Abstract
Genetic engineering can add new capabilities or traits and direct method using biolistic particle delivery holds key for rapid, routine and efficient transformation of chickpea. Regeneration efficiencies of five different explants derived from BAP pre-treated breeders’ chickpea seeds (cv. DCP 92-2) raised in phytohormones combinations (BAP and KIN for shoot primordia induction; GA3 for shoot elongation and NAA for rooting) were compared. Best response was obtained using the embryonic axis (EAX) explants with 86.69% regeneration efficiency followed by epicotyl (EPI) explants (78.69%). Direct genetic transformation were demonstrated in two responding explants by bombarding with pre-treated tungsten, coated with plant expression cassette (harboring Bt and nptII gene) from a distance of 4 cm with 1100 psi helium pressure. Transgenic chickpea lines with multiple and single copy integrations were obtained with transformation frequency of 0.72% for EPI explants and 1.21% for EAX explants, significantly higher than Agrobacterium tumefaciens mediated transformation of same genotype (0.076%). Southern blot based analyses of seven single copy transgenic chickpea lines exhibited presence and transmission to subsequent generations (T1 and T2). Presence of Bt protein were detected in the leaves of transgenic chickpea lines at pre-flowering (6.63–11.95 ng/mg TSP) and post flowering stages (4.85–8.93 ng/mg TSP). Genetic fidelity analysis using genome wide SSR markers of ten independent transgenic lines indicated true to type with original genotype. Taken together, this study describes a protocol that can be adapted for direct genetic transformation of chickpea with high efficiency.
Key message
Routine protocol for direct transformation of grain legume, chickpea.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Chickpea (Cicer arietinum L.) is a climate-smart grain legume of Leguminosae family grown in over 50 countries worldwide. It is globally cultivated on 13.7 mha land accounting for an average annual production of 14.25 m t. India is among the top producer of chickpea accounting for the around 70% of global production (9.9 m t) (FAO 2019). Genetic diversity is the key to genetic improvement; however, the introduction and adaptation of modern high-yielding genetically uniform varieties has gradually resulted in loss of genetic diversity. Crop domestication and post-domestication diversification, together with breeding efforts to meet specific human needs contributed to narrow genetic diversity in the cultivated gene pool of chickpea. Recent reports indicated fourfold reduction in genetic diversity (calculated based on diversity indices π and ω) in chickpea landraces (θπ = 0.86; θω = 0.87) and elite cultivars (θπ = 0.74; θω = 0.74) compared to the wild species (θπ = 3.80; θω = 2.79) (Varshney et al. 2019). Genetic engineering is a novel tool that has the potential for introducing new traits or capabilities in plants. Till date, various traits including insect resistance (Kar et al. 1997; Sanyal et al. 2005; Indurker et al. 2007; Acharjee et al. 2010; Indurker et al. 2010; Mehrotra et al. 2011; Asharani et al. 2011; Ganguly et al. 2014; Khatodia et al. 2014; Chakraborty et al. 2016; Das et al. 2017), and enhanced drought tolerance (Anbazhagan et al. 2015; Das et al. 2021) have been introduced in chickpea employing genetic engineering approaches.
Genetic transformation in chickpea has been reported employing both direct transformation method (DT) (particle delivery via. biolistic method) and indirect transformation method (Agrobacterium tumefaciens/Agrobacterium rhizogenes mediated transformation (ATMT/ARMT) and modification thereof like sonication-assisted Agrobacterium tumefaciens—mediated transformation (SAAT). Although, majority of chickpea transformation reports have employed ATMT, higher transformation efficiency are reported with DT method. On an average basis, transformation efficiency of Agrobacterium-mediated transformation is lower than particle bombardment (Gao and Nielsen 2013). Besides, DT allows overcoming transformation barriers like recalcitrance, including low frequency of response in tissue culture conditions, and amenability for co-transformation strategy and clean gene technology. First report of successful direct transformation in chickpea using EAX explants has been reported with co-transformation frequency of 45.8% (Kar et al. 1997). In another report, hypocotyl (HYP) explant was employed for chickpea transformation using both gus and nptII genes (Husnain et al. 2000). Direct transformation of chickpea using gus gene was reported in four different explants viz. mature zygotic EAX, cotyledonary node, shoot tip and leaf. Comparative efficiencies were also calculated based on transient gus expression in tested explants and gold/tungsten particle as carriers of recombinant DNA (Sanyal et al. 2003). The successful report of use of non-antibiotic AK (aspartate kinase)/LT (lysine threonine) selection system in chickpea was demonstrated using decapitated embryo explants bombarded with desensitized aspartate kinase (AK) gene coated on tungsten particles (Tewari-Singh et al. 2004). The first report of optimized parameters crucial for particle bombardment in chickpea was demonstrated with EPIs explants using gold particles as micro-carriers in combination with helium pressure of 900 psi, resulting in higher transformation frequency (18.0%) (Indurker et al. 2007). Recently direct transformation of leaf epidermal cells of chickpea for transient expression of CAP2 was reported using Helios gene gun with gold particles, helium pressure of 120 psi and distance of 2 cm from tissue (Jain and Chattopadhyay 2013).
In vitro regeneration protocols in chickpea are mostly genotype dependent and affected by various phytohormones. Most of the regeneration studies were reported either using multiple shoot induction from explants (Kar et al. 1997; Indurker et al. 2007; Ganguly et al. 2014; Das et al. 2017) or somatic embryogenesis (Sagare et al. 1993, Shukla et al. 2015). Diverse explants sub-cultured on various combination of phytohormone (cytokinins: auxins) have been reported for chickpea regeneration. Due to high responsiveness to in vitro regeneration, EAX has been the choice of explants for chickpea regeneration and transformation experiments (Das et al. 2020). Various explants employed for chickpea regeneration and transformation studies include EPIs (Indurker et al. 2007), HYPs (Husnain et al. 2000; Aggarwal et al. 2018), plumule (PLU) (Senthil et al. 2004), decapitated embryo with cotyledon (Tewari-Singh et al. 2004), stem (Indurker et al. 2007, 2010), dissected cotyledon with half embryo (Das and Sarmah 2005; Chakraborti et al. 2009) and axillary meristem explants (Jayanand et al. 2003; Das et al. 2017). Despite several successful reports on chickpea regeneration and transformation, an efficient and stable direct genetic transformation technique for transferring agronomically useful trait into chickpea genotype is necessary perquisite.
Here, we report regeneration of five different explants originating from mature chickpea seed using various combinations of phytohormones cytokinin, auxins and gibberellins and its establishment in Containment Facility. We also demonstrated the amenability of two most responding explants for direct transformation (particle gun based) of chickpea using insecticidal Bt-gene. Integration and expression of the transgene in the progenies derived from developed transgenic lines were also demonstrated. Genetic fidelity of the primary transformants was also established using SSR markers spanning all eight linkage groups of chickpea. This is a maiden report on use of five explants and several media combination to develop a routine direct transformation protocol in chickpea.
