Abstract
High-grade astrocytoma with piloid features (HGAP) is a recently recognized glioma type whose classification is dependent on its global epigenetic signature. HGAP is characterized by alterations in the mitogen-activated protein kinase (MAPK) pathway, often co-occurring with CDKN2A/B homozygous deletion and/or ATRX mutation. Experience with HGAP is limited and to better understand this tumor type, we evaluated an expanded cohort of patients (n = 144) with these tumors, as defined by DNA methylation array testing, with a subset additionally evaluated by next-generation sequencing (NGS). Among evaluable cases, we confirmed the high prevalence CDKN2A/B homozygous deletion, and/or ATRX mutations/loss in this tumor type, along with a subset showing NF1 alterations. Five of 93 (5.4%) cases sequenced harbored TP53 mutations and RNA fusion analysis identified a single tumor containing an NTRK2 gene fusion, neither of which have been previously reported in HGAP. Clustering analysis revealed the presence of three distinct HGAP subtypes (or groups = g) based on whole-genome DNA methylation patterns, which we provisionally designated as gNF1 (n = 18), g1 (n = 72), and g2 (n = 54) (median ages 43.5 years, 47 years, and 32 years, respectively). Subtype gNF1 is notable for enrichment with patients with Neurofibromatosis Type 1 (33.3%, p = 0.0008), confinement to the posterior fossa, hypermethylation in the NF1 enhancer region, a trend towards decreased progression-free survival (p = 0.0579), RNA processing pathway dysregulation, and elevated non-neoplastic glia and neuron cell content (p < 0.0001 and p < 0.0001, respectively). Overall, our expanded cohort broadens the genetic, epigenetic, and clinical phenotype of HGAP and provides evidence for distinct epigenetic subtypes in this tumor type.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
High-grade astrocytoma with piloid features (HGAP) is a recently discovered and codified brain tumor type whose classification is dependent on its global DNA methylation signature [22]. These tumors can occur throughout the neuraxis, but are often located within the posterior fossa [21, 22]. The histopathology of HGAPs variably demonstrates lower-grade or higher-grade features, with characteristics overlapping those of pilocytic astrocytoma and glioblastoma [3, 22]. HGAPs are IDH-wildtype and frequently contain alterations in the mitogen-activated protein kinase (MAPK) pathway, in combination with homozygous deletion of CDKN2A/B and/or ATRX mutations [22]. While these genetic alterations may be characteristic, they are not specific to HGAP. What is unique to HGAP is its epigenetic profile; indeed, DNA methylation profiling is required for the definitive diagnosis of HGAP [4, 12]. Definitive diagnosis of HGAP is important for prognostic considerations, where its overall survival is between that of pilocytic astrocytoma (CNS WHO grade 1) and glioblastoma, IDH-wildtype (CNS WHO grade 4) [22]. The discovery of HGAP exemplifies the utility of identifying novel tumor entities through DNA methylation profiling [18, 22, 27]. DNA methylation profiling and clustering analysis are additionally useful for intra-class subtyping across numerous CNS tumor types [4, 16, 19, 28, 30]. Although HGAP was recently incorporated into the 2021 fifth edition of the World Health Organization Classification for Central Nervous System Tumors [12], there is incomplete understanding of this tumor type as there is a general lack of large published HGAP cohort studies [3, 22]. In this study, we sought to validate previously described aspects of HGAP as well as identify potential epigenetic subtypes that may have associated clinical implications.
Materials and methods
Tissue samples and datasets
The use of human subject material was performed in accordance with the World Medical Association Declaration of Helsinki and with the approval of the participating Institutional Review Boards. This study included (1) previously described cases of anaplastic astrocytoma with piloid features [3, 22], (2) cases from the Laboratory of Pathology clinical consult service at the National Institutes of Health (NIH) in Bethesda, MD, USA and (3) other published cases with available methylation array data in the form of raw IDAT files. Referral consult cases were received from the following institutions: Mayo Clinic (Rochester, Minnesota); University of Michigan Medical Center (Ann Arbor, Michigan); University of California San Francisco Medical Center (San Francisco, California); Barnes Jewish Hospital (St. Louis, Missouri); Children’s Hospital of Philadelphia (Philadelphia, Pennsylvania); Hospital of the University of Pennsylvania (Philadelphia, Pennsylvania); Duke University Medical Center (Durham, North Carolina); Laboratorio Bacchi (Sao Paulo, Brazil); Stanford University Medical Center (Palo Alto, California); University of Chicago Medical Center (Chicago, Illinois); Beaumont Hospital (Royal Oak, Michigan); Boston Children’s Hospital (Boston, Massachusetts); Brigham and Women’s Hospital (Boston, Massachusetts); Cleveland Clinic (Cleveland, Ohio); Emory University Hospital (Atlanta, Georgia); Inova Fairfax Hospital (Falls Church, Virginia); Intermountain Central Laboratory (Murray, Utah); Johns Hopkins Hospital (Baltimore, Maryland); Massachusetts General Hospital (Boston, Massachusetts); Mercy General Hospital (Sacramento, California); University of Nebraska Medical Center (Omaha, Nebraska); Northwestern Memorial Hospital (Chicago, IL); Sheba Medical Center (Tel Hashomer, Israel); Tucson Medical Center (Tucson, Arizona); University of California Los Angeles Medical Center (Los Angeles, California); University of Alabama (Birmingham, Alabama); University of Colorado Hospital (Aurora, Colorado); University of Iowa Hospitals (Iowa City, Iowa); ARUP Laboratories (Salt Lake City, Utah); and the University of Virginia Health System (Charlottesville, Virginia). Each of the 61 referral consult cases was reviewed for histopathology as part of the diagnostic process by at least two board-certified neuropathologists (PJC, MQ, DP, KA). When available, we noted clinical features including subject demographics, tumor location, surgical procedure (biopsy versus excision—subtotal versus gross total was not available), and tumor occurrence (primary versus recurrence). Accompanying treatment modalities (chemotherapy and radiation) were not available. We defined progression-free survival as the duration from surgery of initial tumor to the date of clinical-radiographic progression or date of second resection for clinically/radiographically-determined tumor progression. Similarly, overall survival was defined as the duration from surgery of the initial tumor to death. These definitions were applied uniformly throughout our cohort. Cases were also extracted from publicly available methylation datasets included E-MTAB-5552 [14], GSE109381 [4], GSE143843 [11], GSE157397 [7], GSE175543 [6], Children’s Brain Tumor Network (https://cbtn.org), and The Cancer Genome Atlas (https://gdc.cancer.gov).
DNA methylation array profiling
For institutional samples, tumor cellularity was confirmed to comprise at least 60% of the tissue. Genomic DNA was extracted from Formalin-Fixed Paraffin-Embedded (FFPE) tissue and underwent bisulfite-conversion (EZ DNA Methylation Kit, Zymo Research D5001) and restoration (Infinium FFPE DNA Restore kit, Illumina, USA). The DNA was then assayed on the Infinium MethylationEPIC kit (Illumina, USA) according to the Infinium HD FFPE Methylation Assay automated protocol (Illumina, USA). Raw idat files were used for classification using version 11b6 of the Heidelberg methylation-based classifier [4]. Cases which received a primary methylation class designation of anaplastic pilocytic astrocytoma, (the original class designation for the HGAP WHO tumor type) and had other features consistent with this glial tumor (such as astrocytic histomorphology, glial fibrillary acidic protein immunopositivity, and/or Olig2 immunopositivity), were included for analysis in this study. Glial features were confirmed centrally and independently by at least two board-certified neuropathologists (PJC, MQ, DP, KA).
Bioinformatic analyses
Microarray idat files were processed with functional normalization using the meffil package [15]. This R package is available from Bioconductor (https://www.bioconductor.org/) and GitHub (https://github.com/perishky/meffil). Functional normalization was performed with the meffil package, which is based on linear regression modelling [15]. The quantile normalization runs an inbuilt algorithm based on illumina control probes. This model removes control probe variance from the sample quantiles and simultaneously adjusting probe intensities for additional fixed and random effects. Those fixed and random effects variables supplied to normalization function and coded from the metadata we collected for all samples. We choose type of array variables (HumanMethylation450 vs MethylationEPIC BeadChip) and tissue preservation method (Frozen vs FFPE) as fixed effect and also included random effect attributed to individual chip barcode. Random effects are handled in meffil by using the lme4 R package [2]. The meffil package provided automated estimation of functional normalization parameters to compensate for chip to chip (barcode) variance.
For unsupervised hierarchical clustering, the most significant probes were clustered and plotted with ComplexHeatmap package available through Bioconductor [9]. This package allows hierarchical cluster with different methods for columns and rows. We used default “euclidean” metrics for distance matrix and “average” for the dendrogram construction. Normalized beta values were filtered to the most differentially methylated probes calculated by student t-test contrasting each group pairs: Group1 vs Group2, Group1 vs gNF1, Group2 vs gNF1. Bonferroni correction was applied to p-values and 772 probes with p-value below 0.05 were selected for heat map and Uniform Manifold Approximation and Projection (UMAP) plots. UMAP plot was produced with r package uwot: The Uniform Manifold Approximation and Projection (UMAP) Method for Dimensionality Reduction (https://arxiv.org/abs/1802.03426). We used two methods to estimate tumor ratio and cell composition in each sample: meffil package described above and the CIBERSORT package (https://cibersortx.stanford.edu/). Both methods showed good correlation with each other. The meffil package supplied prebuild reference “guintivano dlpfc” for the brain tissue deconvolution with five cell types. Those reference profiles were based on published Bioconductor package FlowSorted.DLPFC.450 k. (https://github.com/perishky/meffil/wiki/Estimating-cellular-composition). Signature matrices were used to determine neoplastic cell and various tumor microenvironment cell fractions through methylCIBERSORT as previously described [25]. The r package methylGSA was used for gene ontology pathway analysis [23].
