Abstract
Phosphoinositide 3-kinase (PI3K) has been considered to be a potential drug target for the treatment of several human body diseases. Nowadays, great efforts have been made on the development of selective PI3Kδ inhibitors because of the FDA approval of idelalisib, which is the first listed PI3K inhibitor. But serious side effects occur during the use of idelalisib that greatly promotes the development of novel PI3Kδ inhibitors. Nevertheless, idelalisib is still an important milestone in the development of selective PI3Kδ inhibitors, but the detailed selective binding mechanisms between idelalisib and PI3Ks have not been well elucidated. Therefore, in this study, an integrated modeling strategy combining molecular docking, molecular dynamics simulation, and free energy calculation was performed to reveal the molecular-level binding mechanisms of idelalisib and class I PI3K. First, molecular docking was carried on to obtain a reasonable binding posture of idelalisib in different PI3K isoforms. Then, key residues for selective inhibition of PI3Kδ were highlighted by molecular dynamics simulation and energy calculations. Finally, idelalisib was also compared with its lead compound, IC87114, to reveal the reason for the higher potency of Idelalisib for PI3Kδ. We hope that this study would provide some guidance for the rational design of selective PI3Kδ inhibitors.

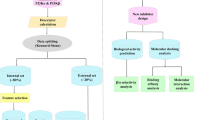
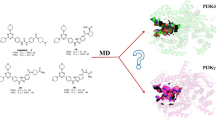
Graphical abstract
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
The extensive research conducted over the past decades has implicated the role of various cell signaling events in numerous malignant, inflammatory, autoimmune, and cardiovascular diseases, out of which the phosphoinositide 3-kinase (PI3K) signaling is one of the most relevant pathways because of its various vital functions, such as cell survival, proliferation, differentiation, and motility [1]. PI3K is an intracellular phosphatidylinositol kinase that is a key nodal protein in the PI3K/Akt signal transduction pathway [2]. PI3Ks belong to a lipid family which phosphorylates the inositol ring of membrane phosphoinositide, producing the second messenger, PIP3, that activates downstream signaling proteins, such as Akt (protein kinase B) [3]. The PI3Ks family is divided into three classes (I, II, and III) based on sequence homology, types of regulatory subunits, and substrates specificity. Among them, class I PI3Ks have been most widely studied, and class I PI3K further can be divided into two groups: IA class, which consists of PI3Kα, β, and δ and IB class has only one member, PI3Kγ [4].
Since the first PI3K inhibitor, LY294002, was reported in 1994 [5], a large number of PI3K inhibitors have been developed and reported over the past few decades, most of these inhibitors are PI3K pan-inhibitors, which reversibly bind the ATP pockets of all isoforms [6]. However, current research has focused on more selective PI3K inhibitors. Comparing PI3K pan-inhibitors, isoform-selective inhibitors contain strong efficacy and less toxic side effects, so they are the focus of research and development of PI3K inhibitors [7]. The approval of idelalisib (CAL-101), a PI3Kδ-selective inhibitor, for the treatment of chronic lymphocytic leukemia and small lymphocytic lymphoma by the U.S. Food and Drug Administration in 2014 was an important milestone for the development of selective PI3K inhibitions [8, 9]. Although idelalisib is considered as an effective drug, the unpredictable immune-mediated toxicity pattern has led to a black box warning in the USA, which limits its current role as an immunomodulator [10]. PI3Kδ expression is primarily restricted to the hematopoietic system, specifically in leukocytes. The accumulated studies have highlighted the association of deregulated PI3Kδ with numerous malignant, inflammatory, autoimmune, and cancer diseases, that depicts PI3Kδ as a significant therapeutic target [11]. Therefore, it is not surprising that a large body of PI3Kδ inhibitors have been developed in recent years, and some of them have entered the clinic trials [12].
Although serious side effects associated with idelalisib limit its clinical application, it still represents a useful tool in the study of structure selective activity relationship between PI3Kδ and inhibitors [13]. But, the detailed selective binding mechanisms between idelalisib and the four PI3K isoforms have not been well elucidated. Therefore, in this study, an integrated modeling strategy, combining molecular docking, molecular dynamics (MD) simulation, and free energy calculations, was conducted to reveal the binding modes of idelalisib for class I PI3Ks at the molecular level. Firstly, molecular docking was used to get the rational binding postures within different isoforms. And then, MD and energy calculations were employed to highlight the key residues critical for PI3Kδ-selective inhibition. Finally, IC87114, the lead compound of idelalisib, was also evaluated to reveal the reason for the higher potency of idelalisib for PI3Kδ. We hope that our work would provide some research assistance for the rational design of novel PI3Kδ-selective inhibitors.
