Abstract
Photosystem II (PS II) of oxygenic photosynthesis is found in the thylakoid membranes of plastids and cyanobacteria. The mature PS II complex comprises a central core of four membrane proteins that bind the majority of the redox-active cofactors. In cyanobacteria the central core is surrounded by 13 low-molecular-weight (LMW) subunits which each consist of one or two transmembrane helices. Three additional hydrophilic subunits known as PsbO, PsbU and PsbV are found associated with hydrophilic loops belonging to the core proteins protruding into the thylakoid lumen. During biogenesis the majority of the LMW subunits are known to initially associate with individual pre-assembly complexes consisting of one or more of the core proteins; however, the point at which the PsbJ LMW subunit binds to PS II is not known. The majority of models for PS II biogenesis propose that the three extrinsic proteins and PsbJ bind in the final stages of PS II assembly. We have investigated the impact of creating the double mutants ∆PsbJ:∆PsbO, ∆PsbJ:∆PsbU and ∆PsbJ:∆PsbV to investigate potential cooperation between these subunits in the final stages of biogenesis. Our results indicate that PsbJ can bind to PS II in the absence of any one of the extrinsic proteins. However, unlike their respective single mutants, the ∆PsbJ:∆PsbO and ∆PsbJ:∆PsbV strains were not photoautotrophic and were unable to support oxygen evolution suggesting a functional oxygen-evolving complex could not assemble in these strains. In contrast, the PS II centers formed in the ∆PsbJ:∆PsbU strain were capable of photoautotrophic growth and could support oxygen evolution when whole-chain electron transport was supported by the addition of bicarbonate.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Photosystem II (PS II) is a multisubunit complex that catalyses the light-driven splitting of water in the thylakoid membrane of oxygenic phototrophs (Shen 2015; Vinyard and Brudvig 2018). High resolution X-ray-derived PS II structures (~ 1.9–2.1 Å) from thermophilic cyanobacteria have shown PS II to be dimeric, with each monomer containing the chlorophyll a-binding core antenna proteins CP43 and CP47 together with the D1/D2 reaction center subunits and at least 16 additional polypeptides and 70 cofactors (Umena et al. 2011; Suga et al. 2015, 2017; Kern et al. 2018). Detailed structures have shown conservation of these intrinsic components of PS II between cyanobacteria, algae and higher plants (Ago et al. 2016; Wei et al. 2016).
Thirteen low-molecular-weight (LMW) subunits have been identified on the periphery of each PS II monomer (Ferreira et al. 2004; Umena et al. 2011). Mutagenesis studies have shown the absence of these subunits frequently results in strains with impaired photoautotrophic growth or PS II assembly (Müh et al. 2008; Shi et al. 2012; Luo et al. 2014; Fagerlud et al. 2020). The assembly of PS II involves the formation of different pre-complexes that contain distinct groups of PS II proteins along with multiple assembly factors that are not present in mature PS II centers (Komenda et al. 2012; Mabbitt et al. 2014). Most of the LMW subunits have been found to be associated with different pre-complexes (Nickelsen and Rengstl 2013; Eaton-Rye and Sobotka 2017).
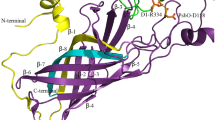
The PsbJ subunit has not been identified in any of the assembly pre-complexes; however, in mature PS II centers, PsbJ is a single transmembrane helix that is located near the PsbE and PsbF proteins that comprise the cytochrome b559 complex (Heinz et al. 2016; Müh and Zouni 2016). In Thermosynechococcus elongatus deletion of PsbJ led to the accumulation of an intermediate monomeric PS II complex which contained the Psb27 and Psb28 assembly factors (Nowaczyk et al. 2012).
In addition to the membrane-spanning subunits of PS II, three hydrophilic extrinsic proteins, PsbO, PsbU and PsbV, are present on the lumenal face of the photosystem (Bricker et al. 2012). The PsbJ protein was absent from a PS II structure obtained by cryo-electron microscopy that accumulated in a Thermosynechococcus vulcanus strain in which the psbV gene, encoding PsbV, had been deleted (Huang et al. 2021). The ∆PsbV strain accumulated PS II complexes that contained the assembly factor Psb27 and these Psb27-PS II complexes formed dimers which exhibited structural changes compared to mature PS II centers. This included conformational shifts associated with PsbE and PsbF that may have resulted in the loss of PsbJ (Huang et al. 2021). These observations are consistent with late addition of PsbJ to PS II complexes.
In Synechocystis sp. PCC 6803 PsbJ is not required for photoautotrophic growth; however, the deletion of PsbJ affected the efficiency of PS II electron flow between the primary and secondary plastoquinone electron acceptors QA and QB and back electron transfer to the oxidized Mn4CaO5 cluster of the oxygen-evolving complex (OEC) (Regel et al. 2001). An additional QC-binding site has also been identified in Thermosynchococcus elongatus (Guskov et al. 2009). The PsbJ protein, along with the cytochrome b559 complex, are located close to the QC site and have been suggested to contribute to a second channel allowing exchange of plastoquinone or plastoquinol with the pool of plastoquinone in the thylakoid membrane (Müh and Zouni 2016).
