Abstract
Intercropping is of interest to farmers and researchers because of the increase in productivity observed in these systems. Beneficial soil microorganisms present in the microbiome could play a role in increased crop yields as they improve plant growth, provide protection from soilborne pathogens, and aid with drought stress. Therefore, research has been done to determine if these advantages are observed in intercropping systems and how it impacts the makeup and function of the soil microbiome. This review covers current findings on how the soil microbiome is impacted by intercropping regarding its roles in nutrient availability, plant stress responses, and if the soil microbiome can be altered to further improve plant success.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Intercropping, the practice of growing two or more crops together, has been of increasing interest to farmers due to improvements in yield and resource use efficiency (Dowling et al. 2021). The research focus of various intercropping systems is on evaluating land equivalent ratio (LER) with an emphasis on productivity (Dowling et al. 2021; Fletcher et al. 2015, 2016). Typically, the LER is greater than 1, showing there was an overall yield increase in the intercropping system compared to monoculture (Duchene et al. 2017; Fletcher et al. 2015, 2016). A meta-analysis on intercropping showed that it increased crop yield by 8.9% compared to monoculture (Chen et al. 2023). Microorganisms present in the soil microbiome could be responsible for this increased productivity, with the soil microbiome being defined as all microorganisms present in this environment and their genes. However, most studies demonstrating benefits from soil microorganisms have been limited to monoculture systems.
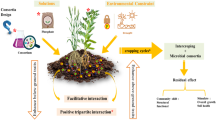
In monoculture, plant growth-promoting rhizobacteria (PGPR) and fungi benefit plants by improving availability of essential plant nutrients (N and P), and mitigating abiotic and biotic stressors such as drought, soil pollution, and pathogens (Ayangbenro and Babalola 2021; Bittencourt et al. 2023; Chepsergon and Moleleki 2023; Gorka et al. 2019). These functions can improve plant growth in addition to producing plant growth regulators and inducing plant hormones (Fig. 1A; Abdelaal et al. 2021; Bittencourt et al. 2023; Wasternack 2014). Protection from abiotic and biotic stressors is caused by the secretion of exopolysaccharides by PGPR in addition to the regulation of soil enzyme activity (Fig. 1A; Bittencourt et al. 2023; Ju et al. 2020; Morcillo and Manzanera 2021). These benefits likely exist in polyculture, but the degree to which they do is still being investigated. There is some evidence that microorganisms are important to the success of intercrops. In the pea-canola system, LER is highest under minimal N fertilizer and fungicide usage (Dowling et al. 2021). Literature reviewed by Dettweiler et al. (2023) found similar detrimental impacts of fertilizer application in casava-legume intercropping systems. This suggests that beneficial microorganisms could be responsible for increased productivity since fungicides kill beneficial fungi, and N and P inputs downregulate relationships with rhizobia and arbuscular mycorrhizal fungi (AMF) (Qin et al. 2015, 2019; Verzeaux et al. 2017). Resource use efficiency by plants in intercrops may also be improved due to shifts in microbial diversity and abundance (Fig. 1B; Duchene et al. 2017).
A Plant-microbe interactions help facilitate growth increases in plants and improve responses and protection from biotic and abiotic stressors. B Shifts in the microbiome under intercropping may lead to increased nutrient availability and stress tolerance due to reductions in pathogens from competition and resource sharing via mycorrhizal networks.
Furthering our understanding of how the soil microbiome contributes to intercropping success will allow us to promote beneficial changes to the microbiome (Duchene et al. 2017). This will have further impacts since decreasing synthetic inputs will improve the sustainability of agricultural practices as they have negative environmental impacts such as eutrophication of aquatic ecosystems and soil acidification (Sharma et al. 2022). Decreased necessity for these synthetic inputs partnered with increased land productivity also leads to larger profit margins for producers (Dowling et al. 2021). This could improve producer wellbeing as socioeconomic pressures negatively impact their mental health, especially with the challenges of climate change (Daghagh Yazd et al. 2020).
