Abstract
According to the recently updated World Health Organization (WHO) classification (2016), grade II–III astrocytomas are divided into IDH-wildtype and IDH-mutant groups, the latter being significantly less aggressive in terms of both progression-free and total survival. We identified a small cohort of WHO grade II–III astrocytomas that harbored the IDH1 R132H mutation, as confirmed by both immunohistochemistry and molecular sequence analysis, which nonetheless had unexpectedly rapid recurrence and subsequent progression to glioblastoma. Among these four cases, the mean time to recurrence as glioblastoma was only 16 months and the mean total survival among the three patients who have died during the follow-up was only 31 months. We hypothesized that these tumors had other, unfavorable genetic or epigenetic alterations that negated the favorable effect of the IDH mutation. We applied genome-wide profiling with a methylation array (Illumina Infinium Human Methylation 450k) to screen for genetic and epigenetic alterations in these tumors. As expected, the methylation profiles of all four tumors were found to match most closely with IDH-mutant astrocytomas. Compared with a control group of four indolent, age-similar WHO grade II–III astrocytomas, the tumors showed markedly increased levels of overall copy number changes, but no consistent specific genetic alterations were seen across all of the tumors. While most IDH-mutant WHO grade II–III astrocytomas are relatively indolent, a subset may rapidly recur and progress to glioblastoma. The precise underlying cause of the increased aggressiveness in these gliomas remains unknown, although it may be associated with increased genomic instability.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Diffuse gliomas are a relatively common primary brain tumor in adults with more than 20,000 cases occurring each year in the United States, approximately 70% of which are astrocytic neoplasms. The most malignant of these tumors, glioblastoma (GBM), represents grade IV out of IV in the World Health Organization (WHO) classification and carries an especially dismal prognosis [1]. The recent update of the WHO classification uses both histological features and mutations in isocitrate dehydrogenase one and two genes (IDH1/2) to classify grade II–IV gliomas [2]. IDH mutations occur in more than 70% of WHO II–III astrocytomas and secondary GBMs (those that evolved from lower-grade tumors). Patients with IDH-mutated tumors are on average 6 years younger than the patients with IDH-wildtype tumors at the time of diagnosis [3–5]. Studies in the past decade have demonstrated that IDH mutations in grade II–III gliomas are a strong independent favorable prognostic marker in terms of mortality and time to progression to secondary GBM. Patients with IDH-mutant grade II astrocytomas have a mean total survival of 151 months compared with a survival of 60 months among patients with IDH-wildtype grade II astrocytomas. The mean total survival among patients with IDH-mutant and IDH-wildtype grade III astrocytomas is 81 and 19 months, respectively [6].
We evaluated four patients with grade II–III astrocytomas that rapidly progressed to glioblastoma despite the presence of the favorable prognostic factor IDH1 R132H mutation [2] in all four cases. A genome-wide analysis of the tumors revealed a general increase of overall copy number alterations compared with more classically behaving, indolent IDH-mutated astrocytomas, but no consistent specific genetic or epigenetic defects were identified. Several of the cases had isolated genetic alterations affecting the p53 system, however without the identification of a common amplification, deletion, or mutation the underlying cause of the unusual aggressiveness remains unclear.
Methods
Case selection and clinical review
We identified a total of four cases that had an initial diagnosis of WHO grade II or III astrocytoma, an IDH mutation, and a subsequent diagnosis of glioblastoma within 3 years of the original diagnosis between 2006 and 2016. All available clinical history, presentation, imaging results, laboratory results, operative reports, subsequent follow-up encounters, and pathologic findings were reviewed (Table 1). The study was performed in accordance with a protocol approved by the Institutional Review Board of the University of Texas Southwestern Medical Center (IRB STU 022011-081).
