Abstract
Quinones serve as redox active cofactors in bacterial photosynthetic reaction centers: photosystem I, photosystem II, cytochrome bc 1, and cytochrome b 6 f. In particular, ubiquinone is ubiquitous in animals and most bacteria and plays a key role in several cellular processes, e.g., mitochondrial electron transport. Their experimentally measured redox potential values for one-electron reduction E m(Q/Q·−) were already reported in dimethylformamide (DMF) versus saturated calomel electrode but not in water versus normal hydrogen electrode (NHE). We calculated E m(Q/Q·−) of 1,4-quinones using a quantum chemical approach. The calculated energy differences of reduction of Q to Q·− in DMF and water for 1,4-quinone derivatives correlated highly with the experimentally measured E m(Q/Q·−) in DMF and water, respectively. E m(Q/Q·−) were calculated to be −163 mV for ubiquinone, −260 mV for menaquinone and phylloquinone, and −154 mV for plastoquinone in water versus NHE.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
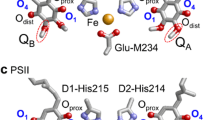
Quinones can accept two electrons and two protons via the initial protonation of semiquinone (Q·− to QH·) and the second protonation of hydroquinone (QH− to QH2). Ubiquinone serves as an electron acceptor at the QA and QB binding sites in reaction centers of purple bacteria (PbRC) from Rhodobacter sphaeroides and serves as an electron donor in cytochrome bc 1. Similarly, menaquinone (vitamin K2) is the acceptor at the QA site in PbRC from Blastochloris viridis, whereas phylloquinone (vitamin K1) is the active center at the A1A and A1B sites in photosystem I (PSI). In reaction centers of green non-sulfur bacteria from Chloroflexus aurantiacus, menaquinones are also located at both QA and QB sites (Hale et al. 1983). It should be noted that phylloquinone and menaquinone have the same head-group structure (Fig. 2). Plastoquinone serves as an electron acceptor at the QA and QB sites in photosystem II (PSII) (Fig. 1) (Robinson and Crofts 1984; Rutherford et al. 1984; Okamura et al. 2000; Brettel and Leibl 2001; Wraight 2004) and serves as an electron donor in cytochrome b 6 f. In PbRC and PSII, both QA and QB are located near the non-heme Fe2+, and the Fe2+ ligands (i.e., His-L190 and His-M217 (or M219) in PbRC and D1-His215 and D2-His214 in PSII) donate an H-bond to the carbonyl O atoms of quinones that are proximal to the Fe complex (Oprox) (Fig. 1a–c). The carbonyl O atoms of quinones at the distal position (Odist) also form H-bonds with the proteins. On the other hand, the non-heme Fe2+ is absent in PSI, but the Fe4S4 cluster FX is located near the two A1 binding sites (Fig. 1d).
Quinones in photosynthetic reaction centers: a menaquinone as QA and ubiquinone as QB in bacterial photosynthetic reaction centers from Blastochloris viridis (Blc. viridis, PDB ID: 2I5N) (Li et al. 2006), b ubiquinone as QA and QB in bacterial photosynthetic reaction centers from Rhodobacter sphaeroides (Rb. sphaeroides, PDB ID: 3I4D), c plastoquinone as QA and QB in PSII (PDB ID: 3ARC) (Umena et al. 2011), and d phylloquinone as A1A and A1B in PSI (PDB ID: 1JB0) (Jordan et al. 2001). Red and blue balls indicate O and N atoms, respectively. In PbRC and PSII, Oprox and Odist stand for O atoms of the quinones at the proximal and distal positions with respect to the non-heme Fe complex, respectively. Note that except for QB in PSII, Oprox is O1 and Odist is O4 in PbRC and PSII
Redox potential values for one-electron reduction, E m(Q/Q·−), for 1,4-quinones, including ubiquinone, menaquinone (phylloquinone), and plastoquinone, were experimentally measured in dimethylformamide (DMF) versus saturated calomel electrode (SCE) by Prince et al. (Prince et al. 1983). E m(Q/Q·−) for 1,4-quinones were also experimentally measured in water versus normal hydrogen electrode (NHE) by Swallow (1982). Since E m values for redox active sites in proteins are often reported as the values measured in water versus NHE, E m(Q/Q·−) for ubiquinone, menaquinone (phylloquinone), and plastoquinone measured in water versus NHE are preferentially required when analyzing interaction between the quinone binding site and the protein environment in PbRC, PSI, PSII, cytochrome bc 1, and cytochrome b 6 f. However, as far as we are aware, experimentally measured E m(Q/Q·−) for ubiquinone, menaquinone (phylloquinone), and plastoquinone in water versus NHE have not been reported (Fig. 2). Here, we report E m(Q/Q·−) for ubiquinone, menaquinone (phylloquinone), and plastoquinone in water versus NHE, obtained using a quantum chemical approach.
