by candidate gene approach after establishing the genetic effects of SNPs (Pinedo et al. 2009a; Kumar et al. 2017, 2020; Gopi et al. 2020).
The SLC11A1 (solute carrier family 11 member 1) gene encoding natural resistance-associated macrophage protein 1 (NRAMP1) provides resistance against Mycobacterium sp. During the initial stage of infection, NRAMP1, prevents bacterial growth in macrophages and thus plays a key role in innate immunity (Paixão et al. 2012). Similarly, Interferon gamma (IFNG), a member of interferon gene family (Shtrichman and Samuel 2001), is an important pro-inflammatory cytokine which favours Th1 mediated immunity and facilitates macrophages in regulating the spread of bacterial infection (Coussens et al. 2004). The ANKRA2 gene encodes for ankyrin repeat family A2 protein, located on the chromosome 20 and regulates the function of major histocompatibility complex class II (MHC-II) genes (Krawczyk et al. 2005). The increased ANKRA2 expression activates MHC-II gene transcription, indicating that ANKRA2 regulates expression of MHC-II genes (Krawczyk et al. 2005). Bovine PGLYRP1 gene encodes for peptidoglycan recognition proteins (PGLYRPs) which play a critical role in innate immunity due to their ability to bind various pathogen-associated molecular patterns (PAMPs), including lipoteichoic acid, lipopolysaccharide and peptidoglycans (Tydell et al. 2006). It recognizes and exert bacteriostatic or bactericidal activity on different microorganisms including Mycobacterium (Tydell et al. 2006).
The crucial role of these genes in the innate immunity to bacterial pathogens is suggestive of their probable involvement in the differences in response to MAP infection. Hence, detection of polymorphisms and exploration of their correlation with the resistance/susceptibility to MAP infection would exhibit interesting results. This study aimed to find SNPs in SLC11A1, ANKRA2, IFNG and PGLYRP1 genes and investigate their association with variable resistance/susceptibility to MAP infection in Indian cattle.
Materials and methods
Experimental animals and site
All the animal experimentation had prior approval of the Institutional Animal Ethics Committee (ICAR-Indian Veterinary Research Institute, Izatnagar, India), which follows the guidelines of CPCSEA (Committee for the Purpose of Control and Supervision of Experimentation in Animals), Government of India. The experimental animals used in this study belong to the same study population which was used in our earlier publication (Gopi et al. 2020). Briefly, 549 cattle, which consisted of 330 Sahiwal, 51 Gir, 26 Tharparkar, 7 Kankrej and 135 Frieswal were tested for identification of MAP infection status of animals. The sampled population belonged to two farms located in Punjab and UP where the incidence of PTB was 23.3 % and 27.58 %, respectively (Narnaware and Tripathi 2017; Singh et al. 2008). The highest probability of testing positive for MAP in cattle is from 2.5 to 4.5 yrs of age (Nielson and Ersbøll 2006). In other study, Chiodini et al. (1984) and Radostits et al. (2006) stated that most cases occur between 3 and 5 yr of age. Hence, cattle above 2 years of age were selected to determine the effect of age on susceptibility to MAP infection.
Phenotypic classification of animals
The physical body condition of animals was classified by visual appraisal in four scales, referred as physical body condition score (PBCS) viz. 4+, 3+, 2+ and 1+. The coding for PBCS was done as described by Kumar et al. (Kumar et al. 2019a).
Blood collection for DNA isolation and serum
Blood collection and isolation of DNA was done according to the procedure described in an earlier study (Gopi et al. 2020). Briefly, 5 ml jugular vein blood sample was collected in a K2EDTA coated vial (BD Vacutainer®) for DNA isolation and 3 ml blood was collected without anticoagulant for serum separation under sterile condition. The tubes were kept at -20 ºC until DNA isolation. Phenol-chloroform-isoamyl alcohol extraction method was used for genomic DNA isolation. The Quality of genomic DNA was assessed through 1 % horizontal agarose gel electrophoresis. NanoDrop 1000 Spectrophotometer was used for assessing the purity and concentration of DNA samples. Samples with A260 /A280 ratio of 1.7–1.9 were considered pure and used for further downstream work.
Collection of faecal samples
Faecal samples were collected per-rectum using sterile glove and transported to laboratory in ice and stored at -20 °C till further use.
Establishment of case-control population
The phenotypically classified animals were tested for MAP infection status using four diagnostic tests viz. ELISA, Johnin test, Faecal microscopy and IS900 blood PCR to establish case-control resource population as described in an earlier study (Gopi et al. 2020).
Johnin PPD test
Johnin PPD skin test for delayed type hypersensitivity was used to screen MAP positive animals. Briefly, Johnin PPD, prepared from Mycobacterium paratuberculosis containing 1 mg PPD per ml and preserved with 0.5 ml phenol, produced in Biological Products Division of Indian Veterinary Research Institute, Bareilly was utilized in this study. Intradermal inoculation of 0.1 ml Johnin PPD was done in the middle of neck as per manufacturer’s protocol. Skin thickness was measured before and after 72 h of inoculation by using vernier callipers. Animals with diffused oedema and increased thickness of about 4 mm and above at the site of injection were considered as positive for MAP infection.
Enzyme linked immunosorbent assay (ELISA)
ELISA was carried out using commercially available PARACHEK® 2 kit (Thermo Fisher Scientific) from the serum samples of all the animals to know the MAP infection status as per manufacturers protocol. The absorbance value of each well was read using a 450 nm filter and 620 nm as reference wavelength. These absorbance values were used to calculate results. The sample % positivity was calculated as per the given formula-
The serum samples which were above or equal to the cut-off of 15 % was considered as positive and below the cut-off was considered as negative for MAP infection.
Faecal microscopy
The faecal samples were processed for microscopic examination. About 2 g of faecal sample was finely grounded with 10-12 ml of distilled water and centrifuged at 4500 rpm for 45 min. at 25ºC. The supernatant was discarded and thin smear was made from the middle layer. Ziehl-Neelsen staining was performed to examine the presence of acid-fast bacilli by microscopic examination (Singh et al. 2013). Slides displaying pink coloured short rods, compatible to MAP were considered as positive for MAP infection.
IS900 blood PCR
About 500 µl of blood sample was taken in an eppendorf tube and 1ml chilled RBC lysis buffer was added. After proper mixing it was centrifuged three to four times at 2500 rpm for 10 minutes until white pellet was obtained. The white leucocyte pellet was used for DNA isolation by phenol-chloroform-isoamyl alcohol extraction method (Gopi et al. 2020). The isolated DNA pellet was dried and 20µl nuclease free water added and stored at -20oC until further use. PCR reaction mixture (25 µl) was prepared using 12.5 µl GoTaq master mix (Promega Corporation), 9.5 µl of nuclease free water (Qiagen), 0.5 µl each of forward and reverse primers and 2 µl of DNA template. PCR amplification of IS900 elements was done by using MAP specific primers, Forward primer 5’-CCGCTAATTGAGAGATGCGATTGG-3’ Reverse primer 5’-AATCAACTCCAGCAGCGCGGCCTCG-3’ (Ellingson et al. 2005). The PCR reaction condition were: initial denaturation at 94 ºC for 10 min for 1 cycle, denaturation for at 94ºC for 30 s, annealing at 61ºC for 30 s and elongation at 72ºC for 30 s for 37 cycle and final extension at 72ºC for 7 min for 1 cycle after that holding at 4ºC. The PCR amplification which shows 229 bp fragments in 2 % agarose gel electrophoresis was considered positive for MAP infection.
