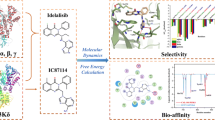
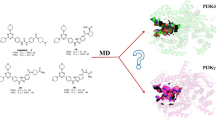
Abstract
Phosphoinositide 3-kinases (PI3Ks) are crucial for cell proliferation, metabolism, motility, and cancer progression. Since the selective PI3Kδ inhibitor, idelalisib, was firstly approved by the FDA in 2014, large numbers of selective PI3Kδ inhibitors have been reported, but the detailed mechanisms of selective inhibition to PI3Kδ for idelalisib or its derivatives have not been well addressed. In this study, 3D-QSAR with COMFA, molecular docking, and molecular dynamic (MD) simulations was used to explore the binding modes between PI3Kδ and idelalisib derivatives. Firstly, a reliable COMFA model (q2 = 0.59, ONC = 8, r2 = 0.966) was built and the contour maps showed that the electrostatic field had more significant contribution to the bioactivities of inhibitors. Secondly, two molecular docking methods including rigid receptor docking (RRD) and induced fit docking (IFD) were employed to predict the docking poses of all the studied inhibitors and revealed the selective binding mechanisms. And then, the results of the MD simulation and the binding free energy decomposition verified that the binding of PI3Kδ/inhibitors was mainly contributed from hydrogen bonding and hydrophobic interactions and some key residues for selective binding were highlighted. Finally, based on the models developed, 14 novel inhibitors were optimized and some showed satisfactory predicted bioactivity. Taken together, the results provided by this study may facilitate the rational design of novel and selective PI3Kδ inhibitors.

.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Since the 1980s, PI3Ks (phosphoinositide 3-kinases) have been regarded as important drug targets in the cancer research field, and they regulate various biological processes, such as cell growth, metabolism, survival, and motility, and play important roles in the progression of cancer [1,2,3,4,5,6,7]. According to the specific structure and lipid substrate specificity, PI3Ks can be divided into three classes (classes I, II, and III), and class I PI3Ks, the most extensively investigated class, can be further divided into class IA (PI3Kα, PI3Kβ, and PI3Kδ) and class IB (PI3Kγ) enzymes. The class IA PI3Ks is mostly activated by RTKs (receptor tyrosine kinases), while the class IB is mainly activated by GPCRs (G protein-coupled receptors) [8]. The activated class I PI3Ks can phosphorylate the 3′OH position of PIP2 to PIP3, which would transmit intracellular signal to downstream signaling proteins with the PH (pleckstrin homology) domain, like AKT and PDK1. PI3Kα and PI3Kβ are widely expressed in tissues, while PI3Kδ and PI3Kγ are mainly expressed in leukocytes and thymus [9, 10].
Accumulated molecular researches have revealed that the PI3K signaling pathway is involved in various cancers, such as breast, colorectal, and lung cancer, and endometrium and hematological malignancies [11,12,13], suggesting that PI3Ks are ideal targets for anti-cancer drug discovery. Up to now, a large number of PI3K inhibitors have been developed, and some of these have been pushed into clinical trials. Moreover, isoform-selective inhibitors show lower toxicity and higher therapeutic efficacy than pan-inhibitors [13,14,15]. Idelalisib (CAL-101), the first listed selective PI3Kδ inhibitor, was approved by the FDA in 2014 for the treatment of chronic lymphocytic leukemia and small lymphocytic lymphoma, which greatly promotes the exploitation of selective PI3Kδ inhibitors [16]. Recently, another selective PI3K inhibitor, duvelisib (IPI-145, δ/γ inhibitor), was approved by the FDA for the treatment of hematological malignancies [17, 18]. Nowadays, many PI3Kδ inhibitors have been reported and some of them have been entered into clinical trials, but the detailed mechanisms of selective inhibition to PI3Kδ for these inhibitors have not been well elucidated.
Computer-aided drug design (CADD) methods, including quantitative structure-activity relationship (QSAR) analysis, molecular docking, molecular dynamics (MD) simulations, and free energy calculations, have been widely used to design or discover novel small molecule inhibitors [14, 19,20,21,22,23,24,25,26,27,28,29,30,31,32,33,34]. Therefore, in this study, 3D-QSAR, molecular docking, and MD simulations were used to reveal the mechanisms of PI3Kδ binding selectivity through exploring the relationship between PI3Kδ and a series of potent PI3Kδ inhibitors which are transformed from the marketed drug idelalisib. Our findings may assist in designing or discovering more novel selective PI3Kδ inhibitors.
Materials and methods
3D-QSAR with COMFA
In this work, the 3D-QSAR model with COMFA was built by Sybyl-X 2.0. All the 41 quinazolinone derivatives as the studied PI3Kδ inhibitors were collected from the literature reported by Leena Patel [35] and the 2D structures with their affinities of PI3Kδ are shown in Table S1. The bioactivity defined as pIC50 (-logIC50) was used as the dependent variable in the QSAR study. The 3D structures of all the inhibitors were sketched in Maestro and then minimized with the Powell method under the Tripos force field with the Gasteiger-Huckel charge. The 41 inhibitors were spilt into a training set with 31 compounds for creating the QSAR model and a test set with 10 compounds for assessing the quality of the model based on the structural feature and the distribution of biological data through the Generate Training and Test Data module of Discovery Studio 3.5 (DS3.5). All the inhibitors were aligned onto the common substructure (Fig. 1a) through the Align Database module and the crystal structure of Cpd24 was selected as the template, and the molecular alignments are shown in Fig. S1. During the calculation of the parameters of COMFA, steric and electrostatic potentials of a compound were evaluated beyond every 2 Å lattice point with Tripos standard force field and other parameters were set default. After adding the parameters of COMFA, a leave-one-out cross-validation was firstly used to determine the optimal number of components (ONC) in the Partial Least Squares Analysis module. The last step was to perform a no-validation analysis with ONC to generate the COMFA model.