Methods and materials
Molecular docking
Molecular docking was performed using the CDOCKER module in Discovery Studio 3.5 (DS3.5) software package. The 3D structure of idelalisib was retrieved from the co-crystal structure of PI3Kδ/idelalisib complex from the PDB database (PDB ID: 4XE0) [14] and then pre-treated through the Ligand Prepare module with the default parameters set. The other isoform structures, PI3Kα (PDB ID: 4OVU) [15], PI3Kβ (PDB ID: 2Y3A) [16], and PI3Kγ (PDB ID: 5EDS) [17], all came from PDB database. All proteins were prepared with the Prepare Protein module in DS3.5 with the default parameter, namely, removed water molecules, assigned bond sequences and protonated states, added hydrogen atoms, and the CHARMm field. Afterward, idelalisib was docked into the binding pockets of PI3Kα/β/γ, respectively. In the CDOCKER protocol, the parameters were set to default. Finally, the top-scored complex poses were collected for the next analysis.
Molecular dynamics simulation
The co-crystal structure of idelalisib/PI3Kδ and the idelalisib/PI3Kα/PI3Kβ/PI3Kγ complexes given from above molecule docking was used as the initial structures for MD simulation using the SANDER program in AMBER18 [18]. In this study, IC87114 (PDB ID: 2X38) [19], a lead compound of idelalisib, was also employed to perform MD, to compare the affinity of idelalisib for PI3Kδ. The AMBER ff14SB force field was used for the protein receptors and the ligands were minimized by the general AMBER force field (gaff) [20]. Each inhibitor was optimized using the semi-empirical AM1 method in Gaussian09 [21]. Then, the atomic partial charge was obtained by fitting the electrostatic potential calculated at the HF/6-31G* level using the RESP (constrained electrostatic potential) fitting technique. The entire system was neutralized with Na+ ions and immersed in a rectangular frame of TIP3P water molecules that extend 10 Å from any solute atom in all three dimensions [22]. The particle grid Ewald (PME) scheme is used to process long-range static electricity and truncated 10 Å for VDW interaction. Before the MD simulations, each system was subjected to three-stage minimizations using the SANDER program [23,24,25]. First of all, 1000 cycles of minimizations (500 cycles of steepest descent and 500 cycles of the conjugate gradient) were conducted with the backbone carbons to constrain the initial structures (50 kcal mol−1 Å−2). Then, 1000 cycles of minimizations with a weaker harmonic potential (10 kcal mol−1 Å−2) were performed. Finally, the whole system was relaxed by 5000 cycles of minimizations (1000 cycles of steepest descent and 4000 cycles of the conjugate gradient) without any restrain. When the minimization is finished, each system was gradually heated from 0 to 300 K over a period of 50 ps in the NVT ensemble and then performed a 50-ps MD simulation in NPT ensemble with a temperature of 300 K and pressure 1 atm. Finally, 20-ns NPT MD simulations were performed. All bonds involving hydrogen atoms were constrained using the SHAKE algorithm, and the time step was set to 2.0 fs. Coordinates were saved every 10 ps.
MM/GBSA free energy calculations and decomposition
The snapshots of each system extracted from the last stable 10-ns MD trace were used to calculate the binding free energy (ΔGbind) using the MM/GBSA method according to Eq. (1): [14, 26, 27]
The protein-ligand interaction profile of each residue was calculated by MM/GBSA free energy decomposition analysis which was calculated according to Eq. (2): [28,29,30]
where molecular mechanical energy (ΔGMM) is the gas phase interaction energy between the ligand and the protein, calculated by electrostatic (ΔGele) and van der Waals interaction (ΔGvdw). The solvation free energy (ΔGsol) consists of polarity (ΔGGB) and nonpolarity (ΔGSA). ΔGGB is calculated from the generalized Born (GB) model developed by Onufriev et al. [31]. ΔGSA is calculated on the basis of SASA by using the fast LCPO algorithm with a probe radius of 1.4 Å. –TΔS is a change in conformational entropy after ligand binding [13].
