Abstract
The high cost of producing conventional hybrid cotton seeds led to more research efforts on cotton male sterility systems. There is a lack of studies on cytology, histology, morphological variation, yield, and altered restorer backgrounds to identify and develop male sterility markers in cotton hybrids. Hybrid cotton can be efficiently produced by exploiting genetic male sterility. Among the 19 Genetic Male Sterility (GMS) genes discovered, the lines with ms5ms6 genes are mostly utilised to establish successful hybrid cotton in India. Molecular markers closely associated with the MS alleles are identified to facilitate the efficient and rapid backcrossing of male-sterility genes into elite lines or cultivars by marker-assisted backcrossing. The majority of the markers which are random DNA markers (RDMs), are probably lost, when recombination occurs. In contradiction, molecular markers (functional markers, or FMs) within the genic region can be identified and employed in crops for diverse traits, if prospective characteristic genes are known. In this review, the mechanism of male sterility, its gene expression level, and the need for functional markers for the male sterility trait in cotton have been put forward.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Economic importance of cotton
Gossypium sp., commonly known as cotton, is a principal cash crop used for fibre, as well as edible oil, and protein worldwide. Among the 50 species reported in the genus, four are predominantly cultivated, and G. hirsutum produces 90% of the total cotton fibre. Although genetic engineering technology has been standardised in cotton to a greater extent [1], the phenomenon of heterosis and exploitation of hybrid vigour is still gaining traction in China and India [3]. Strong heterosis has been reported by several researchers for fibre quality, number of bolls, boll weight and seed cotton yield in upland cotton [1].
Role of male sterility in hybrid production
In cotton, intraspecific and interspecific hybrids are produced conventionally through manual emasculation and pollination. But the disadvantages of traditional hybrids are two-fold. Firstly, the emasculation process is tedious and requires employing numerous skilled labourers, which drives up the cost of hybrid seeds. Secondly, the hand emasculation process damages the female reproductive system, which impairs hybrid seed production. Given the above facts, unearthing male sterility becomes vital in cotton.
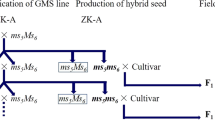
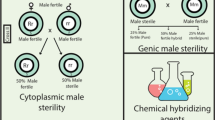
It is an efficacious and effortless way of developing quality hybrid cotton seeds [2]. On the basis of genes responsible for inheritance of male sterility, it can be categorised into three types, namely, nuclear male sterility (also known as genic male sterility; GMS), cytoplasmic male sterility (CMS) and cytoplasmic genetic male sterility (CGMS). In upland cotton, the first line with heritable partial male sterility was discovered by Justus and Leinweber in 1960. Suguna (CPH2), the first GMS-based hybrid in history, was created in 1978 at the Central Institute for Cotton Research (CICR) Regional Station in Coimbatore, India. There are also the CGMS-based hybrids, which include PKV Hy 3, PKV Hy 4, and MECH 4.
Male sterility genes
According to [3], anther development can be categorized into 14 typical stages based on genetic and cytological factors. Based on cell proliferation, development, differentiation, maturation and degradation, the anther and pollen formation process can be fit into four stages namely, premeiotic phase (specification of PMCs and tapetum), meiotic phase (differentiation of tapetal cells), post-meiotic phase (development of pollen wall and mature pollen) and dehiscence of anther and pollen germination phase. An anomaly in any of the stages brings about male sterility in flowering plants (Figure 2). Many transcriptomic, epigenetic and biochemical studies have been conducted to elucidate the mechanism of MS at molecular level. Nevertheless, genes associated with MS have not yet been identified in many species [4,5,6,7]. In GMS cotton, gene identification, mechanism understanding and manipulation of heterosis have been decoded with continual research.
In cotton, the availability of GMS sources have been documented in American, Egyptian, and arboreum species. Both recessive and dominant genes are accountable for governing male sterility in tetraploid cotton. About 19 GMS genes, seven single recessive (ms1, ms2, ms3, ms13, ms14, ms15, and ms16), eight single dominant (Ms4, Ms7, Ms10, Ms11, Ms12, Ms17, Ms18, and Ms19) and four double recessive (ms5ms6 and ms8ms9), have been recorded in tetraploid cotton [8, 9]. All the above genes were discovered in G. hirsutum except Ms11, Ms12, Ms13, Ms18, and Ms19, which were reported in G. barbadense (Table 1). Among the above, only ms14 and ms5ms6 lines have been effectively used in China and India to generate hybrids [10]. Male partial genic sterility has been reported to be conferred by the single recessive genes ms1 and ms3 [8, 11]. Richmond and Kohel, 1961, observed that stable male sterility was imparted by paired recessive genes (ms5 and ms6) and a single recessive gene (ms2) [12]. Justus et al., (1963), reported that, in a greenhouse condition, ms3 plants produced more fertile flowers than in the field. However, recessive genetic mechanisms with advantages of stable and absolute male sterility, particularly by ms5 and ms6, have been frequently exploited in cotton hybrid cultivation since fertile GMS maintainers are needed [13].