Targeted next-generation DNA sequencing
For cases with available DNA, next generation sequencing (NGS) data were obtained as previously described using either a custom amplicon-based brain tumor specific panel [20] or a commercial panel TruSight Oncology 500 (TSO 500; Illumina, San Diego, USA) [18]. The TSO 500 panel genes are available on the manufacturer’s website (https://www.illumina.com/products/by-type/clinical-research-products/trusight-oncology-500.html). Illumina platform libraries were prepared using TSO 500 kit according to the manufacturer’s protocol. Amplified pre-enriched libraries were hybridized to probes specific to the TSO 500 targeted gene panel. Enriched libraries were amplified, quantified, normalized, and sequenced as paired-end reads on a high-output NextSeq 500/550 flow cell. The TSO 500 local app was used for alignment and variant calling. Variants were manually reviewed by visual inspection of the raw sequencing read alignments using the Integrative Genomics Viewer [24]. For a subset of cases, sequencing was obtained from publicly available sources or performed at the outside referring institutions and those results were reported as part of the consultation process (sources of sequencing summarized in Supplemental Table 1). For previously described HGAP cases [22], sequencing data were obtained from the authors.
Statistics
Statistical analyses were performed using R software (Version 3.3.2, RProject for Statistical Computing, http://www.r-project.org/). Kaplan–Meier analysis for survival, correlation matrix analysis, and principal component analysis were performed using GraphPad Prism software (Version 7.02, https://www.graphpad.com/scientific-software/prism). For direct comparisons between groups, Mann–Whitney U test was used unless otherwise indicated. Chi-squared test was performed to compare frequencies across groups.
Results
Expanded HGAP demographics, histologic features, and genetic characteristics
Our total cohort was composed of 148 tumor samples from 144 patients (Supplemental Table 2). The mean and median age of patients at the time of surgery were 44 years and 43 years (range 4–88 years), respectively. Of the 135 patients with known age at time of diagnosis, 12 (8.9%) were in the pediatric age group (< 18 years) (Supplemental Table 2). Tumors occurred in 59 female and 85 male patients (ratio 1:1.44). Disease stage included 74 primary tumors, 38 recurrent tumors, and 36 tumors with unknown designation. Tumor location was known for 130 patients, with 81 located in the posterior fossa, 34 located in the supratentorial region, and 13 in the spinal cord. Histopathology on available cases included glioblastoma-like histomorphology (Fig. 1a–f) and most with an infiltrative growth pattern, in 26 of the 30 (86.7%) evaluable cases. The presence of necrosis and/or microvascular proliferation was observed in 38 of 61 (62.3%) HGAP tumors. The piloid-associated features of Rosenthal fibers and/or eosinophilic granular bodies were present in a minority (23 of 61; 37.7%). Mitotic activity was variable, ranging from 0 to 6 mitoses/mm2. Similarly, the maximum Ki-67 proliferative index had a broad spectrum, ranging 1–30%.
Representative high-grade histologic features frequently present in HGAP. a HGAP tumors often had an infiltrative growth pattern. b, c Tumors repeatedly demonstrated pleomorphism and/or poorly differentiated cytomorphologies. d Increased mitotic activity occurred in a subset of cases. e Palisading necrosis and/or f) microvascular proliferation were present in a majority of HGAP tumors. (Scale bars: a 500 μm, b 100 μm, c 100 μm, d 50 μm, e 500 μm, f 100 μm)
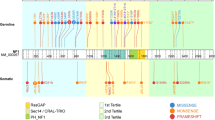
The most frequent genetic alterations were in the Cyclin Dependent Kinase Inhibitor 2A (CDKN2A), Alpha Thalassemia/mental Retardation syndrome, X-linked (ATRX) and Neurofibromin 1 (NF1) genes, consistent with previous findings of HGAP [22] (Supplemental Table 1). CDKN2A was the most commonly altered gene (121/144, 84.1%). ATRX status, as determined either by DNA sequencing or surrogate immunohistochemistry, was known for 111 cases, of which 65 (58.6%) were mutated or lost. NF1 was the most frequently altered gene of the MAPK pathway. NF1 was sequenced in 89 cases, with a mutation identified in 36 (40.4%). FGFR1 was the second most frequently altered MAPK pathway gene, with 17 of the 89 cases sequenced for FGFR1 harboring a mutation (19.1%), and an additional 6 of 43 (14.0%) tested cases harboring fusions of FGFR1 (Supplemental Table 1). BRAF is another MAPK pathway gene for which mutations/fusions were nearly as common as FGFR1 alterations. There were 18 cases of 97 (18.6%) tested which harbored a BRAF fusion, and 2 cases of 86 (2.3%) sequenced were BRAF-V600E mutant. KRAS was a less frequently altered gene of the MAPK pathway, with a mutation identified in 1 of 91 (1.1%) samples tested. The overall composition of MAPK alterations in our HGAP cohort (n = 144), were as follows: n = 79 with one reported MAPK alteration, n = 8 with two or more MAPK alterations, n = 33 without a detected alteration, and n = 24 with no genomic testing results available (Supplemental Table 1). A TERT promoter mutation was identified in 1 of 93 (1.1%) tumors and was mutually exclusive of ATRX loss. Sequencing revealed genetic alterations not yet reported in HGAP, including TP53 mutations and a KANK2::NTRK2 fusion. TP53 mutation status was available for 93 cases, of which 5 (5.4%) contained pathogenic mutations as determined by DNA sequencing. Immunohistochemistry was not used in our evaluation of p53 status.
