Abstract
Sugar beet is an economically important crop and one of the major sources of sucrose around the world. Beet necrotic yellow vein virus (BNYVV) and Beet severe curly top virus (BSCTV) are two widespread viruses in sugar beet that cause severe damage to its performance. Previously, we have successfully produced resistance to BNYVV based on RNA silencing in sugar beet by introducing constructs carrying the viral coat-protein-encoding DNA sequence, CP21, in sense and anti-sense orientations. Yet, the RNA silencing-mediated resistance to a specific virus could be affected by other ones as a part of synergistic interactions. In this study, we assayed the specificity of the induced resistance against BNYVV in two sets of transgenic events, S3 and S6 carrying 5’-UTR with or without CP21-coding sequences, respectively. These events were subjected to viral challenges with either BNYVV, an Iranian isolate of BSCTV (BSCTV-Ir) or both. All the plants inoculated with just BSCTV-Ir displayed curly-leaf symptoms. However, partial resistance was evident in S3 events as shown by mild symptoms and reduced PCR amplification of the BSCTV-Ir coat protein encoding sequence. Based on the presented data, resistance to BNYVV was stable in almost all the transgenic plants co-infected with BSCTV-Ir, except for one event, S3-229. In general, it seems that the co-infection does not affect the resistance to BNYVV in transgenic plants. These findings demonstrated that the introduced RNA silencing-mediated resistance against BNYVV in transgenic sugar beets is specific and is not suppressed after co-infection with a heterologous virus.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
As a primary source of sugar production, sugar beet (Beta vulgaris L.) is one of the most important industrial crops in the world. Due to the increased global demand for sugar, the sustainability of sugar beet production is essential (Stevanato et al. 2019). Rhizomania, caused by Beet necrotic yellow vein virus (BNYVV), is one of the most devastating and widespread diseases of sugar beet that could diminish sugar beet yield by up to 80% (McGrann et al. 2009; Biancardi and Lewellen 2016). The virus is transmitted to sugar beet roots by Polymyxa betae Keskin, a plasmodiophorid vector, which remains viable in the soil for over 15 years by forming resting spores (Pferdmenges 2007; Biancardi and Lewellen 2016). While the use of chemicals is now phased out as part of the Montreal Protocol (McGrann et al. 2009), current resistant cultivars, such as those carrying Rz1 and/or Rz2 genes have been the only solution for cultivation in diseased areas so far. The molecular mechanism that underlies Rz1 resistance is still unclear (Funk et al. 2018). It was identified that the Rz2 gene encodes a coiled-coil nucleotide-binding leucine-rich repeat (CC-NB-LRR) protein (Capistrano-Gossmann et al. 2017). However, the BNYVV resistance conferred by Rz1 and/or Rz2 genes is reported to be prone to break in some regions (for example see Pferdmenges and Varrelmann 2009; Kutluk Yilmaz et al. 2018). Thus, it is necessary to explore other ways to effectively deal with this disease as soon as possible.
The advent of genetic engineering has opened up new ways to control Rhizomania by introducing novel resistance genes resources (Pavli et al. 2011; Dhir et al. 2019). In recent years, several methods based on RNA silencing and predominantly pathogen-induced resistance have emerged to strengthen plant defenses against viral invasions (Palukaitis 2011; Duan et al. 2012; Uslu and Wassenegger 2020). Virus-induced gene silencing (VIGS) is an RNA silencing-based mechanism that innately activates the plant’s natural defense mechanism against viruses (Lu et al. 2003; Duan et al. 2012). In this approach, part of the viral genome is introduced into plant cells which generates double-stranded RNAs (dsRNA) intermediates that trigger the silencing mechanism producing short interfering RNA (siRNA) (Lu et al. 2003; Duan et al. 2012).
To date, RNA silencing-mediated resistance has been effectively applied in various plants (Duan et al. 2012; Jin et al. 2020; Jiang et al. 2022). In particular, through the RNA silencing mechanism, the transgenic N. benthamiana expressing the coat protein (CP) read-through domain of BNYVV revealed very low levels of virus after inoculation (Andika et al. 2005). In another study, an inverted cDNA repeat derived from the BNYVV replicase gene was transferred into the sugar beet genome and showed considerable resistance to the virus (Lennefors et al. 2006). Transgenic hairy roots of sugar beet exhibited a remarkable resistance against Rhizomania through expressing BNYVV-derived dsRNA (Pavli et al. 2010). In our recent publications, we have shown RNA silencing-mediated resistance against Rhizomania in sugar beet in both transient and stable transformation of a number of constructs expressing BNYVV-derived RNA which confirmed the effectiveness of this mechanism in the greenhouse and field experiments (Zare et al. 2015; Safar et al. 2021).