Markers for male sterility trait
Male sterility becomes an essential trait in plant species for the complete success of hybridization with ease. To fulfil the requirements of breeding objectives, male sterility plays a major role in getting rid of the difficulties in manual emasculation, thereby solving the problems of the need for skilled labour, time-consuming efforts, labour wages etc. Marker-assisted selection (MAS) has been proven prominent in this cash crop owing to the accessibility of molecular markers and genetic linkage maps. Markers have the utmost importance in transferring the GMS allele to the elite genetic background by MAS. Despite the uses and significance of molecular markers for improvement of other important traits, its necessity in GMS is inevitable. This is because of the practical difficulty that, the screening of sterile plants is only possible during the reproductive stage of the crop. The molecular markers for GMS help in screening of plants in the seedling stage itself and in decreasing the cost and resources of hybrid seed production. This study aims to investigate the genetic basis of male sterility in cotton and develop reliable markers for its identification.
Types of male sterility in cotton
Genetic male sterility (GMS)
One of the best tools for accelerating hybrid breeding is GMS. It is the term used to describe pollen sterility resulting from nuclear genes. Environmental factors such as temperature (TGMS), photoperiod (PGMS), and humidity (HGMS) can have an impact on male gametogenesis in environment-sensitive GMS (EGMS) [14, 15]. In particular, intentional suppression of autogamy and subsequent promotion of allogamy is done through the transmission of advantageous male sterility genes. Thus, GMS has served the purpose of harnessing the yield linked with hybrid vigour [14, 16]. Numerous genes that regulate gene transcription, protein metabolism, and other physiological processes such as anther dehiscence and pollen germination, have been identified by cytological and biochemical investigations of several GMS mutants [17].
Cytoplasmic genetic male sterility (CGMS)
The combination of sterile cytoplasm and nuclear gene(s) conditioning sterility results in CGMS. In contrast to CMS, this kind of male sterility allows for the restoration of fertility. The R gene (s) located in the nucleus is responsible for restoring fertility. Therefore, the fertility or sterility of such plants is determined by the interaction of both nuclear genes and cytoplasmic factors. The primary bottleneck of CGMS is the laborious process of incorporating R genes into cultivated species or genotypes.
When alien cytoplasm from diploid wild species is introduced into cultivated tetraploid cotton, CMS ensues. Cotton contains the comparable independent restorer genes Rf1 and Rf2. Gametophytic Rf2, which is from the D8 genome, is designated as the D8 restorer gene. Sporophytic Rf1, which is from the D2–2 genome, is designated the D2 restorer gene. The two restorer gene loci are closely linked, with a genetic distance of 0.93 cM on the same chromosome [20].
Male sterility could be produced by interaction of three diploid species namely, G. harknessii, G. arboreum, and G. anomalum, with the G. hirsutum nuclear genes. However, the the stability of cytoplasms of G. arboreum and G. harknessii are affected by heat stress. The G. harknessii cytoplasm and its interaction with G. hirsutum genome produces reliable and stable CMS in cotton across many environments. But this also has some linkage drag causing certain unfavourable effects on micronaire value, ginning outturn and susceptibility to bacterial blight [18].
Chemical hybrid agent (CHA) / male gametocides
The labour-intensive nature of emasculation, currently limits the production of hybrid cotton. It impelled scientists and cotton breeders to employ innovative techniques like chemical hybrid agents (CHA), which have been already shown to be effective in crops such as wheat, rape and rice [19]. The effects of most common CHAs like ethrel and benzotriazole were studied in cotton. It was resulted in a range of 96.8 to 100% pollen sterility by both the agents. Nevertheless, a substantial decline in yield characteristics was associated with it. On the other hand, plants sprayed with lower ethrel concentrations could not exhibit a discernible decrease in yield. Thus, it is conceivable to use ethrel as a hybridizing agent with G. hirsutum. [20].
Furthermore, glyphosate, a significant herbicide, was considered a possible candidate for CHA after it was found to induce male sterility in cotton. [21]. The sterile phenotypes, that resulted in glyphosate application, were having the following typical characteristics: (1) shorter and fused filaments; (2) greater interspace between androecium and stigma; (3) indehiscence of anther; (4) tapetum degradation; (5) enhanced expression of G6-EPSPS (roughly eight times, in pistil) and (6) increase of ascorbate peroxidase (APX) activity in stamens, but decrease in other organs. Compared to manual emasculation, it is reliable in inducing male sterility because of its ability to cause complete pollen abortion and lack of negative effects on F1 yield traits and fibre quality [22]. However, the reproductive portions were observed to suffer significant harm from glyphosate. It has been reported to cause aberrant flowers, along with non-dehiscent anthers with less viable pollen grains having uneven shapes [23].