Selection of SNPs and genotyping
A total of 10 SNPs from IFNG, SLC11A1, ANKRA2 and PGLYRP1 genes were selected from NCBI and animal genome database (www.animalgenome.org) on the basis of available literature and primers were designed using primer3 online software(https://www.bioinformatics.nl/cgi-bin/primer3plus/primer3plus.cgi) to amplify a 200-522 bp fragment containing the desired SNPs. The restriction enzyme (RE) was identified using NEBcutter V2 program (Vincze et al. 2003). The list of SNPs and their details like dbSNP id, HGVS name, location on the gene, type of mutation and reference sequence of all the selected SNPs are given in Table 1. The list of primers, their sequences and other details are depicted in Supplementary material_Table 1. Genotyping of each SNP was determined by PCR-RFLP technique. The genotypes of animals were determined on the basis of size of RE digested fragments resolved on 3.5-4 % MetaPhorTM agarose gel stained with ethidiun bromide. MetaPhorTM Agarose (Lonza biosciences) is the highest resolution agarose, ideal for resolving small PCR fragments. The details of RE used for genotyping of SNPs are listed in Supplementary material_Table 2.
Statistical analysis
The association of PTB susceptibility/resistance to various allelic variants were analysed using SAS 9.3 software. The univariate logisitic regression analysis considered the infection status as categorical response variable whereas, SNPs, breed, herd (farm factor), age and PBCS were included as possible explanatory variables. The univariate logistic model of PROC LOGISTIC procedure of SAS was executed and only one SNP at a time was imputed in the model rather than all SNPs fitted simultaneously, to know the significance of association and to determine the odds ratio (OR) of each genotype. The logistic regression model was used to determine the relative risk of incidence among the genotypes is given below.
This regression model estimates one/two OR for heterozygous (Ht) and homozygous (H0) genotypes where α, β1, β2 and γ were regression coefficients associated with population, heterozygotes, homozygotes and error, respectively. Genotype frequencies were tested for departure from Hardy–Weinberg equilibrium (HWE) by using Exact Test and Chi-square test. The PROCALLELE, PROC FREQ and CHISQ EXACT procedure of SAS were used for testing linkage disequilibrium (LD), Hardy–Weinberg equilibrium (HWE), estimation of heterozygosity and polymorphism information content (PIC) of SNPs markers. The HWE was tested for each SNPS in the whole case-control population. The significance level for all the tests were considered as P<0.05 for significant difference. For adjusting the p-value, Exact statistics was applied in SAS for yate’s correction of genotypes having observations less than 5. The corrected p value was determined by mid p value of Exact conditional analysis.
Results and discussion
PTB is a complex disease, the outcome of which is affected by mutations in genes regulating the immune system and interaction between host and pathogen. Therefore, exploring the causal relationship between genetic variation in innate immune genes viz. SLC11A1, IFNG, ANKRA2 and PGLYRP1 might help in identifying animals with inherent resistance to MAP infection. Association of these genes with MAP infection have been reported in cattle by earlier studies (Pinedo et al. 2009b; Ruiz-Larrañaga et al. 2010; Casas et al. 2011; Vázquez et al. 2014) however, to the best of our knowledge this is the first report on association of ANKRA2, IFNG and SLC11A1 genes with bovine PTB in India.
Establishment of case-control population
Screening of infection status of animals was done by multiple diagnostic tests to establish resource population for further investigation. Accordingly, out of 549 animals tested, we obtained 50 animals which exhibited positive infection status in at least two tests and thus were included in the case population. The result of diagnostic tests for case population is depicted in Supplementary material_Table 3. Other 50 animals with negative results in all the four diagnostic tests were included in control population randomly. The resource population consisted of 27 (25 Sahiwal and 2 Tharparkar) animals from farm 1 and 23 ( Frieswal) from farm 2 in case population, and similarly in control population, 27 animals (25 Sahiwal and 2 Tharparkar) were from farm 1 and 23 ( Frieswal) animals from farm 2. Early infection to MAP may go unnoticed because the overt signs of the disease are not manifested and the bacterial loads are not enough to be diagnosed (Vázquez et al. 2014). This may lead to misclassification of animals and is the major limitation of case-control studies especially for PTB due to lack of a gold standard in vivo diagnostic test (Chiodini et al. 1984). Previous studies have advocated that the basis of classification of individuals should be wisely chosen while working on genetic association studies (Vázquez et al. 2014). Moreover, there is a lot of variability in sensitivity and specificity of different diagnostic tests available and no single test can rule out the possibility of MAP infection (Garg et al. 2015). Therefore, we have used multiple diagnostic tests namely ELISA, intradermal Johnin test, faecal microscopy and IS900 blood PCR to identify MAP infection status of animals. By doing so, we have tried to establish a robust resource population without any false positive in case and false negative in control population. On the one hand it provided strength to our resource population but on the other hand it caused reduction in the size of our resource population and this is the limitation to our study. However, many researchers have used the same approach to establish the case-control resource population. Yadav et al. (2014) has done the association study for PTB resistance with a case-control resource population of 93 cattle by intradermal Johnin PPD test and ELISA. Sadana et al. (2015) reported association study of MAP infection with a case-control resource population of 94 cattle only by ELISA. Kumar et al. (2019a, b) has also reported association study of MAP infection with a case-control resource population of 102 cattle. Similarly, Prakash et al. (2014) had reported the association of SNPs in innate immune genes with bovine brucellosis using a resource population of 83 cattle.
Effect of non-genetic factors on occurrence of PTB
The breed, herd, PBCS and age did not differ significantly (p>0.05) for the MAP infection in the study population as reported in our earlier publications. These findings corroborate several earlier studies that reported, breed, PBCS and age were did not affect the MAP infection outcome in cattle (Chase et al. 2008; Kumar 2015). The OR of Sahiwal and Tharparkar vs. Frieswal was 1.0 (0.438-2.286 ) and 1.0 (0.255-3.926), respectively. These results were in accordance with other reports which did not find significant effect (p>0.05) of breed on the occurrence of disease (Sadana et al. 2015; Yadav et al. 2014; Kumar et al. 2019a). PBCS did not differ significantly (p>0.05) between case and control population. The OR of animals with PBCS 2+ and 3+ vs. 4+ was 4.500 and 1.125, respectively. However, the effect is nonsignificant, OR indicates that animals with poor PBSC (2+ and 3+) were at higher risk of MAP infection than animals with 4+ PBSC. Our study corroborates Sadana et al. (2015) where PBSC had no effect on MAP infection. The effect of herd was also nonsignificant on the MAP infection status. The odds of herd 1 vs. herd 2 was 1.0 (0.455-2.196) at 95 % CI which explains that both farms were similar for MAP infection status. The corrected p value for the effect of herd on MAP infection was 0.9206, indicating that the effect of farm or herd on the MAP infection was not significantly different. The odds for animals with low age group (up to 3 years) compared to high age group (above 3 years) was 1.728 (0.853 – 4.781) at 95 % CI. Despite the non-significant effect of age (p>0.05) lower age group was at a higher risk of infection than higher age group animals. This finding is in accordance with an earlier study of Chase et al. (2008), where they showed that young animals are most susceptible, most likely due to their lower immune competence. Kumar (2015) also reported that lower age group was at a higher risk of infection than higher age group animals.