This mini-review explores our current understanding of the role the soil microbiome plays in intercropping success of legume non-legume systems. We focus on how the soil microbiome (1) changes between monoculture and intercropped systems; (2) improves nutrient availability in intercropped systems; (3) contributes to improved plant stress responses in intercropped systems; and (4) could be altered to improve plant success.
Does the microbiome change significantly between monoculture and intercropping systems?
In various intercropping systems, fungal and bacterial communities have been characterized to address how intercropping changes the microbiome. Overall, bacterial community composition was impacted by intercropping with bacteria found to be important to the decomposition of organic matter, N fixation, denitrification, plant growth promotion, or metal detoxification being enriched (Table 1). Fungal communities were measured less often and were generally found to have higher diversity and greater changes to community composition in intercropping systems, with enriched fungi being saprotrophic, mycorrhizal, or biocontrol (Table 1). However, pathogens were enriched in some intercropping systems (Table 1). Meta-analyses have also confirmed the positive correlation between intercropping and bacterial and fungal abundance as measured by phospholipid fatty acids (Morugán-Coronado et al. 2022).
Changes observed in microbial communities in intercropping systems are correlated with changes in soil physicochemical properties such as pH, nutrient content, and enzyme activity (Chen et al. 2023; Madsen et al. 2022; Malviya et al. 2021). These changes provide a benefit to crops following their planting known as the plant-soil feedback effect (Wang et al. 2020). Therefore, plants in soils that have a history of intercropping can call on a more robust microbiome that can aid in stress responses and plant nutritional needs (Bakker et al. 2018).
Does the microbiome improve nutrient availability in intercropped systems?
Intercropping has been observed to increase soil available C and N in addition to C and N in the microbial biomass (Chen et al. 2023). It is known that intercropping increases the abundance of plant growth promoting-bacteria (PGPB) which impacts soil nutrient content due to their involvement in N fixation, nutrient uptake, plant hormone production, and the regulation of soil enzymes (Table 2; Bittencourt et al. 2023; Malviya et al. 2021; Solanki et al. 2020). A meta-analysis confirmed the positive impacts of intercropping on N-targeting enzymes N-acetyl-glucosaminidase, protease, and urease (Chen et al. 2023). The abundance of saprotrophic and mycorrhizal fungi also increases under intercropping, further enhancing the release and uptake of plant nutrients (Mwakilili et al. 2021; Malviya et al. 2021; Wang et al. 2020; Yang et al. 2022a). The microbiome is thus often credited for improved nutrient availability in intercrops, contributing to overyielding. Intercrops often have increased nutrient uptake of N and Fe, leading to improved crop nutritional quality (Dai et al. 2019; Sun et al. 2022a).
Studies manipulating the microbiome have been done to determine their role in nutrient uptake and availability in intercropping systems. Qiao et al. (2022) manipulated the microbiome and P availability and found that overyielding in relation to P availability was only observed in unsterilized conditions, with the best yields observed in low P environments. They also found that less diverse microbial communities resulted in improved wheat growth and P uptake compared to more complex microbial communities. This could be due to the enrichment of specific microorganisms that thrive in low P environments, as these microorganisms tend to contain genes which are important to P-cycling encoding carbon-phosphorus lyase, phosphonotase degradation pathways, and membrane-bound quinoprotein glucose dehydrogenase encoded by gcd (Oliverio et al. 2020; Wu et al. 2022). Further research found that the abundance of the microbial gene ppa, which encodes inorganic pyrophosphatase that hydrolyses inorganic P into P, and bacteria harboring this gene were increased by intercropping with a legume and AMF inoculation (Liao et al. 2023). Therefore, it is possible that intercropping creates an environment where P-cycling bacteria thrive improving the availability of P and contributing to overyielding.