Immunohistochemistry
Four micrometre thick sections of formalin-fixed, paraffin-embedded tissue underwent heat-induced epitope retrieval using CC1 (Ventana, Tucson, AZ), a tris-based buffer at pH 8–8.5, followed by immunohistochemical staining with a monoclonal mouse antibody to Ki-67/MIB-1 (Dako, Carpinteria, CA) diluted 1:80, polyclonal rabbit antibody to ATRX (Sigma-Aldrich, St. Louis, MO) diluted 1:200, monoclonal mouse antibody to p53 (Ventana) diluted 1:100, monoclonal mouse antibody to IDH1 R132H (Dianova, Hamburg, Germany) diluted 1:20, and polyclonal rabbit antibody to nestin (Sigma-Aldrich) diluted 1:20 on either the Ventana Benchmark XT or Ventana Benchmark Ultra automated stainer, using a Ventana UltraView Universal DAB Detection Kit.
IDH1 and IDH2 sequence analysis
Tumor DNA extracted from deparaffinized tissue sections (QIAamp DNA FFPE Tissue Kit, Qiagen) was tested using Sequenom iPLEX genotyping protocols. IDH1 exon 4 was PCR amplified and subsequently queried at codon 132 by single-base primer extension with products analyzed by MALDI-TOF mass spectrometry (Sequenom MassArray Analyzer 4). Assay primers were designed with MassARRAY Assay Design Suite software. IDH1 genotyping determination on the tumor specimens were made by manual inspection.
DNA extraction
DNA was extracted from formalin-fixed paraffin-embedded tissue (FFPE). Areas with the highest available tumor content were chosen. Extraction was carried out using the automated Maxwell system (Promega, Madison, WI).
Genome-wide methylation profiling
The Illumina Infinium Human Methylation 450 Bead-Chip (450k) array was used to determine the DNA methylation status of 482,421 CpG sites (Illumina, San Diego, CA) according to the manufacturer’s instructions. Methylation profiles were compared to a reference cohort of 2150 cases from 77 tumor entities previously profiled and analyzed at German Cancer Research Center using a random forest algorithm and customized bioinformatics packages. In addition, the array data was used to calculate a low-resolution copy number profile (CNP) as previously described by others [7, 8] and us [9, 10]. Analysis was performed on specimens at the time of initial diagnosis.
Droplet digital PCR for MDM2 gene
Droplet digital PCR was performed on a Bio-Rad QX200 (Bio-Rad, Hercules, CA). Primers were designed against regions of amplification for MDM2 and EGFR, which were not amplified by the array analysis. The RRP30 gene (diploid) was used as control. Primers will be provided upon request. 20 ng of HindIII digested genomic DNA were used per reaction, using the following protocol: 1 cycle at 95 °C for 5 min, 40 cycles at 95 °C for 15 s and 60 °C for 1 min, 1 cycle at 4 °C for 5 min, and 1 cycle at 90 °C for 5 min all at a ramp rate of 2 °C/s on a Bio-Rad T100 thermal cycler was used for the PCR step. Droplets were quantified using the Bio-Rad Quantisoft software. A total of two replicates were used per sample.
Results
Unexpectedly aggressive clinical course in IDH-mutated grade II–III astrocytomas
We identified four grade II–III gliomas that progressed rapidly to glioblastoma in spite of harboring an IDH mutation. Two of the four patients included in this study were initially diagnosed with WHO grade II astrocytoma (Table 1; patients 1 and 4) and two patients were initially diagnosed with WHO grade III anaplastic astrocytoma (Table 1; patients 2 and 3). The radiologic features were consistent with the histologic diagnosis: the patients with WHO grade II tumors at initial diagnosis had non-enhancing masses (Fig. 1), whereas the WHO grade III tumors had patchy heterogeneous enhancement. The MIB-1 proliferation indices were also consistent with the histologic grade (Fig. 2). All four of these tumors were negative for the 1p/19q codeletion but had IDH1 R132H mutations by immunohistochemistry (Table 1; Fig. 2), yet all progressed rapidly to glioblastoma (Figs. 1, 2). The mean time of progression to secondary glioblastoma was 16 months (n = 4) with a mean total survival time of 31 months among the three patients who died during the follow-up (Table 1). Nestin expression, an independent adverse prognostic factor for survival in patients with grade II–III glioma [11], was high in all four cases by immunohistochemistry and computer-assisted image analysis.