Computational procedures
In reduction of the oxidized state (A) to reduced state (A·−) in aqueous solution, the redox potential E m relative to the normal hydrogen electrode (NHE) is defined as
where ΔG aq is the free energy difference between A and A·− [i.e., ΔG aq = G aq(A·−) − G aq(A) − G NHE], n is the number of electron involved in the reaction (i.e., n = 1 in the present case), and F is the Faraday constant. ΔG aq can also be approximated as
where k is the scaling factor, ΔE QM/PCM is the energy difference between A and A·− in aqueous phase [i.e., ΔE QM/PCM = E QM/PCM(A·−) − E QM/PCM(A)], which can be calculated using a quantum chemical approach with the polarizable continuum model (PCM) method, and C is a constant (Matsui et al. 2012; Hasegawa et al. 2017). The Eq. 1 can be written as Eq. 3 using Eq. 2,
where k′ is the scaling factor and C′ is a constant (Matsui et al. 2012; Hasegawa et al. 2017). To determine k′ and C′, we calculated ΔE QM/PCM(DMF) (and ΔE QM/PCM(water)) for ten (nine) 1,4-quinones whose experimentally measured E m(Q/Q·−) are reported for DMF (Prince et al. 1983) [and water (Swallow 1982)].
We employed the unrestricted density functional theory (DFT) method with the B3LYP functional and 6-31g++** basis sets for Q·− (the total spin S = 1/2) and the restricted DFT method for Q (S = 0), using the Gaussian (Frisch et al. 2004) program code with the PCM method. Solvent molecules were considered implicitly, using the SCRF = water option and the SCRF = Dimethylformamide option with the values of 78.3553 for water and 37.219 for DMF for dielectric constant (i.e., default values), respectively. However, it should be noted that it is only one of many internal parameters used to define solvents in the PCM method (Frisch et al. 2004). Thus, simply changing the dielectric constant value will not define a new solvent properly.
Since the isoprene units do not comprised conjugated double bonds, the isoprene side-chain length n (Fig. 2) was set to 1 or 2 for the calculations of ubiquinone, menaquinone (phylloquinone), and plastoquinone similar to previous studies (Hasegawa et al. 2017). This could also reduce the number of possible conformations. In fact, the length of the ubiquinone does not practically affect its energetics, as demonstrated by the similar experimentally measured E m(Q/Q·−) values of ubiquinone-1 and -10 in DMF (−611 and −602 mV versus SCE, respectively) (Prince et al. 1983). It should also be noted that n = 0, which corresponds to 2,3-dimethoxy-5-methyl-1,4-benzoquinone [E m(Q/Q·−) = −539 mV in DMF versus SCE (Prince et al. 1983)], i.e., ubiquinone-0, as presented in Ref. (Cape et al. 2006), is a less relevant representation of E m(Q/Q·−) for quinones in PbRC, PSI, PSII, cytochrome bc 1, and cytochrome b 6 f. Ubiquinone-0 corresponds to 2,3-dimethoxy-5-methyl-1,4-benzoquinone rather than 2,3-dimethoxy-5,6-dimethyl-1,4-benzoquinone [in contrast to the statement in ref. (Prince et al. 1983)].