However, a sugar beet field may be exposed to several kinds of pathogens such that the co-infection of plants by two or more viruses is quite possible (Susi et al. 2015; Moreno and López-Moya 2020). Co-infection often leads to interactions between viruses which can affect disease development in plants both negatively (antagonistic) and positively (synergistic) (Syller 2014; Syller and Grupa 2016; Mascia and Gallitelli 2016). Syller and Grupa (2016) suggested that synergistic interactions within plants mostly occur between unrelated viruses. Such viral interactions have been reported to enhance infection severity, particularly through the suppression of RNA-silencing machinery (Li et al. 2017; Liang et al. 2017; Aulia et al. 2019). For instance, rice tungro disease is caused by the synergistic interaction of Rice tungro bacilliform virus (RTBV) and Rice tungro spherical virus (RTSV). It was shown that combined actions of RTBV ORF-IV and RTSV CP3 proteins play a key role in tungro symptom development by suppressing RNA silencing in rice (Anand et al. 2022). Therefore, some concerns have been raised over the efficiency of RNA silencing-based resistance of transgenic plants under co-infection conditions.
Beet curly top virus (BCTV), a member of the Curtovirus genus, is another common and destructive virus in sugar beet fields around the world. Beet severe curly top virus (BSCTV, recently called BCTV-Svr) is a strain of BCTV named for the severity of curly symptoms in infected sugar beet. Iranian isolate of Beet severe curly top virus (BSCTV-Ir) is one of the main causal agents of the curly top disease in sugar beet farms in Iran. The C2/L2 protein of BCTV has been described as a suppressor of RNA silencing machinery (Yang et al. 2007). Besides, it was recently revealed that V2 of BCTV can also act as an inhibitor of RNA silencing (Luna et al. 2017).
Considering that BNYVV and BSCTV co-infection of sugar beets occurs in most sugar beet growing fields of Iran and perhaps in other parts of the world, the present study was conducted to explore the possible interactions between these viruses and their effects on the resistance against BNYVV in the transgenic plants. We also questioned if the silencing against CP21 BNYVV could inhibit BSCTV-Ir propagation.
Materials and methods
Plant material
Based on our previous studies (Zare et al. 2015), a number of transgenic events carrying intron-hairpin RNA (ihpRNA) construct containing the 5' UTR with or without coding sequence of CP of BNYVV, called IHP-P (S3) and IHP-U (S6), respectively, were selected. Three T1 progenies of S3-12 and one of the S3-13.2 events were chosen named 227, 228, 229, and 219, respectively. Also, two T1 progenies of S6-2 and S6-44 events named 221 and 231 were selected. These events showed high resistance to BNYVV as assessed by ELISA analysis. A diploid monogram cultivar as a wild-type parental plant, named ‘9597’, and a cultivar called ‘Dorothea’ carrying the Rz1 gene, a Holly-based resistant plant, served as the negative and positive controls, respectively, which were kindly provided by Sugar Beet Seed Institute of Iran.
Micropropagation of transgenic plants
Transgenic plants were propagated through tissue culture to obtain a sufficient number of genetically identical individuals. The culture medium was composed of MS salts (Murashige and Skoog 1962) at pH 5.8 and supplemented with 3% (m/v) sucrose, 0.1 mg/l IBA, 1 mg/l BA, and 0.1 mg/l GA3. The root-inducing medium was MS containing 3 mg/l NAA hormone. Clonally propagated plants were transferred into the soil composed of peat, perlite, and vermiculite at a 1:1:1 ratio and adapted under the yellow–white fluorescent bulbs with 16 h of light photoperiod. The temperature was 25–30 °C and the humidity was adjusted to 40–60%.
Molecular analysis for transgenic plants
To select progenies carrying the transgene, a dot blot analysis was performed on virus-free transgenic plants. Genomic DNA was isolated from 50 mg of sugar beet leaves, according to Dellaporta et al. (1983). Genomic DNA (30 µg) was directly spotted on a positively charged nylon membrane (Roche Diagnostics, Germany) using a vacuum-assisted dot blotter tool (Gentaur BVBA, Belgium). Digoxigenin (DIG)-labeled probes were synthesized from the plasmids carrying each construct by PCR reaction using U+1 and U−1 primers (Table 1) and a DIG DNA labeling and detection kit (Roche Biochemical, Germany). The temperature for hybridization was 65 °C and the concentration of salt for the last wash was 0.1 mM NaCl in sodium citrate buffer. Detection was done by NBT/BCIP as instructed and the darkness of dots was inspected visually.