a The alignment of the common template structure which is based on Cpd24. b Plot of the experimental versus predicted pIC50 of training sets and test sets. c Electrostatic and d steric contour maps for the COMFA model of Cpd3. More positive charges near the blue regions increase activity; more negative charges near the red regions increase activity. Bulk groups near the green regions increase activity; bulk groups near the yellow regions decrease activity
Molecular docking
Two different molecular docking methods including rigid receptor docking (RRD) and induced fit docking (IFD) were employed to predict the binding mode between PI3Kδ and the inhibitors. Crystal structure of PI3Kδ/Cpd24 (PDB ID: 5I6U) was used as initial receptor for subsequent molecular docking analysis. The Protein Preparation Wizard module in Maestro was employed to prepare the complex to assign bond orders, add hydrogen atoms, assign protonated states, delete unwanted chains and waters, assign partial charges, and lastly, minimize with the OPLS-2005 force field. All 41 inhibitors were prepared with the LigPrep module with the OPLS-2005 force field. During the period of preparation, each molecule would generate right protonated states at target PH from 5.0 to 9.0, stereochemistry, tautomers, and low-energy ring conformations to output the most stable conformation. After that, a bounding box with the size of 10 Å × 10 Å × 10 Å, which was defined in the studied receptor and centered on the co-crystal ligand to confine the ligands to the enclosing active site, was built through the Receptor Grid Generation Panel of Maestro. Firstly, the RRD was carried on with Glide and the parameters were default except for writing out at most 10 poses for each ligand, where the atoms of ligands were applied to the scaling factor of 0.8 for van der Waals radii with partial atomic charges less than or equal to 0.25 to soften the potential for non-polar parts of the ligands. After finishing the docking calculations by using XP (extra precision) scoring modes, the best docked structure of each ligand was selected for analysis. Secondly, in order to consider the flexibility of both ligand and receptor, IFD in Schrödinger was employed, which allowed the receptor to flexibly move the backbone or side chain to transform its binding site in order that it more closely combined to the shape of the ligand [36]. During the process of IFD, each ligand was firstly docked into the rigid receptor using a softened potential by the Glide model with the default of holding a maximum 20 poses per ligand. Then each receptor-ligand complex was sampled with the freedom of receptor degrees and minimized under the OPLS-2005 force field. Now the structure of the receptor in each pose was induced fit to the structure and conformation of the ligand, and the residues within 5 Å of each docked pose of the best receptor-ligand complex identified by the predicted binding affinities were refined. Finally, each receptor-ligand complex was redocked within a specified energy of the lowest-energy structure (default 30 kcal/mol) where the ligand was strictly docked into the induced fit structure of the receptor. The XP scoring mode was used for all docking calculations.
Molecular dynamics (MD) simulation
The crystal structures PI3Kδ/Cpd24 complex (PDB ID: 5I6U) and PI3Kδ/Cpd3 complex with the worst experimental activity generated by the IFD module were used as initial structures for the next MD simulations. On the other hand, as the template structure for optimization of these studied inhibitors, idelalisib, the first listed PI3Kδ inhibitor, was also taken into account and the PI3Kδ/idelalisib crystal structure (PDB ID: 4XE0) was retrieved from the Protein Data Bank [37]. All three complexes for the MD simulation were carried out by using the SANDER program in AMBER14 [38]. The AMBER ff03 force field [39] and general AMBER force field [40] were applied for protein and ligands, respectively. The semi-empirical AM1 method in Gaussian09 [41] was used to optimize each ligand. After that, the restrained electrostatic potential (RESP) fitting technique was employed to fit the electrostatic potential of the atomic partial charges at HF/6-31G* level [42]. Each complex was neutralized with sodium ions and solvated in an extended 10 Å of rectangular box suffused with the TIP3P water molecules. The PME (particle mesh Ewald) scheme [43] was applied to handle the long-range electrostatics, with a cutoff of 10 Å for the van der Waals interactions. All bonds referring to hydrogen atoms were restricted by the SHAKE algorithm [44], with the time step set to 2.0 fs.
There were three-step minimizations applied for relaxing each system before the MD simulations. Firstly, the backbone carbons of protein were restrained (50 kcal/mol/Å2) during 500 cycles of steepest descent and 500 cycles of conjugate gradient. Secondly, 1000 cycles of minimizations with a weaker harmonic potential (10 kcal/mol/Å2) were performed. Finally, 5000 cycles of minimizations (1000 cycles of steepest descent and 4000 cycles of conjugate gradient) were used for relaxing the whole system without any restrain. In the process of the MD simulation, the temperature of each system was successively up from 0 to 300 K over 50 ps. After a 50-ps simulation with the NPT ensemble (300 K, 1 atm) performed in each system, 30-ns NPT simulations were finally performed.
MM/GBSA free energy calculations and decomposition
The snapshots of each system extracted from the last 10-ns stable MD trajectory were applied for calculating the binding free energy (ΔGbind) using the MM/GBSA method according to Eq. 1 [37, 45,46,47,48,49,50,51,52,53,54,55]:
For each complex, the protein-ligand interaction spectrum on a per-residue basis was calculated by the MM/GBSA free energy decomposition analysis using the MM-PBSA program of AMBER 14, which was computed based on Eq. 2:
where the molecular mechanics energy (ΔGMM) is the gas-phase interaction energy between ligand and protein, calculating by the electrostatic (ΔGele) and the van der Waals interaction (ΔGvdw). The solvation free energy (ΔGsol) is composed of the polar (ΔGGB) and the non-polar (ΔGSA) contributions. ΔGGB is calculated by the generalized Born (GB) model developed by Onufriev et al. [56]. ΔGSA is computed on the basis of SASA (solvent-accessible surface area) by using a fast LCPO (linear combination of the pairwise overlap) algorithm with a probe radius of 1.4 Å [57]. – TΔS is the change in the conformational entropy upon ligand binding [10].