Results and discussion
Structural stability of the studied systems
In order to demonstrate the dynamic interaction patterns of studied complexes, 20-ns conventional MD simulations were conducted for four PI3K isoforms. Firstly, the quality of the MD simulation was assessed by monitoring the root-mean-square deviations (RMSDs) of the backbone atoms along the entire MD trajectory of each complex. As shown in Fig. 1 in the last 10 ns, the RMSD values of the stimulated systems were quite stable, which indicated that the MD simulations reached equilibrium within 10 ns.
Isoform selectivity predicted by MM/GBSA
Subsequently, the MM/GBSA method was used to calculate the binding free energies based on a total of 1000 snapshots extracted from the last 10 ns of the MD trajectories of each system (Table 1). The predicted binding affinities of idelalisib for the four isoforms are − 30.36 (α), − 38.97 (β), − 41.65 (δ), and − 40.36 (γ) kcal/mol, respectively. The predicted free energies and the experimental data highlight the good ranking ability of MM/GBSA calculation. Notably, the MM/GBSA affinities are much stronger than the experimental values because of ignorance of the conformational entropies, accumulated studies illustrated that in most cases the MM/GBSA approach can only give good ranking results rather than accurately predicting the absolute binding free energy [32,33,34,35,36]. Furthermore, the energy components were calculated and the results were summarized in Table 1. As a whole, the van der Waals (ΔEvdw) and electrostatic (ΔEele) terms are quite favorable for inhibitor binding to proteins, while the polar solvation term is unfavorable for binding in all complexes. In particular, the ΔEvdw is the largest contributor to the binding affinities between idelalisib and PI3Kδ. The nonpolar components were defined as ΔEvdw + ΔGSA, and the values are − 47.18 kcal/mol for PI3Kα, − 49.98 kcal/mol for PI3Kβ, − 51.81 kcal/mol for PI3Kδ, and − 49.09 kcal/mol for PI3Kγ, respectively. The nonpolar components are also consistent with the experiment results and it indicates that the nonpolar plays a critical role in determining the binding specificity of PI3Kδ. Considering the hydrophobicity of the ATP-binding pocket of PI3K, it is not surprising to understand why idelalisib has stronger inhibition to PI3Kδ than other isoforms. It is noteworthy that PI3Kδ shows lower ΔEele than PI3Kβ and PI3Kγ, suggesting that the electrostatic term is propitious for PI3Kβ and PI3Kγ. The above results preliminarily showed that increasing the van der Waals interaction for PI3Kδ and decreasing the electrostatic interaction for other isoforms may improve the PI3Kδ-selective binding for inhibitors.
The mechanisms of isoform-selective inhibition
In order to understand the isoform-selective inhibition and reveal the key residues involved in the specific binding process, the binding free energies of the four complexes were decomposed into the individual residue contributions. And, the results were tabulated in Table S1. Here, the non-bonded interaction was defined as ΔGnonpolar = ΔEvdw + ΔGSA and the polar contributions ΔGplar = ΔEele + ΔGGB. The key interaction spectra were illustrated in Figs. 2 and 3. At first glance, the four complexes share three similar strong interaction patterns with the active site residues, as shown in Fig. 2, they are Met (772 for α, 773 for β, 752 for δ, 804 for γ), Ile (800 and 932 for α, 797 and 930 for β, 777 and 910 for δ, 831 and 963 for γ). It indicates that these residues could be critical to the bio-affinity of these inhibitors. Then, the four complexes will be analyzed one by one.