For genotyping of the progenies, DNA extraction of transgenic events was performed from the leaves using a GTP kit (Gene Transfer Pioneers, Iran). The presence of the transgene in each progeny was monitored by PCR amplification using gene-specific pairs of primers (Table 1). The reaction mixture contained 1 µl (50 ng) genomic DNA template, 2 pmol of each primer, 10 µl 2X PCR master mix (Thermo Fisher Scientific, USA), 2 mM MgCl2, 200 µM of each dNTPs, and 5 U Taq DNA polymerase (Cinagen, Iran) in a volume of 30 µl. Amplification cycles were as follows: denaturation cycle at 95 °C for 5 min, 40 cycles of 94 °C, 60 °C, and 72 °C (1 min each) with a final extension step at 72 °C for 10 min. The PCR products were separated on 1% agarose gels, stained with ethidium bromide, and visualized by UV light.
Viral challenges and bioassays
The micropropagated plants with 6–8 leaves were challenged with BNYVV or BSCTV-Ir viruses individually or both. Plants were transplanted into the mixture of BNYVV-infested and sterile soil at a 1:1 ratio. BSCTV-Ir infection was done through agro-injection of sugar beet plants with a full-length recombinant BSCTV-Ir construct (Ebadzad Sahraei et al. 2008) using Agrobacterium tumefaciens strain C58. To this end, the recombinant Agrobacterium was cultured in LB medium supplemented with Rifampicin and Kanamycin at 50 µg/ml and grown to OD600 1.0. The bacteria were pelleted and resuspended in MS medium supplemented with 2% (m/v) sucrose, 10 mM MgCl2, and 150 µM acetosyringone at pH 5.8 and diluted to OD600 0.5. After 3 h of incubation at room temperature, it was injected into the back of the leaves.
After 30 days of BSCTV-Ir infection, the presence of the virus was detected by PCR for an expected band of 761 bp using a pair of primers (Table 1). Total DNA was isolated from 50 mg of sugar beet leaves using the i-Genomic Plant DNA Extraction Mini Kit (Intron Biotechnology, South Korea). The PCR reaction mixture and program were carried out as above.
After 60 days of transfer to infested soil, BNYVV titers for each event were estimated from root samples using the enzyme-linked immunosorbent assay (ELISA) either by a DAS-ELISA kit (BIOREBA, Switzerland) based on the instructions provided by the manufacturer or according to Clark and Adams (1977) using an anti-CP21 antibody kindly provided by Dr. Izadpanah (Shiraz University, Iran). The antibody solution was added to the coating buffer containing 1.59 g Na2CO3, 0.2 g NaN3, and 2.93 g NaHCo3 (pH = 9.6) at 1:1000 dilution. The antibody mixture was added to the wells of the plate (Nunc, Thermo Scientific, US) and incubated at 37 °C for 3.5–4 h. 100 mg of each plant sample were homogenized in extraction buffer composed of 2% (m/v) polyvinylpyrrolidone (PVP) in phosphate-buffered saline (PBS; 137 mM NaCl, 2.7 mM KCl, 8 mM Na2HPO4, and 2 mM KH2PO4). After washing three times with washing buffer (PBS-Tween 20) and drying the plate, the plant extract (100 µl) was added to each well and incubated for 4 h. The plate was completely covered and incubated overnight at 4 °C. After a three-time washing step with the washing buffer, 100 µl of antibody-conjugate carrying Alkaline phosphatase enzyme diluted in conjugating buffer (1:1000) was added to wells of the plate. Then, the covered plate was incubated at 37 °C for 3.5–4 h after which 100 µl of the substrate solution composed of 10 mg of para-nitrophenylphosphate (pNPP) dissolved in 10 mL of 1X Diethanolamine substrate buffer was added to each well. Following overnight incubation of plate at room temperature in dark, the absorbance for each sample was measured at optical density (OD) value of 405 nm. The cut-off value was calculated by formula suggested by Bioreba (2014) which was the mean ± 3 times standard deviation for non-infected wild-type plants. If the absorbance was more than two times the cut-off value, the plant was considered to have high infection, whereas if it was lower than or equal to the cut-off, the plant was assumed healthy, and if between one and two cut-off values, the plant was designated as low infected.