Results and discussion
COMFA model analysis
As we know, reasonable alignment of samples and bioactive conformation selection are two vital factors for obtaining robust and meaningful 3D-QSAR models. So Cpd24 retrieved from the crystal structure (ID: 5I6U) was chosen as the template, and the common substructure for alignment was showed in Fig. 1a. Partial least squares (PLS) regression analysis was used to correlate COMFA fields with the activity value of inhibitors and the reliability of this model is validated by using several especially statistical parameters including q2 (cross-validation coefficient), r2 (non-cross validation coefficient), SEE (standard estimate difference), and F values. When q2 > 0.5, r2 > 0.9, and r2pred > 0.5, the QSAR model has an effective statistical criterion. In this study, all predicted and experimental pIC50 values were listed in the Table S1. The COMFA model provides q2 of 0.59, ONC of 8, r2 of 0.966, SEE of 0.204, and F values of 91.292. As shown in Fig. 1b, the linear correlation r2pred between the experimental and the predicted pIC50 values of the test set is 0.719, indicating that this model is good enough to predict the bioactivities of inhibitors. The contributions of the electrostatic and steric fields are 65.4% and 34.6% respectively (Fig. 1c, d), which suggest that the electrostatic field contributes most to the bioactivity of inhibitors. In the COMFA model, the StDev*Coeff contour map is built to help visualize why some inhibitors have high experimental activities. The Cpd3, with the worst experimental activity, was chosen to illustrate all contour maps of the COMFA models. In Fig. 1c, the blue regions, namely electrostatic contour representing 80% level contributions, mean that importing positive groups near this position favored the activity of inhibitor, while that is unfavorable when negative groups are imported near the red regions representing 20% level contributions. According to Fig. 1c, we can visually see that there is a large blue contour encompassing the N3 position, C4 position, and C2 position of the pyrimidine ring, which could explain why Cpd14 (pIC50 = 10.00) with the positive amino in the C2 position shows higher activity than Cpd15 (pIC50 = 8.37) with the negative chlorine atom, and why Cpd2 (pIC50 = 7.00) with the positive amino in the C4 position shows higher activity than Cpd3 (pIC50 = 5.70) without any positive group. Besides, the blue regions located upon the C2′ position, C3′ position, and C5′ position of the phenyl ring suggest that the positive groups would also enhance the bioactivity. Due to the introduction of the negative fluorine atom into the C3′ position of the phenyl ring, Cpd25 (pIC50 = 8.70) shows lower activity than Cpd14. The red regions map under the C5 position of the pyrimidine ring (showed in Fig. 1c), which could illustrate the higher activity of Cpd5 (pIC50 = 9.40) with negative cyano in comparison with Cpd10 (pIC50 = 7.85) with methyl. On the other hand, the steric contour of the COMFA model is showed in Fig. 1d, where the green (namely positive bulk groups) and yellow (namely negative bulk groups) contours respectively represent 80% and 20% level contributions. A large green region is located under the area between the extension of the C5 position and C4 position of the pyrimidine ring (showed in Fig. 1d), which advises that bulk groups could increase the activity. For example, Cpd4 and Cpd2 both have a cyano or amino substituent at these two locations, respectively, which observably increases the activity (pIC50 = 8.10 and pIC50 = 7.00) in comparison with Cpd3 (pIC50 = 5.77). And Cpd5 shows higher activity (pIC50 = 9.40) than Cpd4 or Cpd2, owing to the replacement of the hydrogen atom on the C4 position and C5 position with two bulk groups. Compared with the activity of Cpd15, we find Cpd14 with the C2 position amino (pIC50 = 10) shows better activity than Cpd15 with the C2 position chlorine atom (pIC50 = 8.37). This is in good agreement with the yellow region mapped near the C2 position (showed in Fig. 1d), which suggests that the introduction of bulk groups may decrease the activity.
From the above analysis of the COMFA model, some key clues about the structure activity relationship of these PI3Kδ inhibitors could be found: positive and small substituents rather than negative and large groups should be introduced into the C2′ position and C3′ position of the phenyl ring; the C2 position and N3 position of the pyrimidine ring are favored with small and positive groups; the C4 position and C5 position of the pyrimidine ring are in favor of large substituents, and it will be better when the negative bulks are introduced into the C5 position.
Molecular docking analysis
Herein, in order to generate the appropriate binding orientations and conformations of the active molecules, molecular docking protocol was employed to predict the docking poses of the studied inhibitors. Rigid receptor docking (RRD) with Glide was firstly employed. To validate whether the performance of the Glide was accurate for studying the interaction between inhibitors and PI3Kδ, Cpd24 was extracted from the crystal structure of PI3Kδ (ID: 5I6U) and then redocked into the binding site of PI3Kδ. As shown in Fig. S2a, the redocked complex was followed aligned with its initial crystal structure, and the RMSD (root mean square deviation) values between the redocked complex and the initial crystal structure are only 0.17 Å for all atoms of PI3Kδ, which means that the Glide program is capable of identifying the active site of the initial receptor and predicting the correct conformations of the inhibitors docked into the PI3Kδ. As a potent PI3Kδ-selective inhibitor, Cpd24 shows stronger interaction with PI3Kδ. As shown in Fig. 2a, in the adenine pocket, known as the hinge pocket, residues 825–827 form the hinge between the C-lobe and the N-lobe of the catalytic domain [58]. The aminopyrimidine Cpd24 can form two hydrogen bonds with the hinge residues Glu826 and Val828, and form an arene-hydrogen bond with another hinge residue Ile825 which is also called as the “Gatekeeper” residue [59]. And the pyrazole ring of Cpd24 forms two hydrogen bonds with Lys779 and Asp787 in the affinity pocket of PI3Kδ. These hydrogen bonds both assist Cpd24 in wrapping into the active pocket of PI3Kδ, all above of which show the importance of hydrophobic interactions for binding and that could explain why Cpd24 owns high experimental inhibition to PI3Kδ. The worst binding modes of bio-effective inhibitor Cpd3 and the highest bio-effective inhibitor Cpd14 are showed in Fig. S3. Around the 2,4-aminopyrimidine ring of Cpd14 (showed in Fig. S3b), the C2-amino and N3 position nitrogen all combine the residue Val828 with a hydrogen bond. Besides, the C4-amino substitution of Cpd14 also forms a hydrogen bond with the residue Glu826. The N3 position nitrogen on the pyrimidine ring of Cpd3 (showed in Fig. S3a) interacts with the residue Val828 with a hydrogen bond. We could find the backbone of hinge residue Val828 establishes hydrogen bonds to all three inhibitors which means the hinge residue Val828 is a foundational residue for binding between PI3Kδ and inhibitors [58, 59]. However, the Glide docking scores correlated with the experimental activities do not show good linear relevancy (r2 = 0.27, showed in Fig. 2b). These could be that most of the inhibitors binding with the rigid receptor are not precisely predicted by the Glide docking, which suggests considering the flexibility of protein into docking analysis.