As in the case of PsbJ, the extrinisic proteins are predicted to be added late to the PS II complex (Bricker et al. 2012; Nickelsen and Rengstl 2013). The extrinsic proteins form a stabilising structure around the catalytic Mn4CaO5 of the OEC protecting it from bulk reductants and contributing to putative channels associated with the access of substrate H2O and the egress of O2 and H+ (Ho and Styring 2008; Roose et al. 2016).
In Synechocystis sp. PCC 6803, strains lacking any one of the PsbO, PsbU or PsbV proteins are able to grow photoautotrophically (Burnap and Sherman 1991; Shen et al. 1995a, 1997) but strains lacking two of these proteins either grow photoautotrophically (∆PsbU:∆PsbV mutant; Eaton-Rye et al. 2003), are unable to grow phototrophically (∆PsbO:∆PsbV mutant; Shen et al. 1995b) or exhibit conditional growth at elevated pH (∆PsbO:∆PsbU mutant; Eaton-Rye et al. 2003; Summerfield et al. 2007). This double mutant approach identified that each of these extrinsic subunits is able to bind in the absence of one other extrinsic protein and that the presence of at least two of the proteins are required for stable assembly of PS II centers (except at elevated pH) (Morris et al. 2016, 2019). Due to the predicted late addition of PsbJ and the extrinsics proteins to PS II, we have employed a similar approach to investigate whether the absence of the extrinsic proteins altered the impact of removing PsbJ in Synechocystis sp. PCC 6803.
Materials and methods
Cyanobacterial strains and culture conditions
The glucose-tolerant GT-O1 substrain of Synechocystis sp. PCC 6803 was used and referred to throughout as wild type (Williams 1988; Morris et al. 2014). The mutant strains were made by transforming the GT-O1 substrain. The ∆PsbJ strain was constructed with the psbJ gene replaced with a spectinomycin-resistance cassette introduced 57 bp after the start codon and 63 bp from the stop codon. The psbO gene was inactivated by inserting a kanamycin-resistance cassette 503 bp after the start codon and 267 bp before the stop codon. Both psbU and psbV were also inactivated by the insertion of a kanamycin-resistance cassette. In the case of psbU, the antibiotic-resistance cassette was inserted 36 bp after the start codon and 151 bp before the stop codon. In the case of psbV, the antibiotic-resistancc cassette was located 68 bp after the start condon and 120 bp from the stop codon. The double mutants ∆PsbJ:∆PsbO, ∆PsbJ:∆PsbU and ∆PsbJ:∆PsbV were made by transforming the ∆PsbJ strain using the corresponding plasmids in which psbO, psbU or psbV had been inactivated.
All strains were grown on BG-11 media agar plates containing 5 mM glucose, 20 μM atrazine, 10 mM TES-NaOH (pH 8.2) and 0.3% sodium thiosulfate. Liquid cultures were grown mixotrophically in unbuffered BG-11 media containing 5 mM glucose. In both solid and liquid media antibiotics were added at 25 µg mL−1 and cultures were maintained at 30 °C under constant illumination at 30 μmol photons m−2 s−1 (Eaton-Rye 2011). Cells in liquid culture were harvested when they reached mid-logarithmic growth by centrifugation at 5000×g for 10 min at room temperature. Photoautotrophic growth curves were carried out as described in Forsman and Eaton-Rye (2021).
Oxygen evolution assays in cells
Cells grown mixotrophically were washed and then resuspended in BG-11 containing 25 mM HEPES–NaOH (pH 7.5) at a chlorophyll concentration of 10 µg mL−1 and 30 °C. Oxygen evolution was measured with a Clark-type electrode (Hansatech, King’s Lynn, U.K.) at 30 °C using 15 mM NaHCO3− or a combination of 0.2 mM 2,5-dimethyl-1,4-benzoquinone (DMBQ) or 2,5-dichloro-1,4-benzoquinone (DCBQ) and 1 mM K3Fe(CN)6. Saturating actinic light (2 mmol photons m−2 s−1) was provided by an FLS1 light source (Hansatech, King’s Lynn, U.K.) passed through a Melis Griot OG 590 sharp cutoff red glass filter.
Variable chlorophyll a fluorescence induction
Cells were washed and resuspended in BG-11 (pH 7.5) at a chlorophyll concentration of 5 μg mL−1 and dark adapted at room temperature for 5 min. Variable chlorophyll a fluorescence was measured with a FL-3000 double modulation kinetic fluorometer (PSI Instruments, Brno, Czech Republic) using a blue 455 nm measuring light. The actinic voltage was set to 50% for all samples and the measuring light voltage was set to 80%. When present, 3-(3,4-dichlorophenyl)-1,1-dimethylurea (DCMU) was at a final concentration 40 µM, and added 1 min prior to measurement. The decay of variable fluorescence following a single actinic flash was measured according to Forsman and Eaton-Rye (2021).