Transgenic male sterility
Biotechnological approaches have been widely employed to develop transgenic plants in several crops, cotton (G. hirsutum L.), soybean (Glycine max (L.) Merr.), canola (Brassica napus L. and B. rapa L.) and maize (Zea mays L.), for exploitation in commercial scale. Agrobacterium-mediated genetic transformation and particle bombardment have been used to develop transgenic male sterility in crops like citrus, wheat, soybean, tobacco, rapeseed, rice, sesamum and mustard. Transgenic male sterility is the term used to describe male sterility caused by genetic engineering. The ‘A’ line incorporates the ‘barnase’ gene, which causes male infertility. Another gene, barstar can be utilized to restore fertility since it inhibits the activity of the male sterility gene barnase. It has been reported that up to 50% of the transgenic plants exhibit stable male sterility, and in a few instances, around 90% of transgenics exhibited stable male sterility in few crops like mustard, rapeseed, and tobacco, but the same is not yet available in cotton. Therefore, in the near future, this approach can be used to produce commercial hybrid cotton [24].
GMS - atop the CGMS
The drawback of GMS is that, in theory, heterozygous pollinators will result in segregation as 1:1 male sterile and fertile plants in the progenies. Accordingly, it is sometimes not economically feasible to score and eliminate the fertile offsprings [25]. Despite that, in comparison, the GMS line is efficient and inexpensive to a greater extent in commercial hybrid cotton seed production. Practically, since every commercial cultivar can act as its own restorer line in GMS, it can promote random mating to generate hybrids, in contrast to CMS. One among the breeding advantages of GMS, contrasted with CMS, is the effortless transfer of the MS trait to an array of genetic backgrounds. It is achieved by consecutive backcrossing and inheritance with complete penetrance in subsequent generations [26].
There also exists evidence regarding the undesirable consequences of the sterile cytoplasm in the case of CGMS. The yield of hybrid cotton is negatively impacted by sterile cytoplasm, which was evident in the study of [27]. The yield reduction of the hybrids produced by employing restorers has been reported to be 4.76 to 15.63%, which could be due to the sterile cytoplasm from G. harknessii. Nonetheless, no adverse impact was detected regarding ginning out turn and further characteristics of fibre quality. Even though, CMS-based hybrids showed an enhanced span length, the hybrid combinations showed the largest reduction in micronaire value, up to 30%. In a similar vein, CMS-based hybrids showed an increase in tenacity value, which was mirrored in the strength/length ratio. To identify the right hybrids with higher yield potential and better fibre quality utilizing CMS system, a larger number of cross combinations with suitable restorer and stable CMS lines are employed. In this manner, the negative yield-reducing effect of sterile cytoplasm could be mitigated [27].
Genesis of MS
Male sterility arises due to varied reasons that differ on the basis of genetic background and type. Numerous loci regulate a series of essential steps involved in microsporogenesis genetically, and a loss-of-function mutation in any one of them could lead to the generation of underdeveloped microspores or non-functioning pollen grains (Fig. 1) and thus, male sterility [28]. The genes responsible for male sterility have been annotated in various species and are still not yet discovered in many species. As far as cotton is concerned, the genes and molecular mechanism of GMS and CMS have been exploited better enough at the transcriptomic, biochemical and epigenetic levels. A majority of the research were executed on cytoplasmic and recessive genic male sterility, whereas only a few on dominant genic male sterility, particularly in cotton. This is owing to the unavailability of such mutants [29].
On grounds of the apprehension in the complex mechanism of male sterility and its responsible genes in the model crops, the germplasms were exploited to discover ms-related genes in other crops [30]. The genes, ms5 and ms6, were hypothesized to have originated either from G. tomentosum or from the progeny of an interspecific hybrid with G. hirsutum [26]. While the precise molecular and biochemical reasons for male sterility produced by ms5ms5 and ms6ms6 remain ambiguous, research on similar species may be able to identify the mechanisms that account for pollen development failure in cotton. Four scenarios were identified as the root causes of male gamete development failure in Arabidopsis: Absence of tryphine, the substance that coats pollen; non-dehiscent anthers; non-functioning stamen that results in malformed anthers; and failure in microsporogenesis [31]. Each of the four cases could be responsible for male sterility in cotton. In view of the above fact, male sterility in cotton is ascribable to obstruction in the post-meiotic phase in GMS and abnormalities in the pre-meiotic phase in CGMS systems [32] (Fig. 2).
The phenomenon of GMS is a consequence of loss of function in the nuclear genes, accountable for male reproductive development. The microspores were vacuolated and nuclei were convoluted resulting in shrinkage of cytoplasm, and finally, disintegration of deformed developing microspores, forming the mechanism of GMS [32] (Fig. 2). Genetic studies of GMS in cotton revealed that non-dehiscent anthers and unviable pollens are the common forms of occurrence, resulting in failure of reproductive development. Furthermore, the metabolism of sugars has a significant role in the male reproduction of flowering plants. Any shortcomings in this metabolic pathway in the time of male reproductive development results in GMS [33]. According to physiological and biochemical studies carried out, male sterile plants have lower starch levels during the entire period of pollen development and lower soluble sugar levels in young stamens [34].