HWE, LD test and polymorphic parameters
The HWE was checked for whole case-control population for all the SNPs and it was found that rs110514940, rs109453173, rs41933863 and rs68268283 SNPs deviated from HWE while all other investigated loci were in HWE. This departure may be possibly due to selection, chance factor induced genotypic error and population under inbreeding with assortative mating. This constitutes a limitation for this study. However, the possibility of population stratification in our study is less because of absence of association with MAP infection in other unlinked markers concurrently. This concern is also reduced due to the fact that the case-control animals belong to the same sub-population means case and control population has equal animals from farm 1 and farm 2 and the farm was not differing significantly in case and control population. The PIC ranged from 17.6 % (rs110217377) to 35.89 % (rs68268283) in our resource population under study. Heterozygosity ranged from 12.0 % (rs110515940) to 47.0 % (rs109453173) in case-control population. The PIC, average heterozygosity and HWE of each SNP locus are shown in Table 2. Chi square probabilities was used for testing LD of different loci and it showed that many of SNP in our study were linked together, but most of these were non-significantly associated with disease. Hence, probability of obtaining haplotypes from the significant loci affecting PTB were rare.
Effect of SNPs in candidate genes on resistance/susceptibility of PTB disease
The information on the association parameters of all SNPs, distribution of allele and genotype frequency, OR, p-Value and corrected p-Value under study are given in Tables 3 and 4, respectively.
Effect of SNPs in SLC11A1 gene
SLC11A1 gene translates into integral membrane protein, NRAMP1, is expressed exclusively in the monocytes and macrophages lysosomal compartment and regulates the level of iron within phagosomes (Canonne-Hergaux et al. 1999). It indicates that SLC11A1 may affect the predisposition to intracellular pathogens, especially Mycobacterium spp. Two SNPs viz. rs110514940 and rs109453173 were amplified with specific primers and digested with RE AciI and BpmI, respectively and the SNPs were found to be polymorphic in our case-control population. SNP rs110514940 yielded two genotypes viz. AC and CC with genotype frequencies 4 and 96 in case and 0 and 100 in control population, respectively. The allele frequency of A and C were 2 and 98 in case and 0 and 100 in control population, respectively. Neither allele nor genotype frequencies differed significantly (p>0.05) between case-control resource population.
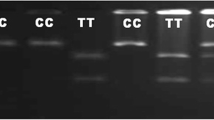
The other SNP rs109453173 yielded two alleles C and G and consequently three genotypes namely CC, CG and GG with their respective frequencies 2, 62 and 36 in case and 2, 32 and 66 in control population (Table 4). The allele frequencies of C and G alleles were 33 and 67 in case and 18 and 82 in control population, respectively (Table 3). The allele and genotype frequency differs significantly (p<0.05) between case-control resource population. The genotypic profile of SNP rs109453173 is depicted in Supplementary material_ Fig. 1. The OR of CC and CG verses GG genotype was 1.833 and 3.552, respectively. It denotes the probability of PTB occurrence in animals having CC and CG genotype were 1.833 and 3.55 times more as compared to GG genotypes, respectively. It was supported by the greater odds of animals having C allele (OR 2.24) as compared to G allele. Here one interesting finding may be noted that animals with CG genotypes were more in case population i.e. heterozygous disadvantage was showed by the genotypes at this locus. If resistance against MAP infection during selection is desired, a breeding programme favouring GG genotype and G allele in SNP rs109453173 may be beneficial. Consequently, G allele will be fixed in the population. However, due to less no. of animals and wide range of OR in CC genotype, the association of this SNP for MAP infection need to be validated in a larger population to warrant the strength of association of the study. Our study is in line with the previous studies who reported the association of SLC11A1 gene with PTB not only in bovine (Pinedo et al. 2009; Ruiz-Larrañaga et al. 2010; Vázquez et al. 2014) but also in sheep and goat (Reddacliff et al. 2005; Abraham et al. 2017). Furthermore, an association between CD, MAP infection and polymorphism in SLC11A1 gene was also reported in humans (Sechi et al. 2006) and mice (Roupie et al. 2008). The polymorphisms of SLC11A1 gene have been associated with resistance to many infectious agents and diseases like tuberculosis (Baqir et al. 2016; Holder et al. 2020) and brucellosis in cattle (Prakash et al. 2014) and buffalo (Ganguly et al. 2011), pneumonia in horses (Halbert et al. 2006), salmonellosis in pig (Ding et al. 2014), leishmaniasis in dog (Sanchez-Robert et al. 2008), inflammatory bowel disease (IBD) (Stewart et al. 2010) and auto immune disease (Awomoyi 2007) in humans, which underlines the importance of this gene in determining the immune traits.
Effect of SNPs in ANKRA2 gene
Four SNPs in ANKRA2 gene viz., rs17871543, rs41933863, rs41933906 and rs17870710 were studied by amplifying with specific primers and digested with RE BsrI, AciI, EcoRV and SmII, respectively. Out of this SNP rs41933863 was polymorphic and other 3 SNPs viz. rs17871543, rs41933906 and rs17870710 were monomorphic with CC, CC and GG genotype, respectively. The rs41933863 yielded two alleles A and G and consequently three genotypes viz. AA, AG and GG with genotypic frequencies 38, 8 and 54 in case and 20, 26 and 54 in control population, respectively (Table 4). The allele frequencies of A and G in case and control population were 42 and 58 and 33 and 67, respectively (Table 3). Genotype frequencies were significantly different (p<0.05) between case-control resource population and hence, it was inferred that a significant association (p<0.05) of ANKRA2 gene exists with resistance to PTB. The genotypic profile of SNP rs41933863 is depicted in Supplementary material_Figure 2. The OR of AA and AG genotypes verses GG genotype was 1.900 and 0.308, respectively, indicating that animals with AG genotypes were three and six times more resistant than animals with GG and AA genotype, respectively (Table 4). This may be a case of transgressive inheritance in which the heterozygotes perform better than both the contemporary homozygotes (Rieseberg et al. 1999). In this condition also the AG genotype is associated with more resistance against MAP infection than both homozygotes and animals with AG genotypes were more in control than in case population. Hence, a selection program favouring AG genotype may be beneficial for developing a MAP infection resistant herd. The odds of allele A versus G was 1.47, which revealed that animal with A allele were at a higher risk of PTB occurrence than cattle with G allele. From the odds of A vs. G allele, it is evident that the G allele is associated with resistance against MAP infection but due to the heterosis and transgressive inheritance AG heterozygote is associated with resistance against MAP infection. Our results are in line with the study by Casas et al. (2011) who found the significant association of SNP in ANKRA2 gene with MAP infection and bovine respiratory disease in Brahman x Angus crossbred cattle (Casas et al. 2011). No other studies have reported the association of ANKRA2 gene to any other diseases in animal or human. The ANKRA2 gene plays a key role in regulating the function of MHC-II genes and mutations in ANKRA2 produce the bare lymphocyte syndrome causing immunodeficiency in humans (Masternak et al. 1998). Thus, ANKRA2 gene play critical role in MHC-II driven immunological response in livestock, bearing implications in the immune response against mycobacteria.