To assure the infection process, P. betae spores were stained with acid fuchsin in lactophenol 0.05% (m/v) (Maneval 1936) and observed microscopically.
Statistical analyses of bioassays
Analysis of variance (ANOVA) for bioassay data was performed in a factorial experiment with a completely randomized design and three replications. The means comparisons were done by Duncan’s multiple range test (P < 0.05). All the statistical analyses were conducted with the use of SPSS software (IBM, USA).
Bioinformatics data analysis
To examine the possible similarity between the coat proteins of BNYVV (GenBank Accession No. AY277887) and BSCTV (GenBank Accession No. X97203), their nucleotide sequences were aligned pairwise with MegAlign software in Lasergene package (DNASTAR, USA).
Results
Following the previous studies (Zare et al. 2015; Safar et al. 2021), six T1 progenies of transgenic events with induced silencing against BNYVV CP21 were selected. As summarized in Table 2, the presence of the transgene and the expected effects on the selected events were verified using dot blot, PCR, and ELISA methods.
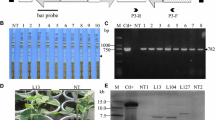
The compiled data for the not-infected or infected plants with BNYVV and BSCTV-Ir are summarized in Table 3. After agro-infection with recombinant BSCTV-Ir DNA constructs, S3 events showed mild curly leaves while severe symptoms were observed in S6 events, Dorothea, and wild-type ‘9597’ cultivar (Fig. 1). The same patterns of symptoms were also observed for the co-infected plants. Accordingly, lower levels of PCR products were detected in S3 plants using BSCTV-Ir primer pairs for the coat protein-encoding DNA sequence (Fig. 2, Tables 1 and 3).
Symptoms of BSCTV-Ir virus 30 days after agro-infection with recombinant viral DNA constructs on sugar beet plants comprise S3 events (a, b), S6 events (c), Dorothea (d), and wild-type ‘9597’ cultivar (e). S3 and S6 are transgenic events carrying 5’-UTR with or without full-length CP21-encoding sequences, respectively. 9597 and Dorothea served as the negative and positive controls, respectively
The level of BSCTV-Ir virus as detected by gel electrophoresis of PCR products in the infected sugar beet plants; 227–229 and 219 events as progenies of S3 events (lanes 1–4); 221 and 231 events as progenies of S6 events (lanes 5 and 6); Dorothea (lane 7); wild-type plant (lane 8). For abbreviations, see Fig. 1 legend
Since partial resistance was observed in some transgenic events infected with BSCTV-Ir, the possible sequence identity between the coding sequence of BNYVV and BSCTV-Ir coat proteins was inspected by pairwise alignment. As shown in Fig. 3, substantial sequence identities were observed in some regions between these nucleotide sequences.
The clonally propagated plants were challenged with BNYVV and BSCTV-Ir viruses, individually or together for 60 days. The BNYVV infection was confirmed as P. betae spores were detected in the roots of all examined plants by microscopic observations (Table 3). Based on the ELISA data, fourteen S3 plants were challenged with only BNYVV, almost all of them were found healthy or with low infection for the duration of the experiment. Among those plants co-infected with BNYVV and BSCTV-Ir, sixplants were healthy and four were slightly infected with BNYVV, while two plants showed high infection when they were exposed to both BNYVV and BSCTV-Ir. For the S6 construct, thirteen plants were either infected by BNYVV or co-infected by both BNYVV and BSCTV-Ir. Just one plant was highly infected to BNYVV when infected with BNYVV only. In all S6 plants, the co-infection of BNYVV and BSCTV-Ir did not affect the symptoms of the latter virus.
To overlook the positional effects of gene insertion and genotype variations, the means of the 6 selected transgenic events (4 events of S3 and 2 from S6) were compared (Fig. 4). In all plants, except S3-229, no significant difference was observed between BNYVV single infection or co-infection with BSCTV-Ir. In S3-229 case, the BNYVV accumulation was significantly higher in co-infection treatments than the single infections. The wild-type cultivar also showed higher BNYVV titers under co-infection conditions, although it was not significant.