77 K chlorophyll a fluorescence emission spectroscopy
Low-temperature (77 K) fluorescence emission spectroscopy was performed as previously described (Jackson et al. 2014). Spectra were normalized to the Photosystem I peak at 725 nm.
Protein analyses
For protein analyses, cultures of the wild type and mutant strains were grown photomixotrophically to mid-exponential phase at 30 °C and 40 μmol of photons m−2 s−1. Isolation of thylakoid membranes, blue-native polyacrylamide gel electrophoresis and western blotting were performed as previously described (Jackson et al. 2014).
Pigment extraction
Chlorophyll concentrations were determined according to MacKinney (1941) and carotenoids were extracted according to Chamovitz et al. (1993).
Results
Removal of PsbJ from strains lacking PsbO, PsbU or PsbV abolishes or reduces photoautotrophic growth
Strains lacking either PsbJ, PsbO, PsbU, PsbV grew photoautotrophically (Fig. 1). The photoautotrophic doubling time for the wild-type and ∆PsbU strains was 12 h and this increased to 17 h in ∆PsbV cells, 18 h in ∆PsbJ cells and was extended to 20 h in the ∆PsbO strain. In contrast, the doubling time in the ∆PsbJ:∆PsbU mutant was slowed to 60 h (Fig. 1b) while removal of PsbO or PsbV in the ΔPsbJ background prevented photoautotrophic growth (Fig. 1a, b).
Photoautotrophic growth of Synechocystis sp. PCC 6803 strains in BG-11 medium was measured by optical density at 730 nm. a Wild type (black circles), ΔPsbJ (blue circles), ΔPsbO (black squares) and ΔPsbJ:ΔPsbO (blue squares). b ΔPsbU (black triangles), ΔPsbJ:ΔPsbU (blue triangles), ΔPsbV (black diamonds) and ΔPsbJ:ΔPsbV (blue diamonds). Error bars represent standard error of the mean for three separate experiments
Oxygen evolution is impaired in strains lacking PsbJ and one of the extrinsic proteins PsbO, PsbU or PsbV
The rates of oxygen evolution observed for the ΔPsbJ strain and wild type were similar in the presence of bicarbonate but were reduced to ~ 59% of the wild-type rate in the ΔPsbJ strain in the presence of either DCBQ or DMBQ which are PS II-specific electron acceptors (Table 1). In contrast, the ΔPsbJ:ΔPsbO strain did not evolve oxygen under any any conditions; however, the ΔPsbJ:ΔPsbU strain exhibited an initial rate in the presence of bicarbonate that was similar to the observed rate for wild-type cells, but had less than 20% the wild-type rate with either PS II-specific electron acceptor (Table 1). In ∆PsbJ:∆PsbV cells, oxygen evolution was reduced to ~ 34% of the wild-type rate when supported by bicarbonate and essentially abolished when supported by either DCBQ or DMBQ (Table 1). This impaired oxygen evolution indicates a specific effect on PS II activity following the removal of PsbJ when an extrinsic protein is also absent.
Chlorophyll a fluorescence is altered in the ∆PsbJ strain in the presence and absence of extrinsic proteins
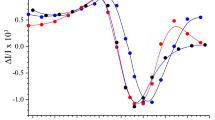
Variable chlorophyll a fluorescence induction assays were performed, in the presence and absence of the PS II-specific herbicide DCMU, to assess the impact of removing PsbJ. The ∆PsbJ strain exhibited chlorophyll a fluorescence induction kinetics with typical O, J, I and P features: O indicates the Fo fluorescence origin, the rise from O to the J peak reflects the photoreduction of QA to QA−, and the rise from J to the intermediary inflection I and the maximum fluorescence (Fm or peak (P)) reflects the reduction of the plastoquinone pool (Govindjee 1995). The O to J and I to P transitions were reduced in the ∆PsbJ strain compared to wild type (Fig. 2a). As previously reported, removal of the extrinsic proteins altered chlorophyll fluorescence induction kinetics (Summerfield et al. 2013). The removal of PsbU flattened the O to J rise, but a substantial I to P rise remained; in contrast, removal of either PsbO or PsbV resulted in a reduced J level and almost no I to P rise. Combined removal of PsbJ and one of the extrinsic proteins further suppressed fluorescence induction including the absence of an I to P rise for the ∆PsbJ:∆PsbO and ∆PsbJ:∆PsbV mutants and a greatly reduced I to P rise in the ∆PsbJ:∆PsbU strain (Fig. 2a, b).