Evidence for N transfer from a legume to non-legume in intercropping systems has been observed in a barley-pea intercropping system, with 11.1% of N symbiotically fixed by pea being transferred to barley in a 1:1 planting system (Chapagain and Riseman 2014). This could occur through direct transfer facilitated by mycorrhizal networks, indirect transfer through root exudates and the decomposition of legume roots and nodules, and indirect transfer from increased N mineralization by mycorrhizae (Fig. 2; Homulle et al. 2022). To determine if N is being transferred directly between the legume and non-legume in intercropping systems, isotopically labeled N (15N) is used and is either provided to the legume or the 15N natural abundance method is used in the field (Homulle et al. 2022; Isaac et al. 2012; Ingraffia et al. 2019; Tsialtas et al. 2018). Evidence for the direct transfer of N via AMF hyphal networks has been observed as inoculation with AMF increases N transfer between soybean and maize, and faba bean and wheat (Wang et al. 2016; Ingraffia et al. 2019). Co-inoculation of AMF and rhizobia was found to produce the highest level of N transfer (Wang et al. 2016). Other studies have demonstrated direct N transfer from the legume to non-legume (Homulle et al. 2022; Isaac et al. 2012). In intercropping systems where crops are spatially close or have direct root contact, indirect transfer via nodule and root decomposition would be possible in addition to direct transfer (Fig. 2).
Mycorrhizal networks may be important to P transfer between plants in intercrops (Dowling et al. 2021; Homulle et al. 2022; Wang et al. 2016). Overyielding cannot be explained solely by direct transfer by AMF since intercrops involving non-mycorrhizal plants still experience increased LERs (Madsen et al. 2022). This suggests that other mechanisms (indirect transfer and changes to nutrient availability) can be important to intercrop success. Further work should be performed to determine the importance of N transfer as it would improve our understanding of the role of plant-plant facilitation via the microbiome and mycorrhizae.
Does the microbiome contribute to improved plant stress responses in intercropped systems?
The soil microbiome may improve performance of intercrops in salt and drought stressed conditions (Homulle et al. 2022; Shi et al. 2022). Under drought stress, plants exchange water via mycorrhizal networks, which can be especially beneficial when intercropping a shallow-rooted plant with a deep-rooted plant. The deep-rooted plant can bring deeper water sources to the shallow-rooted plant through hydraulic lift, with transfer of water between plants occurring via mycorrhizal networks (Homulle et al. 2022). Under salt stressed conditions, total P and NH4+-N were significantly higher in the intercropped peanut-sorghum system compared to monoculture peanut, and peanut pod yield increased (Table 2; Shi et al. 2022). Soil nitrate reductase and soil fructose-1,6-biphosphate aldolase enzyme were the only enzymes found to be significantly more active in the intercrop than in peanut monoculture (Shi et al. 2022). The activities of soil protease and soil polyphenol oxidase were significantly less under salt stress, but the activity of soil urease was increased (Shi et al. 2022) Functional analyses of genes showed the involvement of bacteria and fungi in nutrient cycling processes matching the differences observed in soil nutrient content and enzymatic activity (Shi et al. 2022). Both benefits will become increasingly important due to changing climate conditions and increased issues with saline soils (Dowling et al. 2021; Shi et al. 2022).
The microbiome in intercropping systems could also be responsible for disease suppression. Two studies, one conducted on banana intercropped with five different legumes and one conducted on faba bean intercropped with wheat or maize, showed that intercropping decreased the abundance of Fusarium oxysporum from monoculture soils and decreased the disease incidence improving plant performance (Wang et al. 2020; Yang et al. 2022a). The conclusion was that F. oxysporum’s abundance was likely negatively impacted by changes to soil properties (organic matter and NH4+-N content) induced by intercropping and competition with other fungal species present in the more diverse microbial community (Wang et al. 2020; Yang et al. 2022a). This is supported by literature reviewed by Zhu and Morel (2019) which found that beneficial bacteria attracted by intercrops could be responsible for decreased disease incidence. Literature reviewed by De Corato (2020) also suggested the release of nutrients and competition as other mechanisms of disease suppression. This is supported by Sun et al. (2022b) as they found a decrease in gene copies of F. oxysporum in addition to four bacteria isolated from intercropped soils that demonstrated antagonistic traits towards F. oxysporum, providing further evidence for the role of the microbiome in reducing disease. Additional research should be performed on the role of the intercrop microbiome in suppressing other relevant soilborne pathogens to further our understanding of this mechanism.
Can the microbiome be altered in intercropped systems to improve plant success?