In Photosensitive Genetic Male Sterility (PGMS) system, day length and photoperiod modulate the pollen fertility due to premature tapetum degeneration [35]. In expression profiling studies of sterile mutant and fertile wildtype (WT) anthers, under long-day conditions, induction of ubiquitin-proteasome system mutant uninucleate pollen (UNP) were observed to cause degradation of pollen proteins, that formed the basis of male sterility [36]. Both the reproductive and vegetative sections of plants are impacted by high temperature, although the former is more sensitive [37]. According to [38], the male reproductive system is more susceptible to High Temperature (HT) stress than the female reproductive system. As the anther development is exceptionally organized in comparison to other male reproductive components, it is more sensitive to HT stress [39]. Numerous genes, whose expression and regulation were impacted by HT, are involved in the intricate process of anther and pollen growth [38]. Flower developmental stages, the structure of anther and the cellular morphology of microspore are impacted, even by a single day of high-temperature stress [40].
Being a maternally inherited trait, CMS eventuate due to energy deficiency in mitochondria, CMS protein cytotoxicity and programmed cell death (PCD) of premature tapetum [35]. The transfer of diploid genome (A2) of G. arboreum to G. anomalum cytoplasm, having B1 diploid genome, created the first cytoplasmic male sterility. When G. arboreum pollen was used to pollinate the CMS inducer, male sterile plants were created and when G. anomalum pollen was used to pollinate the sterile lines, fertility was regained [41, 42]. As these CMS lines had undesirable traits and the resultant male sterility was incomplete and unstable, a different native CMS system known as CMS-D2 was developed by transferring the commercial cotton genome (G. hirsutum) to G. harknessii cytoplasm [43, 44]. Tapetum plays a crucial role in case of pollen nourishment, development and maturation. However, premature degeneration of the tapetum in the later stages of pollen mother cell arrested the normal course of meiosis in sterile lines. These pre-meiotic abnormalities seemed to be the root cause of CMS in Gossypium sp [34] (Fig. 2).
A restorative gene, Rf1, from G. harknessii was transferred to G.hirsutum genome to form the CMS-D2 restorer system [44]. Another restorer, Rf2 was also discovered in genome D8 of G. trilobum [45] and its cytoplasm was utilized to build CMS-D8 system [46]. A nonallelic relationship and tight linkage were discovered between Rf1 (sporophytically functioning) and Rf2 (gametophytically functioning) in chromosome LGD08 by [47, 48]. The fertility of both CMS-D2 and CMS-D8 systems could be restored by Rf1, while Rf2 could restore the fertility of CMS-D8 sterile lines only [49].
In CGMS, the flower traits were consequentially different, whereas all other morphological traits were akin in fertile and sterile counterparts. The early stages of microspore development were indistinguishable in fertile and sterile plants. Some of the floral traits like the size of ovary, style, staminal column, anther number and anther filament length were decreased in CGMS lines, whereas their size differences were obscure in GMS lines [32]. Despite the fact that CMS produces nearly 100% sterility, a wider use of CMS has not been practicable because of the cytoplasmic effect on CMS stability and the presence of potential yield drag and limited genetic combinations of CMS and restorer lines [50].
Gene expression in MS
It is essential to comprehend the complex mechanism, expression, regulation, and activity of several genes as well as the quantitative nature of male sterility. [28]. In prior studies, an array-based technique was implemented to compare differentially expressed genes. Presently, RNA-seq, a next-generation sequencing method, is prominently used to reveal the transcriptome between the sterile line and its maintainer. It is because the microarray-based chip is unable to find unique transcripts implicated in targeted pathways. Compared to micro-array, RNA-seq offers expression profiling results with high accuracy and minimum background signal at a faster and less expensive rate. Expression level detection in distinct allelic variants forms its prominent application [51].
The differences in gene expression between anthers of fertile and sterile lines in several species were explored by proteomic studies, that help in decoding the MS mechanism. Numerous proteins and their corresponding transcripts concerned with cytoskeleton formation, energy conversion, stress tolerance and defence mechanisms, have been reported in angiosperm mature pollen.
Down-regulation of proteasome and protein 5B, both of which effectuate in degeneration of tapetum in 7B-1 male sterile mutants of tomato have been reported. They also had high-level of cystatins, which are endogenous proteolytic regulators of programmed cell death (PCD), besides seed development and germination [52]. In addition, another proteomic analysis disclosed that proteins linked to photosynthesis, flavonoid synthesis, energy and carbohydrate metabolism have adequate contributory roles in pollen development. Down-regulation of these proteins caused CMS in Brassica napus [53]. In upland cotton, two enzymes linked to photosynthesis and carbohydrate metabolism, viz., glutaminyl-tRNA synthetase and cytosolic ascorbate peroxidase 1, were found to be low in the mutant anthers of the GMS line implicating their role in development of pollen [54].