Materials and methods
Plant material and explants preparation
Breeders’ seeds of the desi chickpea cultivar, DCP 92-3 employed in the regeneration and transformation experiments were obtained from ICAR-Indian Institute of Pulses Research (IIPR), Kanpur. Procured seeds were washed with 0.1% (v/v) Tween 20 (Himedia Laboratories Pvt. Ltd, Mumbai) and surface sterilized using 70% of ethanol (Himedia Laboratories Pvt. Ltd, Mumbai) for 10 min and 1.0% (v/v) of sodium hypochlorite (Himedia Laboratories Pvt. Ltd, Mumbai) for 5–10 min. Sterilized seeds were intermittently washed thrice with sterile distilled water to completely remove traces of the sterilizing agents on the seed’s surface. Sterilized seeds were kept for overnight imbibition and subsequently seed coats were carefully removed and the obtained intact cotyledonary seeds without the seed coat were inoculated on modified MS medium (Murashige and Skoog 1962) supplemented with 0.5 mg/L BAP, 30 mg/L sucrose, B5 vitamins, and 0.8% agar (pH 5.8) (Himedia Laboratories Pvt. Ltd, Mumbai). Post inoculation, seeds were incubated in a growth chamber for 2 days. During all regeneration studies, growth chamber was maintained at 22 ± 3 °C with 16 h of photoperiod illumination [35–40 µmol m−2 s−1 supplied from cool white fluorescent (Philips lighting, India)] and 8 h dark period.
For comparing the regeneration potential, five types of explants were prepared from 48 h pre-germinated chickpea seeds: Axillary meristems (AMS), EAX, PLU, EPI and HYP. The intact embryonic axes were separated from the cotyledons and were used as explants. Similarly, separate PLU explants were prepared by removing the radicle portion from the isolated EAX. The EPI explant was derived from the shoot portion after the removal of apical meristem, while the HYP explants were prepared from the root portion of the EAX, lying between the EPI and radical portion. AMS were prepared from germinated seedlings after removing axillary buds up to the base and cuts were made to remove the PLU and radicle tips (Supplementary Fig. 1).
The entire regeneration experiment was planned into four distinct phases of shoot induction, shoot elongation, rooting and establishment. Unless otherwise stated, MS media supplemented with B5 vitamins were used as the basal media for all the shoot organogenesis-based regeneration experiments. All the explants-based regeneration experiment was set in RBD with three replications per treatment. Collected variable, were summarized and analyzed in t-Test: Paired Two Sample for Means, using Microsoft excel 2007 (MS office, 2007), comparative analysis were conducted for the significant results using LSD at 0.05 probabilities. Different media combination and explants tested employed are described below:
Shoot induction media (SIM)
A total of four different media comprising of four different phyto-hormones [cytokinins: BAP, KIN, TDZ and 2iP (Duchefa Biochemie, Netherlands)] singly or in combination were assessed for induction of shoot organogenesis pathway in all five different explants. Following were the media combinations tested in the current study: SIM 1 (Modified MS salts + 0.88 mg/L TDZ + 2.03 mg/L 2iP + 0.43 mg/L KIN), SIM 2 (Modified MS salts + 2.20 mg/L TDZ) and SIM 3 (Modified MS salts + 0.5 mg/L BAP + 0.1 mg/L KIN) and Control (modified MS salts without phytohormones). Responsiveness of the explants in the tested media combinations were assessed for number of shoots primordia after 2 weeks of inoculation.
Shoot elongation media (SEM)
All responding multiple shoot primordia were assessed for elongation using four phytohormones viz. TDZ, 2iP, KIN, GA3 (Duchefa Biochemie, Netherlands) singly or in combination for shoot elongation by shoot organogenetic pathway. The different shoot elongation media assessed were: SEM 1 (Modified MS salts + 0.55 mg/L TDZ), SEM 2 (Modified MS salts + 1.02 mg/L 2iP + 0.43 mg/L KIN), SEM 3 (Modified MS salts + 0.73 mg/L GA3) and control (Modified MS salts without phytohormones). Responsiveness of the explants in the tested media combinations were assessed for shoot length after 2 weeks of inoculation in SEM.
Rooting media (RM) and plant establishment
For the assessment of root induction, a total of four phytohormones viz. IAA, KIN, IBA and NAA (Duchefa Biochemie, Netherlands) and inorganic additive (KNO3) were used singly or in combination. Composition of three different rooting media: RM 1 (Modified MS salts + 0.5 mg/L IAA + 0.5 mg/L KIN), RM 2 (Modified MS salts + 949.4 mg/L KNO3 + 1.015 mg/L IBA), RM 3 (Modified MS salts + 0.465 mg/L NAA) and a control (Modified MS media without phytohormones). Healthy green shoots of 3–4 cm in length were separated out from mother clump and inoculated on three media and the rooting efficiencies were compared in terms of average number and average length of roots. The rooted healthy shoots were transferred to soil pots (size: 9–10 cm diameter) containing soil, vermicompost and coco-peat in equal proportions at Transgenic Containment Facility (Plant Bio-safety Level-1), IIPR, Kanpur. Plantlets exhibiting well developed rooting system are carefully removed from the rooting media and transplanted at a depth of 1.5 to 2 inches in the pot containing hardening mixture. Transplanted plantlets were covered with transparent polyethylene bag for a week, for acclimatization. Post acclimatization, polyethylene bags were gradually removed from the established plantlets. Regeneration efficiency of five tested explants (75 nos. each) were calculated based on the number of rooted explants that could be successfully established to maturity using the formula: Regeneration efficiency (%) = [No. of plantlets successfully established till maturity/Total No. of shoot regenerated on shooting media] × 100.
Plant expression vector and particle gun bombardment
For the genetic transformation experiment, modified plant expression vector harboring two foreign genes viz. synthetic Bt gene and plant selectable marker gene, neomycin phosphotransferase II (nptII) was employed (Supplementary Fig. 2). Two highly responsive explants viz. EPI and EAX were evaluated in the direct transformation experiments for development of transgenic chickpea lines. For direct transformation 30 mg of tungsten microcarrier (1.1 μm diameter) (Bio-Rad Laboratories, USA) were surface sterilized using 70% ethanol followed by intensive washing with TE buffer (pH-9.5). Washed microcarriers were suspended in 500 μl of sterile 50% glycerol making the final concentration 60 µg/ml and aliquots of 50 µl were prepared. For coating microcarrier with the modified plant expression vector, following mixture composition was used: 5 µl of plasmid DNA (1 µg/µl), 50 µl of CaCl2 (2.5 M) and 20 µl of (0.1 M) spermidine in TE buffer (pH-9.5) followed by repeated washing with 100% ethanol. Plant expression vector coated microcarrier was suspended in 50 µl of absolute ethanol and 10 µl aliquot of the suspension was spread to dry on sterile macro-carrier disk. The macro-carrier carrying coated micro-carrier was installed in the Biolistic® PDS-1000/He particle delivery system (Bio-Rad Laboratories, USA) assembly as per manufacturer’s instructions for the transformation experiment. All transformation experiments were carried out at helium pressure 1100 psi and the distance between the macro-carrier (containing the coated micro-carriers) and target tissue was chosen to be 4 cm. For efficient particle delivery explants (EAX and EPI) were arranged in a circular pattern at the center of petri-dish containing MS basal media. After 2 days, bombarded explants (EAX and EPI) were transferred to pre-standardized SIM 3 and incubated in the growth chamber having light regime of 16/8-h light and dark at 25 ± 2 °C for 2 weeks. After 2 weeks, explants exhibiting shoot induction were selected and healthy shoot primordia were separated from the mother clump and were transferred to pre-standardized SEM 3 containing 100 mg/L kanamycin (Duchefa Biochemie, Netherlands). After 2 to 3 rounds of antibiotic selection cum regeneration, the elongated shoots were separated and sub-cultured on RM 3 rooting media for root development and establishment. The rooted healthy shoots were transferred to clay pots (size: 9–10 cm diameter) containing matrix composed of soil, vermicompost and cocopeat in equal proportions for hardening and plantlet acclimatization and establishment to mature fertile plants.
