Abstract
Rotaviruses have a worldwide distribution and the infection is associated with diarrhea in young of ruminants as well as children. However, limited data exist on its prevalence and types in Yobe state, Nigeria. Detection of rotavirus A and types in ruminant population in Yobe state was the aim of the study. A total of 470 diarrheic fecal samples were collected and tested for rotavirus and types using serology and molecular techniques respectively. A prevalence rate of 2.98% (14/470) was found in the three species with specific rates of 2.9% (6/202), 3.8% (6/158), and 1.8% (2/110) in goat, sheep, and cattle respectively. The prevalence rates of 3.6% (12/331), 1.2% (1/84), and 1.8% (1/55) were for those aged < 1–3, 4–6, and 7–9 months old, respectively, while 4.9% (9/185) and 1.7% (5/285) were in males and females respectively. Rotavirus genes VP7 and VP4 were detected in 2 (14.3%) out of the 14 ELISA-positive samples while deduced amino acid sequences of the major variable regions revealed the genes to belong to types G3P[11] strain. Significant association was found between the infection and sex (P < 0.05) unlike in the species and age groups of the ruminants. The circulation of rotavirus virus in ruminants and type G3P[11] in cattle has been confirmed in the study. Hence, there is a need for continuous surveillance, awareness campaign, and assessment of the economic losses and public health implications of rotavirus infection in Nigeria.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Rotaviruses are among the disease-causing agents responsible for great public health challenges and high economic losses in livestock industry all over the world (FAO 2000). Nigeria has been ranked third among the 10 countries with the greatest number of rotavirus disease-associated deaths with up to 33,000 annual estimated deaths in children under 5 years old (Aminu et al. 2010). Recently, an 8-year surveillance study on diarrhea diseases in children less than 5 years of age has revealed that 2043 (47%) of the 4377 cases were positive for rotavirus with a case fatality rate of 1.3% (WHO 2018). They belong to the virus family Reoviridae and genus Rotavirus. They consist of eleven segments structural and non-structural proteins and measure 75 nm in diameter (Estes and Kapikian 2007). They are distinguished into eight serotypes (A–H) and several of them have been identified and reported in humans and animals worldwide (Adah et al. 1997a; Andrej et al. 2008; Dzikwi et al. 2008; Waggie et al. 2010). They cause diarrhea in young animal and children with high morbidity and mortality rates (Aminu et al. 2008; Sarma 2009; Dash et al. 2011). The infection has been revealed to be transmitted via the fecal-oral route and by contact with contaminated fomites (Mwenda et al. 2010; Dash et al. 2011).
Data on the epidemiology of the infection and the associated public health impact have not been fully documented (Sarma 2009; Dash et al. 2011; Ayuba 2011). At the present, more than 27 G and 35 P genotypes have been reported to be either restricted in some animal species or humans while some have shown high levels of interspecies transmission and from animals to humans (Gentsch et al. 2005; Rahman et al. 2005; Khamrin et al. 2006; Khamrin et al. 2007; Miyazaki et al. 2011). The reported increase in the isolation rate of G and P types in different parts of the world, like the predominance of G9[P6] strains in India, G8[P6] strains in Malawi, and G5[P8] strains in Brazil, calls for more attention in our environment (Van der Heidel et al. 2005). In Nigeria, there is still gap in literature on infection in animals compared with many report on the prevalence in children (Nimzing et al. 2000; Junaid et al. 2011; Aminu et al. 2014; Alkali et al. 2016). This study, therefore, was aimed at determining the prevalence of rotavirus A infection and detection of strain types in ruminants in Yobe state, Nigeria.
Materials and methods
Study area
Yobe state is located in the northeastern geopolitical zone of Nigeria between latitudes 10° 30′ and 13° 25′ N and longitudes 9° 35′ and 12° 30′ E of the Northern hemisphere. The state shares international boundary with Niger Republic in the extreme north and interstate boundaries with Borno state to the east, Bauchi and Jigawa states to the west, and Gombe state to the south (Oruonye 2009). The state has one of the largest livestock market in West Africa, and animal production in the state is consist of ruminants and local poultry (Oruonye 2009). Pastoralism is practiced by sedentary livestock farmers and nomadic Fulani groups (Oruonye 2009; Ayuba 2011) (Fig. 1).
Study sites
The study was carried out in the 17 local government areas (LGAs) of the state which has been grouped into five agricultural zones (MAWR 2008) as follows: Gashua zone (Bade, Karsuwa, and Jakusko LGAs), Geidam zone (Geidam, Bursari, and Yunusari LGAs), Gujba zone (Gujba, Gulani, Tarmua, and Damaturu LGAs), Nguru zone (Nguru, Machina, and Yusufari LGAs), and Potiskum zone (Potiskum, Fune, Nangere, and Fika LGAs). In all, five LGAs (one selected at random from each of the five agricultural zones) were used (Thrusfield 1997).