Studies based on RNA-seq unlocked the exact mechanism regarding CMS in cotton. Signalling pathway and gene enrichment analysis indicate that abnormal regulation of tubulin, actin and myosin-related genes may be the cause of the early developmental defects in cytokinesis of pollen. It was also found that some fertility restorer genes, such as the PPR family, were downregulated. Enhanced expression of NAC, HSP and WRKY and down-regulation of transcription factors like MYB, bHLH and TCP is associated with pollen development and circadian clock. Furthermore, aberrant pollen development and definitive male sterility could be caused by the down-regulation of genes taking part in cell wall development and energy metabolism in pollen maturation [55]. RNA sequencing could also be used to identify candidate genes for fertility restoration by using bulked segregant analysis (BSA) in F2 individuals of CMS and restorer lines. According to one such report by [56] the genes namely GH_D05G3183, GH_D05G3384, and GH_D05G3490 identified in the restorer line R186, restores fertility in the CMS line, 2074 A.
By using light and electron microscopy, it was possible to determine the primary causes of male sterility phenotype in PGMS mutants with uneven exine, lack of tryphine, and immature anther cuticle. It may be a consequence of the down-regulation of genes, ABA and MYB transcription factors, involved in the assembly of tryphine and anther cuticle. The functional expression of these genes may be related to the conversion of fertility in various photoperiods [57].
Other male reproduction genes include four homoeologous pairs of GhFAD2 gene family (GhFAD2–1 to GhFAD2–4), among which GhFAD2–3 play a major role in anther development in cotton. When its RNA is silenced and expression is inhibited, anther developmental stages after meiosis are affected until the maturation of pollen, resulting in male sterility [58]. Pollen exine is one of the important structures in pollen grains, with role in its viability. The gene GhTKPR1_8 has been identified as a crucial gene in the synthesis of sporopollenin in exine. It plays an integral role in catalyzing the reduction of tetraketone carbonyl to hydroxylated α-pyrone. [59]. According to the authors, expression of GhTKPR1_8 is localized in the endoplasmic reticulum and anther developmental tetrad stage. Dehiscence of anther and viability of pollen are affected by the knockdown of the gene. Apart from this, some flavonoids play a role in pollen fertility and plant growth. The contribution of 4-Coumarate CoA ligase (4CL) and flavonoids in cotton anther remains understudied. A homologous gene pair, Gh4CL20A and Gh4CL20, have been reported to expresses in petals as well as stamens. Mutants for the genes exhibited white petals, aborted pollen in reduced number and indehiscent anthers. This male sterility was caused by tapetum degradation at the tetrad stage and abnormal development of pollen at the maturation stage, which was revealed in histological studies [60].
Since post-meiotic abnormalities are the root cause of male sterility in cotton, microspore development and pollen maturation are the crucial phases to study for male sterility. The ill effects of environmental stresses during the above sensitive stages are overcome by the anther cuticle and pollen exine. Glycerol-3-phosphate acyltransferase (GPAT) is involved in cuticle formation and hence critical for fertility studies. GhGPAT12/25 (a paralogs pair on A12/D12 sub-chromosome of cotton) is engaged in the formation of cuticle, development of exine and degradation of tapetum. The mutant developed by knocking down both the genes, exhibited all the symptoms of male sterility, viz. swollen tapetum, abnormally developed anther cuticle and irregularly shaped inviable microspores with exine defect. It was also reported that GhMYB80s enhance the expression of these genes [61].
Development of molecular markers for GMS in cotton
It is vital to create hybrids by functional male sterility, to increase the financial worth of commercial hybrid seed production. For rapid identification and elimination of individuals in a population, the use of molecular markers becomes inevitable, especially in GMS. The marker development in cotton for the MS trait improves the efficiency of hybrid production by decreasing the time and increasing the accuracy in selection.
The map placements of GMS genes, ms5 and ms6, were determined in backcross populations by [10], by utilising higher-density SSR markers. Molecular markers for each gene, indicating dominant and recessive alleles, were reported by the authors by testing these patterns in male sterile (ms5ms5ms6ms6) and male fertile (Ms5ms5Ms6ms6) progenies. The markers that were linked to the genes were compared to other molecular markers to determine their genetic distance. An alternative strategy for the above is to exploit the populations, that are selective for only one of the two alleles, with the second gene fixed to recessive homozygote [26].
It was anticipated that ms5 and ms6 genes are homeologs and perhaps duplicated during polyploidization. Further genomic polymorphisms from the second homoeologous chromosome in flanking regions can be employed as flanking markers, giving the two haplotypes a higher rate of precision in phenotypic prediction. With 98% accuracy, two haplotypes made up of two SNPs each linked to ms5 and ms6, were found to explain male sterility and fertility [26].