Results and discussion
Correlation of calculated energies with experimentally measured E m(Q/Q·−) for 1,4-quinones in DMF and water
The calculated ΔE QM/PCM for reduction of deprotonated Q to Q·− for ten 1,4-quinones in DMF (ΔE QM/PCM(DMF)) and water (ΔE QM/PCM(water)) were highly associated with the experimentally measured E m(Q/Q·−) in DMF, ranging from −401 to − 751 mV versus SCE (Prince et al. 1983), and the experimentally measured E m(Q/Q·−) in water, ranging from −240 to 99 mV versus NHE (Swallow 1982), which were best fitted to the following equations (Figs. 3a, b):
a Correlation between experimentally measured E m(Q/Q·−) in DMF versus SCE and calculated ΔE QM/PCM(DMF) (coefficient of determination R 2 = 0.96). ΔE QM/PCM(DMF) can be calculated using a quantum chemical approach with the PCM method for DMF. Closed circles indicate ubiquinone, menaquinone (phylloquinone), and plastoquinone, whereas open circles indicate the other ten 1,4-quinones listed in Table 1 (Prince et al. 1983). The solid line was drawn according to Eq. 4 for the ten 1,4-quinones. b Correlation between experimentally measured E m(Q/Q·−) in water versus NHE and calculated ΔE QM/PCM(water) (coefficient of determination R 2 = 0.98). ΔE QM/PCM(water) can be calculated using a quantum chemical approach with the PCM method for water. Open squares indicate the nine 1,4-quinones listed in Table 1 (Swallow 1982). The solid line was drawn according to Eq. 5 for the nine 1,4-quinones. c Correlation between experimentally measured E m(Q/Q·−) in DMF versus SCE and the experimentally measured E m(Q/Q·−) in water versus NHE (coefficient of determination R 2 = 0.97). Open triangles indicate the nine 1,4-quinones listed in Table 1 (Swallow 1982; Prince et al. 1983). The solid line was drawn according to Eq. 6 for the nine 1,4-quinones
Using Eqs. 4 and 5, the calculated E m(Q/Q··−) in DMF versus SCE and E m(Q/Q·−) in water versus NHE for ten and nine 1,4-quinones, respectively, are listed in Table 1. Our results confirm that Eqs. 4 and 5 can reproduce the experimentally measured E m(Q/Q·−) in DMF versus SCE and E m(Q/Q·−) in water versus NHE, respectively. The overall root mean square deviation between the experimentally measured E m(Q/Q·−) in DMF versus SCE and the calculated E m(Q/Q·−) in DMF versus SCE based on Eq. 4 for the ten 1,4-quinones is ±21 mV. The overall root mean square deviation between the experimentally measured E m(Q/Q·−) in water versus NHE and the calculated E m(Q/Q·−) in water versus NHE based on Eq. 5 for the nine 1,4-quinones is ±16 mV. These deviations are sufficiently small with respect to those obtained in other theoretical studies [e.g., ±131 mV (Schmidt am Busch and Knapp 2005)].
Notably, to obtain Eq. 4, E m(Q/Q·−) for ubiquinone, menaquinone (phylloquinone), and plastoquinone in DMF versus SCE were not included [where k = −32.1 (mV mol/kcal), C = −108.5 (mV), excluding ubiquinone, menaquinone (phylloquinone), and plastoquinone]. Nevertheless, the experimentally measured E m(Q/Q·−) in DMF versus SCE (Prince et al. 1983) and the calculated ΔE QM/PCM(DMF) for ubiquinone, menaquinone (phylloquinone), and plastoquinone can also be described by Eq. 4 [where k = −31.8 (mV mol/kcal), C = −108.8 (mV), including ubiquinone, menaquinone (phylloquinone), and plastoquinone] (Fig. 3a), which demonstrates that E m(Q/Q·−) for ubiquinone, menaquinone (phylloquinone), and plastoquinone can be described accurately by Eqs. 4 and 5.
In contrast to other quantum chemical approaches, e.g., (Schmidt am Busch and Knapp 2005), the present approach neither need to calculate the zero-point vibrational energy and the excess vibrational free energy at 298 K for both Q and Q·− nor optimize the atomic radii of Q/Q·− for the solvation energy. Once k′ and C′ are uniquely determined, E m(Q/Q·−) can be accurately calculated based on calculated ΔE QM/PCM, without considering further details of Q/Q·− and solvent. The strong correlation between experimentally measured E m(Q/Q·−) and calculated ΔE QM/PCM (Fig. 3), in turn, suggests that k′ and C′ are similar for these 1,4-quinones.
