Abstract
Toll like receptors (TLRs) are pattern recognition molecules involved in cellular recognition of Mycobacterium avium subspecies paratuberculosis (MAP), the infectious agent causing Paratuberculosis (PTB), a notified disease of domestic and wild ruminants. The present study was undertaken to investigate the presence of single nucleotide polymorphisms (SNPs) in TLR2 and TLR4 gene and to evaluate association of these SNPs with occurrence of PTB in Indian cattle. A total of 213 cattle, were subjected to multiple diagnostic tests viz. Johnin PPD, ELISA test (Indigenous and Parachek kit method), fecal microscopy and fecal culture for detection of MAP infection. Based on screening results 51 animals each were assigned to case and control population. Two SNPs viz. rs55617172, rs41830058 in TLR2 gene and two SNPs viz. rs8193046, rs8193060 in TLR4 gene and were genotyped by PCR-RFLP method. All SNPs were found to be polymorphic except rs41830058 in the case-control population. Both SNPs in TLR4 gene but none in TLR2 genes were significantly associated with the occurrence of PTB in our population. The genotypes in SNP rs8193046 and SNP rs8193060 were significantly (P < 0.01) different in case-control population. These findings suggest that SNPs rs8193046 and rs8193060 are likely a potential marker against MAP infection and a selection programme eliminating AG genotype for rs8193046 and CT genotype for rs8193060 might be beneficial in conferring resistance to MAP infection in Indian cattle population.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Paratuberculosis (PTB) or Johne’s disease (JD), is a chronic disease of domestic and wild ruminants, caused by Mycobacterium avium subspecies paratuberculosis (MAP) (Chaubey et al. 2016; Sun et al. 2015). It causes granulomatous gastroenteritis, profuse diarrhea, reduced reproductive efficiency, emaciation and wasting as clinical disease progresses (Fecteau 2018; McKenna et al. 2006), leading to early death of an animal (Pieper et al. 2015). MAP is facultative intracellular pathogen residing in macrophages and shed sporadically and progressively by subclinically and clinically diseased animals respectively, thus contributing to environmental contamination (Schukken et al. 2015; Kralik et al. 2014; Pillars et al. 2009). Susceptible animals, typically young calves, become infected by ingesting contaminated colostrum, milk, or feces (Sweeney et al. 2012) and then enter a lengthy subclinical period lasting up to 10 years (Sweeney 2011; Magombedze et al. 2013).
PTB is prevalent worldwide causing significant loss to dairy industry due to reduced milk production, decreased fertility, early culling and higher management costs (Smith et al. 2017; Sergeant et al. 2008). This dreaded disease is responsible for annual economic losses of approximately US$ 250 million to the US dairy industry (Wadhwa et al. 2012). MAP infection has also been associated with multiple diseases in humans like: Crohn’s disease (Agrawal et al. 2014), Blau syndrome (Dow and Ellingson 2010), type 1 diabetes (Cossu et al. 2011a), Hashimoto thyroiditis (Masala et al. 2014) and multiple sclerosis (Cossu et al. 2011b). Crohn’s disease is a chronic, painful, diarrhoeal inflammatory disease of the intestinal tract in humans that resembles PTB in ruminants. Thus, MAP also poses a serious threat to public health apart from compromising animal health and welfare.
The sensitivity of available detection methods of MAP is low (Nielsen and Toft 2008) and there is limited knowledge on the host factors controlling immune responses to MAP. Also the lack of effective vaccines and differentiation of infected and vaccinated animals (DIVA) based diagnostics (Bastida and Juste 2011) leads to an underestimation of prevalence and to an “Iceberg effect” whereby the true prevalence in infected herds is greater than the apparent prevalence (Magombedze et al. 2013). As a result, most infected animals, especially thosein the early infection stage and sub-clinically infected animals remain undetected. These factors are supplemented by lack of knowledge concerning genetic factors that control susceptibility or resistance to infection. In this scenario, appropriate control measures for MAP infection to prevent the incidence and prevalence of PTB in cattle is required. Genetic selection for resistant animals is one of the proven strategies to control livestock diseases (Bishop and Woolliams 2014). In order to identify resistant or susceptible animals, the best approach can be defining simple genetic markers that are associated with or linked to the complex genes controlling disease resistance. Additive genetic variance exists in cattle for MAP and heritability estimates for MAP infection ranges from 0.09 to 0.18 (Gonda et al. 2006). Incidence of PTB has genetic variability associated with it, and thus it is possible to identify genomic regions associated with resistance or susceptibility to MAP. The importance of this area has been taken seriously and several workers have identified polymorphism in different candidate genes viz. SLC11A1, NOD2, SP110, TLR2, SP110, CLEC7A, CD209 and TLR4 genes and quantitative trait loci (QTL) associated with susceptibility or resistance to PTB (Kumar et al. 2019; Kumar et al. 2017; Sadana et al. 2015; Pant et al. 2014; Vazquez et al. 2014; Yadav et al. 2014; Ruiz-Larranaga et al. 2010) in different resource population, but there is dearth of study in Indian cattle population.