HGAP epigenetic subtyping with relevant clinical features
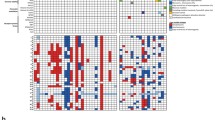
Analyzing whole-genome DNA methylation for multiple glial/glioneuronal tumor types, we confirmed our classifier results in which HGAP forms a distinct tumor type by unsupervised Uniform Manifold Approximation and Projection (UMAP) (Fig. 2A) and unsupervised hierarchical clustering (Supplemental Fig. 1). To identify potential distinct HGAP epigenetic subtypes in an unbiased fashion, we started with unsupervised hierarchical clustering analysis. This clustering led to the identification of three distinct HGAP epigenetic subtypes (Fig. 2b, Supplemental Fig. 2). Cluster subtyping was further supported by unsupervised dimensionality reduction using UMAP analysis (Fig. 2c). The smallest of the three groups (n = 18, 12.5%) was observed to contain a relatively high proportion of patients with a clinical diagnosis of Neurofibromatosis Type 1 (NF1) (p = 0.0008), and therefore this subtype was designated as group NF1 (gNF1) (Figs. 2, 3a). The other two subtypes were designated simply as Group 1 (g1; n = 72, 50.0%) and Group 2 (g2; n = 54, 37.5%). Although the gNF1 was enriched for patients with germline alterations, there was no difference in the frequency of NF1 alterations detected in the tumors across subtypes (Fig. 3b). However, given the fact that hypermethylation of the NF1 promoter region may be an alternate mechanism for NF1 inactivation [10], we evaluated for potential NF1 methylation differences across the three HGAP subtypes. Indeed, 5 of 30 (16.7%) of the CpG probes related to NF1 were differentially methylated in gNF1 when compared to the other subtypes (Fig. 3c).
HGAP resolves into three epigenetically determined subtypes. a Uniform Manifold Approximation and Projection (UMAP) dimensionality reduction confirms that HGAP forms a distinct methylation cluster when compared to other CNS glial/glioneuronal tumors. b Unsupervised hierarchical clustering heatmap of whole-genome CpG methylation for HGAP uncovers the presence of three distinct HGAP epigenetic subtypes. c UMAP dimensionality reduction of HGAP only confirms the presence of three HGAP subtypes. IT Infratentorial, ST supratentorial, PF posterior fossa, SC spinal cord)
NF1 germline status, somatic alterations, and NF1 methylation define a relevant HGAP subtype. a HGAP subtype gNF1 is enriched for patients having Neurofibromatosis Type 1 (NF1) (gNF1 = 33.3%, g1 = 6.9%, g2 = 3.6%) and b) ATRX alterations (gNF1 = 87.5%, g1 = 59.6%, g2 = 47.5%). b CDKN2A/B (gNF1 = 83.3%, g1 = 83.3%, g2 = 85.5%) and NF1 (gNF1 = 62.5%, g1 = 41.3%, g2 = 32.4%) alterations are common across HGAP subtypes. c There are 5 of 30 (16.7%) CpG probes in the NF1 regulatory region that are differentially methylated in the gNF1 subtype, 3 of which are in the gene enhancer region
Several clinical features beyond NF1 status tended to differ across HGAP subtypes. The initial tumors of gNF1, g1, and g2 subtypes had median ages of 43.5 years, 47.0 years, and 32.0 years, respectively (p > 0.05 for all pairwise comparisons, Mann–Whitney U tests, Fig. 4a, b). Only one of the twelve pediatric tumors was gNF1 subtype (Supplemental Table 2). Tumor localization varied among the three HGAP subtypes, with gNF1 having 100% of the tumors located in the posterior fossa (Fig. 4c, d). By contrast, subtypes g1 (35.4% supratentorial, 55.4% posterior fossa, 9.2% spinal cord) and g2 (29.2% supratentorial, 56.2% posterior fossa, 14.6% spinal cord) were found throughout the neuraxis (Fig. 4d). While HGAP subtype was associated with tumor location (chi-square, **p = 0.001) the significance was less than that of subtype association with NF1 syndrome. Furthermore, no association was observed between tumor location and NF1 syndrome (p = 0.359). Tumor location itself was not associated with difference in survival (Supplemental Fig. 3). Progression-free survival trended towards worse outcome for the gNF1 subtype when compared to the other two subtypes (Fig. 4e, f). The gNF1, g1, and g2 subtypes had median progression-free survival times of 252 days, 565 days, and 618 days, respectively. Overall survival appeared similar among all subtypes, although the number of patients with overall survival data was low (Fig. 4g, h). Three of the patients had recurrent tumors, all of which maintained their subtype assignment when compared to their original tumor. The first of these three patients had known Neurofibromatosis Type 1, with their original and two recurrent HGAPs all belonging to the gNF1 HGAP subtype. The other two patients did not have a clinical diagnosis of Neurofibromatosis Type 1. The second patient had paired initial and recurrent HGAP tumor in the g1 subtype while the third patient had paired initial and recurrent HGAP tumor in the g2 subtype.