a The 2D representation of the interactions between PI3Kδ and Cpd24 by Glide docking. b The linear relation between the experimental activities and docking scores showed by Glide docking. c The 2D representation of the interactions between PI3Kδ and Cpd24 by IFD. d The linear relation between the experimental activities and docking scores showed by IFD. The binding modes between the PI3Kδ and e Cpd14 and f Cpd3, using IFD
So IFD was chosen to calculate the binding modes of all inhibitors, which expressed an inspiring result showed in Fig. 2d. The predicted scores obtained by IFD show great linear correlation with the experimental activities (r2 = 0.62) and are obviously greater than RRD (r2 = 0.27), which means the flexibility of protein is crucial for binding between PI3Kδ and inhibitors. Similarly, the RMSD values between the IFD complex of Cpd24 and the initial crystal structure are 0.18 Å for all atoms of PI3Kδ (Fig. S2b), which still exhibits good alignment in active site with the initial crystal structure. Following the movement of PI3Kδ, there are more residues around Cpd24 in active pocket (showed in Fig. 2c), especially residues Met752 and Trp760 in the “specificity” pocket where there is a hydrophobic region sandwiching the inhibitor between Met752 and Trp760, which could improve the binding affinity and selectivity of Cpd24 [10, 37, 60], while the Cpd14 obtains a better IFD score of − 13.639 than − 13.079 of Cpd24, showing a stronger binding mode with the PI3Kδ (showed in Fig. 2e). In the “specificity” pocket, residue Met752 against the chlorine-quinazolinone ring of Cpd14, the side chain residue Trp760 forms arene-hydrogen bonds with the paralleled chlorine-quinazolinone ring of Cpd14, and the chlorine-quinazolinone ring contacts with the residue Thr750 with a hydrogen bond. These three residues induce Cpd14 more sandwiched into the “specificity” pocket, which notably improves the affinitive between Cpd14 and PI3Kδ comparing Cpd24. For Cpd3 with the worst IFD score of − 2.976, there is only one residue Trp760 located upon the chlorine-quinazolinone ring in the “specificity” pocket (showed in Fig. 2f), which could not form normal sandwich between PI3Kδ and the inhibitor. So the “specificity” pocket may be key for binding affinities of PI3Kδ/inhibitor. Besides, as shown in Fig. 2e, the hinge residue Val828 combines with the C2 position amino Cpd14 forming a hydrogen bond, and the C4 position amine forms a sturdy hydrogen bond with the hinge residue Glu826, which is similar with Cpd24 except the “Gatekeeper” residue Ile825 located more closer beside other two hinge residues Glu826 and Val828 at the adenine pocket in contacting with Cpd14 than Cpd24. This difference of the “Gatekeeper” residue Ile825 may be another reason for Cpd14 owning stronger binding affinity with the PI3Kδ. As shown in Fig. 2e, there is a hydrophobic pocket (also known as hydrophobic region II) formed by Met900 and Ile910 binding with Cpd14 [37], which is the same for Cpd3 binding PI3Kδ using hydrogen bonding with the residues Ile910 and Met900 (showed in Fig. 2f). And from Fig. 2e, we could find that there is an acidic amino acid Asp832 near the C2′ position and C3′ position of the phenyl ring of Cpd3, which could explain why the positive and small substituents are welcomed into this area, while Cpd3 obtains a lowest docking score, which could be caused by the vanishment of the interaction with hinge residue Val828. It is interesting to note those important residues binding with compounds are non-polar and hydrophobic, which suggests the importance of non-polar and hydrophobic interactions for PI3Kδ/inhibitor binding.
MD simulations and binding free energy calculations
To discover the binding modes between the inhibitors and PI3Kδ during the dynamic binding process, MD simulations were performed for two crystal structures of the first listed inhibitor idelalisib/PI3Kδ complex and PI3Kδ/Cpd24 complex, and also for the worst experimental activity Cpd3/PI3Kδ prepared by IFD. The RMSD were calculated and exhibited in Fig. S4, which showed the three studied systems almost reached an equilibrium state after 30 ns of the MD simulations. In addition, the RMSD of heavy atoms of ligands were also calculated, as shown in Fig. 3a; all ligands reached equilibrium state in the last 10 ns. In order to investigate the dynamical properties of the ligands, the MD ligands were all aligned onto the corresponding apo-ligands and the results were illustrated in Fig. 3b, c, and d. Among these ligands, Cpd3 showed the largest flexibility in the binding site (Fig. 3d) that may be caused by the lower binding affinity between Cpd3 and PI3Kδ. On the contrary, Cpd24, with the highest bioactivity, exhibited better structural stability, which indicated that Cpd24 may form the strongest interaction with PI3Kδ. These results were in agreement with the experimental data and the docking results, but more details are still needed for further analysis.
Based on the 1000th-ps snapshot extracted from the last 10-ns stable MD trajectory of three systems, the binding free energies were calculated. As shown in Table 1, the predicted binding free energies (ΔGpred) are − 50.71 kcal/mol for PI3Kδ/Cpd24, − 37.42 kcal/mol for PI3Kδ/idelalisib, and − 29.34 kcal/mol for PI3Kδ/Cpd3. PI3Kδ than idelalisib and Cpd3. And the linear relationship (r2) between the predicted binding free energies and experimental activities is 0.97, suggesting the predictions are in good agreement with the experimental activities. To ascertain the prime contributors to inhibitor binding, we also calculated the linear correlation (r2) between different energy components of each complex and their predicted binding free energies (ΔGpred). According to Table 1, ΔGvdw (van der Waals interaction) is the largest contributor to the inhibitor binding (r2 = 0.997). The non-polar contributions (ΔGvdw + ΔGSA) of PI3Kδ/Cpd24, PI3Kδ/idelalisib, and PI3Kδ/Cpd3 are − 61.99, − 47.08, and − 39.79 kcal/mol, respectively, which nearly predominates complex binding (r2 = 0.997) in comparison with the polar contributions (ΔGele + ΔGGB) with r2 = 0.397. Therefore, the ΔGvdw and the non-polar contribution are the foundation of favorable binding free energy, which is positive with our QSAR and the docking analysis. As shown in Fig. 2e and f, Cpd14 with two positive amino substituents and a bulk hydrophobic cyano in the pyrimidine ring shows observable improvement on the experimental activity than Cpd3, which could be due to the high increase of van der Waals interactions and non-polar interactions with some important residues like Val828, Glu826, Ile910, etc.