Accumulation of BNYVV CP21 in the examined plants. Clonally propagated plants at the 6–8 leaf stage were infected with BNYVV alone (black bars) or both BNYVV & BSCTV-Ir (white bars). For abbreviations, see Fig. 1 legend. The titer of CP21 was assayed by ELISA method with three biological and two technical replicates. Different letters represent significant differences among the plants at P < 0.05
Discussion
In order to explore the possible effects of viral co-infection on the efficiency of RNA silencing-mediated resistance, transgenic events were exposed to BNYVV or BSCTV-Ir individually and together. Almost all transgenic events were resistant to single infections of BNYVV. Consistent with our previous studies (Zare et al. 2015; Safar et al. 2021), the reduced propagation of BNYVV in transgenic events indicates the effectiveness of the CP21-based inserts in inducing resistance against Rhizomania. Similarly, other researchers have already shown that the introduction of BNYVV-based constructs can be an effective way to control Rhizomania disease (Mannerlöf et al. 1996; Lennefors et al. 2006, 2008). Considerable resistance against Rhizomania disease was achieved through the expression of dsRNA of BNYVV replicase gene sequence in the transformed sugar beet plants (Pavli et al. 2010). Similarly, transgenic Nicotiana benthamiana plants encoding CP readthrough protein exhibited high resistance to BNYVV (Andika et al. 2005).
As expected, all the transgenic plants carrying S6 constructs produced severe curly top symptoms, when subjected to BSCTV-Ir. However, S3 events with IHP-P construct moderately resisted BSCTV-Ir compared to control and S6 events carrying IHP-U. The inhibition of propagation of a particular virus in transgenic plants containing the insert derived from another virus is commonly referred to as “heterologous resistance” (Dinant et al. 1993). So far, several cases of heterologous resistance in different transgenic plants have been reported (Dinant et al. 1993; Hassairi et al. 1998; Peng et al. 2014; Ali et al. 2019). Medina-Hernández et al. (2013) evaluated the efficiency of Tomato Chino La Paz virus (ToChLPV)-derived construct for resistance against Pepper Golden Mosaic virus (PepGMV) in N. benthamiana plants. It was shown that the severity of PepGMV symptoms was reduced to 45% in transgenic plants. As shown in Fig. 3, there are several regions of sequence identities between genes coding for BNYVV and BSCTV-Ir coat proteins. Therefore, the slight resistance to BSCTV-Ir observed in S3 events could be due to the presence of some BNYVV CP-derived siRNAs.
From the other point of view, almost all transgenic events showed stable resistance to BNYVV compared to wild-type plants under co-infection conditions. Yet, the high titer of BNYVV in 3 out of 28 S3 progenies (Table 3) needs further investigations. It might be due to either interaction of a suppressor protein encoded by the BSCTV-Ir virus, rearrangement of the transgene, or co-infection with other soil-borne viruses. Regardless of this exception, co-infection with BNYVV and BCTV-Ir does not appear to affect the RNA-silencing-based resistance of transgenic sugar beets in general. Similar to our findings, co-infection with heterologous viruses does not always suppress the resistance of transgenic plants as shown by other researchers (Vassilakos 2012). For instance, the co-infection Plum pox virus (PPV) with either Apple chlorotic leaf spot virus (ACLSV) or Prune dwarf virus (PDV) did not suppress RNA silencing against PPV coat protein gene in transgenic plum (Prunus domestica L.) (Singh et al. 2019). The RNA silencing-mediated resistance against BNYVV was not affected by co-infection with either Beet Soil-Borne virus, Beet Virus Q, Beet Mild Yellowing virus or Beet Yellows virus (BYV) in transgenic sugar beets (Lennefors et al. 2008).
In summary, the presented results indicate that RNA-silencing against BNYVV CP21 is highly efficient in hindering BNYVV propagation and provide a powerful mean for breeding programs for the control of Rhizomania disease in sugar beet. Moreover, the siRNAs generated in the S3 transgenic plants can be effective in inducing heterologous resistance to other sugar beet viruses like BSCTV-Ir. It was also demonstrated that the co-infection of BSCTV-Ir with BNYVV does not affect the efficiency of inducing silencing by the constructs producing RNA with hair-pin structures. Overall, the induced RNA silencing-based resistance was stable in transgenic plants under both single and multiple infection conditions and, therefore, it can be a suitable alternative for the conventional breeding cultivars with BNYVV resistance.