Given that the microbiome contributes to the success of intercropping systems, we could manipulate the microbiome to amplify plant success. So far, we know that successful intercrops require selecting crops that fill different niches and have compatible root traits and secondary metabolites (Yu et al. 2022). Information on how to manipulate the microbiome is scarce, although research on inoculums has been done. A study on intercropped black cumin and fenugreek inoculated with AMF (Funneliformis mosseae and Rhizophagus irregularis) or PGPB (Pantoea agglomerans, Pseudomonas putida, and Azotobacter vinelandii) that solubilize P and fix N revealed that crop quality was improved, and LER (1.44) was the highest with bacterial inoculum and a planting ratio of 66:34 (Rezaei-Chiyaneh et al. 2021). Similar results were found in a peanut-maize intercropping system inoculated with Azotobacter chroococcum as inoculation increased the LER by 12–16% compared to uninoculated plots, with the highest LER of 1.70 seen with 100% peanut and 50% maize (Pourjani et al. 2022). This pattern also persisted in a durum wheat-faba bean intercropping system co-inoculated with PDP13 (Rahnella aquatilis) and PS11 (Pseudomonas sp.) showing the best plant performance (Bechtaoui et al. 2019). Aside from the ability of microorganisms to improve nutrient availability through N fixation and P solubilization, it is possible that PGPB are improving plant growth and production by producing plant growth hormones (Pourjani et al. 2022). Microbial communities can also be manipulated through host plant selection (Tosi et al. 2020). Further work on altering the microbiome by inoculation or other agronomic practices such as cover cropping, no-till, or soil amendments is needed. Reduced tillage has been shown to have a positive impact on bacterial and fungal abundance making it especially promising to pair with intercropping practices (Morugán-Coronado et al. 2022). Synthetic communities made up of known PGPB could also prove to be beneficial to intercrop success through their enrichment. Research should be done to explore this further.
Conclusion
The soil microbiome is an important player in the success of our current intercropping systems. It is involved in regulating plant stress responses and nutrient availability and uptake making cropping systems more robust and productive. There are still many unknowns regarding how the soil microbiome functions in intercropping systems, and how we can utilize it to improve agricultural productivity and sustainability. To further our understanding, we need to determine how the soil microbiome (1) is involved in the transfer and availability of N, P, and other essential nutrients; (2) improves the availability of essential plant nutrients in intercropping systems involving non-mycorrhizal plants; and (3) can be managed to further improve intercropping success. Further insights in these areas will allow us to make changes to the soil microbiome and improve its functionality. Looking forward, understanding plant-microbe interactions will help us continue to produce food for a growing population in a world where climate conditions are no longer certain and farmable land is decreasing.
Data availability
There is no data associated with this manuscript.
References
Abdelaal K, AlKahtani M, Attia K et al (2021) The role of plant growth-promoting bacteria in alleviating the adverse effects of drought on plants. Biology 10:520. https://doi.org/10.3390/biology10060520
Ayangbenro AS, Babalola OO (2021) Reclamation of arid and semi-arid soils: the role of plant growth-promoting archaea and bacteria. Curr Plant Biol 25:100173. https://doi.org/10.1016/j.cpb.2020.100173
Bakker PAHM, Pieterse CMJ, De Jonge R, Berendsen RL (2018) The soil-borne legacy. Cell 172:1178–1180. https://doi.org/10.1016/j.cell.2018.02.024