Seeds were harvested from mature fertile chickpea plants in Transgenic Containment Facility. The selfed seeds were sown in Containment Facility and molecular analyses were conducted in the T1 and T2 generation. Details of seeds harvested in each generation were documented and deposited in seed repository for seven selected lines, originating from two different explants tested.
Molecular characterization of transgenic chickpea events
Genomic DNA from the leaves derived from transformed (T1 and T2 progenies) and control (DCP 92-3) chickpea were extracted using DNeasy Plant Kit, (Qiagen, Germany). The presence of Bt and nptII gene in the progenies of individual transgenic lines were confirmed using gene specific primers [Primers A4F: 5′-CCTTGTACAGAAGACCCTTCAATATC-3′ (forward) and A4R: 5′-TCTATTCTGAATGTTATTTCCACTGC-3′ (reverse) for Bt gene and NPTF: 5′-ATGACGCGGGACAAGCCGTT-3′ (forward) and NPTR: 5′-CGCGAGCCCCTGATGCTCTT-3′ (reverse) for nptII gene]. The PCR mixture contained 2.5 µl of 10X Taq buffer, 1.0 µl 2.5 mM dNTP (Thermo Scientific, USA), 200 ng DNA template, 1U Taq DNA polymerase (New England BioLabs Inc., USA) and 1 µl of 10 pM forward and reverse primers in a final volume of 25 µl. PCR for both the primer sets were carried out using following thermal profile: Initial de-naturation at 95 °C for 2 min followed by 35 cycles of denaturation at 95 °C for 30 s; Annealing at 60 °C for 1 min and elongation 72 °C for 1 min after the end of cycle; final elongation was done at 72 °C for 9 min. PCR amplified products were electrophoresed on 1.0% agarose gel containing 0.4 µg/ml ethidium bromide and visualized on Gel documentation system (Gel Doc XR, BioRad Laboratories, USA).
Southern blot hybridization was carried out using extracted genomic DNA from the leaves of PCR positive transgenic chickpea lines and DCP 92-3 (control) plants, as described earlier (Das et al. 2017). 25.0 µg of isolated DNA was digested with HindIII restriction endonuclease (New England BioLabs Inc., USA), having unique restriction site near the terminator of Bt gene and in a separate reaction same quantity of genomic DNA was double digested with HindIII and EcoRI (New England BioLabs Inc., USA) to release the entire Bt expression cassette. The digested products were separately resolved by electrophoresis on 0.8% (w/v) agarose gel. The digested DNA was subsequently transferred to a positively charged nylon membrane (Hybond-N+; Amersham™ Biosciences, UK). A 441 bps Bt specific amplification was digoxygenin (DIG) labeled using DIG labeled dNTPs mix (Roche Diagnostics, Germany) under same set of PCR thermal profile described earlier and used as a probe. Hybridization of the DIG labeled probe was detected by chemiluminescent substrate, CDP-Star® (disodium 2-chloro-5-(4-methoxyspiro {1,2-dioxetane-3,2′-(5′-chloro)tricyclo[3.3.1.1]decan}-4-yl)-1-phenyl phosphate), as per the manufacturer’s instruction (Roche Diagnostics, Germany). Signal was detected by chemiluminiscent documentation imager (Omega Lum™ G-Aplegen, USA) after an X-ray exposure of 15 min.
The transmission of transgene(s) was confirmed based on PCR analysis of T2 progenies. Transformation efficiency was estimated based on the number of PCR positive chickpea progenies (T1 progenies derived from individual T0), derived from original number of explants bombarded using the formula: Transformation Efficiency (%) = (Number of PCR positive chickpea lines/Total number of explants bombarded) × 100. Segregation ratios of seven single locus transgene integration lines were calculated based on standard chi-square (χ2) test.
Quantitative estimation of Bt protein expression was determined using ELISA kit as described earlier (Das et al. 2017). Total protein was estimated using the Bradford Assay as per manufacturer’s instructions (Sigma Aldrich, USA). Detection and estimation of Bt protein was done using Quantitative ELISA kit as per manufacturer’s instructions (QuantiPlate Kit for Cry1Ab/Cry1Ac, EnviroLogix Inc., USA) and the absorbance readings were recorded at 450 nm in ELISA reader (Multiskan EX, Thermo Scientific, USA). Temporal expression of Bt protein was quantified in the pre-flowering stage [65 days of sowing (DAS)] and post flowering stage [115 DAS] in the leaves samples of selected chickpea events at T1 and T2 stage in terms of ng/mg total soluble protein (TSP).
For western blotting, TSP was extracted from young leaves of PCR screened positive transgenic chickpea progenies (T2 stage) and the control (DCP 92-3), 65 days and 115 DAS. Based on standard Bradford assay, 30.0 µg of total protein from individual lines was separated on 10% SDS-PAGE and blotted on to nitrocellulose membrane (BioRad Laboratories, USA) by wet transfer. After blocking, the membrane was probed with Cry1A specific primary antibody (Envirologix, USA). Substrate based detection was done using horse radish peroxidase conjugated secondary antibody (Sigma Aldrich, USA). The blot was developed in X-ray films using chemiluminiscent substrate, luminal (Roche Diagnostics, Germany) in developer and fixer (Kodak, Germany).
Genetic fidelity testing of in vitro regenerants
For clonal fidelity testing, ten randomly chosen in vitro raised independent T0 lines derived from different explants along with the mother plant (cv. DCP 92-3) were selected. Genomic DNA was extracted using DNeasy Plant Kit, (Qiagen, Hilden, Germany). Genetic stability and homogeneity of these lines were assessed using 27 SSR primers (TA and NCPGR series), which are previously mapped on to the eight linkage groups (LG 1 through LG 8) of chickpea genome. PCR amplifications were carried out in 10 µl reaction volume containing 1 µl of 10× PCR buffer, 1 µl (1 mM dNTPs mix), 0.5 µl of each forward and reverse primers (10 pmol), 0.2 µl of Taq DNA polymerase (5U/µl), 1 µl (25 ng) of template DNA and rest of volume adjusted by MilliQ water. Touchdown PCR was performed that employs an initial annealing temperature above the projected melting temperature (Tm) of the primers being used, then progressively transitions to a lower, more permissive annealing temperature over the course of successive cycles. Any difference in Tm between correct and incorrect annealing will produce an exponential advantage of twofold per cycle. The PCR conditions follows: 95 °C for 3 min, followed by 5 cycles of 94 °C for 20 s, decrease 1 °C/cycle from 60 °C for 20 s, 72 °C for 30 s, followed by 35 cycles of 94 °C for 20 s, 55 °C for 20 s, 72 °C for 30 s, and a final extension at 72 °C for 10 min. PCR amplified products were finally resolved on 3% agarose gels using 1× TBE running buffer stained with ethidium bromide (0.5 µg/µl). Finally, gel images were photographed using gel documentation system (Gel Doc XR, Bio-Rad, USA) and scored for clear amplified fragments.
Results
Multiple shoot induction from explants
Shoot induction response of five different explants (AMS, EAX, PLU, EPI and HYP) on three different shoot induction medium and control exhibited variation in average number of shoot primordia per responding explant in response cytokinin treatment (Fig. 1a; Supplementary Table 1). Among the various media combination tested, significantly higher frequency of shoot induction was observed on SIM 3 for all the explants tested, except for PLU cultured on SIM 2 and EAX sub cultured on SIM 1/SIM 2 which exhibited comparable responses with SIM 3 (P < 0.05; two-tail). Highest number of shoot primordia was observed in AMS (11.67 ± 0.49) followed by EPI (4.58 ± 0.67), HYP and PLU (4.50 ± 0.67) explants.