Clinical features of HGAP methylation subtypes. Median age with interquartile range for a all HGAPs and b HGAP subtypes (p > 0.05 for all pairwise comparisons, Mann–Whitney U tests). Localization for c all HGAP tumors and d HGAP by subtype (ST supratentorial, PF posterior fossa, SC spinal cord). Progression-free survival (PFS) with number at risk for e all HGAP subtypes and f gNF1 subtype versus the others. Overall survival (OS) with number at risk for g all HGAP subtypes and h gNF1 subtype versus the others
HGAP subtype specific genetic, epigenetic, and tumor microenvironment features
In addition to NF1 alterations already discussed, we evaluated subtype-specific genetic changes in the other commonly altered genes ATRX and CDKN2A/B, as well as the novel TP53 mutants and NTRK2 gene fusion. ATRX alterations (as determined by immunohistochemistry and/or sequencing) were overrepresented (p = 0.023) in the gNF1 subtype (87.5%) when compared to the g1 and g2 groups (59.6% and 47.5%, respectively) (Fig. 3b). No difference in CDKN2A/B deletion status was observed across HGAP subtypes (p = 0.949), with all subtypes having a high frequency of alterations (gNF1 = 84.2%, g1 = 83.3%, g2 = 85.5%) (Fig. 3b). Of the five novel TP53 mutant cases we found, four were present in the g1 subtype, one was in g2, and none were found in the gNF1 subtype. The sole NTRK2 gene fusion positive HGAP was present in the g2 subtype.
Given the clinical and genetic relevance of the gNF1 subtype so far, we sought to interrogate global epigenetic changes to uncover potential underlying biological pathways specifically related to the gNF1 subtype when compared to the other subtypes. First, we performed differential methylation analysis for two separate comparisons, gNF1 versus g1 and gNF1 versus g2 (Fig. 5a, Supplemental Table 3). Then we filtered for the significantly altered genes shared between both comparison lists. Next, we performed MethylGSA for Gene Ontology (GO) analysis. When we look at all gene probes in all gene structure sites, six of the top ten pathways that are relatively dysregulated in the gNF1 subtype involve RNA processes (RNA methylation, RNA methyltransferase activity, DNA-directed RNA polymerase complex, RNA polymerase complex, nuclear DNA-directed RNA polymerase complex, and RNA polymerase II holoenzyme) while the last four processes involve protein regulation (histone deacetylase complex, protein acetylation, protein deubiquitylation, protein modification by small protein removal) (Fig. 5b). Next, we sought to understand what portion of the gene structure (promoter versus gene body) was responsible for the differential methylation signature related to the RNA pathway dysregulation in gNF1 (Fig. 5c). To do this, we made multiple comparisons of CpG probes that were either relatively hypermethylated or hypomethylated in the gNF1 subtype. Of these comparisons, there were no dysregulated RNA processing pathways in the top ten pathways found in the promoter 1 (TSS1500 + TSS200) or promoter 2 (TSS1500 + TSS200 + 5´ UTR + EXON 1) regions (Fig. 5d, Supplemental Fig. 4). In contrast, three of the top ten pathways in gNF1 hypermethylated gene bodies (coding region), as well as three of the top ten pathways in gNF1 hypomethylated gene bodies, involved RNA processing pathways. Overall, this indicates an association between both hypermethylation and hypomethylation of gene bodies and dysregulation of RNA processing pathways in the gNF1 subtype.
Gene Ontology (GO) pathway analysis of HGAP subtypes. a Schematic for differentially methylated CpG probes between HGAP subtypes and downstream GO analysis. b Top 10 GO pathways that are dysregulated in gNF1 compared to other HGAP subtypes when utilizing all gene structure methylation sites for CpG sites that are either hypermethylated or hypomethylated. c General gene structure of evaluated regions. d RNA processing dysregulation in the gNF1 subtype is a function of hypermethylation or hypomethylation in the gene body region, not promoter region of the associated genes
Given the important contribution of the tumor microenvironment (TME) to tumor biology, we sought to understand the cellular composition of HGAP and its subtypes. To do this, we began by performing methylCIBERSORT [25] for deconvolution of cell type components, both neoplastic and non-neoplastic, from the whole-genome methylation data. For HGAP overall as a tumor type, we found positive correlations between monocytes and B cells (r = 0.56), NK cells and B cells (r = 0.48), and neurons and glia (r = 0.47) (Fig. 6a). These TME cell associations were further confirmed with principal component analysis (Fig. 6b). Next, we analyzed tumor content in a subtype specific manner, beginning with the neoplastic cell component. Subtypes gNF1 and g1 had a lower neoplastic cell content (higher TME cell component) when compared to g2 (p = 0.0055 and p < 0.0001, respectively) (Fig. 6c). When looking at the non-neoplastic cells, g1 and gNF1 appear to have unique signatures. Subtype g1 is enriched for monocytes when compared to gNF1 (p < 0.0001) and g2 (p < 0.0001) (Fig. 6d, e). In partial alignment with the monocytes and B cells being moderately correlated in HGAP, subtype g1 is also enriched in B cells when compared to gNF1 (p = 0.0385), but not when compared to g2 (p = 0.1696). The enriched immune cell component of subtype g1 does not appear associated with primary versus recurrent tumor status, as these occurrences (primary versus recurrence) do not differ across subtypes (Supplemental Table 4). Also consistent with the correlation matrix, subtype gNF1 is enriched for both neurons and glia when compared to subtypes g1 (p < 0.0001 each comparison) and g2 (p < 0.0001 each comparison) (Fig. 6d, e). Summarizing the TME component of HGAP subtypes, gNF1 is enriched for neurons and glia, g1 is enriched for monocytes and B cells, and g2 has the lowest amount of non-neoplastic TME content.