Study population
The study participants and subjects were young of ruminants in study sites. Calves, kids, and lambs with diarrhea in the selected LGAs constituted the target population.
Sampling frame
Five selected LGAs in all were visited for sample collection. Pastoralists’ camps, institutional farms, and household farms in the selected LGAs were identified and sampled.
Sampling methods
A multi-stage and cluster sampling methods were adopted to select sampling units (grazing fields and farms) (Mitchell 2003; USDA 2016; Casburn 2016). All individual target units (calves, kids, and lambs) in selected units (pastoralist’s camps, institutional farms, and household farms) were sampled as described by Putt et al. (1987). Calves, kids, and lambs were grouped according to age (< 1–3, 4–6, and 7–9) months old. Age categories of the ruminant where no records existed were determined by approximation by owner of the animal (Banerjee 1998).
Sample size estimation
The minimum sample size for each species was calculated using the formula n = z2 × p (1 − p) ÷ d2 (Thrusfield 1997) where n = sample size, z = score for confidence interval (95% confidence coefficient was taken at 1.96 score), p = expected prevalence, and d = 5% desired precision.
For cattle, p = 3.2% (Adah et al. 2002) n = 1.962 × 0.032(1 – 0.032) ÷ 0.052 = 48. For sheep, p = 11% (Alkali et al. 2017) n = 1.962 × 0.11(1 – 0.11) ÷ 0.052 = 150. For goat, p = 4% (Alkali et al. 2017) n = 1.962 × 0.04(1 – 0.04) ÷ 0.052 = 59. A minimum of 257 samples was required. However, in order to maximize the ELISA reagents available, 470 samples were used in the study.
Inclusion criteria
Only ruminant owners who consented to request for sample collection were included in the sample collection. Non-diarrheic and older than 12 months calves, kids, and lambs and adult cattle, goat, and sheep were excluded.
Descriptive criteria for diarrhea
Diarrhea is loose, watery stools which is said to have occurred if an animal has loose stools three or more times in 1 day. Acute diarrhea is diarrhea that lasts a short time about 1 or 2 days which is the most common sign of rotavirus infection in young animals (WHO 2009a, b).
Sample collection and storage
About 1–2 g fecal samples was collected per-rectum into a labeled, capped, sterilized, wide mouth, universal plastic container from each of the animals using aseptic disposable wood spatula and hand gloves. The samples were transported cooled using ice to the laboratory and stored at 2–4 °C for less than 48 h and further processed as described by Dash et al. (2011) and Junaid et al. (2011).
Sample preparation
About 0.1 g or 300 μL fecal suspension was added to 1 mL distilled water for each sample. The mixture was vortex for a minute and 1 mL of supernatant was transferred into a fresh labeled tube and used for ELISA test.
Detection of rotavirus group A (RVA) virus
Enzyme-linked immunoassay (ELISA) test kit for antigenic detection of rotavirus was used (Bio K 343/2™, Bio-X Diagnostics, Belgium) on 470 samples while the ELISA-positive samples were subjected to RT-PCR and second round PCR to determine their genotypes (WHO 2009a, b). In addition, deduced amino acids sequencing of the major variable regions of the genes was equally conducted. For the ELISA test, a commercially available ELISA kit was used and the optical density readings were measured using microplate spectrophotometer (Optical Ivymen System®) with a 450-nm filter and calculated according to the manufacturers’ instruction (Yilmaz et al. 2017). RNA extraction, RT-PCR of VP7 and VP4 to complementary DNA (cDNA), and gel electrophoresis (PAGE) were done as described by Herring et al. (1982) and Dash et al. (2011), to confirm the rotavirus group A types using specific primers (Dzikwi et al. 2008; Esona et al. 2010) (Tables 1 and 2).
RNA extraction for RT-PCR
RNA extraction was performed by TRIzol method. Briefly, 500 μL of fecal suspension was transferred into an Eppendorf tube; 1 mL TRIzol reagent was added, vortexed, and incubated for 5 min at room temperature; and 200 μL chloroform was added, vortexed, and kept at room temperature for 3 min. The mixture was centrifuged for 10 min at 20817.16 gravity. The upper aqueous phase containing the dsRNA 500 μL was carefully harvested and placed in clean labeled Eppendorf tubes. To the freshly harvested RNA, 500 μL isopropanol was added and incubated at room temperature for 10 min which allowed the reaction to precipitate. The solution was centrifuged at 14000 rpm for 10 min then the fluid was decanted without disturbing the pellet. The pellets were washed using 500 μL of 75% ethanol and the fluid decanted without disturbing the pellet. The pellet was air-dried and suspended in 50 μL nuclease-free water and incubated at 55 °C for 5 min. The extracted dsRNA was used in the one-step RT-PCR.