The binding mechanism of PI3Kα complex
From Fig. 3 a and c, we found that the displacement of idelalisib in the PI3Kα cavity is greater than that in the PI3Kδ cavity; the predicted binding free energy of idelalisib/PI3Kδ (pIC50 = 5.60) is − 41.65 kcal/mol, which is greater than − 30.36 kcal/mol (pIC50 = 3.09) of PI3Kα. Table 1 shows that both complexes form strong van der Waals interactions with the corresponding residues. Herein, the residues with binding energies below − 2 kcal/mol are identified to the key residues. For PI3Kα, these residues include Arg770, Met772, Pro780, Ile800, Val850, and Ile932, while in PI3Kδ, these residues are Met752, Trp760, Ile777, Glu826, Val827, Val828, Met900, and Ile910 (Fig. 2 a and c). As shown in Fig. 3c, Met900 of PI3Kδ to be parallel to the quinazolinone group of idelalisib, rather than perpendicular to the group like the corresponding Met922 of PI3Kα (Fig. 3a), that led to a stronger van der Waals interaction with Met900 (− 4.54 kcal/mol) than Met922 (− 1.82 kcal/mol) (Fig. 2e). Moreover, comparing with Thr750 of PI3Kδ, Arg770 of PI3Kα would cover the tryptophan surface formed in the PI3Kα cavity to produce a steric blockage, which prevents the inhibitor from socking into the ATP-binding pocket, it is not conducive to the combination of idelalisib and PI3Kα (Fig. 3 a and c) [12].
On the other hand, the polar contributions were taken into consideration, as shown in Table 1, the ΔEele value of PI3Kδ is much higher than PI3Kα, indicating that the hydrogen bond (H-bond) interactions may play an important role in the bonding of idelalisib and PI3Kδ. Therefore, the H-bond occupancy was calculated and the results were summarized in Table 2. Glu826 and Val828 of PI3Kδ both formed H-bonds with the inhibitor, while the corresponding Val851 and Glu849 of PI3Kα lost these H-bonds (Figs. 2f and 3a). As shown in Table 2, the averaged H-bond occupancies between Val828 and Glu826 of PI3Kδ are 60.5% and 29.7%. Val828 and Glu826 of PI3Kδ have been identified two core residues, which could both form characteristic H-bonds in almost all of the PI3Kδ/inhibitors complexes [14, 19].
The binding mechanism of PI3Kβ complex
As shown in Table 1, the nonpolar contribution of idelalisib/PI3Kβ (ΔEvdw + ΔGSA = − 49.98 kcal/mol) complexes is similar to that of idelalisib/PI3Kδ (ΔEvdw + ΔGSA = − 51.81 kcal/mol), suggesting that the nonpolar interactions play an equal role in both systems. For PI3Kβ, the key residues consist of Lys771, Met773, Trp781, Ile797, Tyr833, Glu846, Val847, Val848, Met920, and Ile930. Comparing the ΔGnonpolar of PI3Kβ and δ, Met752 and Ile777 of PI3Kδ exhibited more contribution than the corresponding Met773 and Ile797 of PI3Kβ (− 6.26 and − 5.48 versus − 5.10 and − 4.60 kcal/mol) (Table S1). As shown in Fig. 2e, the contribution of Trp781 of PI3Kβ to ΔEvdw (− 7.78 kcal/mol) is higher than corresponding residues of PI3Kδ (Trp760), which almost dominate the nonpolar contribution in PI3Kδ (− 6.60 kcal/mol, Fig. 2e), it indicates that the residue has a greater effect on the binding of the inhibitor to PI3Kβ. As mentioned in our previous work, these residues could form the “hydrophobic pocket,” and it is significant to the binding affinity rather than the selectivity for this series of compounds [37]. Besides, Met900 and Ile910 of PI3Kδ (corresponding to Met920 and Ile930 of PI3Kβ) have basically the same force, which is why the nonpolar contribution between PI3Kβ/PI3Kδ and the inhibitor are very similar. On the other hand, Val848 of PI3Kβ forms an H-bond with the ligand (Fig. 3b); the occupancy is slightly lower than Val828 of PI3Kδ, while the H-bond occupancy for Glu846 of PI3Kβ falls below 15%, suggesting that this H-bond interaction was almost lost in PI3Kβ system (Table 2).