Effect of SNPs in IFNG gene
Two SNPs viz. rs110853455 and rs382197650 were genotyped by amplifying with specific primers and digested with RE HphI and HpaII, respectively. In our population, rs382197650 was monomorphic with GG genotype and SNP rs110853455 was polymorphic having two alleles A and G and consequently three genotypes viz., GG, AG and AA. The genotype frequencies were 60, 40 and 0 in case and 72, 22 and 6 in control population, respectively. The allele frequencies of A and G in case and control population were 20 and 80 and 17 and 83, respectively. The logistic regression analysis revealed the odds of AG (OR 2.18) genotypes were higher than AA (OR <0.001) and GG genotype (OR 1.0) (Table 4). Similarly, the odds of A allele (OR 1.22) was higher than G allele for MAP infection. The genotypes in rs110853455 were differing significantly (p<0.05) in case-control population against MAP infection. However, due to less no. of animals and wide range of OR at 95 % CI in AA genotype, the association of this SNP for MAP infection need to be validated in a larger population to warrant the strength of association of the study. Our results are in agreement with the earlier reports (Pinedo et al. 2009b; Pant et al. 2011b) who reported the significant association of SNPs in IFNG gene and MAP infection. However, non-significant association of SNPs in INFG gene with PTB were reported in previous studies in cattle (Hinger et al. 2007; Sadana et al. 2015). The possible reason for disagreements in results may be due to different resource population used for the study because herd composition also has significant effect on the prevalence of PTB disease (Pinedo et al. 2009b). Moreover, the polymorphism in IFNG was found to be associated with gastrointestinal nematodes in sheep and goats (Dervishi et al. 2011; Bressani et al. 2014), tuberculosis in buffalo (Iannaccone et al. 2018), resistance to ticks infestation in cattle (Maryam et al. 2012) and tuberculosis, IBD and brucellosis in human (Gonsky et al. 2014; Wu et al. 2019).
Effect of SNPs in PGLYRP1 gene
Two SNPs viz. rs68268283 and rs110217377, were amplified by specific primers and digested with RE PspOMI and BsmI, respectively. Both the SNPs were polymorphic in our case-control population. The rs68268283 yielded two allele C and G and consequently three genotypes viz. GG, CG and CC with genotype frequencies 14, 14 and 72 in case and 12, 10 and 78 in control population, respectively. The allele frequency of C and G allele was 79 and 21 in case and 83 and 17 in control population, respectively. The odds of CG (OR 1.20) genotype were higher than animals with GG (OR 1.0) and CC genotype (OR 0.70). The other SNP, rs110217377 yielded two allele G and T and consequently two genotypes viz. GG and GT with genotypic frequency 76 and 24 in case and 80 and 20 in control population, respectively. The allele frequency of G and T in case and control population were 88 and 12 and 90 and 10, respectively. The odds of animals with GG (OR 0.79) genotypes were less than animals with GT genotype which is supported by the odds of allele where animals with G allele (OR 0.81) were lower than animals with T allele (OR 1.0).
PGLYRP1 gene is highly expressed in M cells of intestinal epithelium and peyer’s patches, which are the portals of entry for Mycobacterium sp.(Lo et al. 2003). The logistic regression analysis revealed that both SNPs in PGLYRP1 gene were not significantly associated with PTB disease and neither allele nor genotypes were significantly different in case-control population. However, Pant et al. (2011a) reported that SNP in PGLYRP1 gene was associated with susceptibility to MAP infection (Pant et al. 2011a). The possible reason for this conflicting result may be different resource population or the different approaches to identify and classify the case-control population. We have used a panel of four diagnostic tests which reduces the chance of misclassification of case-control animals whereas earlier study has used only ELISA for determining infection status of animal. The SNPs in PGLYRP1 gene was found to be associated with mastitis (Wang et al. 2013; Shivashanker et al. 2018) and generalized infection status in cattle (Sablik et al. 2020) rheumatoid arthritis (Fodil et al. 2015), Kawasaki disease (Onoyama et al. 2012), IBD and ulcerative colitis (Zulfiqar et al. 2013) in humans which indicates importance of this gene in innate immune responses against morbid and comorbid conditions. Although PGLYRP1 gene failed to show significant association with MAP infection in our study, the analyses are indicative of a probable association between polymorphisms in the PGLYRP1 gene and PTB infection in cattle.
Our results suggest that the SNPs rs41933863 in ANKRA2, rs109453173 in SLC11A1 and rs110853455 in IFNG genes are associated with the resistance against the MAP infection which were also reported by earlier studies (Pinedo et al. 2009b; Ruiz-Larrañaga et al. 2010; Casas et al. 2011; Vázquez et al. 2014) and hence, could be included in the marker panel for identification of PTB susceptible/resistant animals. A selection program favouring AG genotype of SNP rs41933863, GG genotype and G allele of SNP rs109453173 and against AG genotype for rs110853455 would be beneficial in conferring resistance against PTB. However, this needs further validation in an independent and larger resource population to warrant the strength of association.
Data availability
The data will be available on request to the corresponding author.
Code availability
Not applicable.