PCR amplification of the cDNA
The transcribed dsRNA (cDNA) was subjected to amplification in the following order. To the incubated reaction solution, 39.7 μL (OneTaq and One-Step Enzyme Mix) and 0.3 μL Taq polymerase were added and the reaction was thereafter subjected to the following PCR program as follows: Initial denaturation at 94 °C for 2 min, 30 cycles of denaturation at 95 °C for 30 s, annealing at 42 °C for 2 min, extension at 72 °C for 5 min, final extension at 72 °C for 7 min, and held at 4 °C. The PCR products were loaded into a prepared agarose gel which contained 4 μL mL−1 ethidium bromide and was electrophoresed in Tris-acetic acid-EDTA (TAE) buffer at 100 V for 45 min and the migration size of each sample was captured and visualized to determine the rotavirus full-length VP7 and VP4.
RT-PCR products and genotyping of the gene VP4 and VP7 for rotaviruses
The following PCR protocol was used: Initial denaturation at 94 °C for 2 min, 30 cycles of denaturation at 95 °C for 30 s, annealing at 55 °C for 2 min, elongation at 72 °C for 1 min and final elongation at 72 °C for 7 min, and held at 4 °C. The PCR product was loaded to 2% agarose gel and the genotypes in kilo base pair (bp) was determined from the size of the amplicons.
Interpretation of RT-PCR products and genotyping results
The RT-PCR genotyping results were interpreted by comparing the band (produced migration of sample as captured from the gel electrophoreses) with the expected band of positive control of respective genotypes (G types and P types).
Data analyses and presentation
Data were analyzed for percentages/proportion frequency tables. Logistic regression was used to obtain p values, odds ratios (OR), and 95% confidence interval (95% CI). BioEdit BioStatistical software was used for sequences analysis while GenBank/Blast and MEGA 7 were used for the reference strains.
Results
A total of 470 fecal samples from diarrheic young of three ruminant species (bovine, 110; ovine, 202; caprine, 158) at different farms in the five agricultural zones of Yobe state was collected and analyzed. The overall prevalence rate of rotavirus infection in ruminants in the five agricultural zones of Yobe state as detected with ELISA was 2.98% with species-specific prevalence rates of 3.8% (6/158), 2.9% (6/202), and 1.8% (2/110) for kids, lambs, and calves respectively (Table 3). The odds of the infection for ovine species are almost equal to the odds of the infection for both bovine and caprine species with true population effect between 96.0 and 80.9%. The result was not statistically significant (p = 0.636; OR = 0.932; 95% CI = 0.398–1.907).
The infection was detected in all the age groups studied. Out of the 14 ELISA-positive cases recorded, the age group < 1–3 months showed the highest prevalence of 3.6% (12/331) followed by age groups 7–9 months 1.8% (1/84) and age group 4–6 months 1.2% (1/55). The result was not statistically significant (p = 0.435; OR = 1.943; 95% CI = 0.675–5.764) (Table 4).
The prevalence of rotavirus infection in the males was 4.6% (9/185) which is higher than 1.8% (5/285) observed in the females. The result was statistically significant but the odds of the infection for male was 3.6 times more likely than the odds of the infection for female with true population effect between 90.4 and 14.8% (p = 0.050; OR = 3.648; 95% CI = 0.965–11.475) (Table 4).
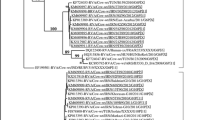
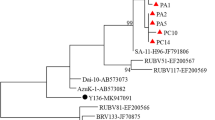
Rotavirus infection was detected in all the five agricultural zones of Yobe state and the distribution of the 14 Rotavirus-positive infection among ruminant species showed that Damaturu zone had the highest prevalence rate 57.1% (8/14) followed by Gashua and Nguru zones 14.3% (2/14) each, while Geidam and Potiskum zones had the least rates of prevalence 7.14% (1/14) (Fig. 2). Amplicons corresponding to full-length VP7 and partial length VP4 genes were detected in 2 (14.3%) of the 14 ELISA-positive samples from ruminants (Figs. 3 and 4). Furthermore, the deduced amino acid sequences of the major variable regions revealed one of the genes to belong to types G3[P11] when compared with sequences from the gene bank (Table 5, Figs. 5 and 6).