Chlorophyll a fluorescence induction transients are altered in strains lacking PsbO, PsbU and PsbV following removal of PsbJ. a, c Wild-type cells (black circles), ΔPsbJ cells (blue circles), ΔPsbO cells (black squares), ΔPsbJ:ΔPsbO cells (blue squares). b, d ΔPsbU cells (black triangles), ΔPsbJ:ΔPsbU cells (blue triangles), ΔPsbV cells (black diamonds), ΔPsbJ:ΔPsbV cells (blue diamonds). Cells have no addition (a, b), or 40 µM DCMU (c, d). Data are the average of at least three independent experiments
In the presence of DCMU, all strains showed a rapid fluorescence rise to Fm due to the inhibition of forward electron transfer from QA−. The variable fluorescence yield in the presence of DCMU is typically indicative of the number of active PS II centers present. The fluorescence yield from the ∆PsbJ, ∆PsbO, ∆PsbU and ∆PsbV mutants was reduced compared to wild type by different amounts, ranging between 96% of the wild-type Fm for ∆PsbU cells (least affected), and 58% of the Fm for the ∆PsbV strain (most affected) (Fig. 2c, d). The The Fm for the ΔPsbO and ΔPsbJ strains was similar at ~68% of the wild-type level (Fig. 2c); however, the ∆PsbJ:∆PsbU double mutant retained only 35% of the wild-type level and the ∆PsbJ:∆PsbO and ∆PsbJ:∆PsbV strains showed a further reduction to ~ 24%. The results in Fig. 2 indicate that the absence of an O–J–I–P transient and, in the presence of DCMU, an Fm value 35% of the wild-type level does not preclude photoautotrophic growth, as the ∆PsbJ:∆PsbU strain grew photoautotrophically.
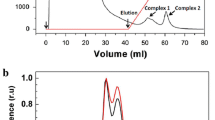
To further investigate PS II assembly, low-temperature fluorescence emission spectra were measured following direct excitation of chlorophyll a at 440 nm (Fig. 3). Following 440 nm excitation, the PS II-specific emission peaks at 685 nm and 695 nm were lower in the ΔPsbJ mutant than in wild type when normalized to the PS I peak at 725 nm. The PS II emission at 685 nm from the ΔPsbO and ΔPsbJ:ΔPsbO strains was increased, compared to wild type, with a shoulder at 695 nm for the ΔPsbO strain but not for the ΔPsbJ:ΔPsbO strain. The ΔPsbU strain had a similar spectrum to wild type but a slightly reduced 695 nm peak, this peak was further reduced in the ΔPsbJ:ΔPsbU strain. The ΔPsbV mutant had a higher 685 nm peak, similar to the ΔPsbO strain but a 695 nm shoulder that was less pronounced than the ΔPsbO strain, and the ΔPsbJ:ΔPsbV strain had a slightly elevated 685 nm peak and no distinct 695 nm shoulder. The reduced 695 nm emission in the ΔPsbJ:ΔPsbU strain and further reduction in the ΔPsbJ:ΔPsbO and ΔPsbJ:ΔPsbV strains is consistent with the reduction of variable fluorescence in the presence of DCMU and indicates reduced accumulation of PS II centers in the ΔPsbJ:ΔPsbU strain and greater reduction in the ΔPsbJ:ΔPsbO and ΔPsbJ:ΔPsbV strains. The reduced levels of assembled PS II in the double mutants was also supported by low-temperature fluorescence emission spectra where the phycobilisomes were excited directly with 580 nm light (Fig. S1).
Low temperature (77 K) fluorescence emission spectra following excitation at 440 nm. a wild type (black solid line), ΔPsbJ (blue solid line), ΔPsbO (black dashed line) and ΔPsbJ:ΔPsbO (blue dashed line), b ΔPsbU (black solid line), ΔPsbV (black dashed line), ΔPsbJ:ΔPsbU (blue solid line), ΔPsbJ:ΔPsbV (blue dashed line). Results are the average of at least three individual biological replicates. All spectra were normalized to the fluorescence emission from PS I at 725 nm
Dimerisation of PS II is reduced in the combined mutants
The abolished or impaired photoautotrophic growth and reduced level of PS II centers in the strains lacking both PsbJ and one of the extrinsic proteins indicated PsbJ may be important for assembly or stability of PS II. Therefore we investigated whether changes to PS II in these mutants included altered dimer abundance. To assess PS II assembly, blue native-polyacrylamide gel electrophoresis (BN-PAGE) followed by immunodetection using D1-specific, D2-specific and CP43-specific antibodies was used to determine the relative abundance of PS II dimers, monomers and other assembly intermediates. Detection of the PS II native complexes in the ΔPsbJ, ΔPsbU and ΔPsbV strains showed a slight reduction in dimers compared to wild type (Fig. 4). The ΔPsbO strain showed greatly decreased dimer abundance, as did all three double mutants. The destabilization of PS II dimer formation in the absence of PsbO is consistent with earlier reports (Bentley and Eaton-Rye 2008; Komenda et al. 2010). Only the ΔPsbJ:ΔPsbU strain had any detectable PS II dimers, this strain also had increased levels of PS II monomers compared to the ΔPsbJ:ΔPsbO and ΔPsbJ:ΔPsbV strains. The three double mutants also showed a shift in the size of the PS II monomer; however, the gel shift was also observed for the PS I trimer band in the BN-PAGE gel consistent with stress-induced accumulation of carotenoids (Paerl 1984; Toth et al. 2015; Vajravel et al. 2017). Elevated levels of carotenoids in the double mutants were confirmed by absorption spectra of N,N-dimethylformamide-extracted pigments (Fig. S2). In addition, each of the double mutants accumulated a putative assembly intermediate complex containing CP43 that was observed at the bottom of the western blot probed with the CP43-specific antibody (Fig. 4). A similar signal was also obtained with an antibody to CP43 in a double mutant lacking the CyanoP and Ycf48 PS II proteins which are both lumenal assembly factors (Jackson and Eaton-Rye 2015).