MRI images from patient 1 in this study demonstrating the rapid progression of a right temporal lobe low-grade glioma (WHO II) to glioblastoma (WHO IV) in a relatively short period of time. T1 post-contrast images (top two rows) show a non-enhancing mass at the time of initial resection with a subsequent nodularly enhancing mass at the time of recurrence (only 383 days later) adjacent to the resection cavity. T2 FLAIR images (row three) demonstrate increased FLAIR signal around the mass at both time points
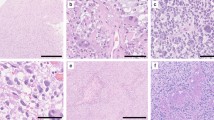
Histologic changes associated with progression from an initial diagnosis of diffuse glioma (WHO II) to a recurrence as glioblastoma (WHO IV) in patient 4. The H&E images in the top rows show progression to a significantly more cellular neoplasm with microvascular proliferation (inset). The initial tumor and recurrence both have the IDH1 R132H mutation by immunohistochemistry. The initial tumor is negative for ATRX and p53 mutations. Immunolabeling indices of both p53 and MIB-1 are elevated in the recurrent tumor. (Original magnification, ×100; scale bar 50 μm.)
Confirmation of IDH mutations by IDH1 and IDH2 sequence analysis
The presence of the IDH1 R132H mutation, originally detected by immunohistochemistry (Fig. 2), was successfully confirmed in all four tumors by sequence analysis based on Sequenom mass spectrometry.
Methylation profiling
Genome-wide methylation profiles obtained using the Illumina Infinium Human Methylation 450k array were automatically compared to a reference cohort of 2150 cases from 77 tumor entities using a DNA methylation-based classification of human brain tumors (http://www.molecularneuropathology.org). The methylation profiles of all four tumors were found to match most closely with IDH-mutated astrocytomas.
IDH-mutant astrocytomas with rapid progression show higher number of copy number changes
Low-grade IDH-mutated astrocytomas at the time of presentation typically show relatively low numbers of copy number changes. These usually include isolated gains or losses of whole chromosomes or large portions of chromosomal arms, while focal amplifications on oncogenes are infrequent (Supplemental Fig. 1) [8]. Overall, 450k methylation arrays suggested a larger degree of genomic instability in rapidly progressive IDH-mutant cases when compared with slow-growing IDH-mutant tumors. Copy number profiles of the initial resection specimens in these four patients revealed a higher level of overall copy number variation across the genome than would be expected in grade II–III IDH-mutant gliomas at the time of initial presentation (Fig. 3). This included large chromosomal gains and losses as well as low-level focal copy number gains. These genomic findings are more consistent with GBMs as GBMs typically show more frequent gains and losses of whole chromosomes or chromosomal arms including chromosome 7, 10, 13, or 19 and focal high level amplifications of tyrosine kinase genes including EGFR, MET and PDGFRA/KIT/VEGFR2 [12]. Patient 1 had low level MYC amplification and Patient 3 had a CDKN2A/B homozygous deletion (Fig. 3). The array data suggested a low level mouse double minute 2 homolog (MDM2) copy number gain in all four tumors; however, the presence of three copies of the MDM2 gene was confirmed by an alternate method in only one tumor (patient 2; Table 1), where sufficient DNA was available by Droplet Digital PCR (ddPCR). Immunohistochemistry for p53 showed a low labeling index (<10%) in all cases, which suggests that the p53 gene was wildtype in all cases at initial diagnosis [13], raising the possibility that an MDM2 copy number gain, if present, could act as an alternative mechanism of TP53 pathway aberration.
Other genetic abnormalities
The MGMT gene promoter was methylated in two of the tumors, unmethylated in one tumor, and equivocal in one tumor (Table 1). Immunohistochemistry for ATRX demonstrated loss of nuclear positivity in tumor cells (indicating ATRX-mutant status) in patients 1–3 (Table 1). Both the initial resection and recurrence specimen in patient 4 showed retained nuclear ATRX immunoreactivity (Fig. 2). In addition, case 4 developed a high nuclear p53 labeling index suggestive of a p53 mutation between the initial diagnosis of grade II diffuse glioma and glioblastoma by IHC (Fig. 2).