Gel picture of rotavirus VP7 gene with RT-PCR products from cattle in Yobe state. Lane 1 indicates 1200 bp DNA marker while lanes 2–16 were samples. Lanes 4 and 5 indicate expected band size for VP7 of 1062 bp. Lane 4 showed faint band while lane 5 showed bold band on the gel as pointed by the thick arrow
Deduced amino acid (15–255) sequences of VP7 antigenic sites of animal rotavirus A detected (BB4_22 consensus) in Yobe compared with G-type reference strains. GenBank/Blast were used for data analysis of the reference strains while MEGA 7 was used for analysis of samples. Reference strains and sequences obtained from GenBank/NCBI are indicated by their accession numbers. Sequences begin from the 15th amino acids (aa). Major variable antigenic sites A (aa 87–101), B (aa 142–152), and C (aa 208–221) were used to deduced genotype
Deduced amino acid (1–214) sequences of VP4 antigenic sites of animal (bovine) rotavirus A (consensus BC4_2) detected in Yobe compared with P[11] reference strains. BioEdit BioStatistical software was used for sequence analysis. Reference strains and sequences obtained from GenBank/NCBI are indicated by their accession number. Sequences begin from the 31st amino acids (aa). Major variable antigenic sites A (aa 87–101), B (aa 142–152), and C (aa 208–221) were used to deduced genotype
Discussion
Rotavirus infection is of both economic and public health importance. Its prevalence in ruminants and the detection of G3 [P11] strain in cattle is a confirmation of the circulation of the virus strain in the study area. The detected strain was different from the G8 genotype previously reported in bovine species in the north-east Nigeria (Adah et al. 2002). Previous studies in Nigeria had also identified strains of G1P[8], G3P[6], G1P[6], and G2P[6] as the common prevalent ones and G3P[8] which could not be typed due to mutation (Adah et al. 1997b; Adah et al. 2001). The discovery of G3 [P11] strain in this study gives an additional insight to the genetic variability and the emergence of rotavirus reassortants in the environment. This unusual diversity among the strains in Nigeria has been associated with mixed infections with rotavirus including those of animal origin (Gouvea and Brantly 1995; Adah et al. 2001). Assessment of evolution and epidemiological pathways of Rotaviruses in human, mammals, and birds has been conducted by genotyping techniques (Matthijnssens and Van Ranst 2012). These include varied mechanisms of evolution by mutation types of one or several combinations of the following: point mutations, genome reassortment, genome rearrangements, and true genome recombination (Kojima et al. 2000; Jere et al. 2011; Matthijnssens and Van Ranst 2012; Luchs and Timenetsky 2014; De Grazia et al. 2014).
Changes in virus virulence resulting from mutation in viral gene products have been reported (Tyler and Fields 1990; Andrej et al. 2008; Midgley et al. 2012). The G3 genotype detected in the study is one of the three rotavirus genotypes which have been reported in humans and animals (Desselberger and Huppertz 2011). This may suggest the expanding landscape of rotavirus genotypes in Yobe state, with a possible reduction in productivity and loss of revenue which can be associated with treatment and death of animals in state which supports production and trade in livestock in Nigeria geographically (Oruonye 2009). The infection in ruminant’ species may not only affect international trade on meat or other livestock products, but there is the risk of acquiring of the infection among other animal species and humans (Santos et al. 1999). Furthermore, it implies that any Rotavirus-control program in cattle alone would not be successful, if not coupled with surveillance and control policies in small ruminants (Gentsch et al. 2005). Therefore, data on circulating rotavirus strains in animals is pertinent in vaccine production for use in developing countries (Aminu et al. 2010).
The overall prevalence (2.98%) recorded in this study can be associated with extremely low viral antigen concentration in the fecal samples as it has been revealed that not less than 109 virus particles per gram are required for detection of Rotaviruses in fecal samples (Grassi et al. 2009). It may also be a reflection of the environmental contamination in the state. Earlier researchers have also observed that rotavirus infection tend to be higher in intensively reared farm animals unlike the extensive system being practiced in the study area (Fenner et al. 1993). The prevalence rate in calves was higher than the zero prevalence reported by Alkali et al. (2017) in the same species in Sokoto, north-western, Nigeria, but was lower than 3.2% and 23.16% prevalence rates in calves as reported by Adah et al. (2002) and Aminu et al. (2014) in Nigeria. Furthermore, the prevalence rates of 3.8% and 2.9% in kids and lambs, respectively, in this study were lower than that of 11.0% in kids and 4.0% in lambs in Sokoto state, Nigeria (Alkali et al. 2017). Discrepancy in prevalence rates of Rotaviruses has been ascribed to many factors including sample size, storage, and screening method used. The detection of the infection in all the age groups studied was in agreement with the findings in earlier studies by Adah et al. (2002) and Aminu et al. (2014) who reported higher prevalence in calves of age group 0–3 months in northern Nigeria. It is believed that severity and susceptibility to rotavirus infection in newborn calves, lambs, kids, or piglets decreases as the animal becomes older (Adah et al. 2002). The high prevalence found in ruminants less than 3 months of age in this study could be due to insufficient antibodies against rotavirus infection in the animals as a result of milk deprivation for economic reasons, feeding on contaminated feed, or contacts with Rotavirus-contaminated environment.