Deconvolution of the tumor microenvironment in HGAP. a Correlation matrix and b principal component analysis of neoplastic and non-neoplastic cell components within HGAP as a single tumor type. c Neoplastic cell contents as a function of HGAP subtype, which is highest in the g2 subtype (error bars as median values with 95% CI). d Non-neoplastic cell contents as a function of HGAP subtype, indicating that glia and neurons are enriched in the gNF1 subtype while monocytes and possibly B cells are enriched in the g1 subtype. e Heatmap summarizing the relative non-neoplastic cell contents in each HGAP subtype
Discussion
The findings from our analysis of a large HGAP cohort has several clinical implications, especially related to tumor classification in general. We have confirmed and helped to solidify the concept that HGAPs are largely infiltrative glial neoplasms, often with glioblastoma-like histology (more so than pilocytic astrocytoma-like histology) and that future WHO classification schemes may consider these features in its nosology [12]. Consideration as to whether the ‘piloid’ designation in HGAP is warranted, as this not reflective of the histology found in most tumors designated as HGAP by DNA methylation profiling. A name more neutral to histological features may be warranted, especially in this tumor type that is currently defined by its DNA methylation profile, not its histological features. DNA methylation profiling has been incorporated into essential or desirable diagnostic criteria for a substantial proportion of CNS tumors in the WHO “blue book” and it is therefore feasible to consider more widespread profiling of cases when HGAP is considered as a diagnostic possibility. DNA methylation profiling to rule out HGAP may be warranted, especially in cases of a cerebellar glial neoplasm that does not fit neatly into another WHO diagnostic category, where the index of suspicion rises in the setting of ATRX loss and/or CDKN2A homozygous deletion. Methylation testing may also be considered when other tumor features are suggestive of HGAP as well, such as a histologically high-grade glioma arising in the setting of Neurofibromatosis Type 1.
Beyond classification, there may also be clinical implications of HGAP subtyping according to distinct epigenetic subtypes. While gNF1 was the least frequent subtype, our data indicates a trend towards a worse outcome than other HGAP subtypes. Furthermore, due to the overrepresented nature of NF1 occurring in the gNF1 subtype, HGAP subtyping analysis may point to the potential need for germline NF1 testing for cases that fall into this class. Our findings are consistent with a recent report describing a larger proportion of HGAPs being found in a series of NF1-associated high-grade gliomas [13]. Methylation findings suggest that the gNF1 subtype may be associated with RNA processing dysregulation, which raises the possibility of this biological process acting as a potential therapeutic vulnerability in gNF1 HGAP, similar to other cancer types [8, 17, 26, 31]. In the future, it is possible that epigenetic subtyping may even be considered for the basis of biomarker-informed clinical trials including HGAP for proper risk stratification [1, 5, 29].
While this study represents a large cohort of patients with HGAP, it has some limitations, including those inherent to a retrospective study. Expanding the patient outcome data would contribute to our understanding of the clinical heterogeneity and understanding of patterns of patient survival for this tumor type. additional helpful data would include surgical extent of resection and adjuvant treatment modalities for these patients. Additionally, next-generation sequencing data were obtained from a variety of different institutions with different gene region coverages, resulting in missing information for some genes. Therefore, it is difficult to know the exact mutational status for all genes of interest for every case in our cohort. We also lacked additional material for orthogonal validation of our biological investigations, such as immunohistochemistry for TME analysis and frozen tissue for high-quality RNA studies of RNA processing dysregulation. Further work is needed to more fully understand the clinical and biologic properties on this recently described tumor type.