In order to understand the binding mode of PI3Kδ/inhibitor, the binding free energy of each complex was decomposed into the inhibitor-residue interaction spectra (Fig. S5), which should be noted that we only concentrated on the important residues within the ATP-binding pocket of PI3Kδ ranging from 540 (Leu740) to 717 (Gly917). In general, the residue-inhibitor interaction spectrum for each inhibitor is similar, especially the residues Met752, Trp760, Ile777, Val827, and Val828. From the binding mode between the PI3Kδ and three inhibitors (Fig. 4a, b, and c), we could find the pyrimidine rings of all the inhibitors are embraced with the hinge residues Val827 and Val828 forming van der Waals interactions, where the adenine pocket is. Besides, the interesting thing is that the quinazolinone rings of the inhibitors are all sandwiched into the specificity hydrophobic pocket between the residues Trp760 or/and Ile777 on one side and P-loop residue Met752 on the other side (Fig. 4a, b, and c). In order to evaluate how the flexibility of the pocket residues affected binding affinity, the surface presentations of PI3Kδ binding pockets were illustrated in Fig. 4d, e, and f, and the alignments of each apo-ligand and corresponding MD ligand could be more convenient to observe. Figure 4b shows that the chlorine-quinazolinone ring of Cpd24 is more parallel with the residues Trp760, Ile777, and Met752, which may be one factor for notable improvement in the experimental activities of Cpd24 (pIC50 = 9.52) compared with Cpd3 (pIC50 = 5.77) and idelalisib (pIC50 = 7.72). These three residues are significant for improving the binding affinities of the PI3Kδ/inhibitor complex. And compared with apo-structure, MD-Cpd24 was soaked deeper into the pocket (Fig. 4e); the yellow surface exhibited that the cavity of PI3Kδ binding pocket was correspondingly distended a little. But because of the highest binding affinity of Cpd24, the dynamic change was not obvious (Fig. 3a and c). The great impacts on the movements of active pocket residues with the binding of a compound could be obviously presented in Cpd3/PI3Kδ complex (Fig. 4f). We could find the whole binding domain displayed a wide range of upward movement, while the residue Val828 and the residue Met900 were more close to the ligand than the apo-Cpd3; it led that the hinge domain and the hydrophobic domain [37] formed by Met900 are linked together, while the hydrophobic region II [37] formed by Ile910 and Met900 is only completed in the Cpd24 system (Fig. 4e), and this region may be too large to keep Cpd3 stabilization, thus resulting in the greater flexibility of Cpd3 (Fig. 3d). This hydrophobic region could be another crucial factor for the improving affinitive of Cpd24 binding with PI3Kδ. As shown in Fig. 4d, the MD-idelalisib was tilted a little to the upper right comparing the initial structure, which accordingly induced the upward movements of Trp760 and Met900 residues that made the pocket enlarge to accommodate the combination of idelalisib. Moreover, we find that Cpd24 contacts with more residues than the other two compounds (Fig. 4b), such as the “Gatekeeper” residue Ile825, and hinge Glu826. Those residues are important for stabilizing the binding of PI3Kδ inhibitors, which help explain why Cpd24 has the best predicted binding free energy, docking score, and experimental data.
The representation of the interactions between PI3Kδ and a idelalisib, b Cpd24, and c Cpd3 (hydrogen bonds colored in red). The surface presentations of movement of importance residues in the binding pockets of PI3Kδ with d apo-idelalisib (colored in yellow) and MD-idelalisib (colored in pink), e apo-Cpd24 (colored in green) and MD-Cpd24 (colored in orange), and f apo-Cpd3 (colored in blue) and MD-Cpd3 (colored in purple)
For better visualization of the difference of these residue-inhibitor interactions, we calculated the binding free energy changes (ΔΔGbind) between Cpd24 and the other two compounds (Fig. 5a). ΔΔGbind were defined as ΔGbind (Cpd24) − ΔGbind (Cpd3 or idelalisib), and thus the negative value of ΔΔGbind represents a favorable contribution for Cpd24, and on the contrary, the positive value represents a favorable contribution for Cpd3 or idelalisib. As shown in Fig. 5a, almost all studied residues of PI3Kδ form more affinitive interactions with Cpd24 than the other two inhibitors, especially hinge residue Glu826 and “Gatekeeper” residue Ile825. These two residues collaborated with other three residues Met752, Trp760, and Ile777, in the specificity hydrophobic pocket and help maintain Cpd24 in the active pocket. The difference between the three compounds could be caused by the introduction of bulk and negative pyrazole ring into the C5 position, which could form stronger non-polar interaction with the “Gatekeeper” residue Ile825 and more hydrophobic interaction with acid hinge residue Glu826 (Fig. 5b, c). In addition to the hydrophobic region II (Fig. 5a), the binding free energy changes of the two residues Met900 and Ile910 are largely different between Cpd24 and the other two compounds, especially the changes of Ile910 between Cpd24 and Cpd3, which may be caused by the introduction of the electronegative pyrazole ring to neutralize the electron density of the pyrimidine ring, resulting in forming more easily non-polar interactions with non-polar residues Met900 and Ile910. Interestingly, the similar effect will be obtained along with the introduction of small and positive substituents into the N3 position of the pyrimidine ring, which could also neutralize the electrons on the pyrimidine ring. But the purine of idelalisib forms stronger hydrophobic interaction with Val827 than the pyrimidine of Cpd24, which may be caused by the stronger non-polar force and the bigger steric structure of purine than pyrimidine.
Design of new compounds
According to the above structure-activity relationship analysis of COMFA, the binding mechanisms of molecular docking, and the binding free energy of MD, we designed 14 new PI3Kδ inhibitors and these inhibitors were prepared by the same method as above; the structures are summarized in Table 2. Firstly, the developed COMFA model was used to predict the activities of those inhibitors. And then, IFD was employed to dock those inhibitors into the active pocket of PI3Kδ and the best docked conformation of each designed inhibitors was chosen to further analysis. The structures and predicted activities and docking scores of designed compounds were all showed in Table 2.
According to the general SAR analysis provided by the publication reported by Leena Patel, the quinazolinone ring was put forward structural optimizations as the mother nucleus [35]. The structure of the N1 and N3 pyrimidine nitrogens and the C4-amino moiety is necessary for selectivity, so the C4-amino moiety in pyrimidine was kept. Besides, with the C5-cyano group on the pyrimidine hinge binder, Cpd5 showed high clearance in human hepatocytes [35] that made us chose Cpd2 without any substituent in the C5-position of the pyrimidine as the basic structure for the following new compound design.