Discussion
The psbJ gene is in the operon encoding cytochrome b559 and PsbL: in cyanobacteria this operon is found in a gene cluster with ycf48 and rubA that both encode assembly factors in PS II biogenesis (Yu et al. 2018; Garcia-Cerdán et al 2019). It has been suggested that cooperation between the LMW and extrinsic subunits of PS II may have arisen in the ancestral photosystem that gave rise to PS II (Cardona 2016). Since the other proteins encoded in the psbEFLJ operon play essential roles in PS II biogenesis (reviewed in Nickelsen and Rengstl (2013)), we probed the role of PsbJ by constructing double mutants inactivating psbJ and the genes encoding the extrinsic proteins of PS II. Consistent with the previous report by Regel et al. (2001), we found the absence of PsbJ impaired PS II-specific oxygen evolution and reduced the number of PS II centers. We also observed a slightly elevated rate of bicarbonate-supported electron transport in our ∆PsbJ and ∆PsbV mutants when compared to wild-type cells; however, the mechanism for this effect is not yet understood but it is possible that alternative pathways for PQH2 oxidation are upregulated in these strains (Lea-Smith et al. 2016).
As expected, removal of any one of the three extrinsic proteins did not prevent photoautotrophic growth but did impact PS II activity and the number of assembled active centers as previously reported (Burnap and Sherman 1991; Shen et al. 1995a, 1997; Eaton-Rye et al. 2003). The combination of removing PsbJ with any one of the three extrinsic proteins, however, prevented or impaired photoautotrophic growth and decreased oxygen evolution, the number of assembled PS II centers, and the detectable level of PS II dimers. The difference in phenotype between the single and double mutants indicates that PsbJ can bind PS II in the absence of any one of the extrinsic proteins. Support for this conclusion is also evident in the chlorophyll a fluorescence decay kinetics obtained after a single actinic flash (Figs. S3 and S4; and Tables S1 and S2).
The observation that PsbJ can bind in the absence of one of the extrinic proteins is in contrast with the reported cryo-EM-derived structure of PS II from a Thermosynechococcus vulcanus strain lacking PsbV where PsbJ was absent (Huang et al. 2021). In the case of the Thermosynechococcus vulcanus strain lacking PsbV, it was suggested that conformational shifts of PsbE and PsbF may result in the loss of PsbJ from the structure (Huang et al. 2021). In addition, in our ∆PsbJ mutant the intensity and mobility of the PS II dimer and monomer bands obtained by BN-PAGE were similar to those observed for wild type. This suggests that there is little or no accumulation of an intermediate PS II complex that retains the Psb27 and Psb28 assembly factors as reported for the ∆PsbJ strain of Thermosynechococcus elongatus (Nowaczyk et al. 2012; Zabret et al. 2021). Our results indicate PsbJ is functionally present in the ΔPsbV strain in vivo in Synechocystis sp. PCC 6803. Similarly, the ΔPsbJ strain had a phenotype distinct from that of the double mutants consistent with the extrinsic proteins binding in the absence of PsbJ. Our results also contrast with the situation in higher plants where deletion of psbJ altered the binding of the PsbP and PsbQ extrinsic proteins and prevented photoautotrophic growth in ∆psbJ plants (Hager et al. 2002; Swiatek et al. 2003; Suorsa et al. 2004).
Conclusion
The PsbJ protein can bind to PS II in cells lacking any one of the PsbO, PsbU and PsbV extrinsic proteins and the extrinsic proteins can bind to PS II in the absence of PsbJ in Synechocystis sp. PCC 6803 cells. Photoautotrophic growth, however, was prevented in the ∆PsbJ:∆PsbO and ∆PsbJ:∆PsbV strains and substantially slowed in the ∆PsbJ:∆PsbU strain. In addition, 77 K fluorescence emission spectra, variable fluoresence induction and oxygen evolution indicate low numbers of functional PS II centers in the double mutants. Taken together, our data suggest that an active OEC could not assemble in the absence of PsbJ when either PsbO or PsbV was missing.