In summary, we have characterized an expanded cohort of patients with HGAP by extending the spectrum of gene alterations to include rare TP53 mutations and an NTRK2 fusion, highlighting that most tumors contain glioblastoma-like histological features, and uncovering the presence of three distinct epigenetic subtypes of HGAP. These HGAP epigenetic subtypes are associated with variations in clinical parameters, genetic alterations, and associated biological processes. Specifically, we identified an HGAP epigenetic subtype (provisionally designated here as gNF1) that is associated with enrichment for NF1 syndrome, increased ATRX alteration, increased methylation in the NF1 enhancer region, evidence of RNA processing dysregulation, increased non-neoplastic glia and neuron cell content, and is confined to the posterior fossa with trends towards worse patient outcome. The goal of our work was to expand the understanding of e clinical and biological features of this recently described and understudied tumor type. Further work will likely elucidate additional biologic and clinical features of this newly defined CNS tumor type.
References
Alexander BM, Trippa L, Gaffey S, Arrillaga-Romany IC, Lee EQ, Rinne ML et al (2019) Individualized Screening Trial of Innovative Glioblastoma Therapy (INSIGhT): a Bayesian adaptive platform trial to develop precision medicines for patients with glioblastoma. JCO Precis Oncol. https://doi.org/10.1200/PO.18.00071
Bates D, Machler M, Bolker B, Walker S (2015) Fitting linear mixe-effects models using lme4. J Stat Softw 67:1–48. https://doi.org/10.18637/jss.v067.i01
Bender K, Perez E, Chirica M, Onken J, Kahn J, Brenner W et al (2021) High-grade astrocytoma with piloid features (HGAP): the Charite experience with a new central nervous system tumor entity. J Neurooncol 153:109–120. https://doi.org/10.1007/s11060-021-03749-z
Capper D, Jones DTW, Sill M, Hovestadt V, Schrimpf D, Sturm D et al (2018) DNA methylation-based classification of central nervous system tumours. Nature 555:469–474. https://doi.org/10.1038/nature26000
Cimino PJ, McFerrin L, Wirsching HG, Arora S, Bolouri H, Rabadan R et al (2018) Copy number profiling across glioblastoma populations has implications for clinical trial design. Neuro Oncol 20:1368–1373. https://doi.org/10.1093/neuonc/noy108
DeSisto J, Lucas JT Jr, Xu K, Donson A, Lin T, Sanford B et al (2021) Comprehensive molecular characterization of pediatric radiation-induced high-grade glioma. Nat Commun 12:5531. https://doi.org/10.1038/s41467-021-25709-x
Dodgshun AJ, Fukuoka K, Edwards M, Bianchi VJ, Das A, Sexton-Oates A et al (2020) Germline-driven replication repair-deficient high-grade gliomas exhibit unique hypomethylation patterns. Acta Neuropathol 140:765–776. https://doi.org/10.1007/s00401-020-02209-8
Ferreira R, Schneekloth JS Jr, Panov KI, Hannan KM, Hannan RD (2020) Targeting the RNA polymerase I transcription for cancer therapy comes of age. Cells. https://doi.org/10.3390/cells9020266
Gu Z, Eils R, Schlesner M (2016) Complex heatmaps reveal patterns and correlations in multidimensional genomic data. Bioinformatics 32:2847–2849. https://doi.org/10.1093/bioinformatics/btw313
Gutmann DH, McLellan MD, Hussain I, Wallis JW, Fulton LL, Fulton RS et al (2013) Somatic neurofibromatosis type 1 (NF1) inactivation characterizes NF1-associated pilocytic astrocytoma. Genome Res 23:431–439. https://doi.org/10.1101/gr.142604.112
Kessler T, Berberich A, Sadik A, Sahm F, Gorlia T, Meisner C et al (2020) Methylome analyses of three glioblastoma cohorts reveal chemotherapy sensitivity markers within DDR genes. Cancer Med 9:8373–8385. https://doi.org/10.1002/cam4.3447
Louis DN, Perry A, Wesseling P, Brat DJ, Cree IA, Figarella-Branger D et al (2021) The 2021 WHO Classification of Tumors of the Central Nervous System: a summary. Neuro Oncol. https://doi.org/10.1093/neuonc/noab106
Lucas CG, Sloan EA, Gupta R, Wu J, Pratt D, Vasudevan HN et al (2022) Multiplatform molecular analyses refine classification of gliomas arising in patients with neurofibromatosis type 1. Acta Neuropathol. https://doi.org/10.1007/s00401-022-02478-5
Mackay A, Burford A, Molinari V, Jones DTW, Izquierdo E, Brouwer-Visser J et al (2018) Molecular, pathological, radiological, and immune profiling of non-brainstem pediatric high-grade glioma from the HERBY Phase II Randomized Trial. Cancer Cell 33(829–842):e825. https://doi.org/10.1016/j.ccell.2018.04.004
Min JL, Hemani G, Davey Smith G, Relton C, Suderman M (2018) Meffil: efficient normalization and analysis of very large DNA methylation datasets. Bioinformatics 34:3983–3989. https://doi.org/10.1093/bioinformatics/bty476