Toll like receptors (TLRs) are germ line encoded pattern recognition receptors that bind pathogen associated molecular patterns (PAMPs) and are intrinsically involved in triggering both the innate and adaptive immune response mechanisms (Bharati et al. 2017). It evolved as the immediate host defense in response to foreign antigens (Cristofaro and Opal 2006) and is found in all animal species. Although most mammalian species share up to 13 TLR genes, only 10 mammalian TLR (TLR1–TLR10) are expressed in the bovine species (McGuire et al. 2006; Menzies and Ingham 2006), which identify molecular patterns from all major classes of pathogens (Oshiumi et al. 2003; Tabeta et al. 2004). TLR2 and TLR4 are involved in the early recognition of MAP (Quesniaux et al. 2004; Buwitt-Beckmann et al. 2006). They are expressed on microglia, schwann cells, monocytes, macrophages, dendritic cells (DCs), polymorphonuclear leukocytes (PMNs or PMLs), B cells and T cells in cattle (Werling et al. 2004, 2006). TLR1-TLR2 heteromers and TLR4 recognize mycobacterial PAMPs by activated macrophages and dendritic cells (Chang et al. 2006; Roura-Mir et al. 2005; Uehori et al. 2005). The critical function of these genes in the immunity against mycobacterium suggests a potential involvement of these genes in the variation in bovine PTB susceptibility. Hence, detection of polymorphisms in TLR2 and TLR4 genes would play a key role in the resistance to MAP infection. The central hypothesis of this study is that, the different arrangements of particular alleles in the TLR2 and TLR4 genes would predominate in a case population as compared to control, suggesting a role in susceptibility to infection. The aim of the present study was to find SNPs in TLR2 and TLR4 genes and to associate these SNPs with high and low susceptibility to bovine PTB disease in Indian cattle population.
Materials and methods
Animals and sampling
A total of 213 cattle from 3 different farms located in western Uttar Pradesh area (65.73% IVRI farm in Bareilly, 19.25% Agarwal Milkey and PG Farm in Bareilly and 15.02% Raghura Dairy, Kashipur) were screened to develop a case-control resource population. Out of 213 animals, 41 each of Sahiwal and Tharparkar, 32 were Holstein Friesian and rest 99 animals were crossbred. Animals were housed under similar environmental conditions and were unvaccinated for PTB. For the purpose of this study, only adult cattle were chosen because they had higher chance of being exposed to MAP than those for younger animals (Ruiz-Larrañaga et al. 2017). All the samples necessary for the screening of PTB viz. blood and fecal samples were collected from each animal aseptically.
Diagnostic tests for screening MAP infection
The resource population was screened using a panel of diagnostic tests viz. Johnin PPD test, fecal microscopy, fecal culture and ELISA (Indigenous and PARACHEK®) test against MAP infection.
Johnin PPD test
The single intra-dermal Johnin test for delayed type hypersensitivity reaction was carried out by the intra-dermal inoculation of 0.1 ml of Johnin PPD on the side of the middle third of the neck. The thickness of skin was measured with vernier calipers before and 72 h after inoculation. Increase in skin thickness of over 4 mm (Fig. 1) was considered as indicative of presence of delayed-type hypersensitivity (Roupie et al. 2018). Along with Johnin PPD, tuberculin intradermal test and Mycobacterium avium PPD test was also done to rule out cross reaction with Mycobacterium bovis and Mycobacterium avium infection respectively (Kumar et al. 2019).
Microscopic examination of fecal smears
About 4–5 g of feacal samples from 213 animals were collected per-rectally in a sterile container from each animal using sterile glove. All these samples were transported to laboratory in ice and stored at −20 °C till further use. Ziehl–Neelsen stained smears of faeces were examined microscopically for presence of acid fast bacilli of MAP (Singh et al. 2013). Slides displaying pink coloured short rods (0.5–1.5 μm), distinguishable to MAP (Fig. 2) were considered as positive for MAP infection.
Cultural examination of fecal samples
Fecal culture examination was also performed in the experimental population which is considered as the gold standard ante-mortem test, and it demonstrates viable bacteria. (Kalis et al. 2002; Whitlock et al. 2000). Herrold’s Egg Yolk medium (HEY medium) was used for culture of MAP and was prepared as per the method of Merkal et al. (1982) with some modifications. Inoculated slants were observed for growth of acid fast MAP after 20 weeks of culture (Fig. 3). They were only confirmed to be acid fast using the microscopic examination of Ziehl-Neelsen stained smears, and that this is a limitation to the study. (Fig. 4).
Detection of MAP-specific antibodies by ELISA
ELISA for the detection of MAP infection was done by commercial ELISA kit (PARACHEK®, Prionics) and indigenous Elisa kit (Singh et al. 2008). Putative cross-reacting antibodies in uninfected animals to other mycobacteria were removed by absorption of serum with M. phlei. Positive and negative sera were included as controls. The absorbance of each sample was recorded at 450 nm by ELISA reader and by using these optical density (OD) values, S/P ratios were calculated as per the formula.
The result was interpreted to provide the infection status of the animal. Samples having S/P ratio more than 0.4 was classified as positive, below 0.1 as negative and value between 0.1 and 0.4 as suspected for MAP infection (Collins 2002).
Based on the screening, 51 animals which were negative in all the tests were designated as control population while 51 animals which were positive in at least two tests were designated as susceptible or case population.
Phenotypic classification of animals
All the 213 animals included in the present investigation were given a score by considering the health and physical condition of animals visualized in four scales called physical body condition score (PBCS). The coding for PBCS was done according to Edmonson et al. (1989) as following:
-
4+: healthy animals with no apparent signs of disease.
-
3+: healthy animals with no apparent signs of disease but somewhat weaker than 4 + .
-
2+: weak and emaciated animals.
-
1+: animals with obvious clinical signs of disease like diarrhea and emaciation.