In the present study, solvent molecules were considered implicitly. This treatment is more appropriate to describe H-bonds between quinones and bulk water/solvent molecules, in which the H-bond patterns are not unique, e.g., bulk solvent. Explicit water/solvent models may be able to describe H-bonds adequately when the H-bond pattern is unique [e.g., water molecules in the well-ordered cluster near the Mn4CaO5 cluster (Saito et al. 2011; Sakashita et al. 2017)] or all possible (H-bond) conformations of water/solvent molecules can be evaluated, e.g., using molecular dynamics simulations; this is not the case for 1,4-quinones investigated in the present study.
We found that the experimentally measured E m(Q/Q·−) for the nine 1,4-quinones in water versus NHE (Swallow 1982) and DMF versus SCE (Prince et al. 1983) correlated strongly (Fig. 3c), which were best fitted to the following equation:
Equation 6 indicates that experimentally measured E m(Q/Q·−) in DMF versus SCE can be practically converted to E m(Q/Q·−) in water versus NHE by adding 480 mV. The E m difference of 480 mV may also contain a liquid junction potential between SCE in DMF and NHE in water. The liquid junction potential can be ignored when E m(Q/Q·−) are compared versus ferrocene (Fc/Fc+); e.g., E m(Q/Q·−) for 1,4-benzoquinone is experimentally measured to be −401 mV in DMF versus SCE, where E m(Fc/Fc+) = 524 mV (Prince et al. 1983). Since E m(Fc/Fc+) = 400 mV in water versus NHE (Koepp et al. 1960), E m(Q/Q·−) for 1,4-benzoquinone is −925 mV in DMF versus Fc/Fc+ and 301 mV in water versus Fc/Fc+, which indicates that E m(Q/Q·−) for 1,4-benzoquinone in DMF and water originally differ by 624 mV in the absence of the liquid junction potential (Table 2). This holds true for all 1,4-quinones investigated. It seems likely that E m(Q/Q·−) for 1,4-benzoquinones already differ by 600 mV even in the absence of the liquid junction potential (Table 2). The presence of H-bond donor to Q·− in water is partly responsible for the E m difference of 600 mV, since the presence of H-bond donor to Q·− stabilizes Q·− and increases E m(Q/Q·−). Nevertheless, the entire difference of 600 mV would not be explained solely by the first sphere water molecules that can directly form an H-bond with Q·−. The surrounding water molecules (e.g., second and third sphere molecules) cannot directly form an H-bond with Q·− but the Q·− stabilization is pronounced by their dipole orientations (Takaoka et al. 2016). The corresponding effect may be ignored in DMF with respect to water.
As far as only E m differences among the redox active cofactors (ΔE m) are discussed in the same proteins, e.g., along the electron transfer chains, E m values of isolated cofactors measured in DMF, which are reported also for chlorophylls (Watanabe and Kobayashi 1991), might possibly be useful. On the other hand, when E m values in the protein environments are discussed, comparison with E m values of isolated cofactors measured in water is recommended, since E m values measured in DMF is originally 600 mV lower than those measured in water even in the absence of the liquid junction potential (Table 2).
E m(Q/Q·−) for ubiquinone, menaquinone, phylloquinone, and plastoquinone in water versus NHE
To the best of our knowledge, experimentally measured E m(Q/Q·−) for ubiquinone, menaquinone (phylloquinone), and plastoquinone in water versus NHE are not reported. By calculating ΔE QM/PCM(water) and using Eq. 5, E m(Q/Q·−) was calculated to be −163 mV for ubiquinone, −260 mV for menaquinone (phylloquinone), and −154 mV for plastoquinone in water versus NHE (Table 1).
In ubiquinone, one of the 2,3-methoxy groups lies outside the quinone ring. Hence, Zhu and Gunner proposed that E m(Q/Q·−) for ubiquinone, a 2,3-dimethoxy-5-methyl-6-isoprenyl benzoquinone, is more similar to the E m(Q/Q·−) for trimethyl-benzoquinone than to the E m(Q/Q·−) for tetramethyl-benzoquinone (Zhu and Gunner 2005). Indeed, the calculated E m(Q/Q·−) = −163 mV for ubiquinone (Table 1) is close to the experimentally measured E m(Q/Q·−) = −165 mV for trimethyl-benzoquinone (Swallow 1982) in water versus NHE, which is consistent with their proposal. Although it was proposed that difference in the 2-methoxy orientation of ubiquinone was responsible for the E m difference of more than 160 mV between QA and QB in PbRC (Taguchi et al. 2013), the similar E m(Q/Q·−) values of trimethyl-benzoquinone and ubiquinone (ref. (Zhu and Gunner 2005) and Table 1) suggest that contributions of methoxy and methyl groups to E m(Q/Q·−) are not significantly different. It should also be noted that estimation by Swallow resulted in a more negative value of E m(Q/Q·−) = −230 ± 20 mV for ubiquinone at pH 7 (Swallow 1982).