According to the SAR results, these new compounds all retained the 4-amino pyrimidine ring and phenyl quinazolinone groups, which could maintain the bioactivities with the hydrogen bonds formed with the amino residues of active pocket. Based on the contour map of COMFA, compounds K1–K2 were both introduced to small and positive groups, methyl, to N3 pyrimidine ring where one yellow region and a large blue contour were mapped, and because of the same reason, K3–K7 were also introduced to methyl or amino to the C2 pyrimidine ring (Table 2). K6 was introduced to another methyl groups, the N3 position of the 4-amino pyrimidine ring, but the predicted activity of K6 (pIC50 = 6.970) was lower than K1 (pIC50 = 9.848) or K5 (pIC50 = 7.456), which contained only one methyl on the 4-amino pyrimidine ring that may be caused by the steric hindrance effect (Table 2). Considering one red contour and one big green region located under the C5 position of the 4-amino pyrimidine ring, K8 (pIC50 = 7.092) introduced one negative and bulk group (CCl3) into the C5-pyrimidine ring, which improved a little compared with Cpd2 (pIC50 = 7.000), and that may lead to the loss of hydrogen bond formed between the C2 position of the 4-amino pyrimidine ring and Val828 residue (Fig. 2e). Thus, K9 (pIC50 = 9.461) was introduced to one amino to the C2 position based on the structure of K8, which had a greatly improved predicted activity compared with K8 (pIC50 = 7.092). In the structures of K10–K14, some different positive and small substituents were introduced to the phenyl ring, because of one blue contour and one yellow region located upon the C2′ position and C3′ position of the phenyl ring. And then, all novel compounds were docked into the pocket of PI3Kδ by IFD. The linear relationship between predicted activities and docking scores of all designed compounds was showed in Fig. 6 a. We could find that K2 (pIC50 = 11.628) and K10 (pIC50 = 9.837) exhibited the higher predicted bioactivities and better docking scores among all designed compounds (the synthetic route schemes shown in Fig. S6 and S7). K2 was introduced one bulk and negative acetylene to the C5-pyrimidine ring and one positive methyl to the N3 position of the 4-amino pyrimidine ring, which improved the binding affinities between K2 and PI3Kδ, especially in the specific hydrophobic pocket formed by the residues Trp760 and Met752 and the hydrophobic region II formed by Met900 and Ile910 (Fig. 6b). K10 was introduced one amino group to the C3′ position of the phenyl ring (Fig. 6c), which formed hydrogen bonds with the hydrophobic residue Ile910 and two hinge residues Glu826 and Val828. In conclusion, based on the analysis discussed above, some new inhibitors, especially K2 and K10, showed higher predicted binding affinity to PI3Kδ. It indicated that our findings may provide some guidance to rational design novel PI3Kδ inhibitors.
Conclusions
In this work, a series of computer-aided drug design strategies, including 3D-QSAR studies, molecular docking, MD simulations, and binding free energy calculations were employed to study the binding mode between inhibitors and PI3Kδ. The developed QSAR model with high linear correlation r2pred provides reliable predictive ability (q2 = 0.59, ONC = 8, r2 = 0.966, the test sets of r2pred = 0.719), which could be effectively used to predict the bioactivities of selective PI3Kδ inhibitors. Meanwhile, the steric and electrostatic contours obtained by COMFA model delicately revealed the structure-activity relationship between those idelalisib derivatives and PI3Kδ, providing valuable information for the rational design of existing selective PI3Kδ inhibitors. Besides, the RRD and IFD suggest that the introduction of the flexibility of PI3Kδ may significantly improve the accuracy of predicting the binding mode between inhibitors and PI3Kδ, and then IFD roughly figured out the ingredient residues of the “specificity” pocket, which may be useful for virtual screening of specific PI3Kδ inhibitors. Besides, MD simulations were performed to deeply understand the dynamic binding mode between PI3Kδ/ligand complexes. The binding energy calculations show that the van der Waals interactions and the non-polar contributions are crucial for maintaining the affinities of complex binding. The introduction of some bulk and negative substituents into the C5 position of pyrimidine could increase the van der Waals interactions and the non-polar interactions with the adenine pocket formed by hinge residues Glu826, Val827, and Val828 and the “Gatekeeper” residue Ile825, and that restrain the ligand from running away from the active pocket with the cooperation between the specific hydrophobic pocket including Met752, Trp760, and Ile777. The MM/GBSA free energy decomposition was simultaneously employed and some key residues for improving the affinities to PI3Kδ were found, including Ile910, Met900, and the “Gatekeeper” residue Ile825, especially the residues Trp760 and Met752 forming the specific hydrophobic pocket. In the end, 14 novel PI3Kδ inhibitors were designed based on our analyses of COMFA, docking, and MD simulation, and two prospective compounds, K2 and K10, were highlighted. In conclusion, all the above results could give some meaningful information for further studies in the design of novel effective selective PI3Kδ inhibitors.