Blood collection for serum and DNA isolation
About 3 ml of venous blood was collected from the jugular vein of the animal into a sterile 15 ml polypropylene centrifuge tube containing EDTA (1.8 mg/ml of blood) under sterile condition for DNA isolation and 3 ml of venous blood from 213 animals was collected in a BD vacutainer under sterile condition from jugular vein of animal without anticoagulant for separation of serum. The samples were kept at −20 °C till the isolation of DNA. Genomic DNA for all experimental cattle was isolated by Phenol: Chloroform extraction method as described by Sambrook and Russel (2001) with minor modifications. The concentration and purity of DNA was determined by Nanodrop 1000 spectrophotometer. The DNA samples showed the OD260:OD280 ratio between 1.7 and 1.9, which were considered as good quality and used for further analysis.
SNP selection and genotyping
Two single nucleotide polymorphisms (SNPs) in TLR2 gene viz. rs55617172, rs41830058 and two SNPs in TLR4 gene viz. rs8193046, rs8193060 were selected from the National Center for Biotechnology Information SNP database (NCBI dbSNP) and suitable primers were designed using online software primer3 and NCBI primer designing software. All the SNPs were found in the exonic region except the SNP rs8193046 which was in the intronic region (Table 1). Primers were designed to amplify a 245–495 bp fragment which includes the targeted SNP. Identification of a restriction enzyme (RE) allowing allele discrimination by in-silico analysis was done by the program designated NEBcutter V2, which has an option for viewing fragments of an in-silico digest (Vincze et al. 2003).
Genotyping of SNPs in the product was done by PCR-RFLP technique. PCR amplification was carried out using thermo cycler (Bio-Rad, USA) in a reaction mixture of 25 μl, comprised of 1.0 μl of forward primer, 1 μl of reverse primer, 2.5 μl of Taq, 0.5 μl of dNTPs mix, 2.0 μl of MgCl2, 0.2 μl of Taq DNA polymerase, 2 μl of DNA template and 15.8 μl of Nuclease Free Water. The optimization of appropriate annealing temperature with respect to each primer was determined by gradient PCR. The PCR conditions involved initial denaturation at 95 °C for 5 min, followed by 35 cycles with denaturation at 95 °C for 30 s, annealing temperature ranged from 56.2 °C to 64.5 °C for 45 s, extension at 72 °C for 45 s followed by a final extension at 72 °C for 10 min.
The restriction enzyme (RE) digestion of PCR product was carried out in 0.2 ml tubes with a total reaction mixture of 23 μl by overnight incubation at a temperature, specified by enzyme manufacturer. The digested products were then kept at −20 °C until further study. The amplified and RE digested PCR products were electrophoresed in 3.5% agarose gel for 2 h at 90 V. About 20 μl of digested product mixed with 4 μl of 6× loading dye were loaded into each well along with 100 bp DNA ladder in a separate lane. After completion of gel electrophoresis the digested product were visualized by UV transilluminator and documented to detect the genotype of each sample. Region covered by each primer, primer sequence, annealing temperature and amplicon size for each primer and information of RE, fragments obtained after digestion and genotyping by RE digestion is shown in Table 2.
Statistical analysis
The effect of nongenetic factors and SNPs in TLR2 and TLR4 genes on the occurrence of PTB disease was analysed by PRO CLOGISTIC procedure of SAS 9.3 software. The univariable analysis was performed for categorical variables with a a Chi-square (χ2) test on the distribution of allele and genotype frequencies between case population (PTB affected) and control population. The differences between case and control were assessed by odds ratio (OR) with 95% confidence intervals 9CIs). The relative risk of incidence among the genotypes was analyzed using a logistic regression model:
Where: log [p/(1 − p)] = Probability of the binomial response phenotype; α, β1, β2 and γ = Regression coefficients associated with population, heterozygotes and error respectively. This model compared heterozygous (Ht) and homozygous (Ho) genotypes and estimated one/two OR, one for Ht and the other for Ho.
The binomial response phenotype (dependent variable) was coded based on the positive (coded as ‘1’) or negative (coded as ‘0’) test for MAP infection. The non-genetic factors like age (2 level), physical body condition score (2 level) and breed (2 level) were fitted. The PROC ALLELE procedure of SAS 9.3 was used for testing of Hardy-Weinberg equilibrium (HWE), estimation of polymorphism information content (PIC), linkage disequilibrium (LD) and heterozygosity of SNPs markers were used in the present investigation.
Results and discussion
PTB is a disease of global economic and potential zoonotic concern (Pinedo et al. 2013). Host genetic variation has been known to influence resistance or susceptibility to infection following exposure to MAP (Casanova and Abel 2002). Bovine macrophages are the primary target cells in which MAP is able to survive and replicate, hence it is possible that, TLR2 and TLR4 function is critically important in PTB infection (Weiss et al. 2008). It has been found that mutations in TLR2 and TLR4 gene causes inefficient recognition of MAP and failure in the subsequent innate and adaptive immune responses which increase risk of mycobacterial infections (Henckaerts et al. 2007; Thuong et al. 2007) and Crohn’s disease (Hong et al. 2007; Lakatos et al. 2005) in humans. Currently, a few studies are involved in exploring association of host-candidate genes with susceptibility to MAP infection in Indian cattle population. The ultimate aim was to find a single SNP or a panel of SNPs that could act as a genetic marker and could be utilized in selection of PTB resistant cattle at early stage.