The binding mechanism of PI3Kγ complex
The predicted binding free energy of PI3Kδ (pIC50 = 5.60) is − 41.65 kcal/mol, which is slightly greater than − 40.36 kcal/mol (pIC50 = 4.05) of PI3Kγ, that is, the selective rate between PI3Kδ/γ for idelalisib is worse than that between α or β. As shown in Fig. 2d, the key residues of PI3Kγ include Met804, Trp812, Ile831, Glu880, Val882, Met953, and Ile963. Figure 3 c and d show that both PI3Kδ and PI3Kγ formed two H-bonds with corresponding residues, but the H-bond occupancies of Glu880 and Val882 of PI3Kγ are both smaller than that of Glu826 and Val828 in PI3Kδ (Table 2). Comparing with PI3Kγ, Ile910 of PI3Kδ shows a stronger van der Waals interaction than Ile963 of PI3Kγ (− 3.44 versus − 1.86 kcal/mol). Similar to PI3Kα, Met900 and Ile910 of PI3Kδ could form hydrophobic regions with equal force, while in PI3Kγ, the nonpolar contribution of Ile963 is much lower than that of Met953, leading to an obvious decrease in inhibitor binding improvement in this “hydrophobic pocket.” In addition, the same as α and β is that Lys802 will cover the surface of the formed tryptophan, and the obstruction of this space is not conducive to the binding of the inhibitor to PI3Kγ (Fig. 3d). Therefore, the corresponding Thr750 plays a very important role in enhancing the selectivity of PI3Kδ. From Fig. 2e, the van der Waals interactions of Met752, Ile 777, and Ile910 of PI3Kδ are stronger than Met804, Ile831, and Ile963 of PI3Kγ. Generally speaking, the main residues of PI3Kδ contribute the more favorable interaction to idelalisib than PI3Kγ, but these binding free energies are not obviously different that is why idelalisib shows the poor selectivity rate to PI3Kγ.
The stronger binding affinity mechanisms of idelalisib to PI3Kδ
IC87114 is the first reported PI3Kδ inhibitor, and idelalisib was developed based on the rational modification of IC87114. As shown in Table 1, the inhibitory activity of idelalisib increases from 3.30 to 5.60 relative to IC87114; therefore, in this study, IC87114, as a lead compound, was also submitted the MD simulation to find the key residues associated with the binding affinity to PI3Kδ. The 2D structures of the two inhibitors and the energy of the key residues are shown in Fig. 4. Compared with IC87114, the thieno[3,2-d]-fused analogues retain the high valence of PI3Kδ for the mother core to a large extent. Previous studies indicated that the small substituent of isoquinolinone is essential for the activity of PI3Kδ that led to the discovery of chloroisoquinolinone motif [38]. The contribution of key amino acids to the energy of the inhibitor shows that Ile777, Val826, Val828, and Met900 are all favorable to idelalisib (Fig. 4d). Comparing the two structures, a hydrophobic group, ethyl, was introduced to idelalisib, which significantly enhances the hydrophobic interaction to the Met900 (Fig. 4 a and b). As illustrated in Fig. 4c, idelalisib and IC87114 both form two H-bonds with Glu826 and Val828, while the H-bond occupancies of IC87114 are lower than that of PI3Kδ (Table 2). These H-bond interactions may make idelalisib stronger binding affinity to PI3Kδ than IC87114.
Conclusion
In the present work, the MD simulations and binding free energy calculations were employed to investigate the selective binding mechanisms between idelalisib and PI3Ks. Firstly, the predicted binding free energies are in good agreement with the bio-activity data. And then, the contribution of individual residue to binding affinity was evaluated through the free energy decomposition analysis. The H-bond interactions between idelalisib and Glu826/Val828 play critical roles in the selective inhibition to PI3Kδ. Besides, the hydrophobic interaction with Met752, Trp760, Ile777, Met900, and Ile910 could improve the stability of the inhibitor at the binding site of PI3Kδ. Comparing with other PI3K isoforms, the residue equivalent to Thr750 of PI3Kδ is lysine or arginine. This unique structure characteristic may account for the high PI3Kδ selectivity. In a word, the specific binding of PI3Kδ is determined by the additive contributions provided by these multiple “key” residues; our studies may provide some guidance for the development of novel selective PI3Kδ inhibitors.