References
Fruman DA, Rommel C (2014) PI3K and cancer: lessons, challenges and opportunities. Nat. Rev. Drug Discov. 13(2):140–156
Cantley LC (2002) The phosphoinositide 3-kinase pathway. Science 296(5573):1655–1657
Carracedo A, Pandolfi PP (2008) The PTEN-PI3K pathway: of feedbacks and cross-talks. Oncogene 27(41):5527–5541
Liu P, Cheng H, Roberts TM, Zhao JJ (2009) Targeting the phosphoinositide 3-kinase pathway in cancer. Nat. Rev. Drug Discov. 8(8):627–644
Courtney KD, Corcoran RB, Engelman JA (2010) The PI3K pathway as drug target in human cancer. J. Clin. Oncol. 28(6):1075–1083
Jiang BH, Liu LZ (2009) PI3K/PTEN signaling in angiogenesis and tumorigenesis. Adv. Cancer Res. 102:19–65
Hennessy BT, Smith D, Ram P, Lu Y (2005) Exploiting the PI3K/AKT pathway for cancer drug discovery. Nat. Rev. Drug Discov. 4(12):988–1004
Zhu J, Wang M, Cao B, Hou T, Mao X (2014) Targeting the phosphatidylinositol 3-kinase/AKT pathway for the treatment of multiple myeloma. Curr. Med. Chem. 21(27):3173–3187
Setti A, Kumar MJ, Babu KR, Rasagna A (2015) Potency and pharmacokinetics of broad spectrum and isoform-specific p110γ and δ inhibitors in cancers. J. Recept. Signal Transduct. Res. 36(1):26–36
Zhu J, Pan P, Li Y, Wang M, Li D, Cao B, Mao X, Hou T (2014) Theoretical studies on beta and delta isoform-specific binding mechanisms of phosphoinositide 3-kinase inhibitors. Mol. BioSyst. 10(3):454–466
Yuan TL, Cantley LC (2008) PI3K pathway alterations in cancer: variations on a theme. Oncogene 27(41):5497–5510
Jabbour E, Ottmann OG, Deininger M, Hochhaus A (2014) Targeting the phosphoinositide 3-kinase pathway in hematologic malignancies. Haematologica 99(1):7–18
Zhu J, Hou T, Mao X (2015) Discovery of selective phosphatidylinositol 3-kinase inhibitors to treat hematological malignancies. Drug Discov. Today 20(8):988–994
Li T, Wang G (2014) Computer-aided targeting of the PI3K/Akt/mTOR pathway: toxicity reduction and therapeutic opportunities. Int. J. Mol. Sci. 15(10):18856–18891
Li M, Sala V (2018) Phosphoinositide 3-kinase gamma inhibition protects from anthracycline cardiotoxicity and reduces tumor growth. Circulation 138(7):696–711
Vyas P, Vohora D (2016) Phosphoinositide-3-kinases as the novel therapeutic targets for the inflammatory diseases: current and future perspectives. Curr. Drug Targets 18(14):1622–1640
Vangapandu HV, Jain N, Gandhi V (2017) Duvelisib: a phosphoinositide-3 kinase δ/γ inhibitor for chronic lymphocytic leukemia. Expert Opin. Investig. Drugs 26(5):625–632
Lannutti BJ, Meadows SA, Herman SE, Kashishian A (2011) CAL-101, a p110δ selective phosphatidylinositol-3-kinase inhibitor for the treatment of B-cell malignancies, inhibits PI3K signaling and cellular viability. Blood 117(2):591–594
Sabbah DA, Vennerstrom JL, Zhong HA (2012) Binding selectivity studies of phosphoinositide 3-kinases using free energy calculations. J. Chem. Inf. Model. 52(12):3213–3224
Kitchen DB, Decornez H, Furr JR, Bajorath J (2004) Docking and scoring in virtual screening for drug discovery: methods and applications. Nat. Rev. Drug Discov. 3(11):935–949
Sabbah DA, Vennerstrom JL, Zhong HZ (2010) Docking studies on isoform-specific inhibition of phosphoinositide-3-kinases. J. Chem. Inf. Model. 50(10):1887–1898
Shao S, Yu R, Yu Y, Li Y (2014) Dual-inhibitors of STAT5 and STAT3: studies from molecular docking and molecular dynamics simulations. J. Mol. Model. 20(8):2399
Zhang C, Du C, Feng Z, Zhu J, Li Y (2015) Hologram quantitative structure activity relationship, docking, and molecular dynamics studies of inhibitors for CXCR4. Chem. Biol. Drug Des. 85(2):119–136
Iqbal S, Krishnan DA, Gunasekaran K (2018) Identification of potential PKC inhibitors through pharmacophore designing, 3D–QSAR and molecular dynamics simulations targeting Alzheimer’s disease. J Biomol Struct Dyn 36(15):4029–4044
Katari SK, Natarajan P, Swargam S, Kanipakam H, Pasala C (2016) Inhibitor design against JNK1 through e-pharmacophore modeling docking and molecular dynamics simulations. J. Recept. Signal Transduct. Res. 36(6):558–571
Rajamanikandan S, Jeyakanthan J, Srinivasan P (2017) Molecular docking, molecular dynamics simulations, computational screening to design quorum sensing inhibitors targeting LuxP of Vibrio harveyi and its biological evaluation. Appl. Biochem. Biotechnol. 181(1):192–218
Zondagh J, Balakrishnan V, Achilonu I, Dirr HW, Sayed Y (2018) Molecular dynamics and ligand docking of a hinge region variant of South African HIV-1 subtype C protease. J Mol Graph Model 82:1–11
Xu C, Ren Y (2015) Molecular modeling studies of [6,6,5] Tricyclic Fused Oxazolidinones as FXa inhibitors using 3D-QSAR, Topomer CoMFA, molecular docking and molecular dynamics simulations. Bioorg. Med. Chem. Lett. 25(20):4522–4528
Tang HJ, Yang L, Li JH, Chen J (2016) Molecular modelling studies of 3,5-dipyridyl-1,2,4-triazole derivatives as xanthine oxidoreductase inhibitors using 3D-QSAR, Topomer CoMFA, molecular docking and molecular dynamic simulations. J Taiwan Inst Chem E 68:64–73
Aksoydan B, Kantarcioglu I, Erol I, Salmas RE, Durdagi S (2018) Structure-based design of hERG-neutral antihypertensive oxazalone and imidazolone derivatives. J Mol Graph Model 79:103–117
Zhao S, Zhu J, Xu L, Jin J (2017) Theoretical studies on the selective mechanisms of GSK3beta and CDK2 by molecular dynamics simulations and free energy calculations. Chem. Biol. Drug Des. 89(6):846–855
Shen M, Zhou S, Li Y, Li D, Hou T (2013) Theoretical study on the interaction of pyrrolopyrimidine derivatives as LIMK2 inhibitors: insight into structure-based inhibitor design. Mol. BioSyst. 9(10):2435–2446
Xu L, Li Y, Li L, Zhou S, Hou T (2012) Understanding microscopic binding of macrophage migration inhibitory factor with phenolic hydrazones by molecular docking, molecular dynamics simulations and free energy calculations. Mol. BioSyst. 8(9):2260–2273