Out of 213 animals screened for MAP infection, we could obtain only 51 animals in case population, since the incidence rate of PTB in cattle is 39.3% in western Uttar Pradesh (Singh et al. 2014) from where our resource population belongs. A panel of diagnostic tests were used for screening MAP infection status and only those animals which showed positive results for two or more than two diagnostic tests were included in case population to rule out any possibility of false positives. However, only those animals were included in control which showed negative results for all the five diagnostic tests to rule out false negative results in the control population. Hence, our resource population got limited to 102 animals (51 cases and 51 controls). Unfortunately, this is one significant limitation to this study. However, it is well accepted that preliminary association studies can be carried out in small population, provided there is true case and control population. Association study of different SNPs with bovine brucellosis with resource population of 83 cattle (38 cases and 45 controls) has been conducted by Prakash et al. (2014). We also studied the effect of non genetic factors like PBCS, age and breed on occurrence of PTB in our resource population. PBCS was significantly (p < 0.01) different among case animals from control animals. The ODDs for animals with poor physical conditions at 95% CI were significantly different from healthy animals which depicts that animals with poor physical conditions were at a higher risk of infection than animals with good physical conditions. Further, we found that although we had diversified breeds and admixed individuals but the effect of breed was not significant (p > 0.05) on the occurrence of PTB disease. The indigenous breeds were compared with exotic and cross bred animals and p value for occurrence of disease between different breed was non-significant. Our results are in agreement with the findings of Sadana et al. (2015) who also reported non-significant effect of breed on the -incidence of bovine PTB. Age also failed to distinguish significantly (p > 0.05) the affected animals from healthy animals in our resource population.
The SNP rs41830058 in TLR2 gene was found to be monomorphic having only GG genotype in case-control population and hence was excluded from association analysis. While the SNP rs55617172 in TLR2 gene was polymorphic having three genotypes viz. AA, AC and CC. These genotypes failed to show significant association between case and control population. The logistic regression analysis revealed that for SNP rs55617172, the animals with AA (OR 0.41) genotypes were at lower risk for occurrence of disease than animals with AC (OR 1.09) and CC (OR 1) genotype which is also reflected by odds of allele A (OR 0.71) versus C (OR 1). Although not significantly different, the allele A confers relative resistance toward PTB at this locus of the gene. The odds of CC and AC are almost equal, indicating the possible dominant effect of C allele with no heterozygote advantage. Sadana et al. (2015) found similar result with SNP rs55617172, as the probability for getting infection in animals with A allele was half as compared to C allele and the odds of susceptibility for AA genotype were only 0.17 as compared to CC genotype (OR 1). The allele and genotype frequency distribution and OR of case and control animals for TLR2 genes are given in Tables 3 and 4 respectively. The PIC, average heterozygosity and HWE of each locus are shown in Table 5.
Bovine TLR2 gene located at the proximal end of BTA17 contains 2 exons and encodes 784 amino acids (White et al. 2003). There are reports that correspond to the association of TLR2 with PTB in cattle (Sadana et al. 2015; Koets et al. 2010; Mucha et al. 2009), however in the present study, no associations were identified between SNPs in TLR2 gene with MAP infections in our resource population. A study by Koets et al. (2010) revealed the significant association of TLR2–1903 T/C SNP with occurrence of PTB in cattle, where CC and CT genotypes were at 1.7 times greater risk than TT genotype for getting MAP infection. The synonymous SNP c.1707 T > C (H569H) in TLR2 gene has been found to be associated with MAP infection in a Dutch Holstein-Friesian population. These conflicting results may be an outcome of the small number of individuals used in this study, due to breed differences or differences in the linkage phase between the SNPs and the actual causative SNPs and due to the use of different panel of diagnostic tests for establishment of case-control population. Previous workers (Sadana et al. 2015; Koets et al. 2010) had used single or a couple of diagnostic test while in our study we used a panel of diagnostic tests, so they might have a biased case-control population, which explains for the difference in results. Most JD control programmes use ELISA to detect potentially infected animals (Lavers et al. 2014) because individual fecal culture is expensive and time-consuming. Another reason for using ELISAs is that the extent of shedding correlates well with the ELISA titres (Dargatz et al. 2001). However, it was reported by Pant et al. (2014) that ELISA test for MAP specific antibodies lacks the desired sensitivity and thus can introduce false negatives in resource population; hence we performed a panel of diagnostic tests viz. Johnin PPD test, ELISA test, fecal microscopy and fecal culture test against MAP so that our screening for resource population are unbiased. We have ensured that for declaring true negative, animals must have negative results for all five tests and thus there is a rare possibility of inclusion of false negatives in control population and false positives in case population.
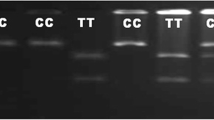
Both SNPs in TLR4 gene rs8193046 and rs8193060 had showed polymorphic pattern in our case-control population and were significantly associated with occurrence of PTB in cattle. The logistic regression analysis revealed that for both SNPs genotype frequency differ significantly (p < 0.01) between case and control population. In SNP rs8193046, three genotypes AA, AG and GG were observed at this locus with frequencies in case and control population were 27.45, 68.63and 3.92 and 50.98, 29.41and 19.61 respectively. The odds of AA and AG genotypes verses GG genotype were 2.69 and 11.67 respectively i.e. probability for getting infection in animals with AA and AG genotype was 2.69 and 11.67 times more as compared to GG genotype. Here we encountered an interesting finding that, animals with AG genotype i.e. heterozygote were more abundant in case than control. Despite the odds of A allele being lower than G allele, the odds of AG were 11.67 times than the odds of GG genotype. It is a possible example of heterozygous disadvantage as the odds of AC were greater than both of the homozygotes. A case of overdominance toward susceptibility side can also be possible reason.
In SNP rs8193060 three genotypes CC, CT and TT were observed at this locus with odds of CC and CT compared with TT were 0.89 and 3.9 respectively i.e. probability for getting infection in animals with CT genotype was 3.9 times more as compared to TT genotype and 4.3 times more as compared to CC. The animals with CT genotype i.e. heterozygote were more abundant in case than control. Despite the odds of C allele being lower than T allele, the ODDs of CT were 3.9 times than the odds of TT genotype. It is also a possible example of heterozygous disadvantage as the odds of CT were greater than both of the homozygotes and a case of overdominance toward susceptibility side.