Discussion
In this study, we performed a genome-wide analysis of gene copy number alterations and methylation profiles in a series of four IDH-mutated, clinically aggressive gliomas using the Illumina Infinium Human Methylation array (Fig. 3) [14, 15]. The 450k methylation array platform is a powerful way to classify brain tumors into clinically relevant diagnostic subgroups. This approach is currently being integrated into standard practice for diagnostic and prognostic purposes [14, 16–20].
We show that IDH-mutant tumors with rapid progression have a higher number of molecular abnormalities than would be expected in grade II–III, IDH-mutated gliomas (Fig. 3), however the overall methylation pattern matched that expected of IDH-mutant astrocytomas. Wakimoto et al. have demonstrated that tertiary oncogenic genetic alterations, including PIK3CA mutation and amplification of the PDGFRA, MET, and N-MYC genes are commonly observed in IDH-mutant tumors at the time of malignant progression [21]. One out of four of these cases also showed an amplification of the MYC gene, an oncogene overexpressed in a number of cancers, perhaps most famously Burkitt’s lymphoma [22–24]. Located on chromosome 8, MYC acts downstream to help advance the cell cycle by interacting with cyclins and p21, promotes pluripotency, and helps maintain global euchromatin patterns, all mechanisms that have been previously implicated in the neoplastic process when its expression is amplified [23, 25, 26]. Previous studies have demonstrated that the MYC locus was amplified in 22% of gliomas progressing to GBM [27]. Identifying MYC amplification in IDH-mutant gliomas is clinically important as it is associated with a more aggressive behavior and malignant transformation. Furthermore, MYC activation has been recently shown as a potential therapeutic avenue in IDH-mutant gliomas by metabolic targeting [28].
In response to stress signals and DNA damage, p53 acts to arrest the cell cycle preventing progression from G1- to S-phase, inducing apoptosis, or activating DNA repair mechanisms [29–31]. Mutations in TP53 occur in 25–30% of primary (de novo) GBMs and 60–70% of secondary GBMs. Nearly 90% of GBMs have an abnormality at some level of the p53 pathway, making it the most common genetic alteration seen in these tumors overall [32]. While these tumors were negative for nuclear accumulation of p53 at the time of diagnosis, indicating a wildtype p53 status (Fig. 2), the copy number analysis showed genomic abnormalities that affect p53 signaling in these tumors, including alterations involving the MDM2 gene (Fig. 3). MDM2 is part of the core regulation of the p53 pathway [29], and its amplification is seen in a minority of lower-grade gliomas and secondary glioblastomas [33, 34], however when present this amplification is an effective silencer of wild-type p53. Transcription of the MDM2 gene is upregulated by p53 protein and the MDM2 protein in turn inactivates p53, effectively tempering the effect of p53 [35]. The MDM2 gene product acts as an E3 ubiquitin ligase, marking p53 for proteasomal degradation and preventing it from halting the cell cycle prior to S-phase [36–40]. Overexpression of the MDM2 oncogene, which occurs in 7–52% of primary GBMs and 0–11% of secondary GBMs, silences wildtype p53 in gliomas [33, 34, 41]. These MDM2-amplified GBM patients have a worse progression-free and total survival and decreased response to therapy [41]. In the subset of tumors with MDM2 amplification, MDM2 could potentially be targeted with inhibitor compounds, allowing the otherwise normal p53 to function [42–46].
Additionally, one out of the four tumors in this study had a homozygous deletion of CDKN2A/B on chromosome 9 (Fig. 3), seen in approximately 44% of low-grade tumors prior to progression to GBM [27] and up to 76% of secondary GBMs [47]. The CDKN2A gene produces 2 proteins by alternative splicing, including the tumor suppressor p14ARF, which binds MDM2, preventing it from inhibiting p53 function. Silencing CDKN2A either by methylation or by homozygous deletion results in the loss of this tumor suppressor function and a further push toward cell cycle dysregulation [47, 48].
All four tumors had high levels of the stem cell marker nestin as assessed by computer assisted image analysis. Nestin expression by both immunohistochemistry and mRNA levels has been demonstrated to correlate directly with tumor grade and inversely with overall survival in patients with WHO grade II–III astrocytomas [11].