The present study shows that E m(Q/Q·−) is −260 mV for menaquinone (phylloquinone) in water versus NHE (Table 1); the calculated E m(Q/Q·−) can be confirmed by Eq. 6, which can be reproduced by adding 480 mV to E m(Q/Q·−) in DMF versus SCE. Previously, Ptushenko et al. considered that E m(Q/Q·−) was −800 mV for phylloquinone in DMF versus NHE by considering a liquid junction potential between SCE in DMF and NHE in water (Ptushenko et al. 2008). Using the low E m(Q/Q·−) value of −800 mV for phylloquinone in DMF versus NHE, they obtained E m(A1A) = −671 mV and E m(A1B) = −844 mV (Ptushenko et al. 2008), and were able to reproduce the reported low E m(A1) in PSI [e.g., −810 mV (Vos and van Gorkom 1990), −754 mV (Iwaki and Itoh 1994), and lower than −700 mV (Brettel and Leibl 2001)]. This, in turn, suggests that the electrostatic interaction of the PSI protein environment at the A1 site is remarkably weak in their computational model. If this is the case, then E m(QA) of −150 mV for the same quinone species (menaquinone) would be regarded as being “unusually high” in PbRC from Blastochloris viridis (Brettel and Leibl 2001), and the PbRC protein environment must dramatically increase E m(Q/Q·−) for menaquinone by more than 600 mV at the QA site in their computational model; obviously this is not the case for the PbRC protein environment, as already demonstrated in theoretical studies (Rabenstein et al. 1998; Ishikita and Knapp 2004; Zhu and Gunner 2005). E m(Q/Q·−) = −260 mV for menaquinone (phylloquinone) in water versus NHE (Table 1) suggests that the PSI protein environment (e.g., the presence of negatively charged FX near A1 (Ishikita and Knapp 2003)) is responsible for low E m(A1) in PSI. When E m(Q/Q·−) = −800 mV for phylloquinone in DMF versus NHE is used, the resulting E m(A1) should contain the E m downshift of ca. 600 mV with respect to water versus NHE as an artifact (Table 2), since the PSI is not solvated in DMF but in water in the thylakoid membrane. One can directly focus on the influence of the PSI protein environment on E m(A1) when using E m(Q/Q·−) = −260 mV in water versus NHE. It seems plausible that using E m values measured in water is more recommended to analyze E m values for the redox active groups in proteins unless the proteins are solvated in DMF.
This fact would be more obvious when considering E m of heme proteins or flavin-binding proteins. E m of heme (Harbury and Loach 1960; Wilson 1983) and flavin (Draper and Ingraham 1968; Anderson 1983) were experimentally measured in water. These cofactors are often largely exposed to the protein bulk surface [e.g., heme (Kerfeld et al. 2003; Clarke et al. 2011) and flavin-binding (Ludwig et al. 1997; Watt et al. 1991) proteins]. As these cofactors are released away from the binding site toward the bulk region, the E m values must be close to those experimentally measured in water; this is exactly the case for QB in PbRC and PSII, which is located near the protein bulk surface. Using spectroelectrochemistry, Kato et al. directly determined E m(QB) to be +90 mV in PSII from Thermosynechococcus elongates versus NHE (Kato et al. 2016). E m(Q/Q·−) is −154 mV for plastoquinone in water versus NHE (Table 1) and would be −750 mV in DMF versus NHE (assuming E m downshift of ca. 600 mV, Table 2). If E m(Q/Q·−) measured in DMF were relevant, the PSII protein environment would need to increase E m(Q/Q·−) for plastoquinone by 840 mV at the QB site. In addition, E m(QA) was determined to be −145 mV in spinach PSII versus NHE, using spectroelectrochemistry (Brinkert et al. 2016); the PSII protein environment would also need to increase E m(Q/Q·−) for plastoquinone by 600 mV even at the QA site, which is less exposed to the protein bulk surface. It seems likely that E m values for quinones measured in water are more recommended when comparing with E m(Q/Q·−) in the protein environments. This would also hold true for the quinone binding sites in cytochrome bc 1 and cytochrome b 6 f, at which quinones from PbRC and PSII can bind, respectively.