The PIC ranged from 35.55% for rs8193046 to 37.42% for rs8193060 in TLR4 gene revealing a moderate polymorphism at these SNPs. Heterozygosity have moderate estimate with 32.35% for rs8193060 to 49.02% for rs8193046 in TLR4 gene. The allele and genotype frequency distribution and OR of case and control animals for TLR4 genes are given in Table 3 and Table 4 respectively. The PIC, average heterozygosity and HWE of each locus are shown in Table 5. The genotypic profile of SNP rs8193046 and rs8193060 is depicted in Figs. 5 and 6 respectively.
Bovine TLR4 gene located at the proximal end of BTA8 contains 3 exons and encodes 841 amino acids. In a multibreed Bos taurus population, SNPs in the TLR4 gene were found to be associated with MAP infection (Mucha et al. 2009), but Pinedo et al. (2009) were unable to confirm the significant association between SNPs in TLR4 gene and PTB infection in multibreed sample composed of Bos taurus and crossbred of Bos taurus and Bos indicus breeds. The significant association between TLR4 gene and MAP infection as found in our study was also observed by Mucha et al. (2009) and Ruiz-Larrañaga et al. (2011) in Holstein-Friesian cattle. However, the SNPs they reported significant were different from our SNPs. In humans, genetic polymorphisms in the TLR4 gene have been associated with differences in LPS responsiveness (Poltorak et al. 1998; Schwartz 2002), sensitivity to mycobacterial antigens (Bulut et al. 2005) and are associated with increased risk of Crohn’s disease (Hong et al. 2007; Lakatos et al. 2005). Our results are in agreement with earlier reports by Sharma et al. (2015), where a strong association was found between different allelic variants in TLR4 gene and occurrence of PTB in Canadian Holsteins. They revealed a strong association between c.-226G > C, and c.2021C > T SNPs and PTB disease. Taken together, this study provides evidence of likelihood of SNPs rs8193046 and rs8193060 loci in TLR4 gene in determining host susceptibility to MAP infection.
Conclusion
This is the first report on the association study of TLR4 gene with the occurrence of PTB disease in India. SNPs rs8193046 and rs8193060 loci would be valuable for genotyping the animals for selection against PTB infection in cattle. Also, further analyses through extended sampling in larger population and comprehensive studies can offer more reliable explanations of association of examined SNPs related to resistance to PTB.
References
Agrawal G, Borody TJ, Chamberlin W (2014) Global warming’to Mycobacterium avium subspecies paratuberculosis. Future Microbiol 9(7):829–832
Bastida F, Juste RA (2011) Paratuberculosis control: a review with a focus on vaccination. J Immune Based Ther Vaccines 9(1):8
Bharati J, Dangi SS, Mishra SR, Chouhan VS, Verma V, Shankar O, Bharti MK, Paul A, Mahato DK, Rajesh G, Singh G (2017) Expression analysis of toll like receptors and interleukins in Tharparkar cattle during acclimation to heat stress exposure. J Therm Biol 65:48–56
Bishop SC, Woolliams JA (2014) Genomics and disease resistance studies in livestock. Livest Sci 166:190–198
Bulut Y, Michelsen KS, Hayrapetian L, Naiki Y, Spallek R, Singh M, Arditi M (2005) Mycobacterium tuberculosis heat shock proteins use diverse toll-like receptor pathways to activate pro-inflammatory signals. J Biol Chem 280(22):20961–20967
Buwitt-Beckmann U, Heine H, Wiesmuller KH, Jung G, Brock R, Akira S, Ulmer AJ (2006) TLR1- and TLR6-independent recognition of bacterial lipopeptides. J Biol Chem 281:9049–9057
Casanova JL, Abel L (2002) Genetic dissection of immunity to mycobacteria: the human model. Annu Rev Immunol 20(1):581–620
Chang JS, Huggett JF, Dheda K, Kim LU, Zumla A, Rook GA (2006) Myobacterium tuberculosis induces selective up-regulation of TLRs in the mononuclear leukocytes of patients with active pulmonary tuberculosis. J Immunol 176:3010–3018
Chaubey KK, Gupta RD, Gupta S, Singh SV, Bhatia AK, Jayaraman S, Kumar N, Goel A, Rathore AS, Sahzad, Sohal JS (2016) Trends and advances in the diagnosis and control of paratuberculosis in domestic livestock. Vet Q 36(4):203–227
Collins MT (2002) Interpretation of a commercial bovine paratuberculosis enzyme-linked immunosorbent assay by using likelihood ratios. Clin Diagn Lab Immunol 9(6):1367–1371
Cossu A, Rosu V, Paccagnini D, Cossu D, Pacifico A, Sechi LA (2011a) MAP3738c and MptD are specific tags of Mycobacterium avium subsp. paratuberculosis infection in type I diabetes mellitus. Clin Immunol 141(1):49–57
Cossu D, Cocco E, Paccagnini D, Masala S, Ahmed N, Frau J, Marrosu MG, Sechi LA (2011b) Association of Mycobacterium avium subsp. paratuberculosis with multiple sclerosis in Sardinian patients. PLoS One 6(4):e18482