Three of the four patients have died during the follow-up, at 29–34 months after the initial diagnosis of a grade II–III, IDH-mutated tumor, which is much shorter total survival than the typical range of 81–151 months reported for grade II–III, IDH-mutated tumors in the literature [6]. Their rapid progression to GBM (mean time to progression = 16 months) is also very unusual considering their IDH-mutated status [2]. Taken together, these cases show that although these tumors had the IDH R132H mutation, this does not always guarantee a better prognostic outcome; some tumors with this mutation may in fact have extremely poor outcomes with rapid progression to GBM and very short mortality intervals. Although this cohort is small, our data suggest that an overall increase in both large and focal copy number alterations such as MDM2 and MYC gains and CDKN2A/B losses may be associated with a more aggressive behavior and shorter survival.
Conclusions
In the majority of cases, grade II–III gliomas harboring an IDH1 or IDH2 mutation have a much more favorable clinical course than grade II–III gliomas lacking an IDH mutation. Here we have identified a subset of tumors within this IDH-mutant glioma group with a significantly worse clinical outcome than would be expected based the 2016 WHO classification system [2]. Genomic instability as evidenced by an increased number of large and focal copy number aberrations at the time of initial diagnosis seem to be associated with a poor outcome in IDH-mutant gliomas.
References
Dolecek TA, Propp JM, Stroup NE, Kruchko C (2012) CBTRUS statistical report: primary brain and central nervous system tumors diagnosed in the United States in 2005–2009. Neuro Oncol 14(Suppl 5):v1–v49
Louis DN, Perry A, Reifenberger G, von Deimling A, Figarella-Branger D et al (2016) The 2016 World Health Organization classification of tumors of the central nervous system: a summary. Acta Neuropathol 131:803–820
Yan H, Parsons DW, Jin G, McLendon R, Rasheed BA et al (2009) IDH1 and IDH2 mutations in gliomas. N Engl J Med 360:765–773
Hartmann C, Meyer J, Balss J, Capper D, Mueller W et al (2009) Type and frequency of IDH1 and IDH2 mutations are related to astrocytic and oligodendroglial differentiation and age: a study of 1,010 diffuse gliomas. Acta Neuropathol 118:469–474
Watanabe T, Nobusawa S, Kleihues P, Ohgaki H (2009) IDH1 mutations are early events in the development of astrocytomas and oligodendrogliomas. Am J Pathol 174:1149–1153
Sanson M, Marie Y, Paris S, Idbaih A, Laffaire J et al (2009) Isocitrate dehydrogenase 1 codon 132 mutation is an important prognostic biomarker in gliomas. J Clin Oncol 27:4150–4154
Sturm D, Witt H, Hovestadt V, Khuong-Quang DA, Jones DT et al (2012) Hotspot mutations in H3F3A and IDH1 define distinct epigenetic and biological subgroups of glioblastoma. Cancer Cell 22:425–437
Wiestler B, Capper D, Sill M, Jones DT, Hovestadt V et al (2014) Integrated DNA methylation and copy-number profiling identify three clinically and biologically relevant groups of anaplastic glioma. Acta Neuropathol 128:561–571
Orillac C, Thomas C, Dastagirzada Y, Hidalgo ET, Golfinos JG et al (2016) Pilocytic astrocytoma and glioneuronal tumor with histone H3 K27M mutation. Acta Neuropathol Commun 4:84
Huse JT, Snuderl M, Jones DT, Brathwaite CD, Altman N et al (2017) Polymorphous low-grade neuroepithelial tumor of the young (PLNTY): an epileptogenic neoplasm with oligodendroglioma-like components, aberrant CD34 expression, and genetic alterations involving the MAP kinase pathway. Acta Neuropathol 133:417–429
Hatanpaa KJ, Hu T, Vemireddy V, Foong C, Raisanen JM et al (2014) High expression of the stem cell marker nestin is an adverse prognostic factor in WHO grade II–III astrocytomas and oligoastrocytomas. J Neurooncol 117:183–189