The higher prevalence of rotavirus in male 4.6% (9/185) than in female 1.8% (5/285) in this study was in agreement with other report findings that reported male calves as highly susceptible to rotavirus infection than the females (Adah et al. 2002; Sarma et al. 2009; Aminu et al. 2014). However, more studies are needed to prove that the sex susceptibility is not by chance (Junaid et al. 2011). Rotavirus infection as detected in ruminant species in all the five agricultural zones of Yobe state is a revelation of widespread distribution of rotaviruses in various communities of the state. Kapikian and Chanock (1996) observed that distribution of rotaviruses in a community is shown by acquisition of antibodies to pathogens in infected animals. The observed difference in rates of infection across zones may be due to difference in spread of animal Rotaviruses or the level of herd immunity of ruminants or environmental contamination of the agricultural zones in the state. It has been reported that sample sources from healthy or non-diarrheic animals usually have low concentrations of rotavirus (Tate et al. 2013); however, the detection of the virus at low level from diarrheic fecal samples with the use of both ELISA and RT-PCR further confirms the low level of environmental contamination in the area. On the other hand, the reason for the low rate of detection using molecular technique in this study could be that the nucleic acid in most of the ELISA-positive samples had been degraded during long storage (Halstead et al. 2013) or were insufficient to permit genotyping (Adah et al. 1997a). Factor like point mutation in the genotyping regions of the VP7 and VP4 isolates has equally been implicated (Manuja et al. 2008; Sarma 2009). Furthermore, RT-PCR procedure with the two most commonly used primer pair sets are among other factors to be considered (Cunliffe et al. 1999; Esona et al. 2010; Gouvea et al. 1990; Fischer et al. 2003).
Sequence analysis has become the standard for both confirmation and identification of “non typable” strains. Confirmatory sequencing is usually performed either on the genotype-specific products or on a fragment of the VP7 or VP4 gene after amplification (WHO 2009a, b) and by compares of the sequences using the genotype-specific variable regions (Kobayashi et al. 2007). The results described herein indicate a 2.98% prevalence of rotavirus infection in ruminant species and a new virus strain G3[P11] in cattle, revealing the diversity of rotavirus strains in the animal population in the dispersed agricultural zones of Yobe state, Nigeria. This could be a pointer to potentials for genetic reassortment among rotavirus strains circulating in the state and other public health and economic implications, thus highlighting the urgent need for continuous surveillance, awareness campaign, and measures to control the infection spread.
References
Adah, M. I., Rohwedder, A. Olaleye, O. D. and Werchau, H., 1997b. Nigerian rotavirus serotype G8 could not be typed by PCR due to nucleotide mutation at the 3_ end of the primer binding site, Archives of Virology, 142,1881–1887.
Adah, M.I., Rohwedder, A., Olaleye, O.D., Durojaiye, O.A. and Werchau, H., 1997a. Serotypes of Nigerian rotavirus Strains, Tropical Medicine and International Health, 2(3), 363-370.
Adah, M.I., Wade, A. and Taniguchi K., 2001. Molecular Epidemiology of Rotaviruses in Nigeria, Detection of Unusual Strains with G2P[6] and G8P[1] Specificities, Journal of Clinical Microbiology, 3969–3975.
Adah, M.I., Jaji, Z., Agwazim, B.F., El-yuguda, A.D. and Mani, A.U., 2002. Detection of human rotavirus in faeces from diarrhoeic calves in north-east Nigeria, Tropical Animal Health and Production, 34, 1-6.
Alkali, B.R., Daneji, A.I., Magaji., Magaji, A.A., Bilbis, L.S. and Bande, F., 2016. Molecular characterization of human rotavirus from children with diarrheal disease in Sokoto state, Nigeria, Molecular Biology International Volume 2016, article I. D 1876065, 9 pages. Hindawi Publishing Corporation.
Alkali, B.R., Daneji A.I., Magaji AA, Ibis, L.S. and Yabo Y.A., 2017. Molecular detection of animal rotaviruses in Sokoto, Nigeria. Scholarly Journal of Biological Science, 6 (1): 1-5.
Aminu, M., Ahmad, A.A., Umoh, J.U., DeWar, J., Esona, M.D. and Steele, A.D., 2008. Epidemiology of rotavirus infection in North-western Nigeria, Journal of Tropical Pediatrics, 54-(5), 340-342.
Aminu, M., Page, N. A., Ahmad, A. A., Umoh, J. U., Dewar, J. and Steele, A. D., 2010. Diversity of Rotavirus VP7 and VP4 Genotypes in Northwestern Nigeria, Journal of Infectious Diseases, 202(S1), S198–S204
Aminu, M., Amupitan, E., Umoh, J.U., Dzikwi, A. and Esona, M.D., 2014. Detection of rotavirus antigens from calves in Zaria, Nigeria,’ Eleventh international rotavirus symposium, held on 3-5 September, New Delhi, India.
Andrej, S., Mateja, P., Darja, B.M. and Jozica, M., 2008. Human, porcine and bovine rotaviruses in Slovenia: evidence of interspecies transmission and genome reassortment, Journal of General Virology, 89, 1690-1698.
Ayuba, H.K., 2011. Baseline study on livestock-wildlife environment interface in the Nigerian sector of the Lake Chad basin, Being an IUCN consultancy Project undertaken, 9-17.