Cristofaro P, Opal SM (2006) Role of toll-like receptors in infection and immunity. Drugs 66(1):15–29
Dargatz DA, Byrum BA, Barber LK, Sweeney RW, Whitlock RH, Shulaw WP, Jacobson RH, Stabel JR (2001) Evaluation of a commercial ELISA for diagnosis of paratuberculosis in cattle. J Am Vet Med Assoc 218(7):1163–1166
Dow CT, Ellingson JL (2010) Detection of Mycobacterium avium ss. Paratuberculosis in Blau syndrome tissues. Autoimmune Dis 2010:127692
Edmonson AJ, Lean IJ, Weaver LD, Farver T, Webster G (1989) A body condition scoring chart for Holstein dairy cows. J Dairy Sci 72:68–78
Fecteau ME (2018) Paratuberculosis in cattle. Vet Clin North Am Food Anim Pract 34:209–222
Gonda MG, Chang YM, Shook GE, Collins MT, Kirkpatrick BW (2006) Genetic variation of Mycobacterium avium ssp. paratuberculosis infection in US Holsteins. J Dairy Sci 89:1804–1812
Henckaerts L, Pierik M, Joossens M, Ferrante M, Rutgeerts P, Vermeire S (2007) Mutations in pattern recognition receptor genes modulate seroreactivity to microbial antigens in patients with inflammatory bowel disease. Gut 56:1536–1542
Hong J, Leung E, Fraser AG, Merriman TR, Vishnu P, Krissansen GW (2007) TLR2, TLR4 and TLR9 polymorphisms and Crohn’s disease in a New Zealand Caucasian cohort. J Gastroenterol Hepatol 22:1760–1766
Kalis CHJ, Barkema HW, Hesselink JW, Van Maanen C, Collins MT (2002) Evaluation of two absorbed ELISAs and a complement fixation test as replacements for fecal culture in the detection of cows shedding Mycobacterium avium subspecies paratuberculosis. Prev Vet Med 14:219–224
Koets A, Santema W, Mertens H, Oostenrijk D, Keestra M, Overdijk M, Labouriau R, Franken P, Frijters A, Nielen M, Rutten VPMG (2010) Susceptibility to paratuberculosis infection in cattle is associated with single nucleotide polymorphisms in toll-like receptor 2 which modulate immune responses against Mycobacterium avium subspecies paratuberculosis. Prev Vet Med 93(4):305–315
Kralik P, Pribylova-Dziedzinska R, Kralova A, Kovarcik K, Slana I (2014) Evidence of passive faecal shedding of Mycobacterium avium subsp. paratuberculosis in a Limousin cattle herd. Vet J 201:91–94
Kumar S, Kumar S, Singh R, Chauhan A, Agrawal S, Kumar A, Singh S (2017) Investigation of genetic Association of Single Nucleotide Polymorphisms in SP110 gene with occurrence of Paratuberculosis disease in cattle. Int J Livest Res 7(3):81–88
Kumar S, Singh RV, Chauhan A, Kumar A, Bharati J, Singh SV (2019) Association of Bovine CLEC7A gene polymorphism with host susceptibility to paratuberculosis disease in Indian cattle. Res Vet Sci 123:216–222
Lakatos PL, Lakatos L, Szalay F, Willheim-Polli C, Österreicher C, Tulassay Z, Molnar T, Reinisch W, Papp J, Mozsik G, Ferenci P (2005) Toll-like receptor 4 and NOD2/CARD15 mutations in Hungarian patients with Crohn’s disease: phenotype-genotype correlations. World J Gastroenterol 11(10):1489–1495
Lavers CJ, Barkema HW, Dohoo IR, McKenna SLB, Keefe GP (2014) Evaluation of milk ELISA for detection of Mycobacterium avium subspecies paratuberculosis in dairy herds and association with within-herd prevalence. J Dairy Sci 97(1):299–309
Magombedze G, Ngonghala CN, Lanzas C (2013) Evalution of the “iceberg phenomenon” in Johne’s disease through mathematical modelling. PLoS One 8:e76636N
Masala S, Cossu D, Palermo M, Sechi LA (2014) Recognition of zinc transporter 8 and MAP3865c homologous epitopes by Hashimoto's thyroiditis subjects from Sardinia: a common target with type 1 diabetes? PLoS One 9(5):e97621
McGuire K, Jones M, Werling D, Williams JL, Glass EJ, Jann O (2006) Radiation hybrid mapping of all 10 characterized bovine toll-like receptors. Anim Genet 37(1):47–50
McKenna SLB, Keefe GP, Tiwari A, VanLeeuwen J, Barkema HW (2006) Johne’s disease in Canada part II: disease impacts, risk factors, and control programs for dairy producers. Can Vet J 47:1089–1099
Menzies M, Ingham A (2006) Identification and expression of toll-like receptors 1–10 in selected bovine and ovine tissues. Vet Immunol Immunopathol 109(1–2):23–30
Merkal RS, Miller JM, Hintz AM, Bryner JH (1982) Intrauterine inoculation of Mycobacterium paratuberculosis into Guinea pigs and cattle. Am J Vet Res 43:676–678
Mucha R, Bhide MR, Chakurkar EB, Novak M, Mikula I (2009) Toll-like receptors TLR1, TLR2 and TLR4 gene mutations and natural resistance to Mycobacterium avium subsp. paratuberculosis infection in cattle. Vet Immunol Immunopathol 128(4):381–388
Nielsen SS, Toft N (2008) Ante mortem diagnosis of paratuberculosis: a review of accuracies of ELISA, interferon-g assay and faecal culture techniques. Vet Microbiol 129:217–235
Oshiumi H, Matsumoto M, Funami K, Akazawa T, Seya T (2003) TICAM-1, an adaptor molecule that participates in toll-like receptor 3-mediated interferon-beta induction. Nat Immunol 4:161–167