Bechtaoui N, El Alaoui A, Raklami A et al (2019) Impact of intercropping and co-inoculation with strains of plant growth-promoting rhizobacteria on phosphorus and nitrogen concentrations and yield of durum wheat (Triticum durum) and faba bean (Vicia faba). Crop Pasture Sci 70:649. https://doi.org/10.1071/CP19067
Bittencourt PP, Alves AF, Ferreira MB et al (2023) Mechanisms and applications of bacterial inoculants in plant drought stress tolerance. Microorganisms 11:502. https://doi.org/10.3390/microorganisms11020502
Chapagain T, Riseman A (2014) Barley–pea intercropping: effects on land productivity, carbon and nitrogen transformations. Field Crops Res 166:18–25. https://doi.org/10.1016/j.fcr.2014.06.014
Chen X, Chen J, Cao J (2023) Intercropping increases soil N-targeting enzyme activities: a meta-analysis. Rhizosphere 26:100686. https://doi.org/10.1016/j.rhisph.2023.100686
Chepsergon J, Moleleki LN (2023) Rhizosphere bacterial interactions and impact on plant health. Curr Opin Microbiol 73:102297. https://doi.org/10.1016/j.mib.2023.102297
Daghagh Yazd S, Wheeler SA, Zuo A (2020) Understanding the impacts of water scarcity and socio-economic demographics on farmer mental health in the Murray-Darling Basin. Ecol Econ 169:106564. https://doi.org/10.1016/j.ecolecon.2019.106564
Dai J, Qiu W, Wang N et al (2019) From Leguminosae/Gramineae intercropping systems to see benefits of intercropping on iron nutrition. Front Plant Sci 10:605. https://doi.org/10.3389/fpls.2019.00605
De Corato U (2020) Soil microbiota manipulation and its role in suppressing soil-borne plant pathogens in organic farming systems under the light of microbiome-assisted strategies. Chem Biol Technol Agric 7:17. https://doi.org/10.1186/s40538-020-00183-7
Dettweiler M, Wilson C, Maltais-Landry G, MacDonald G (2023) Cassava-legume intercropping is more beneficial in low-input systems: a meta-analysis. Field Crops Res 300:109005. https://doi.org/10.1016/j.fcr.2023.109005
Dowling A, O Sadras V, Roberts P et al (2021) Legume-oilseed intercropping in mechanised broadacre agriculture – a review. Field Crops Res 260:107980. https://doi.org/10.1016/j.fcr.2020.107980
Duchene O, Vian J-F, Celette F (2017) Intercropping with legume for agroecological cropping systems: complementarity and facilitation processes and the importance of soil microorganisms. A review. Agric Ecosyst Environ 240:148–161. https://doi.org/10.1016/j.agee.2017.02.019
Fletcher A, Peoples M, Kirkegaard J et al (2015) A review of annual intercrops in rainfed farming systems of Southern Australia. Building Productive, Diverse and Sustainable Landscapes, 17th Australian Agronomy Conference, 20-24 September 2015, Hobart, Australia. Conference Proceedings 486–489. Australian Society of Agronomy Inc
Fletcher AL, Kirkegaard JA, Peoples MB et al (2016) Prospects to utilise intercrops and crop variety mixtures in mechanised, rain-fed, temperate cropping systems. Crop Pasture Sci 67:1252. https://doi.org/10.1071/CP16211
Gorka S, Dietrich M, Mayerhofer W et al (2019) Rapid transfer of plant photosynthates to soil bacteria via ectomycorrhizal hyphae and its interaction with nitrogen availability. Front Microbiol 10:168. https://doi.org/10.3389/fmicb.2019.00168
Homulle Z, George TS, Karley AJ (2022) Root traits with team benefits: understanding belowground interactions in intercropping systems. Plant Soil 471:1–26. https://doi.org/10.1007/s11104-021-05165-8
Ingraffia R, Amato G, Frenda AS, Giambalvo D (2019) Impacts of arbuscular mycorrhizal fungi on nutrient uptake, N2 fixation, N transfer, and growth in a wheat/faba bean intercropping system. PLoS ONE 14:e0213672. https://doi.org/10.1371/journal.pone.0213672
Isaac ME, Hinsinger P, Harmand JM (2012) Nitrogen and phosphorus economy of a legume tree-cereal intercropping system under controlled conditions. Sci Total Environ 434:71–78. https://doi.org/10.1016/j.scitotenv.2011.12.071