References
Abraham A, Naicy T, Raghavan KC, Siju J, Aravindakshan T (2017) Evaluation of the association of SLC11A1 gene polymorphism with incidence of paratuberculosis in goats. J Genet 96(4):641–646. https://doi.org/10.1007/s12041-017-0820-9
Awomoyi AA (2007) The human solute carrier family 11 member 1 protein (SLC11A1): Linking infections, autoimmunity and cancer? FEMS Immunol Med Microbiol 49(3):324–329. https://doi.org/10.1111/j.1574-695X.2007.00231.x
Baqir M, Bhushan S, Kumar A et al (2016) Association of polymorphisms in SLC11A1 gene with bovine tuberculosis trait among Indian cattle. J Appl Anim Res 44(1):380–383. https://doi.org/10.1080/09712119.2015.1091333
Bastida F, Juste RA (2011) Paratuberculosis control: A review with a focus on vaccination. J Immune Based Ther Vaccines 9(1):1–7. https://doi.org/10.1186/1476-8518-9-8
Beard PM, Daniels MJ, Henderson D, Pirie A, Rudge K, Buxton D, Rhind S, Greig A, Hutchings MR, McKendrick I, Stevenson K (2001a) Paratuberculosis infection of nonruminant wildlife in Scotland. J Clin Microbiol 39(4):1517–1521. https://doi.org/10.1128/FJCM.39.4.1517-1521.2001
Beard PM, Rhind SM, Buxton D, Daniels MJ, Henderson D, Pirie A, Rudge K, Greig A, Hutchings MR, Stevenson K, Sharp JM (2001b) Natural paratuberculosis infection in rabbits in Scotland. J Comp Pathol 124(4):290–299. https://doi.org/10.1053/jcpa.2001.0466
Bhutediya JM, Dandapat P, Chakrabarty A, Das R, Nanda PK, Bandyopadhyay S, Biswas TK (2017) Prevalence of paratuberculosis in organized and unorganized dairy cattle herds in West Bengal, India. Vet World 10(6):574–579. https://doi.org/10.14202/vetworld.2017.574-579
Bishop SC, MacKenzie KM (2003) Genetic management strategies for controlling infectious diseases in livestock populations. Genet Sel Evol 35(1):1–5. https://doi.org/10.1186/1297-9686-35-s1-s3
Bressani FA, Tizioto PC, Giglioti R et al (2014) Single nucleotide polymorphisms in candidate genes associated with gastrointestinal nematode infection in goats. Genet Mol Res 13(4):8530–8536. https://doi.org/10.4238/2014.October.20.29
Canonne-Hergaux F, Gruenheid S, Govoni G, Gros P (1999) The Nramp1 protein and its role in resistance to infection and macrophage function. Proc Assoc Am Physicians 111(4):283-289. https://doi.org/10.1046/j.1525-1381.1999.99236.x
Casas E, Garcia MD, Wells JE, Smith TPL (2011) Association of single nucleotide polymorphisms in the ANKRA2 and CD180 genes with bovine respiratory disease and presence of Mycobacterium avium subsp. paratuberculosis. Anim Genet 42(6):571–577. https://doi.org/10.1111/j.1365-2052.2011.02189.x
Chase CCL, Hurley DJ, Reber AJ (2008) Neonatal immune development in the calf and its impact on vaccine response. Vet Clin North Am - Food Anim Pract 24(1):87–104. https://doi.org/10.1016/j.cvfa.2007.11.001
Chiodini RJ, Van Kruiningen HJ, Merkal RS (1984) Ruminant paratuberculosis (Johne’s disease): the current status and future prospects. Cornell Vet 74:218–262
Cossu D, Masala S, Cocco E, Paccagnini D, Tranquilli S, Frau J, Marrosu MG, Sechi LA (2013) Association of Mycobacterium avium subsp. paratuberculosis and SLC11A1 polymorphisms in Sardinian multiple sclerosis patients. J Infect Dev Ctries 7(03):203-207. https://doi.org/10.3855/jidc.2737
Coussens PM, Verman N, Coussens MA, Elftman MD, McNulty AM (2004) Cytokine gene expression in peripheral blood mononuclear cells and tissues of cattle infected with Mycobacterium avium subsp. paratuberculosis: Evidence for an inherent proinflammatory gene expression pattern. Infect Immun 72(3):1409–1422. https://doi.org/10.1128/IAI.72.3.1409-1422.2004
Dervishi E, Uriarte J, Valderrábano J, Calvo JH (2011) Structural and functional characterisation of the ovine interferon gamma (IFNG) gene: Its role in nematode resistance in Rasa Aragonesa ewes. Vet Immunol Immunopathol 141(1-2):100–108. https://doi.org/10.1016/j.vetimm.2011.02.013
Ding X, Zhang X, Yang Y, Ding Y, Xue W, Meng Y, Zhu W, Yin Z (2014) Polymorphism, expression of natural resistance-associated macrophage protein 1 encoding gene (NRAMP1) and its association with immune traits in pigs. Asian-Australasian J Anim Sci 27(8):1189–1195. https://doi.org/10.5713/ajas.2014.14017
Ellingson JLE, Anderson JL, Koziczkowski JJ, Radcliff RP, Sloan SJ, Allen SE, Sullivan NM (2005) Detection of viable Mycobacterium avium subsp. paratuberculosis in retail pasteurized whole milk by two culture methods and PCR. J Food Prot 68(5):966–972. https://doi.org/10.4315/0362-028X-68.5.966
Fodil M, Teixeira VH, Chaudru V, Hilliquin P, Bombardieri S, Balsa A, Westhovens R, Barrera P, Alves H, Migliorin P, Bardin T (2015) Relationship between SNPs and expression level for candidate genes in rheumatoid arthritis. Scand J Rheumatol 44(1):2–7. https://doi.org/10.3109/03009742.2014.918175
Fox NJ, Smith LA, Stevenson K, Davidson RS, Marion G, Hutchings MR (2020) Infection of non-ruminant wildlife by Mycobacterium avium subsp. paratuberculosis. In Behr MA, Stevenson K, Kapur V (eds). Paratuberculosis 2020:200-212
Ganguly I, Sharma A, Mitra A, Kumar N, Ganguly A (2011) Analysis of genetic variations of complete tm4 of buffalo (Bubalus bubalis) SLC11A1 gene. J Appl Anim Res 39(4):324–327. https://doi.org/10.1080/09712119.2011.607943
Garcia AB, Shalloo L (2015) Invited review: The economic impact and control of paratuberculosis in cattle. J Dairy Sci 98(8):5019–5039. https://doi.org/10.3168/jds.2014-9241
Garg R, Patil PK, Singh SV, Sharma S, Gandham RK, Singh AV, Filia G, Singh PK, Jayaraman S, Gupta S, Chaubey KK (2015) Comparative evaluation of different test combinations for diagnosis of Mycobacterium avium subspecies paratuberculosis infecting dairy herds in India. Biomed Res Int :Article ID 983978. https://doi.org/10.1155/2015/983978
Gonda MG, Chang YM, Shook GE, Collins MT, Kirkpatrick BW (2006) Genetic variation of Mycobacterium avium ssp. paratuberculosis infection in US Holsteins. J Dairy Sci 89(5):1804–1812. https://doi.org/10.3168/jds.S0022-0302(06)72249-4
Gonsky R, Deem RL, Landers CJ, Haritunians T, Yang S, Targan SR (2014) IFNG rs1861494 polymorphism is associated with IBD disease severity and functional changes in both IFNG methylation and protein secretion. Inflamm Bowel Dis 20(10):1794-1801. https://doi.org/10.1097/MIB.0000000000000172