Pant SD, Verschoor CP, Schenkel FS, You Q, Kelton DF, Karrow NA (2014) Bovine CLEC7A genetic variants and their association with seropositivity in Johne's disease ELISA. Gene 537(2):302–307
Pieper L, Sorge US, DeVries TJ, Godkin A, Lissemore K, Kelton DF (2015) Evaluation of the Johne’s disease risk assessment and management plan on dairy farms in Ontario, Canada. J Dairy Sci 98(10):6792–6800
Pillars RB, Grooms DL, Kaneene JB (2009) Longitudinal study of the distribution of Mycobacterium avium subsp. paratuberculosis in the environment of dairy herds in the Michigan Johne’s disease control demonstration herd project. Can Vet J 50:1039–1046
Pinedo PJ, Buergelt CD, Donovan GA, Melendez P, Morel L, Wu R, Langaee TY, Rae DO (2009) Candidate gene polymorphhisms (BoIFNG, TLR4, SLC11A1) as risk factors for paratuberculosis infection in cattle. Prev Vet Med 91(2):189–196
Pinedo PJ, Galvão KN, Seabury CM (2013) Innate immune gene variation and differential susceptibility to uterine diseases in Holstein cows. Theriogenology 80:384–390
Poltorak A, He X, Smirnova I, Liu MY, Van Huffel C, Du X, Birdwell D, Alejos E, Silva M, Galanos C, Freudenberg M, Ricciardi-Castagnoli P, Layton B, Beutler B (1998) Defective LPS signaling in C3H/ HeJ and C57BL/10ScCr mice: mutations in TLR4 gene. Science 282:2085–2088
Prakash O, Kumar A, Sonwane A, Rathore R, Singh RV, Chauhan A, Kumar P, Renjith R, Yadav R, Bhaladhare A, Baqir M (2014) Polymorphism of cytokine and innate immunity genes associated with bovine brucellosis in cattle. Mol Biol Rep 41(5):2815–2825
Quesniaux V, Fremond C, Jacobs M, Parida S, Nicolle D, Yeremeev V, Bihl F, Erard F, Botha T, Drennan M, Soler M, Le Bert M, Schnyder B, Ryffel B (2004) Toll-like receptor pathways in the immune responses to mycobacteria. Microbes Infect 6:946–959
Roupie V, Alonso-Velasco E, Van Der Heyden S, Holbert S, Duytschaever L, Berthon P, Van Dosselaer I, Van Campe W, Mostin L, Biet F, Roels S (2018) Evaluation of mycobacteria-specific gamma interferon and antibody responses before and after a single intradermal skin test in cattle naturally exposed to M. Avium subsp. paratuberculosis and experimentally infected with M. Bovis. Vet Immunol Immunopathol 196:35–47
Roura-Mir C, Wang L, Cheng TY, Matsunaga I, Dascher CC, Peng SL, Fenton MJ, Kirschning C, Moody DB (2005) Mycobacterium tuberculosis regulates CD1 antigen presentation pathways through TLR-2. J Immunol 175:1758–1766
Ruiz-Larranaga O, Garrido JM, Manzano C, Iriondo M, Molina E, Gil A, Koets AP, Rutten VPMG, Juste RA, Estonba A (2010) Identification of single nucleotide polymorphisms in the bovine solute carrier family 11 member 1 (SLC11A1) gene and their association with infection by Mycobacterium avium subspecies paratuberculosis. J Dairy Sci 93(4):1713–1721
Ruiz-Larrañaga O, Manzano C, Iriondo M, Garrido JM, Molina E, Vazquez P, Juste RA, Estonba A (2011) Genetic variation of toll-like receptor genes and infection by Mycobacterium avium ssp. paratuberculosis in Holstein-Friesian cattle. J Dairy Sci 94(7):3635–3641
Ruiz-Larrañaga O, Vázquez P, Iriondo M, Manzano C, Aguirre M, Garrido JM, Juste RA, Estonba A (2017) Evidence for gene-gene epistatic interactions between susceptibility genes for Mycobacterium avium subsp. paratuberculosis infection in cattle. Livest Sci 195:63–66
Sadana T, Singh RV, Singh SV, Saxena VK, Sharma D, Singh PK, Kumar N, Gupta S, Chaubey KK, Jayaraman S, Tiwari R (2015) Single nucleotide polymorphism of SLC11A1, CARD15, IFNG and TLR2 genes and their association with Mycobacterium avium subspecies paratuberculosis infection in native Indian cattle population. Indian J Biotechnol 14:469–475
Sambrook J, Russel DW (2001) Molecular cloning- a laboratory manual, 3rd edn. Cold Spring Harbor laboratory Press, Cold Spring Harbor
Schukken YH, Whitlock RH, Wolfgang D, Grphn Y, Beaver A, VanKessel J, Zurakowski M, Mitchell R (2015) Longitudinal data collection of Mycobacterium avium subspecies paratuberculosis infections in dairy herds: the value of precise feld data. Vet Res 46:65
Schwartz DA (2002) The genetics of innate immunity. Chest 121(3):62S–68S
Sergeant ESG, Nielsen SS, Toft N (2008) Evaluation of test-strategies for estimating probability of low prevalence of paratuberculosis in Danish dairy herds. Prev Vet Med 85(1-2):92–106
Sharma BS, Abo-Ismail MK, Schenkel FS, You Q, Verschoor CP, Pant SD, Karrow NA (2015) Association of TLR4 polymorphisms with Mycobacterium avium subspecies paratuberculosis infection status in Canadian Holsteins. Anim Genet 46(5):560–565