References
Ali I, Khurshid M, Iqbal Z, Shafiq M, Amin I, Mansoor S, Briddon R (2019) The antisense 5’end of the V2 gene confers enhanced resistance against the monopartite begomovirus cotton leaf curl Kokhran virus-Burewala strain. Acta Virol 63(1):26–35. https://doi.org/10.4149/av_2019_101
Anand A, Pinninti M, Tripathi A, Mangrauthia SK, Sanan-Mishra N (2022) Coordinated action of RTBV and RTSV proteins suppress host RNA silencing machinery. Microorganisms 10(2):197. https://doi.org/10.3390/microorganisms10020197
Andika IB, Kondo H, Tamada T (2005) Evidence that RNA silencing-mediated resistance to Beet necrotic yellow vein virus is less effective in roots than in leaves. Mol Plant Microbe Interact 18(3):194–204. https://doi.org/10.1094/MPMI-18-0194
Aulia A, Andika IB, Kondo H, Hillman BI, Suzuki N (2019) A symptomless hypovirus, CHV4, facilitates stable infection of the chestnut blight fungus by a coinfecting reovirus likely through suppression of antiviral RNA silencing. Virology 533:99–107. https://doi.org/10.1016/j.virol.2019.05.004
Biancardi E, Lewellen RT (2016) Introduction. In: Biancardi E, Tamada T (eds) Rhizomania. Springer, Cham, pp 3–28. https://doi.org/10.1007/978-3-319-30678-0_1
Bioreba (2014) ELISA data analysis. Bioreba AG. https://www.bioreba.ch/catalog/Haupt-Navigation-ELISA.aspx. Accessed 20 July 2022
Capistrano-Gossmann GG, Ries D, Holtgräwe D, Minoche A, Kraft T, Frerichmann SL, Rosleff Soerensen T, Dohm JC, González I, Schilhabel M (2017) Crop wild relative populations of Beta vulgaris allow direct mapping of agronomically important genes. Nat Commun 8(1):15708. https://doi.org/10.1038/ncomms15708
Clark MF, Adams A (1977) Characteristics of the microplate method of enzyme-linked immunosorbent assay for the detection of plant viruses. J Gen Virol 34(3):475–483. https://doi.org/10.1099/0022-1317-34-3-475
Dellaporta SL, Wood J, Hicks JB (1983) A plant DNA minipreparation: version II. Plant Mol Biol Rep 1(4):19–21. https://doi.org/10.1007/BF02712670
Dhir S, Srivastava A, Yoshikawa N, Khurana S (2019) Plant viruses as virus induced gene silencing (VIGS) vectors. In: Khurana SMP, Gaur RK (eds) Plant biotechnology: progress in genomic era. Springer, Singapore, pp 517–526. https://doi.org/10.1007/978-981-13-8499-8_22
Dinant S, Blaise F, Kusiak C, Astier-Manifacier S, Albouy J (1993) Heterologous resistance to potato virus Y in transgenic tobacco plants expressing the coat protein gene of lettuce mosaic potyvirus. Phytopathology 83(8):818–824. https://doi.org/10.1094/Phyto-83-818
Duan C-G, Wang C-H, Guo H-S (2012) Application of RNA silencing to plant disease resistance. Silence 3:5. https://doi.org/10.1186/1758-907X-3-5
Ebadzad Sahraei G, Behjatnia SAA, Izadpanah K (2008) Infectivity of the cloned genome of Iranian isolate of Beet severe curly top virus in experimental hosts. Iran J Plant Pathol 44(2):176–183
Funk A, Galewski P, McGrath JM (2018) Nucleotide-binding resistance gene signatures in sugar beet, insights from a new reference genome. Plant J 95(4):659–671. https://doi.org/10.1111/tpj.13977
Hassairi A, Masmoudi K, Albouy J, Robaglia C, Jullien M, Ellouz R (1998) Transformation of two potato cultivars ‘Spunta’and ‘Claustar’(Solanum tuberosum) with lettuce mosaic virus coat protein gene and heterologous immunity to potato virus Y. Plant Sci 136(1):31–42. https://doi.org/10.1016/S0168-9452(98)00078-8
Jiang L, Du Z, Zhang G, Wang T, Jin G (2022) Advances in RNA-silencing-related resistance against viruses in potato. Genes 13(5):731. https://doi.org/10.3390/genes13050731
Jin F, Song J, Xue J, Sun H, Zhang Y, Wang S, Wang Y (2020) Successful generation of anti-ToCV and TYLCV transgenic tomato plants by RNAi. Biol Plant 64:490–496. https://doi.org/10.32615/bp.2020.069