Gopi B, Singh RV, Kumar S, Kumar S, Chauhan A, Kumar A, Singh SV (2020) Single-nucleotide polymorphisms in CLEC7A, CD209 and TLR4 gene and their association with susceptibility to paratuberculosis in Indian cattle. J Genet 99(1):1–10. https://doi.org/10.1007/s12041-019-1172-4
Gupta A, Rani M, Agrawal S, Kumar Gupta P (2012) Sero-prevalence of paratuberculosis (Johne’s Disease) in cattle population of south-western bangalore using ELISA kit. Open J Vet Med 02(4):196–200. https://doi.org/10.4236/ojvm.2012.24031
Halbert ND, Cohen ND, Slovis NM, Faircloth J, Martens RJ (2006) Variations in equid SLC11A1 (NRAMP1) genes and associations with Rhodococcus equi pneumonia in horses. J Vet Intern Med 20(4):974–979. https://doi.org/10.1892/0891-6640(2006)20[974:VIESNG]2.0.CO;2
Hinger M, Brandt H, Horner S, Erhard G (2007) Short communication: Association analysis of microsatellites and Mycobacterium avium subspecies paratuberculosis antibody response in German holsteins. J Dairy Sci 90(4):1957–1961. https://doi.org/10.3168/jds.2006-510
Holder A, Garty R, Elder C, Mesnard P, Laquerbe C, Bartens MC, Salavati M, Shabbir MZ, Tzelos T, Connelly T, Villarreal-Ramos B (2020) Analysis of genetic variation in the bovine SLC11A1 Gene, Its influence on the expression of NRAMP1 and potential association with resistance to bovine tuberculosis. Front Microbiol 11:1420. https://doi.org/10.3389/fmicb.2020.01420
Hruska K, Bartos M, Kralik P, Pavlik I (2005) Mycobacterium avium subsp. paratuberculosis in powdered infant milk: paratuberculosis in cattle-the public health problem to be solved. Vet Med (Praha) 50(8):327–335
Hruska K, Slana I, Kralik P, Pavlik I (2011) Mycobacterium avium subsp. paratuberculosis in powdered infant milk: F57 competitive real time PCR. Vet Med (Praha) 56(5):226–230
Iannaccone M, Cosenza G, Pauciullo A, Martino G, Capparelli R (2018) The SNP g.4667G>A at 3′-UTR of IFNG gene is associated with susceptibility to bovine tuberculosis in Mediterranean water buffalo (Bubalus bubalis). Anim Genet 49(5):496–497. https://doi.org/10.1111/age.12698
Juste RA, Perez V (2011) Control of paratuberculosis in sheep and goats. Vet Clin North Am - Food Anim Pract 27(1):127–138. https://doi.org/10.1016/j.cvfa.2010.10.020
Juste RA, Vazquez P, Ruiz-Larrañaga O, Iriondo M, Manzano C, Agirre M, Estonba A, Geijo MV, Molina E, Sevilla IA, Alonso-Hearn M (2018) Association between combinations of genetic polymorphisms and epidemiopathogenic forms of bovine paratuberculosis. Heliyon 4(2):e00535. https://doi.org/10.1016/j.heliyon.2018.e00535
Krawczyk M, Masternak K, Zufferey M, Barras E, Reith W (2005) New functions of the major histocompatibility complex class II-specific transcription factor RFXANK revealed by a high-resolution mutagenesis study. Mol Cell Biol 25(19):8607–8618. https://doi.org/10.1128/mcb.25.19.8607-8618.2005
Kumar S (2015) Single nucleotide polymorphism in candidate genes and their association with occurrence of paratuberculosis in cattle. IVRI (Deemed University), Izatnagar Bareilly, UP, India. Accessed on 20-03-2021. http://krishikosh.egranth.ac.in/handle/1/5810096974
Kumar S, Kumar S, Singh R, Chauhan A, Agrawal S, Kumar A, Singh S (2017) Investigation of genetic association of single nucleotide polymorphisms in SP110 gene with occurrence of paratuberculosis disease in cattle. Int J Livest Res 7:81–88. https://doi.org/10.5455/ijlr.20170223021951
Kumar S, Kumar S, Singh RV, Chauhan A, Kumar A, Bharati J, Singh SV (2020) Association of genetic variability in CD209 gene with bovine paratuberculosis disease: a case–control study in the Indian cattle population. Anim Biotechnol 25:1–8. https://doi.org/10.1080/10495398.2020.1823400
Kumar S, Kumar S, Singh RV, Chauhan A, Kumar A, Bharati J, Singh SV (2019a) Association of Bovine CLEC7A gene polymorphism with host susceptibility to paratuberculosis disease in Indian cattle. Res Vet Sci 123:216–222. https://doi.org/10.1016/j.rvsc.2019.01.016
Kumar S, Kumar S, Singh RV, Chauhan A, Kumar A, Sulabh S, Bharati J, Singh SV (2019b) Genetic association of polymorphisms in bovine TLR2 and TLR4 genes with Mycobacterium avium subspecies paratuberculosis infection in Indian cattle population. Vet Res Commun 43(2):105–114. https://doi.org/10.1007/s11259-019-09750-2
Larsen AB, Moon HW, Merkal RS (1971) Susceptibility of swine to Mycobacterium paratuberculosis. Am J Vet Res 32(4):589–595
Lo D, Tynan W, Dickerson J, Mendy J, Chang HW, Scharf M, Byrne D, Brayden D, Higgins L, Evans C, O’Mahony DJ (2003) Peptidoglycan recognition protein expression in mouse Peyer’s Patch follicle associated epithelium suggests functional specialization. Cell Immunol 224(1):8–16. https://doi.org/10.1016/S0008-8749(03)00155-2
Losinger WC (2005) Economic impact of reduced milk production associated with Johne’s disease on dairy operations in the USA. J Dairy Res 72(4):425–432. https://doi.org/10.1017/S0022029905001007
Maryam J, Babar ME, Nadeem A, Hussain T (2012) Genetic variants in interferon gamma (IFN-c) gene are associated with resistance against ticks in Bos taurus and Bos indicus. Mol Biol Rep 39(4):4565–4570. https://doi.org/10.1007/s11033-011-1246-8
Masternak K, Barras E, Zufferey M, Conrad B, Corthals G, Aebersold R, Sanchez JC, Hochstrasser DF, Mach B, Reith WA (1998) A gene encoding a novel RFX-associated transactivator is mutated in the majority of MHC class II deficiency patients. Nat Genet 20(3):273–277. https://doi.org/10.1038/3081
Miller MA, Davey SC, van Helden LS, Kettner F, Weltan SM, Last R, Grewar JD, Botha L, Van Helden PD (2017) Paratuberculosis in a domestic dog in South Africa. J S Afr Vet Assoc 88(1):1–5. https://doi.org/10.4102/jsava.v88i0.1441
Narnaware SD, Tripathi BN (2017) Seroepidemiology of paratuberculosis in cattle population of organized and unorganized farms of India. Indian J Anim Sci 87(1):21–24. https://www.researchgate.net/profile/Shirish-Narnaware/publication/313859367_Seroepidemiology_of_paratuberculosis_in_cattle_population_of_organized_and_unorganized_farms_of_India/links/58abcfb2aca27206d9bd1b24/Seroepidemiology-of-paratuberculosis-in-cattle-population-of-organized-and-unorganized-farms-of-India.pdf