Singh SV, Singh AV, Singh R, Sharma S, Shukla N, Misra S, Singh PK, Sohal JS, Kumar H, Patil PK, Misra P (2008) Sero-prevalence of bovine Johne's disease in buffaloes and cattle population of North India using indigenous ELISA kit based on native Mycobacterium avium subspecies paratuberculosis ‘Bison type’genotype of goat origin. Comp Immunol Microbiol Infect Dis 31(5):419–433
Singh SV, Dhama K, Chaubey KK, Kumar N, Singh PK, Sohal JS, Mahima CS, Deb R (2013) Impact of host genetics on susceptibility and resistance to Mycobacterium avium subspecies paratuberculosis in domestic ruminants. Pak J Biol Sci 16:251–266
Singh SV, Singh PK, Singh AV, Sohal JS, Kumar N, Chaubey KK, Gupta S, Rawat KD, Kumar A, Bhatia AK, Srivastav AK (2014) ‘Bio-Load’and bio-type profiles of Mycobacterium avium subspecies paratuberculosis infection in the domestic livestock population endemic for Johne's disease: a survey of 28 years (1985–2013) in India. Transbound Emerg Dis 61(s1):43–55
Smith RL, Al-Mamun MA, Gröhn YT (2017) Economic consequences of paratuberculosis control in dairy cattle: a stochastic modeling study. Prev Vet Med 138:17–27
Sun WW, Lv WF, Cong W, Meng QF, Wang CF, Shan XF, Qian AD (2015) Mycobacterium avium subspecies paratuberculosis and bovine leukemia virus seroprevalence and associated risk factors in commercial dairy and beef cattle in northern and northeastern China. Biomed Res Int 2015:1–7. https://doi.org/10.1155/2015/315173
Sweeney RW (2011) Pathogenesis of paratuberculosis. Vet Clin North Am Food Anim Pract 27:537–546
Sweeney RW, Collins MT, Koets AP, McGuirk SM, Roussel AJ (2012) Paratuberculosis (Johne’s disease) in cattle and other susceptible species. J Vet Intern Med 26:1239–1250
Tabeta K, Georgel P, Janssen E, Du X, Hoebe K, Crozat K, Mudd S, Shamel L, Sovath S, Goode J, Alexopoulou L, Flavell RA, Beutler B (2004) Toll-like receptors 9 and 3 as essential components of innate immune defense against mouse cytomegalovirus infection. Proc Natl Acad Sci U S A 101:3516–3521
Thuong NTT, Hawn TR, Thwaites GE, Chau TTH, Lan NTN, Quy HT, Hieu NT, Aderem A, Hien TT, Farrar JJ, Dunstan SJ (2007) A polymorphism in human TLR2 is associated with increased susceptibility to tuberculous meningitis. Genes Immun 8(5):422–428
Uehori J, Fukase K, Akazawa T, Uematsu S, Akira S, Funami K, Shingai M, Matsumoto M, Azuma I, Toyoshima K, Kusumoto S, Seya T (2005) Dendritic cell maturation induced by muramyl dipeptide (MDP) derivatives: monoacylated MDP confers TLR2/TLR4 activation. J Immunol 174:7096–7103
Vazquez P, Ruíz-Larrañaga O, Garrido JM, Iriondo M, Manzano C, Agirre M, Estonba A, Juste RA (2014) Genetic Association Analysis of Paratuberculosis Forms in Holstein-Friesian Cattle. Vet Med Int 2014:321327
Vincze T, Posfai J, Roberts RJ (2003) NEBcutter: a program to cleave DNA with restriction enzymes. Nucleic Acids Res 31(13):3688–3691
Wadhwa A, Foote RS, Shaw RW, Eda S (2012) Bead-based microfluidic immunoassay for diagnosis of Johne’s disease. J Immunol Methods 382(2):196–202
Weiss DJ, Souza CD, Evanson OA, Sanders M, Rutherford M (2008) Bovine monocyte TLR2 receptors differentially regulate the intracellular fate of Mycobacterium avium subsp. paratuberculosis and Mycobacterium avium subsp. avium. J Leukoc Biol 83(1):48–55
Werling D, Hope JC, Howard CJ, Jungi TW (2004) Differential production of cytokines, reactive oxygen and nitrogen by bovine macrophages and dendritic cells stimulated with toll-like receptor agonists. Immunology 111:41–52
Werling D, Piercy J, Coffey TJ (2006) Expression of toll-like receptors (TLR) by bovine antigen-presenting cells: potential role in pathogen discrimination? Vet Immunol Immunopathol 112:2–11
White SN, Kata SR, Womack JE (2003) Comparative fine maps of bovine toll-like receptor 4 and toll-like receptor 2 regions. Mamm Genome 14(2):149–155
Whitlock RH, Wells SJ, Sweeney RW, Van Tiem J (2000) ELISA and fecal culture for paratuberculosis (Johne’s disease): sensitivity and specificity of each method. Vet Microbiol 77:387–398
Yadav R, Sharma AK, Singh R, Sonwane A, Kumar A, Chauhan A, Kumar S, Kumar T, Renjith R, Bhaladhare A, Prakash O (2014) An association study of SNPs with susceptibility to bovine paratuberculosis infection in cattle. Indian J Anim Sci 84(5):490–493
Acknowledgements
We thank Director, Indian Veterinary Research Institute, Izatnagar, India for funding this work.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors do not have any conflict of interest.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Kumar, S., Kumar, S., Singh, R.V. et al. Genetic association of polymorphisms in bovine TLR2 and TLR4 genes with Mycobacterium avium subspecies paratuberculosis infection in Indian cattle population. Vet Res Commun 43, 105–114 (2019). https://doi.org/10.1007/s11259-019-09750-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11259-019-09750-2