Pajtler KW, Witt H, Sill M, Jones DT, Hovestadt V, Kratochwil F et al (2015) Molecular classification of ependymal tumors across all CNS compartments, histopathological grades, and age groups. Cancer Cell 27:728–743. https://doi.org/10.1016/j.ccell.2015.04.002
Parker KA, Robinson NJ, Schiemann WP (2022) The role of RNA processing and regulation in metastatic dormancy. Semin Cancer Biol 78:23–34. https://doi.org/10.1016/j.semcancer.2021.03.020
Pratt D, Abdullaev Z, Papanicolau-Sengos A, Ketchum C, Panneer Selvam P, Chung HJ et al (2022) High-grade glioma with pleomorphic and pseudopapillary features (HPAP): a proposed type of circumscribed glioma in adults harboring frequent TP53 mutations and recurrent monosomy 13. Acta Neuropathol 143:403–414. https://doi.org/10.1007/s00401-022-02404-9
Pratt D, Lucas CG, Selvam PP, Abdullaev Z, Ketchum C, Quezado M et al (2022) Recurrent ACVR1 mutations in posterior fossa ependymoma. Acta Neuropathol 144:373–376. https://doi.org/10.1007/s00401-022-02435-2
Raffeld M, Abdullaev Z, Pack SD, Xi L, Nagaraj S, Briceno N et al (2020) High level MYCN amplification and distinct methylation signature define an aggressive subtype of spinal cord ependymoma. Acta Neuropathol Commun 8:101. https://doi.org/10.1186/s40478-020-00973-y
Reinhardt A, Stichel D, Schrimpf D, Koelsche C, Wefers AK, Ebrahimi A et al (2019) Tumors diagnosed as cerebellar glioblastoma comprise distinct molecular entities. Acta Neuropathol Commun 7:163. https://doi.org/10.1186/s40478-019-0801-8
Reinhardt A, Stichel D, Schrimpf D, Sahm F, Korshunov A, Reuss DE et al (2018) Anaplastic astrocytoma with piloid features, a novel molecular class of IDH wildtype glioma with recurrent MAPK pathway, CDKN2A/B and ATRX alterations. Acta Neuropathol 136:273–291. https://doi.org/10.1007/s00401-018-1837-8
Ren X, Kuan PF (2019) methylGSA: a bioconductor package and Shiny app for DNA methylation data length bias adjustment in gene set testing. Bioinformatics 35:1958–1959. https://doi.org/10.1093/bioinformatics/bty892
Robinson JT, Thorvaldsdottir H, Wenger AM, Zehir A, Mesirov JP (2017) Variant review with the integrative genomics viewer. Cancer Res 77:e31–e34. https://doi.org/10.1158/0008-5472.CAN-17-0337
Singh O, Pratt D, Aldape K (2021) Immune cell deconvolution of bulk DNA methylation data reveals an association with methylation class, key somatic alterations, and cell state in glial/glioneuronal tumors. Acta Neuropathol Commun 9:148. https://doi.org/10.1186/s40478-021-01249-9
Stanley RF, Abdel-Wahab O (2022) Dysregulation and therapeutic targeting of RNA splicing in cancer. Nat Cancer 3:536–546. https://doi.org/10.1038/s43018-022-00384-z
Sturm D, Orr BA, Toprak UH, Hovestadt V, Jones DTW, Capper D et al (2016) New brain tumor entities emerge from molecular classification of CNS-PNETs. Cell 164:1060–1072. https://doi.org/10.1016/j.cell.2016.01.015
Sturm D, Witt H, Hovestadt V, Khuong-Quang DA, Jones DT, Konermann C et al (2012) Hotspot mutations in H3F3A and IDH1 define distinct epigenetic and biological subgroups of glioblastoma. Cancer Cell 22:425–437. https://doi.org/10.1016/j.ccr.2012.08.024
Tanguturi SK, Trippa L, Ramkissoon SH, Pelton K, Knoff D, Sandak D et al (2017) Leveraging molecular datasets for biomarker-based clinical trial design in glioblastoma. Neuro Oncol 19:908–917. https://doi.org/10.1093/neuonc/now312
Thomas C, Sill M, Ruland V, Witten A, Hartung S, Kordes U et al (2016) Methylation profiling of choroid plexus tumors reveals 3 clinically distinct subgroups. Neuro Oncol 18:790–796. https://doi.org/10.1093/neuonc/nov322
Yang B, Wang JQ, Tan Y, Yuan R, Chen ZS, Zou C (2021) RNA methylation and cancer treatment. Pharmacol Res 174:105937. https://doi.org/10.1016/j.phrs.2021.105937
Acknowledgements
This work utilized the computational resources of the NIH HPC Biowulf cluster.
Funding
This work was supported by the Intramural Research Program at the Center for Cancer Research, National Cancer Institute and the National Institute of Neurological Disorders and Stroke of the National Institutes of Health.
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare that there are no conflicts of interest.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Cimino, P.J., Ketchum, C., Turakulov, R. et al. Expanded analysis of high-grade astrocytoma with piloid features identifies an epigenetically and clinically distinct subtype associated with neurofibromatosis type 1. Acta Neuropathol 145, 71–82 (2023). https://doi.org/10.1007/s00401-022-02513-5
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00401-022-02513-5