Conclusion
Experimentally measured E m(Q/Q·−) in DMF versus SCE (Prince et al. 1983) and E m(Q/Q·−) in water versus NHE (Swallow 1982) correlated highly with the quantum chemically calculated energy differences (ΔE QM/PCM) between neutral and reduced states (Figs. 3a, b) and can be best fitted to Eqs. 4 and 5, respectively. It seems likely that E m(Q/Q·−) for 1,4-benzoquinones differ by 600 mV even in the absence of the liquid junction potential between DMF and NHE (versus Fc/Fc+). E m(Q/Q·−) was calculated to be −163 mV for ubiquinone, −260 mV for menaquinone (phylloquinone), and −154 mV for plastoquinone in water versus NHE (Table 1). In particular, E m(Q/Q·−) = −260 mV for phylloquinone in water versus NHE unambiguously demonstrates that remarkably low E m(A1) in PSI does not originate from E m(Q/Q·−) for phylloquinone but from interaction with the PSI protein environment, as suggested previously (Ishikita and Knapp 2003). These E m(Q/Q·−) are prerequisite for analyzing the E m(Q/Q·−) shift caused by electrostatic interactions within the protein environment in photosynthetic reaction centers.
References
Anderson RF (1983) Energetics of the one-electron reduction steps of riboflavin, FMN and FAD to their fully reduced forms. Biochim Biophys Acta 722:158–162
Brettel K, Leibl W (2001) Electron transfer in photosystem I. Biochim Biophys Acta 1507:100–114
Brinkert K, De Causmaecker S, Krieger-Liszkay A, Fantuzzi A, Rutherford AW (2016) Bicarbonate-induced redox tuning in Photosystem II for regulation and protection. Proc Natl Acad Sci USA 113(43):12144–12149. doi:10.1073/pnas.1608862113
Cape JL, Bowman MK, Kramer DM (2006) Computation of the redox and protonation properties of quinones: towards the prediction of redox cycling natural products. Phytochemistry 67(16):1781–1788. doi:10.1016/j.phytochem.2006.06.015
Clarke TA, Edwards MJ, Gates AJ, Hall A, White GF, Bradley J, Reardon CL, Shi L, Beliaev AS, Marshall MJ, Wang Z, Watmough NJ, Fredrickson JK, Zachara JM, Butt JN, Richardson DJ (2011) Structure of a bacterial cell surface decaheme electron conduit. Proc Natl Acad Sci USA 108(23):9384–9389. doi:10.1073/pnas.1017200108
Draper RD, Ingraham LL (1968) A potentiometric study of the flavin semiquinone equilibrium. Arch Biochem Biophys 125:802–808
Frisch MJ, Trucks GW, Schlegel HB, Scuseria GE, Robb MA, Cheeseman JR, Montgomery JA Jr, Vreven T, Kudin KN, Burant JC, Millam JM, Iyengar SS, Tomasi J, Barone V, Mennucci B, Cossi M, Scalmani G, Rega N, Petersson GA, Nakatsuji H, Hada M, Ehara M, Toyota K, Fukuda R, Hasegawa J, Ishida M, Nakajima T, Honda Y, Kitao O, Nakai H, Klene M, Li X, Knox JE, Hratchian HP, Cross JB, Bakken V, Adamo C, Jaramillo J, Gomperts R, Stratmann RE, Yazyev O, Austin AJ, Cammi R, Pomelli C, Ochterski JW, Ayala PY, Morokuma K, Voth GA, Salvador P, Dannenberg JJ, Zakrzewski VG, Dapprich S, Daniels AD, Strain MC, Farkas O, Malick DK, Rabuck AD, Raghavachari K, Foresman JB, Ortiz JV, Cui Q, Baboul AG, Clifford S, Cioslowski J, Stefanov BB, Liu G, Liashenko A, Piskorz P, Komaromi I, Martin RL, Fox DJ, Keith T, Al-Laham MA, Peng CY, Nanayakkara A, Challacombe M, Gill PMW, Johnson B, Chen W, Wong MW, Gonzalez C, Pople JA (2004). Gaussian 03. Gaussian, Inc., Wallingford CT
Hale MB, Blankenship RE, Fuller RC (1983) Menaquinone is the sole quinone in the facultatively aerobic green photosynthetic bacterium Chloroflexus aurantiacus. Biochim Biophys Acta 723(3):376–382. doi:10.1016/0005-2728(83)90044-0
Harbury HA, Loach PA (1960) Oxidation-linked proton functions in heme octa- and undecapeptides from mammalian cytochrome c. J Biol Chem 235:3640–3645