Kutluk Yilmaz ND, Uzunbacak H, Arli-Sokmen M, Kaya R (2018) Distribution of resistance-breaking isolates of beet necrotic yellow vein virus differing in virulence in sugar beet fields in Turkey. Acta Agricult Scand Sect B Soil Plant Sci 68(6):546–554. https://doi.org/10.1080/09064710.2018.1441432
Lennefors B-L, Savenkov EI, Bensefelt J, Wremerth-Weich E, van Roggen P, Tuvesson S, Valkonen J, Gielen J (2006) dsRNA-mediated resistance to beet necrotic yellow vein virus infections in sugar beet (Beta vulgaris L. ssp. vulgaris). Mol Breed 18(4):313–325. https://doi.org/10.1007/s11032-006-9030-5
Lennefors B-L, van Roggen PM, Yndgaard F, Savenkov EI, Valkonen J (2008) Efficient dsRNA-mediated transgenic resistance to Beet necrotic yellow vein virus in sugar beets is not affected by other soilborne and aphid-transmitted viruses. Transgenic Res 17(2):219–228. https://doi.org/10.1007/s11248-007-9092-0
Li S, Zhang T, Zhu Y, Zhou G (2017) Co-infection of two reoviruses increases both viruses accumulation in rice by up-regulating of viroplasm components and movement proteins bilaterally and RNA silencing suppressor unilaterally. Virol J 14(1):1–8. https://doi.org/10.1186/s12985-017-0819-0
Liang Q-l, Wei L-x, Xu B-l, Calderón-Urrea A, Xiang D (2017) Study of viruses co-infecting white clover (Trifolium repens) in China. J Integr Agric 16(9):1990–1998. https://doi.org/10.1016/S2095-3119(16)61606-4
Lu R, Martin-Hernandez AM, Peart JR, Malcuit I, Baulcombe DC (2003) Virus-induced gene silencing in plants. Methods 30(4):296–303. https://doi.org/10.1016/s1046-2023(03)00037-9
Luna AP, Rodríguez-Negrete EA, Morilla G, Wang L, Lozano-Durán R, Castillo AG, Bejarano ER (2017) V2 from a curtovirus is a suppressor of post-transcriptional gene silencing. J Gen Virol 98(10):2607–2614. https://doi.org/10.1099/jgv.0.000933
Maneval W (1936) Lacto-phenol preparations. Stain Technol 11:9–11. https://doi.org/10.3109/10520293609111316
Mannerlöf M, Lennerfors B-L, Tenning P (1996) Reduced titer of BNYVV in transgenic sugar beets expressing the BNYVV coat protein. Euphytica 90(3):293–299. https://doi.org/10.1007/BF00027479
Mascia T, Gallitelli D (2016) Synergies and antagonisms in virus interactions. Plant Sci 252:176–192. https://doi.org/10.1016/j.plantsci.2016.07.015
McGrann GR, Grimmer MK, Mutasa-Göttgens ES, Stevens M (2009) Progress towards the understanding and control of sugar beet rhizomania disease. Mol Plant Pathol 10(1):129–141. https://doi.org/10.1111/j.1364-3703.2008.00514.x
Medina-Hernández D, Rivera-Bustamante RF, Tenllado F, Holguín-Peña RJ (2013) Effects and effectiveness of two RNAi constructs for resistance to Pepper golden mosaic virus in Nicotiana benthamiana plants. Viruses 5(12):2931–2945. https://doi.org/10.3390/v5122931
Moreno AB, López-Moya JJ (2020) When viruses play team sports: mixed infections in plants. Phytopathology 110(1):29–48. https://doi.org/10.1094/PHYTO-07-19-0250-FI
Murashige T, Skoog F (1962) A revised medium for rapid growth and bioassays with tobacco tissue cultures. Physiol Plant 15(3):473–497. https://doi.org/10.1111/j.1399-3054.1962.tb08052.x
Palukaitis P (2011) The road to RNA silencing is paved with plant-virus interactions. Plant Pathol J 27(3):197–206. https://doi.org/10.5423/PPJ.2011.27.3.197
Pavli OI, Panopoulos NJ, Goldbach R, Skaracis GN (2010) BNYVV-derived dsRNA confers resistance to rhizomania disease of sugar beet as evidenced by a novel transgenic hairy root approach. Transgenic Res 19(5):915–922. https://doi.org/10.1007/s11248-010-9364-y
Pavli OI, Stevanato P, Biancardi E, Skaracis GN (2011) Achievements and prospects in breeding for rhizomania resistance in sugar beet. Field Crop Res 122(3):165–172. https://doi.org/10.1016/j.fcr.2011.03.019