References
Vanhaesebroeck B, Stephens L, Hawkins P (2012) PI3K signalling: the path to discovery and understanding. Nat Rev Mol Cell Biol 13(3):195–203. https://doi.org/10.1038/nrm3290
Zhu J, Hou T, Mao X (2015) Discovery of selective phosphatidylinositol 3-kinase inhibitors to treat hematological malignancies. Drug Discov Today 20(8):988–994. https://doi.org/10.1016/j.drudis.2015.03.009
Li K, Zhu J, Xu L, Jin J (2019) Rational design of novel phosphoinositide 3-kinase gamma (PI3Kgamma) selective inhibitors: a computational investigation integrating 3D-QSAR, molecular docking and molecular dynamics simulation. Chem Biodivers 16(7):e1900105. https://doi.org/10.1002/cbdv.201900105
Zhu J, Wang M, Cao B, Hou T, Mao X (2014) Targeting the phosphatidylinositol 3-kinase/AKT pathway for the treatment of multiple myeloma. Curr Med Chem 21(27):3173–3187. https://doi.org/10.2174/0929867321666140601204513
Knight ZA (2010) Small molecule inhibitors of the PI3-kinase family. Curr Top Microbiol Immunol 347:263–278. https://doi.org/10.1007/82_2010_44
Elmenier FM, Lasheen DS, Abouzid KAM (2019) Phosphatidylinositol 3 kinase (PI3K) inhibitors as new weapon to combat cancer. Eur J Med Chem 183:111718. https://doi.org/10.1016/j.ejmech.2019.111718
Wang X, Ding J, Meng LH (2015) PI3K isoform-selective inhibitors: next-generation targeted cancer therapies. Acta Pharmacol Sin 36(10):1170–1176. https://doi.org/10.1038/aps.2015.71
Zhang Z, Liu J, Wang Y, Tan X, Zhao W, Xing X, Qiu Y, Wang R, Jin M, Fan G, Zhang P, Zhong Y, Kong D (2018) Phosphatidylinositol 3-kinase beta and delta isoforms play key roles in metastasis of prostate cancer DU145 cells. FASEB J 32(11):5967–5975. https://doi.org/10.1096/fj.201800183R
Lannutti BJ, Meadows SA, Herman SE, Kashishian A, Steiner B, Johnson AJ, Byrd JC, Tyner JW, Loriaux MM, Deininger M, Druker BJ, Puri KD, Ulrich RG, Giese NA (2011) CAL-101, a p110delta selective phosphatidylinositol-3-kinase inhibitor for the treatment of B-cell malignancies, inhibits PI3K signaling and cellular viability. Blood 117(2):591–594. https://doi.org/10.1182/blood-2010-03-275305
Okkenhaug K, Burger JA (2016) PI3K signaling in normal B cells and chronic lymphocytic Leukemia (CLL). Curr Top Microbiol Immunol 393:123–142. https://doi.org/10.1007/82_2015_484
Nunes-Santos CJ, Uzel G, Rosenzweig SD (2019) PI3K pathway defects leading to immunodeficiency and immune dysregulation. J Allergy Clin Immunol 143(5):1676–1687. https://doi.org/10.1016/j.jaci.2019.03.017
Perry MWD, Abdulai R, Mogemark M, Petersen J, Thomas MJ, Valastro B, Westin Eriksson A (2019) Evolution of PI3Kgamma and delta inhibitors for inflammatory and autoimmune diseases. J Med Chem 62(10):4783–4814. https://doi.org/10.1021/acs.jmedchem.8b01298
Zhu J, Ke K, Xu L, Jin J (2019) Theoretical studies on the selectivity mechanisms of PI3Kdelta inhibition with marketed idelalisib and its derivatives by 3D-QSAR, molecular docking, and molecular dynamics simulation. J Mol Model 25(8):242. https://doi.org/10.1007/s00894-019-4129-x
Somoza JR, Koditek D, Villasenor AG, Novikov N, Wong MH, Liclican A, Xing W, Lagpacan L, Wang R, Schultz BE, Papalia GA, Samuel D, Lad L, McGrath ME (2015) Structural, biochemical, and biophysical characterization of idelalisib binding to phosphoinositide 3-kinase delta. J Biol Chem 290(13):8439–8446. https://doi.org/10.1074/jbc.M114.634683
Miller MS, Schmidt-Kittler O, Bolduc DM, Brower ET, Chaves-Moreira D, Allaire M, Kinzler KW, Jennings IG, Thompson PE, Cole PA, Amzel LM, Vogelstein B, Gabelli SB (2014) Structural basis of nSH2 regulation and lipid binding in PI3Kalpha. Oncotarget 5 (14):5198-5208. doi:10.18632/oncotarget.2263