References
Ago H, Adachi H, Umena Y et al (2016) Novel features of eukaryotic photosystem II revealed by its crystal structure analysis from a red alga. J Biol Chem 291:5676–5687. https://doi.org/10.1074/jbcM115.711689
Bentley FK, Eaton-Rye JJ (2008) The effect of removing Photosystem II extrinsic proteins on dimer formation and recovery from photodamage in Synechocystis sp. PCC 6803. In: Allen JF, Gantt E, Golbeck JH, Osmond B (Eds) Photosynthesis. Energy from the sun: 14th International Congress on Photosynthesis, Springer, Dordrecht, pp 715–717. https://doi.org/10.1007/978-1-4020-6709-9_159
Bricker TM, Roose JL, Fagerlund RD et al (2012) The extrinsic protiens of photosystem II. Biochim Biophys Acta 1817:121–142. https://doi.org/10.1016/j.bbabio.2011.07.006
Burnap RL, Sherman LA (1991) Deletion mutagenesis in Synechocystis sp. PCC 6803 indicates that the Mn-stabilizing protein of photosystem II is not essential for oxygen evolution. Biochemistry 30:440–446. https://doi.org/10.1021/bi00216a020
Cardona T (2016) Reconstructing the origin of oxygenic photosynthesis: do assembly and photoactivation recapitulate evolution? Front Plant Sci 7:257. https://doi.org/10.3389/fpls.2016.00257
Chamovitz D, Sandmann G, Hirschberg J (1993) Molecular and biochemical characterization of herbicide-resistant mutants of cyanobacteria reveals that phytoene desaturation is a rate-limiting step in carotenoid biosynthesis. J Biol Chem 268:17348–17353. https://doi.org/10.1016/S0021-9258(19)85341-3
Eaton-Rye JJ (2011) Construction of gene interruptions and gene deletions in the cyanobacterium Synechocystis sp. strain PCC 6803. Meth Mol Biol 684:295–312. https://doi.org/10.1007/978-1-60761-925-3_22
Eaton-Rye JJ, Sobotka R (2017) Editorial: assembly of the photosystem II membrane-protein complex of oxygenic photosynthesis. Front Plant Sci 8:884. https://doi.org/10.3389/fpls.2017.00884
Eaton-Rye JJ, Shand JA, Nicoll WS (2003) pH-dependent photoautotrophic growth of specific Photosystem II mutants lacking lumenal extrinsic polypeptides in Synechocystis PCC 6803. FEBS Lett 543:148–153. https://doi.org/10.1016/s0014-5793(03)00432-0
Fagerlund RD, Forsman JA, Biswas S et al (2020) Stabilization of photosystem II by the PabT protein impacts photodamage, repair amd biogenesis. Biochim Biophys Acta Bioenerg 1861:148234. https://doi.org/10.1016/j.bbabio.2020.148234
Ferreira KN, Iverson TM, Maghlaoui K et al (2004) Architecture of the photosynthetic oxygen-evolving center. Science 303:1831–1838. https://doi.org/10.1126/science.1093087
Forsman JA, Eaton-Rye JJ (2021) The interaction between PsbT and the DE loop of D1 in Photosystem II stabilizes the quinone-iron electron acceptor complex. Biochemistry 60:53–63. https://doi.org/10.1021/acs.biochem.0c00668
Garcia-Cerdán JG, Furst AL, McDonald KL et al (2019) A thylakoid membrane-bound and redox-active rubredoxin (RBD1) functions in de novo assembly and repair of photosystem II. Proc Natl Acad Sci USA 116:16631–16640. https://www.pnas.org/content/116/33/16631
Govindjee (1995) Sixty-three years since Kautsky: chlorophyll a fluorescence. Aust J Plant Physiol 22:131–160. https://doi.org/10.1071/PP9950131
Guskov A, Kern J, Gabdulkhakov A et al (2009) Cyanobacterial photosystem II at 2.9-Å resolution and the role of quinones, lipids, channels and chloride. Nat Struct Mol Biol 16:334–342. https://doi.org/10.1038/nsmb.1559
Hager M, Hermann M, Biehler K et al (2002) Lack of the small plastid-encoded PsbJ polypeptide results in a defective water-splitting apparatus of Photosystem II, reduced Photosystem I levels, and hypersensitivity to light. J Biol Chem 277:14031–14039. https://doi.org/10.1074/jbc.M112053200
Heinz S, Liauw P, Nickelsen J, Nowaczyk M (2016) Analysis of photosystem II biogenesis in cyanobacteria. Biochim Biophys Acta 1857:274–287. https://doi.org/10.1016/j.bbabio.2015.11.007
Ho FM, Styring S (2008) Access channels and methanol binding to the CaMn4 cluster in photosystem II based on solvent accessibility simulations, with implications for substrate water access. Biochim Biophys Acta 1777:140–153. https://doi.org/10.1016/j.bbabio.2007.08.009
Huang G, Xiao Y, Pi X et al (2021) Structural insights into a dimeric Psb27-photosystem II complex from a cyanobacterium Thermosynechococcus vulcanus. Proc Natl Acad Sci USA 118(5):e2018053118. https://doi.org/10.1073/pnas.2018053118
Jackson SA, Eaton-Rye JJ (2015) Characterization of a Synechocystis sp. PCC 6803 double mutant lacking the CyanoP and Ycf48 proteins of photosystem II. Photosynth Res 124:217–229. https://doi.org/10.1007/s11120-015-0122-0