Peng J-C, Chen T-C, Raja JA, Yang C-F, Chien W-C, Lin C-H, Liu F-L, Wu H-W, Yeh S-D (2014) Broad-spectrum transgenic resistance against distinct tospovirus species at the genus level. PLoS ONE 9(5):e96073. https://doi.org/10.1371/journal.pone.0096073
Pferdmenges F (2007) Occurrence, spread and pathogenicity of different Beet necrotic yellow vein virus (BNYVV) isolates. Cuvillier Verlag, Göttingen
Pferdmenges F, Varrelmann M (2009) Breaking of Beet necrotic yellow vein virus resistance in sugar beet is independent of virus and vector inoculum densities. Eur J Plant Pathol 124(2):231–245. https://doi.org/10.1007/s10658-008-9408-9
Safar S, Bazrafshan M, Khoshnami M, Behrooz AA, Hedayati F, Maleki M, Mahmoudi SB, Malboobi MA (2021) Field evaluation for rhizomania resistance of transgenic sugar beet events based on gene silencing. Can J Plant Path 43(1):179–188. https://doi.org/10.1080/07060661.2020.1783575
Singh K, Neubauerová T, Kundu JK (2019) Quantitative analysis of the interaction of heterologous viruses with Plum pox virus in C5 HoneySweet transgenic plums. J Integr Agric 18(10):2302–2310. https://doi.org/10.1016/S2095-3119(18)62136-7
Stevanato P, Chiodi C, Broccanello C, Concheri G, Biancardi E, Pavli O, Skaracis G (2019) Sustainability of the sugar beet crop. Sugar Tech 21(5):703–716. https://doi.org/10.1007/s12355-019-00734-9
Susi H, Barrès B, Vale PF, Laine A-L (2015) Co-infection alters population dynamics of infectious disease. Nat Commun 6:5975. https://doi.org/10.1038/ncomms6975
Syller J (2014) Biological and molecular events associated with simultaneous transmission of plant viruses by invertebrate and fungal vectors. Mol Plant Pathol 15(4):417–426. https://doi.org/10.1111/mpp.12101
Syller J, Grupa A (2016) Antagonistic within-host interactions between plant viruses: molecular basis and impact on viral and host fitness. Mol Plant Pathol 17(5):769–782. https://doi.org/10.1111/mpp.12322
Uslu VV, Wassenegger M (2020) Critical view on RNA silencing-mediated virus resistance using exogenously applied RNA. Curr Opin Virol 42:18–24. https://doi.org/10.1016/j.coviro.2020.03.004
Vassilakos N (2012) Stability of transgenic resistance against plant viruses. In: Çiftçi YO (ed) Transgenic plants: advances and limitations. IntechOpen, London, pp 219–236. https://doi.org/10.5772/33133
Yang X, Baliji S, Buchmann RC, Wang H, Lindbo JA, Sunter G, Bisaro DM (2007) Functional modulation of the geminivirus AL2 transcription factor and silencing suppressor by self-interaction. J Virol 81(21):11972–11981. https://doi.org/10.1128/JVI.00617-07
Zare B, Niazi A, Sattari R, Aghelpasand H, Zamani K, Sabet M, Moshiri F, Darabie S, Daneshvar M, Norouzi P, Kazemi-Tabar SK, Khoshnami M, Malboobi MA (2015) Resistance against rhizomania disease via RNA silencing in sugar beet. Plant Pathol 64(1):35–42. https://doi.org/10.1111/ppa.12239
Acknowledgements
This research was supported and funded by the National Institute of Genetic Engineering and Biotechnology (NIGEB) of Iran (Grant Nos. 167 and 102M) and Green Transgene Technology Development Company (Tehran, Iran).
Author information
Authors and Affiliations
Contributions
MK, MAM, and FR designed the research. MK, BZ, and HMM performed the experiment. MK and MAB analyzed the data and wrote the manuscript. MAM and MAB reviewed and edited the manuscript. All authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Conflict of interest
The authors have no relevant financial or non-financial interests to disclose.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Khoshnami, M., Zare, B., Mardani-Mehrabad, H. et al. Assessment of co-infection with BNYVV and BSCTV on resistance against Rhizomania disease in transgenic sugar beet plants. Transgenic Res 32, 475–485 (2023). https://doi.org/10.1007/s11248-023-00364-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11248-023-00364-8