Banerjee, G.C., 1998. A textbook of animal husbandry, (Oxford and IBH publishing company Pvt limited, 113B Shahpur Jat, Asian games village side, New Delhi 11004, India)
Casburn, G., 2016. How to tell the age of sheep, Primefact 1481, second edition, (Department of Primary Industries, Wagga, Wagga, New South Wales, UK), 1-2.
Cunliffe, N.A., Kilgore, P.E. and Bresee, J.S., 1999. Epidemiology of rotavirus diarrhea in Africa: a review to assess the need for rotavirus immunization, Bulletin of World Health Organization, 76, 525-537.
Dash, S.K., Tewari, A., Kumar, K., Goel, A. and Bhatia, A.K., 2011. Detection of rotavirus from diarrhoeic cow calves in Mathura, India, Veterinary World, 4 (12), 554-556.
De Grazia, S.,Bonura, F., Colomba, C., Cascio, A,. Di Bernardo, F., Collura, A.,Terranova, D.M., Martella, V.,GiammancoG.M., 2014.Data mining from a 27-years rotavirus surveillance in Palermo, Italy. Infection, Genetics and Evolution, S1567-1348(14)00083-5
Desselberger, U. and Huppertz, H.I. (2011). Immune response to rotavirus infection and vaccination and associated correlates of protection, Journal of Infectious Diseases, 203(2), 188-195
Dzikwi, A.A., Umoh, J.U., Kwaga, J.K.P., Ahmad, A.A., deBeer, M. and Steele, A. D. 2008. Electropherotypes and subgroups of group a rotaviruses circulating among diarrhoeic children in Kano, Nigeria, Annals of African Medicine 7 (4), 163 – 167.
Esona, M.D., Mijatopic-Rustempasic, S., Confardy C., Tong, S., Kuzmin, I.V., Agwanda., Breiman, R.F., Banyai K., Niezgoda M., Rupprecht C.E., Gentsch J.R. and Bowen M.D., 2010. Reassortant group A rotavirus from straw-colored fruit bat (Eidolon helvum), Emerging Infectious Disease, 16, 1844-1852.
Estes, M. and Kapikian A., 2007. Rotaviruses,’ In knipe, D., Howley, P., Griffin, D., Lamo, R., Martin, M., Roizman, B., Straus, S., Eds: Fields’ Virology. Wolters kluwer Health, Lippincort Williams and Wilkins, Philadelphia, 1917 – 1974.
FAO, 2000. FAO animal health manual 10’ manual on participatory epidemiology - method for the collection of action-oriented epidemiological intelligence, Food and Agriculture Organization of the United Nations, Rome, Accessed on 26/07/2016 at hhpp://www.fao.org/ …/livestock_statistics_ concept
Fenner, J.F., Paul, E., Gibbs, J., Murphy, F.A., Rott, R., Studart, M.J. and White D.O., 1993. Veterinary virology, 2nd ed. Academic Press Inc., U.S.A. 538, 550-552.
Fischer, T.K., Page, N.A, Griffin, D.D., Eugen-Olsen, J., Pedersen, A.G., Valentiner-Branth, P., Molbak K., Sommerfelt, H. and Neilsen N.M., 2003. Characterization of incompletely typed rotavirus strains from Guinea-Bissau: identification of G8 and G9 types and a high frequency of mixed infections, Journal of Virology, 311,125–133.
Gentsch, J.R., Laird, A.R., Bielfeld, B., Griffin, D.D., Banyai, K., Ramachandran, M., Virek J., Nigel A.C., Nakagomi C., Kirkwood C.P., Fisher T.K., Parashar U.D., Bresse J.S., Jiang, B. and Glass R.I., 2005. Serotype diversity and reassortment between human and animal rotavirus strains: implications for rotavirus vaccine programs, Journal of Infectious Diseases, 192, 146–159.
Gouvea, V., and Brantly, M., 1995. Is rotavirus a population of reassortants? Trends in Microbiology, 3, 150–162.
Gouvea, V., Glass, R.I., Woods, P., Taniguchi, K., Clark, H.F., Forrester, B. and Fang, Z.Y., 1990. Polymerase chain reaction amplification and typing of rotavirus nucleic acids from stool specimens, Journal of Clinical Microbiology, 28, 278-282.
Grassi, T., Bagordo, F., Idolo, A., Lugoli, F.,Gabutti, G. and De Donno, A., 2009. Rotavirus detection in environmental water samples by tangential flow ultrafiltration and RT-nested PCR, Environ. Monit. Assess, 1661-1669.
Halstead, F., Lee, A.V., Couto-padara, X., Polley, S.D., Ling, C., Jenkins, C., Rachel M Chalmers, Kristin Elwin, Gray J. J., Iturriza-gómara M., Wain J., Clark D.A., Bolton F.J., Manuel R.J. and Olympics G.I. group 2013. Universal extraction method for gastrointestinal pathogens, Journal of Medical Microbiology, 62, 1535-1539.