Ju W, Jin X, Liu L et al (2020) Rhizobacteria inoculation benefits nutrient availability for phytostabilization in copper contaminated soil: drivers from bacterial community structures in rhizosphere. Appl Soil Ecol 150:103450. https://doi.org/10.1016/j.apsoil.2019.103450
Liao X, Zhao J, Yi Q et al (2023) Metagenomic insights into the effects of organic and inorganic agricultural managements on soil phosphorus cycling. Agric Ecosyst Environ 343:108281. https://doi.org/10.1016/j.agee.2022.108281
Madsen IJ, Parks JM, Friesen ML, Clark RE (2022) Increasing biodiversity and land-use efficiency through pea (Pisum aestivum)-canola (Brassica napus) intercropping (peaola). Front Soil Sci 2:818862. https://doi.org/10.3389/fsoil.2022.818862
Malviya MK, Solanki MK, Li C-N et al (2021) Sugarcane-legume intercropping can enrich the soil microbiome and plant growth. Front Sustain Food Syst 5:606595. https://doi.org/10.3389/fsufs.2021.606595
Morcillo R, Manzanera M (2021) The effects of plant-associated bacterial exopolysaccharides on plant abiotic stress tolerance. Metabolites 11:337. https://doi.org/10.3390/metabo11060337
Morugán-Coronado A, Pérez-Rodríguez P, Insolia E et al (2022) The impact of crop diversification, tillage and fertilization type on soil total microbial, fungal and bacterial abundance: a worldwide meta-analysis of agricultural sites. Agric Ecosyst Environ 329:107867. https://doi.org/10.1016/j.agee.2022.107867
Mwakilili AD, Mwaikono KS, Herrera SL et al (2021) Long-term maize-Desmodium intercropping shifts structure and composition of soil microbiome with stronger impact on fungal communities. Plant Soil 467:437–450. https://doi.org/10.1007/s11104-021-05082-w
Oliverio AM, Bissett A, McGuire K et al (2020) The role of phosphorus limitation in shaping soil bacterial communities and their metabolic capabilities. mBio 11:e01718–e01720. https://doi.org/10.1128/mBio.01718-20
Pourjani S, Aminpanah H, Vishkaei MNS (2022) Increasing the productivity of intercropping corn and peanuts by inoculation with Azotobacter chroococcum. RAR 39:327–336. https://doi.org/10.59665/rar3930
Qiao X, Bei S, Wang G et al (2022) Soil biota is decisive for overyielding in intercropping under low phosphorus conditions. J Appl Ecol 59:1804–1814. https://doi.org/10.1111/1365-2664.14187
Qin H, Lu K, Strong PJ et al (2015) Long-term fertilizer application effects on the soil, root arbuscular mycorrhizal fungi and community composition in rotation agriculture. Appl Soil Ecol 89:35–43. https://doi.org/10.1016/j.apsoil.2015.01.008
Qin M, Zhang Q, Pan J et al (2019) Effect of arbuscular mycorrhizal fungi on soil enzyme activity is coupled with increased plant biomass. Eur J Soil Sci ejss.12815. https://doi.org/10.1111/ejss.12815
Rezaei-Chiyaneh E, Battaglia ML, Sadeghpour A et al (2021) Optimizing intercropping systems of black cumin (Nigella sativa L.) and fenugreek (Trigonella foenum‐graecum L.) through inoculation with bacteria and mycorrhizal fungi. Adv Sustain Syst 5:2000269. https://doi.org/10.1002/adsu.202000269
Sharma GK, Khan SA, Shrivastava M et al (2022) Phycoremediated N-fertilization approaches on reducing environmental impacts of agricultural nitrate leaching. J Clean Prod 345:131120. https://doi.org/10.1016/j.jclepro.2022.131120
Shi X, Zhou Y, Guo P et al (2022) Peanut/sorghum intercropping drives specific variation in peanut rhizosphere soil properties and microbiomes under salt stress. Land Degrad Dev 34:736–750. https://doi.org/10.1002/ldr.4490
Solanki MK, Wang F-Y, Li C-N et al (2020) Impact of sugarcane–legume intercropping on diazotrophic microbiome. Sugar Tech 22:52–64. https://doi.org/10.1007/s12355-019-00755-4
Sun L, Dong X, Wang Y et al (2022a) Tea-soybean intercropping improves tea quality and nutrition uptake by inducing changes of rhizosphere bacterial communities. Microorganisms 10:2149. https://doi.org/10.3390/microorganisms10112149