Jackson SA, Hervey JRD, Dale AJ, Eaton-Rye JJ (2014) Removal of both Ycf48 and Psb27 in Synechocystis sp. PCC 6803 disrupts photosystem II assembly and alters QA- oxidation in the mature complex. FEBS Lett 588:3751–3760. https://doi.org/10.1016/j.febslet.2014.08.024
Kern J, Chatterjee R, Young ID et al (2018) Structures of the intermediates of Kok’s photosynthetic water oxidation clock. Nature 563:421–425. https://doi.org/10.1038/s41586-018-0681-2
Komenda J, Knoppová J, Krynická V et al (2010) Role of FtsH2 in the repair of photosystem II in mutants of the cyanobacterium Synechocystis PCC 6803 with impaired assembly or stability of the CaMn4 cluster. Biochim Biophys Acta 1797:566–575. https://doi.org/10.1016/j.bbabio.2010.02.006
Komenda J, Sobotka R, Nixon PJ (2012) Assembling and maintaining the photosystem II complex in chloroplasts and cyanobacteria. Curr Opin Plant Biol 15:245–251. https://doi.org/10.1016/j.pbi.2012.01.017
Lea-Smith DJ, Bombelli P, Vasudevan R, Howe CJ (2016) Photosynthetic, respiratory and extracellular electron transport pathways in cyanobacteria. Biochim Biophys Acta 1857:247–255. https://doi.org/10.1016/j.bbabio.2015.10.007
Luo H, Jackson SA, Fagerlund RD et al (2014) The importance of the hydrophilic region of PsbL for the plastoqunone electron acceptor complex of photosystem II. Biochim Biophys Acta 1837:1435–1446. https://doi.org/10.1016/j.bbabio.2014.02.015
Mabbitt PD, Wilbanks SM, Eaton-Rye JJ (2014) Structure and function of the hydrophilic Photosystem II assembly proteins: Psb27, Psb28 and Ycf48. Plant Physiol Biochem 81:96–107. https/doi.org/https://doi.org/10.1016/j.plaphy.2014.02.013
MacKinney G (1941) Absorption of light by chlorophyll solutions. J Biol Chem 140:315–322. https://doi.org/10.1016/S0021-9258(18)51320-X
Morris JN, Crawford TS, Jeffs A et al (2014) Whole genome re-sequencing of two ‘wild-type’ strains of the model cyanobacterium Synechocystis sp. PCC 6803. N Z J Bot 52:36–47. https://doi.org/10.1080/0028825X.2013.846267
Morris JN, Eaton-Rye JJ, Summerfield TC (2016) Environmental pH and the requirement for the extrinsic proteins of Photosystem II in the function of cyanobacterial photosynthesis. Front Plant Sci 7:1135. https://doi.org/10.3389/fpls.2016.01135
Morris JN, Kovács S, Vass I et al (2019) Environmental pH and a Glu364 to Gln mutation in the chlorophyll-binding CP47 protein affect reox-active TyrD and charge recombination in Photosystem II. FEBS Lett 593:163–174. https://doi.org/10.1002/1873-3468.13307
Müh F, Renger T, Zouni A (2008) Crystal structure of cyanobacterial photosystem II at 3.0 Å resolution: a closer look at the antenna system and the small membrane-intrinsic subunits. Plant Physiol Biochem 46:238–264. https://doi.org/10.1016/j.plaphy.2008.01.003
Müh F, Zouni A (2016) Cytochrome b559 in photosystem II. In: Cramer WA, Kallas T (eds) Cytochrome complexes: evolution, structures, energy transduction, and signaling, Advances in Photosynthesis and Respiration, Vol 22, Springer, Dordrecht, pp 143–175. https://doi.org/10.1007/978-94-017-7481-9_8
Nickelsen J, Rengstl B (2013) Photosystem II assembly: from cyanobacteria to plants. Annu Rev Plant Biol 64:609–635. https://doi.org/10.1146/annurev-arplant-050312-120124
Nowaczyk MM, Krause K, Mieseler M et al (2012) Deletion of psbJ leads to accumulation of Psb27–Psb28 Photosystem II complexes in Thermosynechococcus elongatus. Biochim Biophys Acta 1817:1339–1343. https://doi.org/10.1016/j.bbabio.2012.02.017
Paerl HW (1984) Cyanobacterial carotenoids: their roles in maintaining optimal production among aquatic bloom forming genera. Oecologia 61:143–149. https://doi.org/10.1007/BF00396752
Regel RE, Ivleva NB, Zer H et al (2001) Deregulation of electron flow within photosystem II in the absence of the PsbJ protein. J Biol Chem 276:41473–41478. https://doi.org/10.1074/jbc.M102007200
Roose JL, Frankel LK, Mummadisetti MP, Bricker TM (2016) The extrinsic proteins of photosystem II: update. Planta 243:889–908. https://doi.org/10.1007/s00425-015-2462-6
Shen J-R (2015) The structure of photosystem II and the mechanism of water oxidation in photosynthesis. Annu Rev Plant Biol 66:23–48. https://doi.org/10.1146/annurev-arplant-050312-120129
Shen J-R, Vermaas W, Inoue Y (1995a) The role of cytochrome c-550 as studied through reverse genetics and mutant characterization in Synechocystis sp. PCC 6803. J Biol Chem 270:6901–6907. https://doi.org/10.1074/jbc.270.12.6901