Herring, A.J., Inglis, N.T., Ojeh, C.K., Snodgrass, D.R. and Menzies, J. D., 1982. Rapid diagnosis of rotavirus infection by direct detection of viral nucleic acid in silver stained polyacrylamide gels, Journal of Clinical Microbiology, 16, 473-477.
Jere, K.C., Mlera, L., Page, N.A., van Dijk, A.A., O’Neill, H.G. , 2011. Whole genome analysis of multiple rotavirus strains from a single stool specimen using sequence-independent amplification and 454® pyrosequencing reveals evidence of intergenotype genome segment recombination. Infection, Genetics and Evolution, 11 (8), 2072-2082
Junaid, S.A., Umeh, C., Olabode, A.O. and Banda, J.M., 2011. Incidence of rotavirus infection in children with gastroenteritis attending Jos University Teaching Hospital, Nigeria, Journal of Virology, 8, 233:1-12.
Kapikian, A.Z. and Chanock, R.M., 1996. Rotaviruses, In: Fields virology. Fields BN, Knipe DM, Howley PM, Chanock RM, Melnick JL, Monath TP, Roizman B, Stanus (eds). Vol. II 3rd ed. Philadelpia: Lippincostt-Raven; 1657-1708.
Khamrin, P., Maneekarn, N., Peerakome, S., Chan-it, W., Yagyu, F., Okitsu, S. Ushijima, H., 2006, ‘Molecular characterization of a rare G3P[3] human rotavirus reassortant strain reveals evidence for multiple human-animal interspecies transmissions, Journal of Medical Virology 78, 986–994.
Khamrin, P., Maneekarn, N., Peerakome, S., Chan-it, W., Yagyu, F., Okitsu, S. and Ushijima H., 2007. Novel porcine rotavirus of genotype P[27] shares new phylogenetic lineage with G2 porcine rotavirus strain, Journal of Virology, 361, 243–252.
Kobayashi, N., Ishino, M., Wang, Y-H., Chawla-Sarkar, M., Krishnan, T., and Naik, T.N. 2007. Diversity of G-type and P-type of human and animal rotaviruses and its genetic background, Communicating Current Research and Educational Topics and Trends in Applied Microbiology, 847-858.
Kojima, K., Taniguchi, K., Kawagishi-Kobayashi, M., Matsuno, S. Urasawa, S., 2000. Rearrangement generated in double genes, NSP1 and NSP3, of viable progenies from a human rotavirus strain. Virus Research, 67 (2), 163-171
Luchs, A., and Timenetsky, M.DOC., 2014. G8P[6] rotaviruses isolated from Amerindian children in Mato Grosso do Sul, Brazil, during 2009, close relationship of the G and P genes with those of bovine and bat strains. Journal of General Virology, 95 (Pt 3), 627-641
Manuja, B., Minakshi, K., Manuja, A., Gulat, B.R. and Prasad, G., 2008. A novel genomic constellation (G10P[3]) of group A rotavirus detected from buffalo calves in Northern India, Virus Research, 138, 36-42
Matthijnssens, J., and Van Ranst, M., 2012. Genotype constellation and evolution of group A rotaviruses infecting humans, Current Opinion in Virology, 2 (4), 426-433
MAWR, 2008. Report of the ministry of agriculture and water resources. In Anonymous, (2008). Proceedings of the Yobe state economic summit; edited by Ngama Y. L, Bunu Z and Saidu S.A. (Published by spectrum books limited, spectrum house, ring road, Ibadan, Oyo state, Nigeria in association with safari books (export) limited, 1st floor, 17 bond street, st Helier Jersey JE2 3NP, Channel Islands United kingdom), 37-64.
Midgley, S.E., Banyai, K., Buesa, J., Halaihel, N., Hjulsager, C.K., Jakab, F.J., Kaplon, J., Larsen Le, Monini M., Poljšak-Prijatelj, M., Pothier, P., Ruggeri, F.M., Steyer, A., Koopmans, M. and Bottiger, B., 2012. Diversity and zoonotie potential of rotaviruses in swine and cattle across Europe, Veterinary Microbiology, 156(3-4), 238-245.
Mitchell, T., 2003. How to tell the age of goats,’ (AGFACTS publication of division of animal production, Dubbo, New South Wales, UK), 1-2.
Miyazaki, A. Kuga, K. Suzuki, T. Kohmoto, M. Katsuda, K. and Tsunemitsu, H., 2011 Genetic diversity of group A rotaviruses associated with repeated outbreaks of diarrhea in a farrow-to-finish farm: identification of a porcine rotavirus strain bearing a novel VP7 genotype, G26. Veterinary Research 42 (1):112
Mwenda, J.M., Ntoto, K.M., Abebe, A., Enweronu-Laryea, C., Amina, I., Mchomvu, J., Kisakye, A., Mpabalwani, E.M., Pazvakavambwa, I., Armah, G.E., Seheri, L.M., Kiulia, N.M., Page N., Widdowson, M.A. and Steele A.D., 2010. Burden and epidemiology of rotavirus diarrhea in selected African countries; preliminary results from the African rotavirus surveillance network, Journal of Infectious Diseases, 202 Supplemary (1), 5-11.