Sun X, Zhang C, Bei S et al (2022b) High bacterial diversity and siderophore-producing bacteria collectively suppress Fusarium oxysporum in maize/faba bean intercropping. Front Microbiol 13:972587. https://doi.org/10.3389/fmicb.2022.972587
Tang X, Jiang J, Huang Z et al (2021) Sugarcane/peanut intercropping system improves the soil quality and increases the abundance of beneficial microbes. J Basic Microbiol 61:165–176. https://doi.org/10.1002/jobm.202000750
Tosi M, Mitter EK, Gaiero J, Dunfield K (2020) It takes three to tango: the importance of microbes, host plant, and soil management to elucidate manipulation strategies for the plant microbiome. Can J Microbiol 66:413–433. https://doi.org/10.1139/cjm-2020-0085
Tsialtas IT, Baxevanos D, Vlachostergios DN et al (2018) Cultivar complementarity for symbiotic nitrogen fixation and water use efficiency in pea-oat intercrops and its effect on forage yield and quality. Field Crops Res 226:28–37. https://doi.org/10.1016/j.fcr.2018.07.005
Verzeaux J, Hirel B, Dubois F et al (2017) Agricultural practices to improve nitrogen use efficiency through the use of arbuscular mycorrhizae: Basic and agronomic aspects. Plant Sci 264:48–56. https://doi.org/10.1016/j.plantsci.2017.08.004
Wang G, Sheng L, Zhao D et al (2016) Allocation of nitrogen and carbon is regulated by nodulation and mycorrhizal networks in soybean/maize intercropping system. Front Plant Sci 7. https://doi.org/10.3389/fpls.2016.01901
Wang G, Bei S, Li J et al (2020) Soil microbial legacy drives crop diversity advantage: linking ecological plant–soil feedback with agricultural intercropping. J Appl Ecol 58:496–506. https://doi.org/10.1111/1365-2664.13802
Wasternack C (2014) Action of jasmonates in plant stress responses and development — applied aspects. Biotechnol Adv 32:31–39. https://doi.org/10.1016/j.biotechadv.2013.09.009
Wu X, Rensing C, Han D et al (2022) Genome-resolved metagenomics reveals distinct phosphorus acquisition strategies between soil microbiomes. mSystems 7:e01107–e01121. https://doi.org/10.1128/msystems.01107-21
Yang J, Duan Y, Liu X et al (2022a) Reduction of banana Fusarium wilt associated with soil microbiome reconstruction through green manure intercropping. Agric Ecosyst Environ 337:108065. https://doi.org/10.1016/j.agee.2022.108065
Yang Z, Zhang Y, Wang Y et al (2022b) Intercropping regulation of soil phosphorus composition and microbially-driven dynamics facilitates maize phosphorus uptake and productivity improvement. Field Crops Res 287:108666. https://doi.org/10.1016/j.fcr.2022.108666
Yu R-P, Yang H, Xing Y et al (2022) Belowground processes and sustainability in agroecosystems with intercropping. Plant Soil 476:263–288. https://doi.org/10.1007/s11104-022-05487-1
Zhu S, Morel J-B (2019) Molecular mechanisms underlying microbial disease control in intercropping. MPMI 32:20–24. https://doi.org/10.1094/MPMI-03-18-0058-CR
Funding
The publication of this manuscript was supported by WSARE Graduate Student Grant GW21-228. This article is based upon work that is supported by the U.S. Department of Agriculture (USDA) NIFA project WNP03157 and Hatch project 1014527.
Author information
Authors and Affiliations
Contributions
All authors contributed to the production of this manuscript. The paper was written, and the figures were made by Janice Parks. Editing, feedback, and contributions to figure design were done by Maren Friesen. All authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors have no relevant financial or non-financial interests to disclose.
Additional information
Responsible Editor: Long Li.
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Parks, J.M., Friesen, M.L. The role of plant-microbe interactions in legume non-legume intercropping success. Plant Soil (2024). https://doi.org/10.1007/s11104-024-06550-9
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s11104-024-06550-9