Snuderl M, Fazlollahi L, Le LP, Nitta M, Zhelyazkova BH et al (2011) Mosaic amplification of multiple receptor tyrosine kinase genes in glioblastoma. Cancer Cell 20:810–817
Takami H, Yoshida A, Fukushima S, Arita H, Matsushita Y et al (2015) Revisiting TP53 mutations and Immunohistochemistry—a comparative study in 157 diffuse gliomas. Brain Pathol 25:256–265
Bibikova M, Le J, Barnes B, Saedinia-Melnyk S, Zhou L et al (2009) Genome-wide DNA methylation profiling using Infinium® assay. Epigenomics 1:177–200
Baylin SB, Ohm JE (2006) Epigenetic gene silencing in cancer - a mechanism for early oncogenic pathway addiction? Nat Rev Cancer 6:107–116
Bibikova M, Barnes B, Tsan C, Ho V, Klotzle B et al (2011) High density DNA methylation array with single CpG site resolution. Genomics 98:288–295
Feinberg AP (2010) Epigenomics reveals a functional genome anatomy and a new approach to common disease. Nat Biotechnol 28:1049–1052
Feinberg AP (2010) Genome-scale approaches to the epigenetics of common human disease. Virchows Arch 456:13–21
Petronis A (2010) Epigenetics as a unifying principle in the aetiology of complex traits and diseases. Nature 465:721–727
Rakyan VK, Down TA, Balding DJ, Beck S (2011) Epigenome-wide association studies for common human diseases. Nat Rev Genet 12:529–541
Wakimoto H, Tanaka S, Curry WT, Loebel F, Zhao D et al (2014) Targetable signaling pathway mutations are associated with malignant phenotype in IDH-mutant gliomas. Clin Cancer Res 20:2898–2909
Finver SN, Nishikura K, Finger LR, Haluska FG, Finan J et al (1988) Sequence analysis of the MYC oncogene involved in the t(8;14)(q24;q11) chromosome translocation in a human leukemia T-cell line indicates that putative regulatory regions are not altered. Proc Natl Acad Sci USA 85:3052–3056
Nilsson JA, Cleveland JL (2003) Myc pathways provoking cell suicide and cancer. Oncogene 22:9007–9021
Sheiness D, Bister K, Moscovici C, Fanshier L, Gonda T et al (1980) Avian retroviruses that cause carcinoma and leukemia: identification of nucleotide sequences associated with pathogenicity. J Virol 33:962–968
Cotterman R, Knoepfler PS (2009) N-Myc regulates expression of pluripotency genes in neuroblastoma including lif, klf2, klf4, and lin28b. PLoS ONE 4:e5799
Cotterman R, Jin VX, Krig SR, Lemen JM, Wey A et al (2008) N-Myc regulates a widespread euchromatic program in the human genome partially independent of its role as a classical transcription factor. Cancer Res 68:9654–9662
Bai H, Harmanci AS, Erson-Omay EZ, Li J, Coskun S et al (2016) Integrated genomic characterization of IDH1-mutant glioma malignant progression. Nat Genet 48:59–66
Tateishi K, Iafrate AJ, Ho Q, Curry WT, Batchelor TT et al (2016) Myc-driven glycolysis is a therapeutic target in glioblastoma. Clin Cancer Res 22:4452–4465
Harris SL, Levine AJ (2005) The p53 pathway: positive and negative feedback loops. Oncogene 24:2899–2908
Haupt Y, Barak Y, Oren M (1996) Cell type-specific inhibition of p53-mediated apoptosis by mdm2. EMBO J 15:1596–1606
Haupt Y, Oren M (1996) p53-mediated apoptosis: mechanisms and regulation. Behring Inst Mitt 97:32–59
England B, Huang T, Karsy M (2013) Current understanding of the role and targeting of tumor suppressor p53 in glioblastoma multiforme. Tumour Biol 34:2063–2074
Biernat W, Kleihues P, Yonekawa Y, Ohgaki H (1997) Amplification and overexpression of MDM2 in primary (de novo) glioblastomas. J Neuropathol Exp Neurol 56:180–185
Ohgaki H, Kleihues P (2009) Genetic alterations and signaling pathways in the evolution of gliomas. Cancer Sci 100:2235–2241
Lukashchuk N, Vousden KH (2007) Ubiquitination and degradation of mutant p53. Mol Cell Biol 27:8284–8295