Nimzing, L., Geyer, A., Sebata, T. and Steele A.D. 2000. Epidemiology of adenoviruses and rotaviruses identified in young children in Jos, Nigeria, South Africa Journal of Epidemiology and Infections, 15, 40 -42.
Oruonye, E.D., 2009. Geographical aspects of Yobe state of Nigeria, (Published by Afab educational books, Shukura house, katako junction, Jos, Plateau state, Nigeria), 45-69.
Putt, S.N.H., Shaw, A.P.M., Woods, A.J., Tyler L. and James, A.D. 1987. Veterinary epidemiology and economics in Africa’: A manual for use in the design and appraisal of livestock health policy (Department of Agriculture, University of Reading, ILCA Manual No.3, Berkshire, Reading, England), 27-45.
Rahman, M., Matthijnssens J. and Nahar S., 2005. Characterization of a novel P [25], G11 human group a rotavirus, Journal of Clinical Microbiology, 43, 3208- 3212.
Santos, N., Lima, R.C., Nozawa, C.M., Linhares, R.E. and Gouvea, V., 1999. Detection of porcine rotavirus type G9 and of a mixture of types G1 and G5 associated with Wa-like VP4 specificity: evidence for natural human – porcine genetic reassortment, Journal of Clinical Microbiology, 37, 2734 -2736.
Sarma D.K., 2009. ‘In: A Rotavirus infections in bovine and swine,’ Virology and viral diseases, 2nd edition; 295-298, (Kalyan; publishers, B-15, Sector 8, Noida (U.P), New Delhi-110002, India)
Tate, J.E., Mijatovic-Rustempasic, S., Tam, K.I., Lyde, F.C., Payne, D.C., Zilagyi, P. S. Edwards, K., Staat, M.A., Weinberg, G.A., Hall, C.B., Chappell, J., Mcneal, M., Gentsch, J.R., Bowen, M.D. and Parashar, U.D., 2013. Comparison of 2 Assays for Diagnosing Rotavirus And Evaluating Vaccine Effectiveness in Children With Gastroenteritis, Emerging Infectious Diseases, 19, 1245–1252.
Thrusfield M., 1997. ‘Veterinary Epidemiology, 2ndedition, Blackwell, 18CowleyRoad, Oxford, 183-187.
Tyler, K.L., and Fields B.N.,1990. Pathogenesis of viral infection. In: Fields, BN. Knipe DM, editors. Fields Virology 2nd ed. (Raven Press, New York) 191-239
USDA, 2016, ‘Using dentition to age cattle. Accessible at http//www.fsis.usda.gov/about_FSIS/Technical_Service_Centre/idex. Last modified on 23/03/2015. Accessed 9/03/2016
Van der Heidel, R., Koopmans, M.P.G., Shekary, N., Houwers, D.J., Van Duynhoven, Y. T. H. P. and Van der Poel, W.H.M. 2005. Molecular Characterizations of Human and Animal Group A Rotaviruses in The Netherlands, Journal of Clinical Microbiology, 43, 2669-2675.
Waggie, Z., Hawkridge, A. and Hussey, G.D., 2010. Review of rotavirus studies in Africa: 1976- 2006, Journal of Infectious Diseases, 202 (Supplementary), 23 -33.
WHO, 2009a. Diarrhoea: why children are still dying and what can be done. United Nations Children Fund (UNICEF)/World Health Organization (WHO), WHO press, World Health Organization,20 Avenue Appia,1211 Geneva 27, Switzerland. 9.
WHO, 2009b. Manual of rotavirus detection and characterization methods, Immunization, Vaccines and Biologicals, WHO/IVB/08.17.
WHO, 2018. Nigeria to avert over 160,000 deaths in children yearly, with introduction of rotavirus vaccine into immunization schedule https:www.afro.who.int/news/nigeria_avert, Accessed on 17th February, 2020.
Yilmaz, V., Timurkan, M.O., Coskun, N. and Yildirim, Y., 2017. Investigation of Rotavirus infection in sheep using serological and molecular techniques, Indian Journal of Animal Research, 51(3), 525-530
Acknowledgments
The authors would like to thank the management and staff of the Department of Veterinary Public Health and Microbiology Laboratories, Usmanu Danfodiyo University Sokoto, Nigeria, for their kind assistance during the sample analysis.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
The manuscript does not contain clinical studies or patient data.
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Garba, J., Faleke, O.O., Magaji, A.A. et al. Prevalence of rotavirus A infection and the detection of type G3P[11] strain in ruminants in Yobe state, Nigeria. Trop Anim Health Prod 52, 2905–2915 (2020). https://doi.org/10.1007/s11250-020-02291-w
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11250-020-02291-w