Shen J-R, Burnap RL, Inoue Y (1995b) An independent role of cytochrome c-550 in cyanobacterial Photosystem II as revealed by double-deletion mutagenesis of the psbO and psbV genes in Synechocystis sp. PCC 6803. Biochemistry 34:12661–12668. https://doi.org/10.1021/bi00039a023
Shen J-R, Ikeuchi M, Inoue Y (1997) Analysis of the psbU gene encoding the 12-kDa extrinsic protein of Photosystem II and studies on its role by deletion mutagenesis in Synechocystis sp. PCC 6803. J Biol Chem 272:17821–17826. https://doi.org/10.1074/jbc.272.28.17821
Shi L-X, Hall M, Funk C, Schröder WP (2012) Photosystem II, a growing complex: updates on newly discovered components and low molecular mass proteins. Biochim Biophys Acta 1817:13–25. https://doi.org/10.1016/j.bbabio.2011.08.008
Suga M, Akita F, Hirata K et al (2015) Native structure of photosystem II at 1.95 Å resolution viewed by femtosecond X-ray pulses. Nature 517:99–103. https://doi.org/10.1038/nature13991
Suga M, Akita F, Sugahara M et al (2017) Light-induced structural changes and the site of O=O bond formation in PS II caught by XFEL. Nature 543:131–135. https://doi.org/10.1038/nature21400
Summerfield TC, Eaton-Rye JJ, Sherman LA (2007) Global gene expression of a ∆PsbO:∆PsbU mutant and a spontaneous revertant in the cyanobacterium Synechocystis sp. strain PCC 6803. Photosynth Res 94:265–274. https://doi.org/10.1007/s11120-007-9237-2
Summerfield TC, Crawford TS, Young RD et al (2013) Environmental pH affects photoautotrophic growth of Synechocystis sp. PCC 6803 strains carrying mutations in the lumenal proteins of photosystem II. Plant Cell Physiol 54:859–874. https://doi.org/10.1093/pcp/pct036
Suorsa M, Regel RE, Paakkarinen V et al (2004) Protein assembly of photosystem II and accumulation of subcomplexes in the absence of low molecular mass subunits PsbL and PsbJ. Eur J Biochem 271:96–107. https://doi.org/10.1046/j.1432-1033.2003.03906.x
Swiatek M, Regel RE, Meurer J et al (2003) Effects of selective inactivation of individual genes for low-molecular-mass subunits on the assembly of Photosystem II, as revealed by chloroplast transformation: the psbEFLJ operon in Nicotiana tabacum. Mol Gen Genom 268:699–710. https://doi.org/10.1007/s00438-002-0791-1
Tóth TN, Chukhutsina V, Domonkos I et al (2015) Carotenoids are essential for the assembly of cyanobacterial photosynthetic complexes. Biochim Biophys Acta 1847:1153–1165. https://doi.org/10.1016/j.bbabio.2015.05.020
Umena Y, Kawakami K, Shen JR, Kamiya N (2011) Crystal structure of oxygen-evolving photosystem II at a resolution of 1.9 Å. Nature 473:55–60. https://doi.org/10.1038/nature09913
Vajravel S, Kis M, Kłodawska K et al (2017) Zeaxanthin and echinenone modify the structure of photosystem I trimer in Synechocystis sp. PCC 6803. Biochim Biophys Acta Bioenerg 1858:510–518. https://doi.org/10.1016/j.bbabio.2017.05.001
Vinyard DJ, Brudvig GW (2018) Progress toward a molecular mechanism of water oxidation in photosystem II. Annu Rev Phys Chem 68:101–116. https://doi.org/10.1146/annurev-physchem-052516-044820
Wei X, Su X, Cao P et al (2016) Structure of spinach photosystem II – LHCII supercomplex at 3.2 Å resolution. Nature 534:69–74. https://doi.org/10.1038/nature18020
Williams JGK (1988) Construction of specific mutations in photosystem II photosynthetic reaction center by genetic engineering methods in Synechocystis 6803. Methods Enzymol 167:766–778
Yu J, Knoppová J, Michoux F et al (2018) Ycf48 involved in the biogenesis of the oxygen-evolving photosystem II complex is a seven-bladed beta-propeller protein. Proc Natl Acad Sci USA 115:E7824–E7833. https://doi.org/10.1073/pnas.1800609115
Zabret J, Bohn S, Schuller SK et al (2021) Structural insights into photosystem II assembly. Nat Plants. 7:524–538. https://doi.org/10.1038/s41477-021-00895
Acknowledgements
PC and JAF contributed equally to this project and are listed as joint first authors in alphabetical order. We thank Dr Sandeep Biswas for assistance with the construction of the plasmids used to delete the genes encoding the PsbO, PsbU and PsbV extrinsic proteins.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Choo, P., Forsman, J.A., Hui, L. et al. The PsbJ protein is required for photosystem II activity in centers lacking the PsbO and PsbV lumenal subunits. Photosynth Res 151, 103–111 (2022). https://doi.org/10.1007/s11120-021-00862-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11120-021-00862-y