In order to uncover chromosomal variations at ms5 and ms6 loci, between GMS and wild-type fertile inbred lines, sequence analysis was performed. Sequence polymorphism showed an association of ms5 and ms6 on A12 and D12, respectively. Four SNPs, MOGH583971, MOGH582973, MOGH211275, MOGH493571, that target the ms5 and ms6 genic regions, made up of a haplotype marker-set was developed by [26] and it was validated for association with the GMS trait in cotton. Over 99% of the GMS trait phenotype could be predicted with this haplotype SNP collection [26]. Using this marker set, molecular breeding at a high-throughput level could be attained to select sterile individuals and increase the reliability of hybrid production. It also encourages introducing male-sterility genes into elite lines or cultivars quickly and efficiently, through marker-assisted backcrossing. A rapid approach to find GMS gene markers in the interspecific biparental population using bulk segregant analysis (BSA) has also been reportrd in cotton. Recently, the Ms5 and Ms6 genes have also been discovered to be associated with four SNP markers. The Ms5 gene associated with markers i23493Gh and i46470Gh and Ms6 gene associated with i08605Gh and i08573Gh were reported [62]. These markers would be helpful for rapid screening of numerous individuals and facilitate precise marker-assisted selection (MAS), contributing to accelerated hybrid cotton seed production (Table 2).
Functional marker development
The selection of morphological features of interest in precision breeding is an unambiguous method of plant breeding. It is accomplished accurately only by locating a marker region, that is derived directly from the trait-controlling genic sequences. The above-stated marker regions are designated as functional markers (FMs), or DNA markers, originating from sequence motifs that are functionally characterized [71]. Random DNA markers (RDMs) form the majority of the available marker types like RAPD, RFLP, AFLP, SSR, and SNP, which can be lost during recombination. On the other hand, FMs are impervious to recombination events and evolutionary changes and hence are highly advantageous.
Consequently, compared to genic molecular markers (GMMs) and Random DNA markers (RDMs), SNPs as FMs are more helpful in plant breeding. Even though GMMs might prevail functionally within a desirable gene, they might not be associated with the desired phenotypic character, resulting in false selection in MAS. Wherefore, RDMs have been referred to as “non-perfect markers,” whereas FMs have been termed as “perfect markers”, “precision markers” or “diagnostic markers” [72].
Finding a gene of interest that influences a phenotypic feature, determining its nucleotide sequence, and characterizing its function are the initial steps in the construction of FMs. Many methods, including QTL mapping, expression profiling, positional cloning, and transposon tagging, can be utilised to identify the genes [73]. To functionally characterize a potential gene, plants must be transformed for overexpression or knockdown study [74]. The investigation of allelic variation within identified genes is the second stage in the development of FMs. Then, in order to determine which polymorphism causes variance of the phenotypic trait, allele sequencing between genotypes must be carried out. The allelic or genic sequence, associated with the desirable trait, must be understood critically in order to develop FMs [74].
Need of the hour – FMs
Random DNA markers do not always result in predicted selection for desirable traits as they are produced from the polymorphic regions surrounding the gene of interest. Nevertheless, the selection by FMs can now solve the issues related to the usage of RDMs, by virtue of 100% predictivity of the appropriate phenotype [75].
Owing to the fact that FMs are found inside the target genes (Fig. 3), they are well connected to the morphological characteristics, that makes it highly effective and reliable. In numerous genetic backgrounds, functional sequence motifs (FSMs) fix the alleles without further calibration once genetic effects have been attributed to them. This marker application is useful for selecting parental materials to create segregating populations and for selecting advanced breeding lines. They are resistant to recombination, which acts as the major reason to accurately and quickly characterize germplasm for allele diversity [73]. They also make it possible to lessen the probablity of information loss and incorrect selection in employing MAS.
Furthermore, QTL validation in MAS is required when exploited in diverse genetic backgrounds, whereas, FMs do not need validation. Breeders can detect unusual recombinants by using FMs to help select remarkable phenotypic features in a large population.
Unearthing the FM discovery in plant breeding
The FM development is thus made possible by finding SNPs and indels associated with a variety of economically significant traits. Large-scale genomic impacts caused by alleles have the potential to eliminate phenotypic variation through natural selection. SNP-derived FMs are advantageous compared to indel-derived markers, because, FMs are extensively dispersed across the genome. They have been generated for an array of agronomic, qualitative and stress resistance traits, to exploit MAS, MARS, MABB and GS methods.
However, for the male sterility gene, the FMs discovered are not ample in many crop species (Table 3) and unavailable in the case of cotton. Despite that, functional markers were developed in cotton for cellulose synthase genes, that play a crucial role in fibre development and architecture of plant type. Identification of SNPs was done in silico via Primer-BLAST gene-specific marker design. As a result, 10 SNP markers and 82 gene-specific markers were created in total by [76] These markers were capable of recognizing genotypes from several species [76].