Ekhteiari Salmas R, Unlu A, Bektas M, Yurtsever M, Mestanoglu M, Durdagi S (2017) Virtual screening of small molecules databases for discovery of novel PARP-1 inhibitors: combination of in silico and in vitro studies. J. Biomol. Struct. Dyn. 35(9):1899–1915
Patel L, Chandrasekhar J, Evarts J (2016) 2,4,6-Triaminopyrimidine as a novel hinge binder in a series of PI3Kδ selective inhibitors. J. Med. Chem. 59(7):3532–3548
Sherman W, Beard H, Farid R (2010) Use of an induced fit receptor structure in virtual screening. Chem. Biol. Drug Des. 67(1):83–84
Somoza JR, David K (2015) Structural, biochemical, and biophysical characterization of idelalisib binding to phosphoinositide 3-kinase δ. J Bio Chem 290(13):8439–8446
Case DA, Cheatham TE, Darden T, Gohlke H (2005) The Amber biomolecular simulation programs. J. Comput. Chem. 26(16):1668–1688
Duan Y, Wu C, Chowdhury S, Lee MC, Xiong GM (2003) A point-charge force field for molecular mechanics simulations of proteins based on condensed-phase quantum mechanical calculations. J. Comput. Chem. 24(16):1999–2012
Wang JM, Wolf RM, Caldwell JW, Kollman PA, Case DA (2004) Development and testing of a general amber force field. J. Comput. Chem. 25(9):1157–1174
Stewart JPJ (2010) Optimization of parameters for semiempirical methods I. Method. J Comput Chem 10(2):221–264
Bayly CI, Cieplak P, Cornell WD, Kollman PA (1993) A well-behaved electrostatic potential based method using charge restraints for deriving atomic charges: the resp model. J. Phys. Chem. 97(40):10269–10280
Darden T, York D, Pedersen L (1993) Particle mesh Ewald: an N·log(N) method for Ewald sums in large systems. J. Phys. Chem. 98(12):10089–10092
Vincent K, Van Gunsteren W, Hünenberger P (2001) A fast SHAKE algorithm to solve distance constraint equations for small molecules in molecular dynamics simulations. J. Comput. Chem. 22(5):501–508
Kollman PA, Massova I, Reyes C, Kuhn B, Huo S, Chong L, Lee M (2000) Calculating structures and free energies of complex molecules: combining molecular mechanics and continuum models. Cheminform 32(10):889–897
Hou T, Li Y, Wang W (2011) Prediction of peptides binding to the PKA RIIα subunit using a hierarchical strategy. Bioinformatics 27(13):1814–1821
Sun H, Li Y, Shen M, Tian S, Xu L, Pan P, Guan Y, Hou T (2014) Assessing the performance of MM/PBSA and MM/GBSA methods. 5. Improved docking performance using high solute dielectric constant MM/GBSA and MM/PBSA rescoring. Phys. Chem. Chem. Phys. 16(40):22035–22045
Chen F, Liu H, Sun H, Pan P, Li Y, Li D, Hou T (2016) Assessing the performance of the MM/PBSA and MM/GBSA methods. 6. Capability to predict protein–protein binding free energies and re-rank binding poses generated by protein–protein docking. Phys. Chem. Chem. Phys. 18(18):22129–22139
Pan P, Yu H, Liu Q, Kong X, Chen H, Chen J, Liu Q, Li D, Kang Y, Sun H (2017) Combating drug-resistant mutants of anaplastic lymphoma kinase with potent and selective type-I1/2 inhibitors by stabilizing unique DFG-shifted loop conformation. Acs Cent Sci 3(11):1208–1220
Hou T, Li Y, Wang W (2011) Prediction of peptides binding to the PKA RIIalpha subunit using a hierarchical strategy. Bioinformatics 27(13):1814–1821
Sun H, Li Y, Li D, Hou T (2013) Insight into crizotinib resistance mechanisms caused by three mutations in ALK tyrosine kinase using free energy calculation approaches. J. Chem. Inf. Model. 53(9):2376–2389
Sun H, Li Y, Tian S, Xu L, Hou T (2014) Assessing the performance of MM/PBSA and MM/GBSA methods. 4. Accuracies of MM/PBSA and MM/GBSA methodologies evaluated by various simulation protocols using PDBbind data set. Phys. Chem. Chem. Phys. 16(31):16719–16729
Sun H, Duan L, Chen F, Liu H, Wang Z, Pan P, Zhu F (2018) Assessing the performance of MM/PBSA and MM/GBSA methods. 7. Entropy effects on the performance of end-point binding free energy calculation approaches. Phys. Chem. Chem. Phys. 20(21):14450–14460
Chen F, Liu H, Sun H, Pan P, Li Y, Li D, Hou T (2016) Assessing the performance of the MM/PBSA and MM/GBSA methods. 6. Capability to predict protein-protein binding free energies and re-rank binding poses generated by protein-protein docking. Phys. Chem. Chem. Phys. 18(32):22129–22139
Chen F, Sun H, Wang J, Zhu F, Liu H (2018) Assessing the performance of MM/PBSA and MM/GBSA methods. 8. Predicting binding free energies and poses of protein-RNA complexes. RNA 24(9):1183–1194
Onufriev A, Donald Bashford A, Case DA (2000) Modification of the generalized born model suitable for macromolecules. J. Phys. Chem. B 104(15):3712–3720
Weiser J, Shenkin P, Still W (1999) Approximate atomic surfaces from linear combinations of pairwise overlaps (LCPO). J. Comput. Chem. 20(2):217–230
Berndt A, Miller S, Williams O, Le D (2010) The p110δ crystal structure uncovers mechanisms for selectivity and potency of novel PI3K inhibitors. Nat. Chem. Biol. 6(2):117–124
Williams R, Berndt A, Miller S, Hon W, Zhang X (2009) Form and flexibility in phosphoinositide 3-kinases. Biochem. Soc. Trans. 37(4):615–626
Safina BS, Sweeney ZK, Li J, Chan BK (2013) Identification of GNE-293, a potent and selective PI3Kδ inhibitor: navigating in vitro genotoxicity while improving potency and selectivity. Bioorg. Med. Chem. Lett. 23(17):4953–4959
Funding
The study was financially supported by the National Natural Science Foundation of China (No. 21807049, 81803430) and the Fundamental Research Funds for the Central Universities (JUSRP11892).
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
ESM 1
(DOCX 2327 kb)
Rights and permissions
About this article
Cite this article
Zhu, J., Ke, K., Xu, L. et al. Theoretical studies on the selectivity mechanisms of PI3Kδ inhibition with marketed idelalisib and its derivatives by 3D-QSAR, molecular docking, and molecular dynamics simulation. J Mol Model 25, 242 (2019). https://doi.org/10.1007/s00894-019-4129-x
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s00894-019-4129-x