Histogram depicting response on five different explants in different media combinations. a Average number of shoot primodia per responding explants on SIM. b Average shoot length per responding explants on SEM. c Average number of root per responding explants on RM. d Average length of root per responding explant in RM
Elongation of regenerated shoots
Elongation potential of shoot primordia obtained from five different responding explants (AMS, EAX, PLU, EPI and HYP) raised on SIM 3 for 2 weeks were assessed in three different SEM for maximum shoot elongation and growth. Shoot elongation response of the tested explants on three different SEM exhibited variation in the average length of shoot per responding explants (Fig. 1b; Supplementary Table 2). Significantly higher shoot length was observed on SEM 3 for most of the explants tested except for the AMS and PLU that exhibited better/comparable response on SEM 2 (P < 0.005). Highest shoot length among the tested explants was recorded for EAX (5.46 ± 0.82 cm) followed by PLU (3.91 ± 0.43 cm) and EPI (3.45 ± 0.41 cm) cultured on SEM 3.
Rooting and plant establishment
Rooting was observed in most of the explants within the first week of culturing on RM and fully grown roots were developed within 2–3 weeks. Root induction response of all the five explants (AMS, EAX, PLU, EPI and HYP) on three different rooting medium and a control exhibited variability in terms average number of roots generated per explants and average root length (Fig. 1c, d; Supplementary Table 3). Significantly higher numbers of root per explants and correspondingly higher root length were observed for all the explants sub-cultured on RM 3 (P < 0.05). Highest number of roots was recorded for the EPI (18.6 ± 0.9) explants followed by EAX (18.4 ± 1.2) and AMS (16.6 ± 2.0) explants. Highest root length was recorded for EPI (13.1 ± 2.3 cm) followed by EAX (12.4 ± 0.7 cm) and AMS (11.3 ± 2.3) explants. Comparable rooting response in terms of average root length per explants were observed among explants cultured on control, RM 1 and RM 2 (P > 0.05). Healthy rooted plantlets from all the tested explants were hardened on the soil matrix (soil: vermi-compost: cocopeat in equal proportion) and mature fertile plants were obtained. Overall, regeneration efficiency ranged from 62.72 to 86.69% were observed, with highest efficiency for shoots derived from EAX (86.69%) followed by EPI (76.69%) (Table 1). The overall regeneration system from two responding explants viz. EAX and EPI till rooting has been depicted (Supplementary Fig. 3a–h).
Genetic transformation
Genetic transformation of chickpea cv. DCP 92-3 was carried out using expression vector harboring synthetic Bt gene and selectable marker gene nptII. Two explants EPI and EAX were independently bombarded and regenerants were selected on antibiotic, kanamycin monosulphate (100 mg/L). For genetic transformation experiments, 2653 EPI and 2553 EAX explants were bombarded in batches and transferred to SIM 3 and cultured for 2 weeks. Proliferated shoots were then transferred to SEM 3 containing Kanamycin monosulphate (100 mg/L) and sub-cultured every 2 weeks during which only transformed shoots proliferated. Healthy green elongated shoots, surviving kanamycin selection were transferred to RM 3 for rooting and finally established as plants in Containment facility. A total of 29 and 46 putative primary plants (T0) could be established from EPI and EAX explants, respectively. Seeds harvested from T0 plants were advanced for two successive generations (T1 and T2) and details of seed harvested were documented and deposited in seed repository.
Molecular characterization of transgenic lines
The presence and transmission of the transgene(s) were confirmed by PCR in the progenies (T1 and T2) derived from T0 plants. PCR screening using gene specific primer (A4F and A4R) indicated presence of 441 bp amplification product specific to Bt gene (Fig. 2a).
Presence of Bt gene in transgenic chickpea lines. a PCR amplification of Bt gene segment [L1: 1 kb DNA ladder, L2: AS 18.2, L3: AS18.4, L4:AS32.12, L5: AS38.1, L6: AS44.6, L7: JA17.1, L8: JA17.2, L9: JA18.2, L10: JA18.4, L11: JA20.4, L12: JA20.6, L13: Non-transgenic (DCP 92-3) chickpea line, Transgenic chickpea lines, L14: No template control (NTC), L15 Positive control (PC- Expression vector harboring optimized Vip3Aa gene). b Genomic Southern blotting after single digestion (HindIII) [L1: DNA ladder VII (DIG-labeled), L2: AS18, L3: AS32, L4: AS38, L5: AS44, L6: JA17, L7: JA18, L8:JA20, L9: Control DCP 92-3, L10: Positive control. c Genomic Southern blotting after double digestion (HindIII and EcoRI) [L1: DNA ladder VII (DIG-labeled), L2: AS18, L3: AS32, L4: AS38, L5: AS44, L6: JA17, L7: JA18, L8:JA20, L9: Control DCP 92-3, L10: Positive control]
Southern blot hybridization from pooled progenies (T1 generation) indicated stable integration of Bt gene in the selected transgenic chickpea events. Several single and multiple loci integration were detected in different chickpea lines tested. Genomic DNA digested with the unique restriction endonuclase HindIII exhibited single hybridization signal in events corresponding a unique location in the genome (Events AS18, AS32, AS38, AS44, JA17, JA18 and JA20) (Fig. 2b; Supplementary Fig. 4). Double digestion with EcoRI and HindIII endonucleases exhibited release of 2.62 kb recombinant Bt gene cassette from the genome of same transgenic chickpea lines, indicated intactness of the delivered and integrated copy (Fig. 2c; Supplementary Fig. 5).
Based on the PCR results, T1 progenies derived from 19 primary transformants (EPI explants) and 31 primary transformants (EAX explants) were found PCR positive with transformation efficiency of 0.72% and 1.21% respectively. Details of seeds harvested from seven selected lines have been documented (Supplementary Table 4). Chi-square test of all seven selected lines indicated segregation of transgene following Mendelian segregation pattern of 3:1, except for two lines JA17 and JA18 (Table 2).
Quantitative ELISA confirmed the expression of Bt protein in all the PCR positive chickpea progenies but were absent in the control and negative segregants. Quantity of TSP of the pooled leaf samples from transgenic progenies varied between 0.03 and 0.18 mg/ml. Variation in the temporal expressions during the pre-flowering and post flowering stages were observed in all the transgenic progenies tested. During the pre-flowering stage (65 DAS), Bt expression ranged between 6.63 and 11.95 ng/mg of TSP, while in the post flowering stage, expression dipped to 4.85–8.93 ng/mg of TSP (Fig. 3a).Western blot analysis with Bt-specific antibody indicated presence of 66 kDa band in transgenic leaf samples harvested post-flowering, without any degradation, corresponding to expressed Bt protein (Fig. 3b; Supplementary Fig. 6).
Expression of Bt gene from pooled PCR positive progenies. a Concentration (ng/mg of TSP) variation depicting Bt gene expression in leaves of transgenic chickpea line during the pre-flowering and post-flowering stages. b Western blot depicting expression of Bt gene in leaves of transgenic chickpea line [L1: AS18, L2: AS32, L3: AS38, L4: AS44, L5: JA17, L6: JA18, L7:JA20, L8: Control DCP 92-3, L9: Positive control]
Genetic fidelity of in vitro regenerants
For genetic fidelity testing, a total of 18 genome-wide chickpea SSR primers were screened on ten in vitro raised T0 lines along with the mother plant (cv. DCP 92-3). Out of 18 primers used for screening, clear amplification with 16 primers were observed with expected product size, yielding a total of 19 alleles with average of 1.19 of alleles per primer (Table 3). However, two primers namely, TA203 and TA76 did not show any amplification. The allele size for these sixteen primers ranged from 130 bp (TA 80) to 287 bp (TA 3). All the primers amplified single allele except three primers with two alleles, namely TA 113 (243/220 bp), TA110 (220/190 bp) and TA116 (182/160 bp) based on LG 1, 2 and 5, respectively (Fig. 4). It was very important to note that, for all the SSR primers the amplified allele sizes were mono-morphic across the ten in vitro raised T0 lines in comparison to mother plant.