Nielsen SS, Ersbøll AK (2006) Age at occurrence of Mycobacterium avium subspecies paratuberculosis in naturally infected dairy cows. J Dairy Sci 89(12):4557–4566
Nielsen SS, Toft N (2008) Ante mortem diagnosis of paratuberculosis: A review of accuracies of ELISA, interferon-γ assay and faecal culture techniques. Vet Microbiol 129(3-4):217–235. https://doi.org/10.1016/j.vetmic.2007.12.011
OIE (2020) World organisation for animal health. https://www.oie.int/en/animal-health-in-the-world/animal-diseases/PTB/. Accessed 11-06-2020
Onoyama S, Ihara K, Yamaguchi Y, Ikeda K, Yamaguchi K, Yamamura K, Hoshina T, Mizuno Y, Hara T (2012) Genetic susceptibility to Kawasaki disease: Analysis of pattern recognition receptor genes. Hum Immunol 73(6):654–660. https://doi.org/10.1016/j.humimm.2012.03.011
Paixão TA, Martinez R, Santos RL (2012) Polymorphisms of the coding region of SLC11A1 (Nramp1) gene associated to natural resistance against bovine brucellosis. Arq Bras Med Vet e Zootec 64(4):1081–1084. https://doi.org/10.1590/S0102-09352012000400041
Pant SD, Verschoor CP, Schenkel FS, You Q, Kelton DF, Karrow NA (2011a) Bovine PGLYRP1 polymorphisms and their association with resistance to Mycobacterium avium ssp. Paratuberculosis. Anim Genet 42(4):354–360. https://doi.org/10.1111/j.1365-2052.2010.02153.x
Pant SD, Verschoor CP, Skelding AM, Schenkel FS, You Q, Biggar GA, Kelton DF, Karrow NA (2011b) Bovine IFNGR2, IL12RB1, IL12RB2, and IL23R polymorphisms and MAP infection status. Mamm Genome 22(9):583–588. https://doi.org/10.1007/s00335-011-9332-8
Pavlik I, Jahn P, Dvorska L, Bartos M, Novotny L, Halouzka R (2004) Mycobacterial infections in horses: a review of the literature. Vet Med 49(11):427. https://doi.org/10.17221/5733-VETMED
Pinedo PJ, Buergelt CD, Donovan GA, Melendez P, Morel L, Wu R, Langaee TY, Rae DO (2009a) Association between CARD15/NOD2 gene polymorphisms and paratuberculosis infection in cattle. Vet Microbiol 134(3-4):346–352. https://doi.org/10.1016/j.vetmic.2008.09.052
Pinedo PJ, Buergelt CD, Donovan GA, Melendez P, Morel L, Wu R, Langaee TY, Rae DO (2009b) Candidate gene polymorphisms (BoIFNG, TLR4, SLC11A1) as risk factors for paratuberculosis infection in cattle. Prev Vet Med 91(2-4):189-196. https://doi.org/10.1016/j.prevetmed.2009.05.020
Pithua P, Godden SM, Wells SJ, Stabel JR (2011) Evaluation of the risk of paratuberculosis in adult cows fed Mycobacterium avium subsp paratuberculosis DNA-positive or-negative colostrum as calves. Am J Vet Res 72(11):1456–1464. https://doi.org/10.2460/ajvr.72.11.1456
Prakash O, Kumar A, Sonwane A, Rathore R, Singh RV, Chauhan A, Kumar P, Renjith R, Yadav R, Bhaladhare A, Baqir M (2014) Polymorphism of cytokine and innate immunity genes associated with bovine brucellosis in cattle. Mol Biol Rep 41(5):2815–2825. https://doi.org/10.1007/s11033-014-3136-3
Pritchard TC, Coffey MP, Bond KS, Hutchings MR, Wall E (2017) Phenotypic effects of subclinical paratuberculosis (Johne’s disease) in dairy cattle. J Dairy Sci 100(1):679–690. https://doi.org/10.3168/jds.2016-11323
Radostits OM, Gay CC, Hinchcliff KW, Constable PD (eds) (2006) Veterinary Medicine E-Book: A textbook of the diseases of cattle, horses, sheep, pigs and goats. Elsevier Health Sciences, Amsterdam
Rawat KD, Chaudhary S, Kumar N, Gupta S, Chaubey KK, Singh SV, Dhama K, Deb R (2014) Economic losses in a commercial dairy farm due to the outbreak of Johne’s disease in India. Res J Vet Pract 2:73–77. https://doi.org/10.14737/journal.rjvp/2.5.73.77
Reddacliff LA, Beh K, McGregor H, Whittington RJ (2005) A preliminary study of possible genetic influences on the susceptibility of sheep to Johne’s disease. Aust Vet J 83(7):435–441. https://doi.org/10.1111/j.1751-0813.2005.tb13087.x
Rieseberg LH, Archer MA, Wayne RK (1999) Transgressive segregation, adaptation and speciation. Heredity 83(4):363–372
Roupie V, Rosseels V, Piersoel V, Zinniel DK, Barletta RG, Huygen K (2008) Genetic resistance of mice to Mycobacterium paratuberculosis is influenced by SLC11A1 at the early but not at the late stage of infection. Infect Immun 76(5):2099–2105. https://doi.org/10.1128/IAI.01137-07
Ruiz-Larrañaga O, Garrido JM, Manzano C, Iriondo M, Molina E, Gil A, Koets AP, Rutten VP, Juste RA, Estonba A (2010) Identification of single nucleotide polymorphisms in the bovine solute carrier family 11 member 1 (SLC11A1) gene and their association with infection by Mycobacterium avium subspecies paratuberculosis. J Dairy Sci 93(4):1713–1721
Sablik P, Klenowicz A, Szewczuk M, Olszewski A, Dybus A (2020) The Effect of Polymorphism in PGLYRP1 Gene on the Productivity and Health Traits in Holstein-Friesian Cattle. Russ J Genet 56:333–338. https://doi.org/10.1134/S1022795420030138
Sadana T, Singh RV, Singh SV, Saxena VK, Sharma D, Singh PK, Kumar N, Gupta S, Chaubey KK, Jayaraman S, Tiwari R, Dhama K, Bhatia AK, Sohal JS (2015) Single nucleotide polymorphism of SLC11A1, CARD15, IFNG and TLR2 genes and their association with Mycobacterium avium subspecies paratuberculosis infection in native Indian cattle population. Indian J Biotechnol 14:469-475. Accessed on 20-03-2021. http://nopr.niscair.res.in/bitstream/123456789/33994/1/IJBT%2014(4)%20469-475.pdf
Sanchez-Robert E, Altet L, Utzet-Sadurni M, Giger U, Sanchez A, Francino O (2008) SLC11A1 (formerly Nramp1) and susceptibility to canine visceral leishmaniasis. Vet Res 39(3):36 https://doi.org/10.1051/vetres:2008013
Sechi LA, Dow CT (2015) Mycobacterium avium ss. paratuberculosis Zoonosis - The Hundred Year War - Beyond Crohn’s Disease. Front Immunol 6:96. https://doi.org/10.3389/fimmu.2015.00096
Sechi LA, Gazouli M, Sieswerda LE, Molicotti P, Ahmed N, Ikonomopoulos J, Scanu AM, Paccagnini D, Zanetti S (2006) Relationship between Crohn’s disease, infection with Mycobacterium avium subspecies paratuberculosis and SLC11A1 gene polymorphisms in Sardinian patients. World J Gastroenterol. 12(44):7161-7164. https://doi.org/10.3748/wjg.v12.i44.7161
Shankar H, Singh SV, Singh PK, Singh AV, Sohal JS, Greenstein RJ (2010) Presence, characterization, and genotype profiles of Mycobacterium avium subspecies paratuberculosis from unpasteurized individual and pooled milk, commercial pasteurized milk, and milk products in India by culture, PCR, and PCR-REA methods. Int J Infect Dis 14(2):e121–e126. https://doi.org/10.1016/j.ijid.2009.03.031