Hasegawa R, Saito K, Takaoka T, Ishikita H (2017) pK a of ubiquinone, menaquinone, phylloquinone, plastoquinone, and rhodoquinone in aqueous solution. Photosynth Res 133(1):297–304. doi:10.1007/s11120-017-0382-y
Ishikita H, Knapp E-W (2003) Redox potential of quinones in both electron transfer branches of photosystem I. J Biol Chem 278:52002–52011
Ishikita H, Knapp E-W (2004) Variation of Ser-L223 hydrogen bonding with the QB redox state in reaction centers from Rhodobacter sphaeroides. J Am Chem Soc 126:8059–8064
Iwaki M, Itoh S (1994) Reaction of reconstituted acceptor quinone and dynamic equilibration of electron transfer in the photosystem I reaction center. Plant Cell Physiol 35:983–993
Jordan P, Fromme P, Witt HT, Klukas O, Saenger W, Krauss N (2001) Three-dimensional structure of cyanobacterial photosystem I at 2.5 Å resolution. Nature 411:909–917
Kato Y, Nagao R, Noguchi T (2016) Redox potential of the terminal quinone electron acceptor QB in photosystem II reveals the mechanism of electron transfer regulation. Proc Natl Acad Sci USA 113(3):620–625. doi:10.1073/pnas.1520211113
Kerfeld CA, Sawaya MR, Bottin H, Tran KT, Sugiura M, Cascio D, Desbois A, Yeates TO, Kirilovsky D, Boussac A (2003) Structural and EPR characterization of the soluble form of cytochrome c-550 and of the psbV2 gene product from cyanobacterium Thermosynechococcus elongatus. Plant Cell Physiol 44:697–706
Koepp HM, Wendt H, Stkehlow H (1960) Der Vergleich der Spannungsreihen in verschiedenen Solventien. II. Zeitschrift für Elektrochemie. Berichte der Bunsengesellschaft für physikalische Chemie 64 (4):483–491. doi:10.1002/bbpc.19600640406
Li L, Mustafi D, Fu Q, Tereshko V, Chen DL, Tice JD, Ismagilov RF (2006) Nanoliter microfluidic hybrid method for simultaneous screening and optimization validated with crystallization of membrane proteins. Proc Natl Acad Sci USA 103(51):19243–19248. doi:10.1073/pnas.0607502103
Ludwig ML, Pattridge KA, Metzger AL, Dixon MM, Eren M, Feng Y, Swenson RP (1997) Control of oxidation-reduction potentials in flavodoxin from Clostridium beijerinckii: the role of conformation changes. Biochemistry 36:1259–1280
Matsui T, Baba T, Kamiya K, Shigeta Y (2012) An accurate density functional theory based estimation of pK a values of polar residues combined with experimental data: from amino acids to minimal proteins. Phys Chem Chem Phys 14(12):4181–4187
Okamura MY, Paddock ML, Graige MS, Feher G (2000) Proton and electron transfer in bacterial reaction centers. Biochim Biophys Acta 1458:148–163
Prince RC, Dutton PL, Bruce JM (1983) Electrochemistry of ubiquinones: menaquinones and plastoquinones in aprotic solvents. FEBS Lett 160:273–276
Ptushenko VV, Cherepanov DA, Krishtalik LI, Semenov AY (2008) Semi-continuum electrostatic calculations of redox potentials in photosystem I. Photosynth Res 97(1):55–74. doi:10.1007/s11120-008-9309-y
Rabenstein B, Ullmann GM, Knapp E-W (1998) Energetics of electron-transfer and protonation reactions of the quinones in the photosynthetic reaction center of Rhodopseudomonas viridis. Biochemistry 37:2488–2495
Robinson HH, Crofts AR (1984) Kinetics of proton uptake and the oxidation-reduction reactions of the quinone acceptor complex of photosystem II from pea chloroplasts. In: Sybesma C (ed) Advances in photosynthesis research, vol 1. Martinus Nijhoff/Dr. W. Junk Publishers, The Hague, pp 477–480
Rutherford AW, Renger G, Koike H, Inoue Y (1984) Thermoluminescence as a probe of photosystem II. The redox and protonation states of the secondary acceptor quinone and the O2-evolving enzyme. Biochim Biophys Acta 767(3):548–556