Discussion
Genetic improvement of chickpea using genetic engineering approaches holds promise in light of low genetic variability available in cultivated chickpea gene pool. Biolistic particle bombardment is powerful method of introducing nucleic acid into plants using helium pressure to drive micro-carriers through plant cell walls. The technique is easier and time efficient than Agrobacterium based methods, and can be used for transient or stable expression of foreign genes. Genetic transformation technique in chickpea is largely genotype dependent (Indurker et al. 2007). The desi chickpea cultivar, DCP92-3 was used in the present study, owing to its responsiveness to multiple shoot organogenesis pathways, in presence of suitable hormonal combination. Five different explants viz. AMS, EAX, PLU, EPI and HYP derived from mature chickpea seeds responded to regeneration. Morphogenetic responses of two explants viz. EAX and EPI were better in the terms of shoot primordia initiation, shoot elongation and rooting and establishment as mature fertile plant. Overall regeneration efficiency of both the explants ranged from 78.67 to 86.67%. This is the first attempt to access the potential of five different explants from a tissue culture responsive chickpea genotype. Low regeneration responses accompanied by poor rooting phenomenon are the major challenges in the recovery of transgenic chickpea lines. Despite of various successful reports of chickpea regeneration and transformation, the reported transformation frequency is highly variable (Ganguly et al. 2020). It is imperative to compare the available regeneration protocols for generating an efficient regeneration system in chickpea. Plant regeneration from five different explants were compared on three pre-standardized shoot induction media, shoot elongation and rooting media, as reported earlier (Fontana et al. 1993; Senthil et al. 2004; Bhatnagar-Mathur et al. 2009).
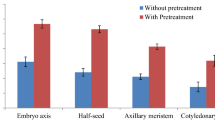
Hormones play a crucial role in morphogenetic response of plant tissues/explants. In the current study, cytokinin like BAP, KIN, TDZ, 2iP, auxins like IAA, IBA, NAA; gibberellins like GA3 and inorganic additives KNO3 in various combinations were tested on all the explants during in vitro regeneration to plant establishment stages. Notably, pre-treatment of the explants with BAP enhanced the regenerative potential, as described earlier (Sanyal et al. 2005; Srivastava et al. 2012). BAP and KIN together augmented shoot primordial initiation, while GA3 alone could effectively elongate majority of the shoots. In rooting best response were observed with NAA with overall establishment rate of > 80%. BAP in a concentration range of 0.5–1 mg/L is the most extensively used cytokinin employed for multiple shoot induction in chickpea. Among the various combination of cytokinin tested for multiple shoot induction, modified MS salts supplemented with 0.5 mg/L BAP and 0.1 mg/L KIN exhibited better response for all the tested explants. Our observation agrees with the earlier reports of shoot induction in diverse chickpea genotypes employing different explants (viz. cotyledon with half EAX, stem, EPI, and EAX) using combination of BAP and KIN in variable proportion (Sarmah et al. 2004; Indurker et al. 2007; Ganguly et al. 2014). The EAX explants turned out to be best explant for multiple shoot induction producing 11.67 ± 0.49 shoots per responding explant followed by EPI. In the shoot induction media containing TDZ, shoots were vitrified and low regeneration frequency was witnessed for EPI and HYP explants. This may be due the prolonged exposure of explants to TDZ that may cause hyperhydracity and abnormal shoot growth (Jayanand et al. 2003; Kumari et al. 2018). For the shoot elongation studies modified MS salts supplemented with GA3 affected 5.46 ± 0.82 cm long shoot with EAX shoot primordia followed by PLU and EPI shoots. Earlier reports have suggested the use of GA3 for increasing internodal length and improving leaf morphology (Jayanand et al. 2003). Effects of different shoot elongation media (SEMs) and the role of GA3 on shoot length of proliferated shoots originated from axillary meristem explants has depicted that the shoot length can be increased up to 2× in the presence of GA3 (Kumari et al. 2018). In our investigation minimum shoot elongation response accompanied with drying of shoot tips was recorded on modified MS salts supplemented with TDZ in all five tested explants. This may be due the prolong effect of TDZ that is reported to cause abnormalities in the in vitro generated shoots at high concentration. Rooting efficiency observed in the study ranged between 78.67 and 94.67%, higher than the earlier report where 50–60% rooting was reported using a pulse treatment of 10 mM IBA for 30 s (Indurker et al. 2007). In the present study, rooting potential was found best in MS salts supplemented with NAA for all the tested explants. NAA at various concentrations 0.186–2 mg/L have been reported to exhibit rooting response in the in vitro generated EAX, PLU, and EPI explants (Altinkut et al. 1997; Senthil et al. 2004; Polowick et al. 2004; Singh et al. 2009). Interestingly, during the initial phase of rooting experiment, modified MS salts supplemented with KNO3 and IBA exhibited higher number of elongated roots compared to the other rooting media combination tested, but in the later stage NAA exhibited a more prominent rooting response in terms of root length and root number in all the tested explants. Taken together, the protocol described here has been significant owing to its high reproducibility and higher recovery of regenerated plantlets in a relatively short period (70–80 days) as compared to Agrobacterium tumefaciens mediated genetic transformation.
Earlier reports of direct transformation in chickpea have demonstrated the potential of gold and tungsten micro-carriers for generating transient/stable transformation in recombinant lines. In the first report of direct transformation in chickpea using gold micro-carriers of 1 µm size have been successfully employed for development of insect resistant chickpea lines harboring Cry1Ac. Later on direct transformation of chickpea HYPs explants employing 1.1 µm tungsten micro-carriers at a burst pressure of 853 psi and distance of 29 cm between the mico-carrier and the target tissue was demonstrated for successful expression of gus gene in 58% of the tested explants using a homemade gene gun (Husnain et al. 2000). In a comparative study to test the efficacy of gold (1 µm) and tungsten (1.1 µm) micro-carriers, result indicates high transformation efficiency with both type of micro-carrier at 900 psi pressure and at a bombardment distance of 6 cm between the micro-carrier and target tissue although higher efficiency (around 3% more) with gold micro-carrier was witnessed (Indurker et al. 2007). In the present study, particle bombardment was done on two best explants (EAX and EPI) and transformation frequency obtained are 0.72 and 1.21% using 1.1 µm tungsten micro-carrier bombarded at a burst pressure of 1100 psi maintain a distance of 4 cm between the micro-carrier and target tissue. Tungsten particles have long been used as microcarriers in biolistic bombardment because of their cost-effectiveness compared to alternative gold particles—even if the former is reported for DNA-degrading activity. Adoption of DNA/tungsten adsorption employing TE buffers at alkaline pH (> 9.0) of the mixture, in which tungsten-bound plasmid DNA cleavage was suppressed was adopted in the study, as described earlier (Yosimitshu et al. 2009).