Shivashanker JM, Nagaraja R, Yathiraj S, Rajeshwari YB, Isloor S (2018) Genetic Polymorphism of PGLYRP-1 Gene and its Association with Mastitis in Deoni and Holstein Friesian Crossbred Cows. Rumin Sci 7:199–202. Accessed on 20-03-2021. https://www.anandpub.com/archive-june-volume-2018-7-1-issue6/
Shtrichman R, Samuel CE (2001) The role of gamma interferon in antimicrobial immunity. Curr Opin Microbiol 4(3):251–259. https://doi.org/10.1016/S1369-5274(00)00199-5
Singh SV, Singh PK, Gupta S, Chaubey KK, Singh B, Kumar A, Singh AV, Kumar N (2013) Comparison of microscopy and blood-PCR for the diagnosis of clinical Johne’s disease in domestic ruminants. Iran J Vet Res 14(4):345-349. Accessed on 20-03-2021. https://ijvr.shirazu.ac.ir/article_1833_d43044a2bd892c904f91ef1040a56ace.pdf
Singh SV, Singh AV, Singh R, Sharma S, Shukla N, Misra S, Singh PK, Sohal JS, Kumar H, Patil PK, Misra P (2008) Sero-prevalence of Bovine Johne’s disease in buffaloes and cattle population of North India using indigenous ELISA kit based on native Mycobacterium avium subspecies paratuberculosis “Bison type” genotype of goat origin. Comp Immunol Microbiol Infect Dis 31(5):419–433. https://doi.org/10.1016/j.cimid.2007.06.002
Stewart LC, Day AS, Pearson J, Barclay ML, Gearry RB, Roberts RL, Bentley RW (2010) SLC11A1 polymorphisms in inflammatory bowel disease and Mycobacterium avium subspecies paratuberculosis status. World J Gastroenterol 16(45):5727–5731. https://doi.org/10.3748/wjg.v16.i45.5727
Tydell CC, Yuan J, Tran P, Selsted ME (2006) Bovine peptidoglycan recognition protein-S: Antimicrobial activity, localization, secretion, and binding properties. J Immunol 176(2):1154–1162. https://doi.org/10.4049/jimmunol.176.2.1154
Vázquez P, Ruiz-Larrañaga O, Garrido JM, Iriondo M, Manzano C, Agirre M, Estonba A, Juste RA (2014) Genetic association analysis of paratuberculosis forms in Holstein-Friesian cattle. Vet Med Int Article ID 321327. https://doi.org/10.1155/2014/321327
Vincze T, Posfai J, Roberts RJ (2003) NEBcutter: A program to cleave DNA with restriction enzymes. Nucleic Acids Res 31(13):3688–3691. https://doi.org/10.1093/nar/gkg526
Wang HL, Li ZX, Wang LJ, He H, Yang J, Chen L, Niu FB, Liu Y, Guo JZ, Liu XL (2013) Polymorphism in PGLYRP-1 gene by PCR-RFLP and its association with somatic cell score in Chinese Holstein. Res Vet Sci 95(2):508–514. https://doi.org/10.1016/j.rvsc.2013.06.005
Whittington R, Donat K, Weber MF, Kelton D, Nielsen SS, Eisenberg S, Arrigoni N, Juste R, Sáez JL, Dhand N, Santi A, … De Waard JH (2019) Control of paratuberculosis: Who, why and how. A review of 48 countries. BMC Vet Res 15(1):1–29. https://doi.org/10.1186/s12917-019-1943-4
Whittington R, Tizard M, Reddacliff L (2004) Mycobacterium paratuberculosis: pain in the gut and challenge for science. Microbiol Aust 25(4):36–37. https://doi.org/10.1071/ma04436
Wu S, Liu X, Wang Y, Zhang M, Wang M, He JQ (2019) Genetic polymorphisms of IFNG and IFNGr1 with latent tuberculosis infection. Dis Markers 2019. Article ID 8410290 https://doi.org/10.1155/2019/8410290
Yadav R, Sharma AK, Singh R, Sonwane AR, Kumar A, Chauhan A, Kumar S, Kumar T, Renjith R, Bhaladhare A, Prakash O (2014) An association study of snps with susceptibility to bovine paratuberculosis infection in cattle. Indian J Anim Sci 84(5):490–493
Zulfiqar F, Hozo I, Rangarajan S, Mariuzza RA, Dziarski R, Gupta D (2013) Genetic association of peptidoglycan recognition protein variants with inflammatory bowel disease. PLoS One 8(6):e67393. https://doi.org/10.1371/journal.pone.0067393
Acknowledgements
Authors are thankful to Director, ICAR-Indian Veterinary Research Institute, Izatnagar, India for providing necessary facilities for conducting this work.
Funding
The work was supported by the Director, ICAR-Indian Veterinary Research Institute, Izatnagar, India for providing necessary facilities and funding of this work with grant no. IXX09774 to Dr. Ran Vir Singh.
Author information
Authors and Affiliations
Contributions
RVS, SAK and AC conceived and designed the study and provided resources. BG, SAK, SUK and AC screened the animals and performed genotyping experiments. BG and SVS performed IS900 blood PCR. AK, JB and SAK analysed the data. BG and SAK wrote the original draft. SAK and JB reviewed and edited the original draft. All authors discussed the results, edited the manuscript and approved the final version of the manuscript for publication.
Corresponding authors
Ethics declarations
Conflicts of interest/Competing interests
The authors do not have any conflict of interest.
Ethics approval
All the animal experimentation had prior approval of the Institutional Animal Ethics Committee (ICAR-Indian Veterinary Research Institute, Izatnagar, India), which follows the guidelines of CPCSEA (Committee for the Purpose of Control and Supervision of Experimentation in Animals), Government of India. The approval of IAEC was given for the experiments conducted in the project vide IXX09774.
Consent to participate
Not applicable.
Consent for publication
All authors had approved the final version of the manuscript for publication.
Additional information
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Supplementary material Figure 1
Genotypic profile of SNP rs109453173 in SLC11A1 gene by PCR-RFLP with 100-bp marker (M represents marker lane). (DOCX 161 kb)
Supplementary material Figure 2
Genotypic profile of SNP rs41933863 in ANKRA2 gene by PCR-RFLP with 100-bp marker (M represents marker lane). (DOCX 68 kb)
Supplementary material Table 1
(DOCX 16 kb)
Supplementary material Table 2
(DOCX 15 kb)
Supplementary material Table 3
(DOCX 13 kb)
Rights and permissions
About this article
Cite this article
Gopi, B., Vir Singh, R., Kumar, S. et al. Effect of selected single nucleotide polymorphisms in SLC11A1, ANKRA2, IFNG and PGLYRP1 genes on host susceptibility to Mycobacterium avium subspecies paratuberculosis infection in Indian cattle. Vet Res Commun 46, 209–221 (2022). https://doi.org/10.1007/s11259-021-09849-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11259-021-09849-5