Saito K, Shen J-R, Ishida T, Ishikita H (2011) Short hydrogen-bond between redox-active tyrosine YZ and D1-His190 in the photosystem II crystal structure. Biochemistry 50:9836–9844
Sakashita N, Watanabe HC, Ikeda T, Saito K, Ishikita H (2017) Origins of water molecules in the photosystem II crystal structure. Biochemistry 56(24):3049–3057. doi:10.1021/acs.biochem.7b00220
Schmidt am Busch M, Knapp E-W (2005) One-electron reduction potential for oxygen- and sulfur-centered organic radicals in protic and aprotic solvents. J Am Chem Soc 127:15730–15737
Swallow AJ (1982) Physical chemistry of semiquinones. In: Trumpower BL (ed) Function of quinones in energy conserving systems. Academic Press, New York, pp 59–72
Taguchi AT, Mattis AJ, O’Malley PJ, Dikanov SA, Wraight CA (2013) Tuning cofactor redox potentials: the 2-methoxy dihedral angle generates a redox potential difference of >160 mV between the primary (QA) and secondary (QB) quinones of the bacterial photosynthetic reaction center. Biochemistry 52(41):7164–7166. doi:10.1021/bi4011896
Takaoka T, Sakashita N, Saito K, Ishikita H (2016) pK a of a proton-conducting water chain in photosystem II. J Phys Chem Lett 7:1925–1932. doi:10.1021/acs.jpclett.6b00656
Umena Y, Kawakami K, Shen J-R, Kamiya N (2011) Crystal structure of oxygen-evolving photosystem II at a resolution of 1.9 Å. Nature 473:55–60
Vos MH, van Gorkom HJ (1990) Thermodynamical and structural information on photosynthetic systems obtained from electroluminescence kinetics. Biophys J 58(6):1547–1555. doi:10.1016/S0006-3495(90)82499-1
Watanabe T, Kobayashi M (1991) Electrochemistry of chlorophylls. In: Scheer H (ed) Chlorophylls. CRC Press, Boca Raton, pp 287–303
Watt W, Tulinsky A, Swenson RP, Watenpaugh KD (1991) Comparison of the crystal structures of a flavodoxin in its three oxidation states at cryogenic temperatures. J Mol Biol 218:195–208
Wilson GS (1983) Electrochemical studies of porphyrin redox reactions as cytochrome models. Bioelectrochem Bioenerg 1:172–179
Wraight CA (2004) Proton and electron transfer in the acceptor quinone complex of photosynthetic reaction centers from Rhodobacter sphaeroides. Front Biosci 9:309–337
Zhu Z, Gunner MR (2005) Energetics of quinone-dependent electron and proton transfers in Rhodobacter sphaeroides photosynthetic reaction centers. Biochemistry 44:82–96
Acknowledgements
This research was supported by JST CREST (JPMJCR1656), JSPS KAKENHI (JP26800224 to K.S., JP16H06560 to K.S and H.I., JP17H05721 to Y.K., and JP26105012 to H.I.), Japan Agency for Medical Research and Development (AMED), Materials Integration for engineering polymers of Cross-ministerial Strategic Innovation Promotion Program (SIP), and Interdisciplinary Computational Science Program in CCS, University of Tsukuba.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
Open Access This article is distributed under the terms of the Creative Commons Attribution 4.0 International License (http://creativecommons.org/licenses/by/4.0/), which permits unrestricted use, distribution, and reproduction in any medium, provided you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made.
About this article
Cite this article
Kishi, S., Saito, K., Kato, Y. et al. Redox potentials of ubiquinone, menaquinone, phylloquinone, and plastoquinone in aqueous solution. Photosynth Res 134, 193–200 (2017). https://doi.org/10.1007/s11120-017-0433-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11120-017-0433-4