Earlier reports of direct transformation in chickpea have tested/compared the transformation and regeneration potential of EAX, decapitated EAX, HYP, cotyledonary node, shoot tip, leaf, stem and EPI explants (Kar et al. 1997; Husnain et al. 2000; Sanyal et al. 2003; Tewari-singh et al. 2004; Indurker et al. 2007). Transformation efficacy for direct transformation method of EAX (6 ± 1.15%), stem (9 ± 0.58%), and EPI (16 ± 0.33%) explants were compared in chickpea and result indicated higher suitability of EPI explant over the other tested explants (Indurker et al. 2007). Another report comparing transformation efficiency of chickpea EAX, cotyledonary node, shoot tip and leaf explants depicted highest number of GUS foci per responding cotyledonary node explants, indicating the suitability of cotyledonary explants for direct transformation (Sanyal et al. 2003). In the present study, transformation efficiency of 0.72% for EPI explants and 1.21% for EAX explants were obtained using the formula: Transformation efficiency = [No. of PCR positive lines (T1)/Total No. of explants bombarded] X 100. In an earlier report higher transformation efficiency of around 16% using EPIs explants was calculated based on the percentage of explants producing de novo shoot regeneration in kanamycin medium after 2 months (Indurker et al. 2007).
In the present study, potential for direct transformation of two most responding explants for in vitro regeneration (EAX and EPI) were tested using synthetic Bt gene. Particle gun bombardment resulted in single and multiple copies of transgene, however in the current study only singe copy lines were used in the study. The levels of Bt protein ranged from 6.63 to 11.95 ng/mg of TSP in the pre-flowering stage and 4.85 to 8.93 ng/mg of TSP in the post- flowering stage comparable to our earlier reports (Das et al. 2017). A wide difference in the levels of Bt protein in different transgenic plants (T1 & T2) and various segregation patterns in T2 progenies could have been due to difference in the site of its integration. Efficacy of the transgene was reported earlier in our reports (% larval mortality ranged: 75–100%).
Many researchers suggested using more than one marker systems for genetic fidelity testing (Singh et al. 2013, Rohela et al. 2019, Sadhu et al. 2020). However, the sensitivity, reproducibility and strong discriminatory power of microsatellite simple sequence repeat (SSR) markers make them particularly suitable for determining genetic stability and the rejecting possibility of somaclonal variants in vitro regenerated plants (Parida et al. 2009). In our study, we have selected and deployed the SSR markers which are mapped on to the eight linkage groups of chickpea (Flandez-Galvez et al. 2003). Compared to multi-locus profiling techniques e.g., RAPD, ISSR, AFLP and SCoT etc. which generates multiple bands with unknown chromosome locations with lesser reproducibility, SSR marker can be used to differentiate homozygotic and heterozygotic alleles between the lines from the same origin. Although SSRs markers have the ability to detect the highest levels of polymorphism, the suitability of SSR markers for evaluation of clonal fidelity was described (Wanmei et al. 2009). The genome-wide profiling of SSR markers with known chromosome locations were found highly appropriate and correct method for genetic fidelity testing in many other crops e.g., pigeonpea (Dutta et al. 2011), rice (Nachimuthu et al. 2015), lentil (Andeden et al. 2015), soybean (Kumar et al. 2015) etc. Further, we reconfirmed that all the SSR primers deployed in our study generated expected allele sizes across chickpea samples as reported earlier (Winter et al. 1999; Parida et al. 2015). In our study, three SSR primers TA 113, TA110 and TA 116 produced two alleles and remaining thirteen primers produce single alleles yielding total 19 alleles with mono-morphic patterns across the ten T0 lines and mother plant. The lack of genetic variation at SSR loci strongly confirmed that the genetically true to-type, stable nature and homogeneity of all the in vitro raised chickpea plantlets tested here. These results also suggested the feasibility of developed in vitro regeneration protocol for carrying out direct transformation using biolistic particle delivery method in chickpea.
Conclusions
EAX and EPI explants are best suited for regeneration and direct genetic transformation of chickpea. The regenerants were true-to-type and fertile, opening avenues for incorporation of new traits in chickpea.
Data availability
All data generated or analyzed during this study are included in this published article. Transgenic chickpea materials reported in the study are available in Seed Repository, ICAR-Indian Institute of Pulses Research, Kanpur, INDIA (https://iipr.icar.gov.in/).
Code availability
Not Applicable.
Abbreviations
- DT:
-
Direct transformation
- Bt :
-
Bacillus thuringiensis gene
- nptII :
-
Neomycin phosphotransferase II
- BAP:
-
6-Benzyl amino purine
- KIN:
-
Kinetin
- TDZ:
-
Thidiazuron
- 2iP:
-
2-Isopentenyladenine
- GA3 :
-
Gibberellic acid
- IAA:
-
Indole-3-acetic acid
- IBA:
-
Indole-3-butyric acid
- NAA:
-
1-Naphthalene acetic acid
- KNO3 :
-
Potassium nitrate
- SSR:
-
Simple sequence repeat
References
Acharjee S, Sarmah BK, Kumar PA, Olsen K, Mahon R, Moar WJ, Moore A, Higgins TJV (2010) Transgenic chickpeas (Cicer arietinum L.) expressing a sequence modified cry2Aa gene. Plant Sci 178:333–339. https://doi.org/10.1016/j.plantsci.2010.02.001
Aggarwal PR, Nag P, Choudhary P, Chakraborty N, Chakraborty S (2018) Genotype-independent Agrobacterium rhizogenes-mediated root transformation of chickpea: a rapid and efficient method for reverse genetics studies. Plant Methods 14:55. https://doi.org/10.1186/s13007-018-0315-6
Altinkut A, Gozukirmiz N, Bajrovic K, Gozukirmizi N (1997) High percentage of regeneration and transformation in chickpea. Acta Hortic 447:319–320. https://doi.org/10.17660/ActaHortic.1997.447.63
Anbazhagan K, Bhatnagar-Mathur P, Vadez V, Dumbala SR, Kishor PBK, Sharma KK (2015) DREB1A over expression in transgenic chickpea alters key traits influencing plant water budget across water regimes. Plant Cell Rep 34:199–210. https://doi.org/10.1007/s00299-014-1699-z
Andeden EE, Baloch FS, Çakır E, Toklu F, Özkan H (2015) Development, characterization and mapping of microsatellite markers for lentil (Lens culinaris Medik.). Plant Breed 134:589–598
Asharani BM, Ganeshaiah KN, Raja A, Kumar V, Makarla U (2011) Transformation of chickpea lines with cry1X using in planta transformation and characterization of putative transformants T1 lines for molecular and biochemical characters. J Plant Breeding Crop Sci 3(16):413–423. https://doi.org/10.5897/JPBCS11.074
Bhatnagar-Mathur P, Vadez V, Jyostna Devi M, Lavanya K, Vani G, Sharma KK (2009) Genetic engineering of chickpea (Cicer arietinum L.) with the P5CSF129A gene for osmoregulation with implications on drought tolerance. Mol Breeding 23:591–606. https://doi.org/10.1007/s11032-009-9258-y