Zhang X, Vadas O, Perisic O, Anderson KE, Clark J, Hawkins PT, Stephens LR, Williams RL (2011) Structure of lipid kinase p110beta/p85beta elucidates an unusual SH2-domain-mediated inhibitory mechanism. Mol Cell 41(5):567–578. https://doi.org/10.1016/j.molcel.2011.01.026
Shin Y, Suchomel J, Cardozo M, Duquette J, He X, Henne K, Hu YL, Kelly RC, McCarter J, McGee LR, Medina JC, Metz D, San Miguel T, Mohn D, Tran T, Vissinga C, Wong S, Wannberg S, Whittington DA, Whoriskey J, Yu G, Zalameda L, Zhang X, Cushing TD (2016) Discovery, optimization, and in vivo evaluation of benzimidazole derivatives AM-8508 and AM-9635 as potent and selective PI3Kdelta inhibitors. J Med Chem 59(1):431–447. https://doi.org/10.1021/acs.jmedchem.5b01651
Wang J, Wolf RM, Caldwell JW, Kollman PA, Case DA (2004) Development and testing of a general amber force field. J Comput Chem 25(9):1157–1174. https://doi.org/10.1002/jcc.20035
Berndt A, Miller S, Williams O, Le DD, Houseman BT, Pacold JI, Gorrec F, Hon WC, Liu Y, Rommel C, Gaillard P, Ruckle T, Schwarz MK, Shokat KM, Shaw JP, Williams RL (2010) The p110 delta structure: mechanisms for selectivity and potency of new PI(3)K inhibitors. Nat Chem Biol 6(2):117–124. https://doi.org/10.1038/nchembio.293
Case DA, Cheatham 3rd TE, Darden T, Gohlke H, Luo R, Merz Jr KM, Onufriev A, Simmerling C, Wang B, Woods RJ (2005) The Amber biomolecular simulation programs. J Comput Chem 26(16):1668–1688. https://doi.org/10.1002/jcc.20290
Stewart JJ (2004) Optimization of parameters for semiempirical methods IV: extension of MNDO, AM1, and PM3 to more main group elements. J Mol Model 10(2):155–164. https://doi.org/10.1007/s00894-004-0183-z
Stewart JJ (2013) Optimization of parameters for semiempirical methods VI: more modifications to the NDDO approximations and re-optimization of parameters. J Mol Model 19(1):1–32. https://doi.org/10.1007/s00894-012-1667-x
Zhu J, Wu Y, Xu L, Jin J (2020) Theoretical studies on the selectivity mechanisms of glycogen synthase kinase 3beta (GSK3beta) with pyrazine ATP-competitive inhibitors by 3DQSAR, molecular docking, molecular dynamics simulation and free energy calculations. Curr Comput Aided Drug Des 16(1):17–30. https://doi.org/10.2174/1573409915666190708102459
Zhu J, Li K, Xu L, Jin J (2019) Insight into the selective mechanism of phosphoinositide 3-kinase gamma with benzothiazole and thiazolopiperidine gamma-specific inhibitors by in silico approaches. Chem Biol Drug Des 93(5):818–831. https://doi.org/10.1111/cbdd.13469
Zhu J, Ke K, Xu L, Jin J (2019) Discovery of a novel phosphoinositide 3-kinase gamma (PI3Kγ) inhibitor against hematologic malignancies and theoretical studies on its PI3Kγ-specific binding mechanisms. RSC Advances 9(35):20207–20215. https://doi.org/10.1039/c9ra02649e
Xu L, Sun H, Li Y, Wang J, Hou T (2013) Assessing the performance of MM/PBSA and MM/GBSA methods. 3. The impact of force fields and ligand charge models. J Phys Chem B 117(28):8408–8421. https://doi.org/10.1021/jp404160y
Sun H, Li Y, Shen M, Tian S, Xu L, Pan P, Guan Y, Hou T (2014) Assessing the performance of MM/PBSA and MM/GBSA methods. 5. Improved docking performance using high solute dielectric constant MM/GBSA and MM/PBSA rescoring. Phys Chem Chem Phys 16(40):22035–22045. https://doi.org/10.1039/c4cp03179b
Wang E, Sun H, Wang J, Wang Z, Liu H, Zhang JZH, Hou T (2019) End-point binding free energy calculation with MM/PBSA and MM/GBSA: strategies and applications in drug design. Chem Rev 119(16):9478–9508. https://doi.org/10.1021/acs.chemrev.9b00055