Leach FS, Tokino T, Meltzer P, Burrell M, Oliner JD et al (1993) p53 mutation and MDM2 amplification in human soft tissue sarcomas. Cancer Res 53:2231–2234
Momand J, Zambetti GP, Olson DC, George D, Levine AJ (1992) The mdm-2 oncogene product forms a complex with the p53 protein and inhibits p53-mediated transactivation. Cell 69:1237–1245
Oliner JD (1993) Discerning the function of p53 by examining its molecular interactions. Bioessays 15:703–707
Oliner JD, Pietenpol JA, Thiagalingam S, Gyuris J, Kinzler KW et al (1993) Oncoprotein MDM2 conceals the activation domain of tumour suppressor p53. Nature 362:857–860
Olson DC, Marechal V, Momand J, Chen J, Romocki C et al (1993) Identification and characterization of multiple mdm-2 proteins and mdm-2-p53 protein complexes. Oncogene 8:2353–2360
Rayburn E, Zhang R, He J, Wang H (2005) MDM2 and human malignancies: expression, clinical pathology, prognostic markers, and implications for chemotherapy. Curr Cancer Drug Targets 5:27–41
Costa B, Bendinelli S, Gabelloni P, Da Pozzo E, Daniele S et al (2013) Human glioblastoma multiforme: p53 reactivation by a novel MDM2 inhibitor. PLoS ONE 8:e72281
Shangary S, Wang S (2009) Small-molecule inhibitors of the MDM2-p53 protein-protein interaction to reactivate p53 function: a novel approach for cancer therapy. Annu Rev Pharmacol Toxicol 49:223–241
Vassilev LT, Vu BT, Graves B, Carvajal D, Podlaski F et al (2004) In vivo activation of the p53 pathway by small-molecule antagonists of MDM2. Science 303:844–848
Vu BT, Vassilev L (2011) Small-molecule inhibitors of the p53-MDM2 interaction. Curr Top Microbiol Immunol 348:151–172
Yu S, Qin D, Shangary S, Chen J, Wang G et al (2009) Potent and orally active small-molecule inhibitors of the MDM2-p53 interaction. J Med Chem 52:7970–7973
Nakamura M, Watanabe T, Klangby U, Asker C, Wiman K et al (2001) p14ARF deletion and methylation in genetic pathways to glioblastomas. Brain Pathol 11:159–168
Biernat W, Tohma Y, Yonekawa Y, Kleihues P, Ohgaki H (1997) Alterations of cell cycle regulatory genes in primary (de novo) and secondary glioblastomas. Acta Neuropathol 94:303–309
Acknowledgements
The authors would like to thank Niccole Williams and Agatha Villegas for administrative professional services. We also thank Ping Shang for some of the immunohistochemical stains. The authors would like to thank Stefan Pfister, David T. W. Jones, Martin Sill and Volker Hovestadt for their help with optimization of the 450 k Illumina Infinium methylation profiling for copy number analysis. A.A.H. was supported, in part, by a grant from the Department of Veteran’s Affairs (I01BX002559). The study was supported by the Friedberg Charitable Foundation grant to M.S. and M.A.K.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare no conflict of interest.
Additional information
Timothy E. Richardson and Matija Snuderl have contributed equally to this work.
Electronic supplementary material
Below is the link to the electronic supplementary material.

11060_2017_2431_MOESM1_ESM.jpg
Supplemental Figure 1. Copy number analysis derived from the Illumina Infinium Human Methylation 450k array data, showing four conventional, indolent IDH-mutated astrocytomas with no subsequent progression. These four tumors show relatively little copy number changes across all chromosomes compared with the four tumors in our study group. (JPG 827 KB)
Rights and permissions
About this article
Cite this article
Richardson, T.E., Snuderl, M., Serrano, J. et al. Rapid progression to glioblastoma in a subset of IDH-mutated astrocytomas: a genome-wide analysis. J Neurooncol 133, 183–192 (2017). https://doi.org/10.1007/s11060-017-2431-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11060-017-2431-y