Chakraborti D, Sarkar A, Mondal HS, Das S (2009) Tissue specific expression of protein insecticidal, Allium sativum leaf agglutinin (ASAL) in important pulse crop, chickpea (Cicer arietinum L.) to resist the phloem feeding Aphis craccivora. Transgenic Res 18:529–544. https://doi.org/10.1007/s11248-009-9242-7
Chakraborty J, Sen S, Ghosh P, Sengupta A, Basu D, Das S (2016) Homologous promoter derived constitutive and chloroplast targeted expression of synthetic cry1Ac in transgenic chickpea confers resistance against Helicoverpa armigera. Plant Cell Tissue Org Cult 125:521–535. https://doi.org/10.1007/s11240-016-0968-7
Das A, Sarmah BK (2005) Genetic transformation of chickpea (Cicer arietinum L.). Res J Contem Concerns 1:1–6
Das A, Datta S, Thakur S, Shukla A, Ansari J, Sujayanand GK, Kumar PA, Chaturvedi SK, Singh NP (2017) Expression of a chimeric gene encoding insecticidal crystal protein Bt gene of Bacillus thuringiensis in chickpea (Cicer arietinum L.) confers resistance to gram pod borer (Helicoverpa armigera Hubner.). Front Plant Sci 8:1423. https://doi.org/10.3389/fpls.2017.01423
Das A, Thakur S, Shukla A, Singh P, Ansari J, Singh NP (2020) Genetic transformation. In: Singh M (ed) Chickpea: crop wild relatives for enhancing genetic gains. Academic Press, London, pp 205–224. https://doi.org/10.1016/b978-0-12-818299-4.00008-7
Das A, Basu PS, Kumar M et al (2021) Transgenic chickpea (Cicer arietinum L.) harbouring AtDREB1a are physiologically better adapted to water deficit. BMC Plant Biol 21:39. https://doi.org/10.1186/s12870-020-02815-4
Dutta S, Kumawat G, Singh BP et al (2011) Development of genic-SSR markers by deep transcriptome sequencing in pigeonpea [Cajanus cajan (L.) Millspaugh]. BMC Plant Biol 11:17. https://doi.org/10.1186/1471-2229-11-17
Flandez-Galvez H, Ford R, Pang ECK et al (2003) An intraspecific linkage map of the chickpea (Cicer arietinum L.) genome based on sequence tagged microsatellite site and resistance gene analog markers. Theor Appl Genet 106:1447–1456. https://doi.org/10.1007/s00122-003-1199-y
Fontana GS, Santini L, Caretto S, Frugis G, Mariotti D (1993) Genetic transformation in the grain legume Cicer arietinum L. Plant Cell Rep 12:194–198. https://doi.org/10.1007/BF00237052
Food and Agriculture Organization (FAO) (2019) FAOSTAT Statistical Database of the United Nation Food and Agriculture Organization (FAO) statistical division, Rome. Accessed on 15 June 2021
Ganguly M, Molla KA, Karmakar S, Datta K, Datta SK (2014) Development of pod borer-resistant transgenic chickpea using a pod specific and a constitutive promoter driven fused cry1Ab/Ac gene. Theor Appl Genet 127:2555–2565. https://doi.org/10.1007/s00122-014-2397-5
Ganguly S, Ghosh G, Ghosh S, Purohit A, Chaudhuri RK, Das S, Chakraborti D (2020) Plumular meristem transformation system for chickpea: an efficient method to overcome recalcitrant tissue culture responses. Plant Cell Tiss Organ Cult 142:493–504. https://doi.org/10.1007/s11240-020-01873-8
Gao C, Nielsen KK (2013) Comparison between Agrobacterium mediated and direct gene transfer using the gene gun. Methods Mol Biol 940:3–16. https://doi.org/10.1007/978-1-62703-110-3_1
Husnain T, Fatima T, Islam R, Riazuddin S (2000) Plant regeneration and expression of beta-glucorinidase gene in hypocotyls tissue of chickpea (Cicer arietinum L.). Pak J Biol Sci 3(5):842–845. https://doi.org/10.3923/pjbs.2000.842.845
Indurker S, Misra HS, Eapen S (2007) Genetic transformation of chickpea (Cicer arietinum L.) with insecticidal crystal protein gene using particle gun bombardment. Plant Cell Rep 26:755–763. https://doi.org/10.1007/s00299-006-0283-6
Indurker S, Mishra HS, Eapen S (2010) Agrobacterium-mediated transformation in chickpea (Cicer arietinum L.) with an insecticidal protein gene: optimization of different factors. Physiol Mol Biol Plants 16(3):273–284. https://doi.org/10.1007/s12298-010-0030-x
Jain D, Chattopadhyay D (2013) Promoter of CaZF, a chickpea gene that positively regulates growth and stress tolerance, is activated by an AP2-family transcription factor CAP2. PLoS One 8(2):e56737. https://doi.org/10.1371/journal.pone.0056737
Jayanand B, Sudarsanam G, Sharma KK (2003) An efficient protocol for the regeneration of whole plants of chickpea (Cicer arietinum L.) by using axillary meristem explants derived from in vitro-germinated seedlings. In Vitro Cell Dev Biol Plant 39:171. https://doi.org/10.1079/IVP2002387
Kar S, Johnson TM, Nayak P, Sen SK (1997) Efficient transgenic plant regeneration through Agrobacterium-mediated transformation of chickpea (Cicer arietinum L.). Plant Cell Rep 16:32–37. https://doi.org/10.1007/BF01275444
Khatodia S, Kharb S, Barta P, Chowdhury VK (2014) Development and characterization of transgenic chickpea (Cicer arietinum L.) plants with cry1Ac gene using tissue culture independent protocol. Int J Adv Res 2(8):323–331
Kumar V, Rani A, Rawal R, Mourya V, Putrevu S, Jhawar J, Dixit A (2015) Genetic diversity of soybean genotypes differing in isoflavones content as revealed by HPLC and SSR markers. Aust J Crop Sci 9(9):844–852
Kumari P, Singh S, Yadav S, Tran LSP (2018) Pretreatment of seeds with thidiazuron delimits its negative effects on explants and promotes regeneration in chickpea (Cicer arietinum L.). Plant Cell Tiss Organ Cult 133:103–114. https://doi.org/10.1007/s11240-017-1365-6
Mehrotra M, Singh AK, Sanyal I, Altosaar I, Amla DV (2011) Pyramiding of modified cry1Ab and cry1Ac genes of Bacillus thuringiensis in transgenic chickpea (Cicer arietinum L.) for improved resistance to pod borer insect Helicoverpa armigera. Euphytica 182:87–102. https://doi.org/10.1007/s10681-011-0501-3
Murashige T, Skoog F (1962) A revised medium for rapid growth and bioassays with tobacco tissue cultures. Physiol Plant 15:473–497. https://doi.org/10.1111/j.1399-3054.1962.tb0805
Nachimuthu VV, Muthurajan R, Duraialaguraja S et al (2015) Analysis of population structure and genetic diversity in rice germplasm using SSR markers: an initiative towards association mapping of agronomic traits in Oryza Sativa. Rice 8:30. https://doi.org/10.1186/s12284-015-0062-5
Parida SK, Dalal V, Singh NK, Mohapatra T (2009) Genic non-coding microsatellites in the rice genome: characterization, marker design and use in assessing genetic and evolutionary relationships among domesticated groups. BMC Genom. https://doi.org/10.1186/1471-2164-10-140
Parida SK, Verma M, Yadav SK, Ambawat S, Das S, Garg R, Jain M (2015) Development of genome-wide informative simple sequence repeat markers for large-scale genotyping applications in chickpea and development of web resource. Front Plant Sci 21(6):645. https://doi.org/10.3389/fpls.2015.00645
Polowick PL, Baliski DS, Mahon JD (2004) Agrobacterium tumefaciens-mediated transformation of chickpea (Cicer arietinum L.): gene integration, expression and inheritance. Plant Cell Rep 23:485–491. https://doi.org/10.1007/s00299-004-0857-0
Rohela GK, Jogam P, Bylla P, Reuben C (2019) Indirect regeneration and assessment of genetic fidelity of acclimated plantlets by SCoT, ISSR, and RAPD markers in Rauwolfia tetraphylla L.: an endangered medicinal plant. BioMed Res Int. https://doi.org/10.1155/2019/3698742