Concluding remarks and future perspectives
Research on male sterility has a wider scope as the mechanisms are complicated in each of the crop species. In cotton, the male sterile and fertile anthers undergo a similar microsporogenesis process. But, shortly after tetrad formation, the sterile anthers start to differ in pollen development by resisting the release of pollen from the callose wall. During the developmental process, the pollen wall collapses in sterile anthers. Consequently, the malformed microspores disintegrate, resulting in GMS. It has been discovered that the cause of sterility in case of CMS is tapetum abnormalities. The meiocytes starve and their normal path of development is interrupted when early degeneration of the tapetum occurs in the sterile lines. As such, it becomes ever more crucial to explore the genes underlying these processes and to develop markers for the rapid production of hybrid seeds.
Despite availability of many random markers, FMs have been widely employed in many crops due to their accuracy in selection. As FMs are mostly produced by coding DNA within genes, it is possible to use these markers to directly choose a phenotypic trait. FMs can also be transferred across species and hence can be applied to those for which there are insufficient genetic resources. They are also used in phylogenetic investigations, QTL mapping, diversity analysis, germplasm screening and gene discovery. Furthermore, the creation of affordable FMs is an added advantage for their successful application in precision breeding. This versatile tool needs to be exploited for complex traits like male sterility, for the effortless hybrid development. The future outlook is the creation of allele-specific FMs, which will further enhance the effectiveness of direct selection. In addition, the full potential of next-generation breeding methods such as GWAS and GS could be achieved through the development and application of functional markers. In summary, considering the critical role of GMS and CGMS in exploitation of hetersis in cotton, research focus on development of functional markers for male sterility needs intensification.
Data availability
No datasets were generated or analysed during the current study.
References
Liu D et al (2015) Construction of a high-density genetic map and lint percentage and cottonseed nutrient trait QTL identification in upland cotton (Gossypium hirsutum L). Mol Genet Genomics 290:1683–1700
Saeidnia F, Hamid R (2023) Heterosis in cotton: challenges and opportunities of hybrid cultivars production. Iran J Cotton Researches 11(1):77–96
Zhang D, Luo X, Zhu L (2011) Cytological analysis and genetic control of rice anther development. J Genet Genomics 38(9):379–390
Fan Y et al (2016) PMS1T, producing phased small-interfering RNAs, regulates photoperiod-sensitive male sterility in rice, Proceedings of the National Academy of Sciences, vol. 113, no. 52, pp. 15144–15149
Li J, Nadeem M, Sun G, Wang X, Qiu L (2019) Male sterility in soybean: occurrence, molecular basis and utilization. Plant Breeding 138(6):659–676
Liu J et al (2022) Wheat male-sterile 2 reduces ROS levels to inhibit anther development by deactivating ROS modulator 1. Mol Plant 15(9):1428–1439
Nie H, Cheng C, Kong J, Hua J (2023) Plant non-coding RNAs function in pollen development and male sterility. Front Plant Sci 14:1109941
Justus N, Leinwebr C (1960) A heritable partially male-sterile character in cotton
Bowman DT, Weaver JB (1979) Analyses of a Dominant Male-Sterile Character in Upland Cotton. II. Genetic Studies 1, Crop Science, vol. 19, no. 5, pp. 628–630, Sep. https://doi.org/10.2135/cropsci1979.0011183X001900050020x
Chen D, Ding Y, Guo W, Zhang T (2009) Molecular mapping of genic male-sterile genes ms15, ms5 and ms6 in tetraploid cotton, Plant Breeding, vol. 128, no. 2, pp. 193–198
Justus N, Meyer JR, Roux JB (1963) A partially male-sterile character in upland cotton
Richmond T, Kohel R (1961) Analysis of a completely male-sterile character in American upland cotton
Basu A (1996) Current genetic research in cotton in India. Genetica 97(3):279–290
Chen L, Liu Y-G (2014) Male sterility and fertility restoration in crops. Annu Rev Plant Biol 65:579–606
Xue Z et al (2018) Deficiency of a triterpene pathway results in humidity-sensitive genic male sterility in rice. Nat Commun 9(1):604
Bohra A et al (2017) Novel CMS lines in pigeonpea [Cajanus cajan (L.) Millspaugh] derived from cytoplasmic substitutions, and their effective restoration and deployment in hybrid breeding. Crop J 5(1):89–94