Sun H, Duan L, Chen F, Liu H, Wang Z, Pan P, Zhu F, Zhang JZH, Hou T (2018) Assessing the performance of MM/PBSA and MM/GBSA methods. 7. Entropy effects on the performance of end-point binding free energy calculation approaches. Phys Chem Chem Phys 20(21):14450–14460. https://doi.org/10.1039/c7cp07623a
Xie T, Yu J, Fu W, Wang Z, Xu L, Chang S, Wang E, Zhu F, Zeng S, Kang Y, Hou T (2019) Insight into the selective binding mechanism of DNMT1 and DNMT3A inhibitors: a molecular simulation study. Phys Chem Chem Phys 21(24):12931–12947. https://doi.org/10.1039/c9cp02024a
Chohan TA, Chen JJ, Qian HY, Pan YL, Chen JZ (2016) Molecular modeling studies to characterize N-phenylpyrimidin-2-amine selectivity for CDK2 and CDK4 through 3D-QSAR and molecular dynamics simulations. Mol Biosyst 12(4):1250–1268. https://doi.org/10.1039/c5mb00860c
Bharadwaj VS, Dean AM, Maupin CM (2013) Insights into the glycyl radical enzyme active site of benzylsuccinate synthase: a computational study. J Am Chem Soc 135(33):12279–12288. https://doi.org/10.1021/ja404842r
Kong X, Sun H, Pan P, Tian S, Li D, Li Y, Hou T (2016) Molecular principle of the cyclin-dependent kinase selectivity of 4-(thiazol-5-yl)-2-(phenylamino) pyrimidine-5-carbonitrile derivatives revealed by molecular modeling studies. Phys Chem Chem Phys 18(3):2034–2046. https://doi.org/10.1039/c5cp05622e
Zhao S, Zhu J, Xu L, Jin J (2017) Theoretical studies on the selective mechanisms of GSK3beta and CDK2 by molecular dynamics simulations and free energy calculations. Chem Biol Drug Des 89(6):846–855. https://doi.org/10.1111/cbdd.12907
Hou T, Wang J, Li Y, Wang W (2011) Assessing the performance of the MM/PBSA and MM/GBSA methods. 1. The accuracy of binding free energy calculations based on molecular dynamics simulations. J Chem Inf Model 51(1):69–82. https://doi.org/10.1021/ci100275a
Xue W, Liu H, Yao X (2012) Molecular mechanism of HIV-1 integrase-vDNA interactions and strand transfer inhibitor action: a molecular modeling perspective. J Comput Chem 33(5):527–536. https://doi.org/10.1002/jcc.22887
Zhu J, Pan P, Li Y, Wang M, Li D, Cao B, Mao X, Hou T (2014) Theoretical studies on beta and delta isoform-specific binding mechanisms of phosphoinositide 3-kinase inhibitors. Mol Biosyst 10(3):454–466. https://doi.org/10.1039/c3mb70314b
Wei M, Wang X, Song Z, Jiao M, Ding J, Meng LH, Zhang A (2015) Targeting PI3Kdelta: emerging therapy for chronic lymphocytic leukemia and beyond. Med Res Rev 35(4):720–752. https://doi.org/10.1002/med.21341
Funding
The study was supported by the National Natural Science Foundation of China (No. 21807049), the Fundamental Research Funds of Changzhou Vocational Institute of Engineering (11130300117010), the Fundamental Research Funds for the Central Universities (JUSRP51703A), and the Top-notch Academic Programs Project of Jiangsu Higher Education Institutions (PPZY2015B146).
Author information
Authors and Affiliations
Contributions
Z.J., L.H., and J.J. developed the study concept and design. Z.H. and S.H. performed the modeling studies. Z.H., Y.L., and C.Y. carried out the data analysis. Z.J., Z.H., and Y.L. drafted the manuscript, C.Y., L.H., and J.J. approved the manuscript.
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Jingyu Zhu, Haoer Zhang and Li Yu are Equivalent authors
Electronic supplementary materials
ESM 1
(DOCX 19 kb)
Rights and permissions
About this article
Cite this article
Zhu, J., Zhang, H., Yu, L. et al. Computational investigation of the selectivity mechanisms of PI3Kδ inhibition with marketed idelalisib: combined molecular dynamics simulation and free energy calculation. Struct Chem 32, 699–707 (2021). https://doi.org/10.1007/s11224-020-01643-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11224-020-01643-4