Sadhu S, Jogam P, Thampu RK et al (2020) High efficiency plant regeneration and genetic fidelity of regenerants by SCoT and ISSR markers in chickpea (Cicer arietinum L.). Plant Cell Tiss Organ Cult 141:465–477. https://doi.org/10.1007/s11240-020-01804-7
Sagare AP, Suhasini K, Krishnamurthy KV (1993) Plant regeneration via somatic embryogenesis in chickpea (Cicer arietinum L.). Plant Cell Rep 12:652–655. https://doi.org/10.1007/BF00232818
Sanyal I, Singh AK, Amla DV (2003) Agrobacterium tumefaciens-mediated transformation of chickpea (Cicer arietinum L) using mature embryonic axes and cotyledonary nodes. Indian J Biotechnol 2:524–532. https://doi.org/10.1007/s12298-010-0030-x
Sanyal I, Singh A, Kaushik M, Amla DV (2005) Agrobacterium mediated transformation of chickpea (Cicer arietinum L.) with Bacillus thuringiensis cry1Ac gene for resistance against Pod borer insect Helicoverpa armigera. Plant Sci 168:1135–1146. https://doi.org/10.1016/j.plantsci.2004.12.015
Sarmah BK, Moore A, Tate W, Molvig L, Morton RL, Rees DP, Chiaieste P, Chrispeels MJ, Tabe LM, Higgins TJV (2004) Transgenic chickpea seeds expressing high levels of a bean a-amylase inhibitor. Mol Breed 14:73–82. https://doi.org/10.1023/B:MOLB.0000037996.01494.12
Senthil G, Williamson B, Dinkins RD, Ramsay G (2004) An efficient transformation system for chickpea (Cicer arietinum L.). Plant Cell Rep 23:297–303. https://doi.org/10.1007/s00299-004-0854-3
Shukla A, Das A, Ansari J, Datta S (2015) In vitro regeneration of chickpea (Cicer arietinum L.) via somatic embryogenesis. J Food Legumes 28(3):199–202
Singh R, Singh NP, Datta S, Yadav IS, Singh AP (2009) Agrobacterium mediated transformation of chickpea using shoot meristem. Indian J Biotechnol 8:78–84
Singh SR, Dalal S, Singh R, Dhawan AK, Kalia RK (2013) Evaluation of genetic fidelity of in vitro raised plants of Dendrocalamus asper (Shult. & Shult. F.) Backer ex K. Heyne using DNA-based markers. Acta Physiol Plant 35:419–430
Srivastava J, Das A, Soren KR, Chaturvedi SK, Nadarajan N, Datta S (2012) Ontogeny of in vitro shoot organogenesis from axillary meristem explants in chickpea (Cicer arietinum L.). J Crop Sci Biotechnol 15:245–250. https://doi.org/10.1007/s12892-012-0032-z
Tewari-Singh S, Sen J, Kiesecker H, Reddy VS, Jacobsen HJ, Guha-Mukherjee S (2004) Use of a herbicide or leucine plus threonine for non-antibiotic selection of transgenic chickpea. Plant Cell Rep 22:576–583. https://doi.org/10.1007/s00299-003-0730-6
Varshney RK, Thudi M, Roorkiwal M et al (2019) Resequencing of 429 chickpea accessions from 45 countries provides insights into genome diversity, domestication and agronomic traits. Nat Genet 51:857–864. https://doi.org/10.1038/s41588-019-0401-3
Wanmei J, Dong J, Wang Y, Mao H, Xiao Z, Chen M (2009) Genetic fidelity of regeneration adventitious shoots in grape through organogenesis. Mol Plant Breed 2:375–379
Winter P, Paff T, Udupa SM, Huttel B, Sharma PC, Sahi S, Espinoza RA, Weigand F, Kahl G (1999) Characterization and mapping of sequence-tagged microsatellite sites in the chickpea (Cicer arietinum L.) genome. Mol Gen Genet 262:90–101
Yoshimitsu Y, Tanaka K, Tagawa T, Nakamura Y, Matsuo T, Okamoto S (2009) Improvement of DNA/metal particle adsorption in tungsten-based biolistic bombardment; alkaline pH is necessary for DNA adsorption and suppression of DNA degradation. J Plant Biol 52:524–532. https://doi.org/10.1007/s12374-009-9068-0
Acknowledgements
The authors are thankful to ICAR-National Institute of Plant Biotechnology, New Delhi, for providing the plant expression vector harboring the Bt gene; Institutional Biosafety Committee (IBSC), IIPR, Kanpur for approving the activities for development of experimental transgenic chickpea lines in Plant Tissue Culture Facility and Transgenic Containment Facility (PBSL1), as per extant Indian Biosafety Regulations (Rules, 1989 and Regulations and Guidelines for Recombinant DNA Research and Bio-Containment, 2017). Thanks are due to Scientific Technical Staffs: Mr. Ajeet Pratap Singh, Mr. Malkhan Singh and Mr. Ravi Ranjan Singh, Division of Plant Biotechnology, ICAR-Indian Institute of Pulses Research, Kanpur, INDIA, for overall help in maintaining the experiments in Containment Facility, chemicals and seed inventory maintenance.
Funding
Research was partially supported by Network Project on Transgenics in Crops (NPTC), National Agricultural Science Fund (NASF) and Indian Council of Agricultural Research-Indian Institute of Pulses Research, Kanpur, India.
Author information
Authors and Affiliations
Contributions
AD, PGP, MR, OPV, NPS: Conception and designing of the experiments; PS, JA, AD, ST, MR: Standardization of in vitro regeneration parameters for particle bombardment of chickpea, JA, AS, AD, ST, NPS: Particle bombardment of chickpea with Bt gene, molecular analyses of transgenic lines, interpreted and compiled the results; NNT, PGP, AD, ST: Conducted genetic fidelity and interpreted the data related to the experiments. All authors compiled the text presented in the manuscript, proof-read and approved the final manuscript.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that there are no competing interests. ICAR-Indian Institute of Pulses Research, Kanpur is the sole owner and holds all rights pertaining to transgenic chickpea materials developed in the study.
Ethical approval
Not Applicable.
Consent to participate
All authors consented to the research work reported in the study.
Consent for publication
All the authors consented to publication of the research article.
Additional information
Communicated by Sergio J. Ochatt.
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
11240_2022_2230_MOESM1_ESM.ppt
Supplementary Figure 1: Five different explants derived from mature chickpea seeds (DCP 92-3). Supplementary Figure 2: Map of plant expression vector harboring insecticidal Bt gene and selectable marker nptII. Supplementary Figure 3. Regeneration and establishment of EAX (a-d) and EPI (e-h) explant (a) Shoot primordial in SIM 3, EAX explants inset (b) Elongated multiple shoots on SEM 3 (c) Regenerated rooting system on RM 3 (d) Establishment of in-vitro regenerated plantlet (e) Shoot primordial in SIM 3, EPI explants inset (f) Elongated multiple shoots on SEM 3 (g) Regenerated rooting system on RM 3 (h) Establishment of in-vitro regenerated plantlet. Supplementary Figure 4: Original blot of genomic Southern blotting after single digestion (HindIII). Supplementary Figure 5: Original blot of genomic Southern blotting after double digestion (HindIII and EcoRI). Supplementary Figure 6: Original western blot. Supplementary file1 (PPT 2854 KB)
Rights and permissions
About this article
Cite this article
Singh, P., Shukla, A., Tiwari, N.N. et al. Routine and efficient in vitro regeneration system amenable to biolistic particle delivery in chickpea (Cicer arietinum L.). Plant Cell Tiss Organ Cult 148, 699–711 (2022). https://doi.org/10.1007/s11240-022-02230-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11240-022-02230-7