Abbas A et al (2021) Exploiting genic male sterility in rice: from molecular dissection to breeding applications. Front Plant Sci 12:629314
Singh SB, Hebbar J, John S (2002) Registration of Genetic male sterile line of cotton. Indian J Plant Genetic Resour 15(3):302–303
Fu D et al (2014) Utilization of crop heterosis: a review. Euphytica 197:161–173
Chauhan S, Gupta H (2005) Detergent-induced male sterility in chickpea (Cicer arietinum L). INDIAN J Genet PLANT Breed 65(03):215–216
Feng PC et al (2014) Improving hybrid seed production in corn with glyphosate-mediated male sterility. Pest Manag Sci 70(2):212–218
Mei L et al (2019) Characterizations of male sterility in a glyphosate-tolerant upland cotton (Gossypium hirsutum L.) induced by glyphosate and its assessments on safety utilization. Ind Crops Prod 134:318–327
Yasuor H, Riov J, Rubin B (2007) Glyphosate-induced male sterility in glyphosate-resistant cotton (Gossypium hirsutum L.) is associated with inhibition of anther dehiscence and reduced pollen viability. Crop Prot 26(3):363–369
Ray K, Bisht NC, Pental D, Burma PK (2007) Development of barnase/barstar transgenics for hybrid seed production in Indian oilseed mustard (Brassica juncea L. Czern & Coss) using a mutant acetolactate synthase gene conferring resistance to imidazolinone-based herbicide’pursuit’. Curr Sci, pp. 1390–1396
Kaul ML, Kaul ML (1988) Gene-cytoplasmic male sterility, Male sterility in higher plants, pp. 97–192
Feng X et al (2015) Development of molecular markers for genetic male sterility in Gossypium hirsutum. Mol Breeding 35:1–9
Tuteja O, Verma S, Singh M (2008) Effect of G. Harknessii based cytoplasmic male sterility on seed cotton yield and fibre quality traits in upland cotton (Gossypium hirsutum L). Indian J Genet Plant Breed 68(03):288–295
Morales KY, Bridgeland AH, Hake KD, Udall JA, Thomson MJ, Yu JZ (2022) Homology-based identification of candidate genes for male sterility editing in upland cotton (Gossypium hirsutum L). Front Plant Sci 13:1006264
Cheng X-Q et al (2020) Characterization and transcriptome analysis of a dominant genic male sterile cotton mutant. BMC Plant Biol 20:1–14
Farinati S et al (2023) Current insights and advances into plant male sterility: new precision breeding technology based on genome editing applications. Front Plant Sci 14:1223861
Okada K, Shimura Y (1994) Genetic analyses of signalling in flower development using Arabidopsis. Plant Mol Biol 26:1357–1377
Khadi B, Kulkarni V, Katageri I, Soddi R (1998) Male sterility-New Frontiers in Cotton Breeding, presented at the Proceedings of the world cotton research conference-2, Anthens, Greece, pp. 246–249
Liu S, Li Z, Wu S, Wan X (2021) The essential roles of sugar metabolism for pollen development and male fertility in plants. Crop J 9(6):1223–1236
Wu Y et al (2015) Defective pollen wall contributes to male sterility in the male sterile line 1355A of cotton. Sci Rep 5(1):9608
Liu J et al (2014) Proteomic analysis of anthers from wild-type and photosensitive genetic male sterile mutant cotton (Gossypium hirsutum L). BMC Plant Biol 14:1–16
Ma J et al (2013) Selection and characterization of a Novel photoperiod-sensitive male sterile line in Upland Cotton. J Integr Plant Biol 55(7):608–618
Ahmad M et al (2021) Adaptation strategies to improve the resistance of oilseed crops to heat stress under a changing climate: an overview. Front Plant Sci 12:767150
Zhang R et al (2022) Rapid identification of pollen-and anther-specific genes in response to high-temperature stress based on transcriptome profiling analysis in cotton. Int J Mol Sci 23(6):3378
Khan AH et al (2020) High day and night temperatures distinctively disrupt fatty acid and jasmonic acid metabolism, inducing male sterility in cotton. J Exp Bot 71(19):6128–6141
Zinn KE, Tunc-Ozdemir M, Harper JF (2010) Temperature stress and plant sexual reproduction: uncovering the weakest links. J Exp Bot 61(7):1959–1968
Meyer VG (1969) Some effects of genes, cytoplasm, and Environment on male sterility of cotton (Gossypium) 1. Crop Sci 9(2):237–242
Meyer VG, Meyer JR (1965) Cytoplasmically controlled male sterility in cotton
Meyer VG (1973) A study of reciprocal hybrids between upland cotton (Gossypium hirsutum L.) and experimental lines with cytoplasms from seven other species 1. Crop Sci 13(4):439–444
Meyer VG (1975) Male sterility from Gossypium harknessii
Zhang J, Stewart JM (2001) Inheritance and genetic relationships of the D8 and D2-2 restorer genes for cotton cytoplasmic male sterility. Crop Sci 41(2):289–294
Stewart J (1992) A new cytoplasmic male sterile and restorer for cotton
Yin J, Guo W, Yang L, Liu L, Zhang T (2006) Physical mapping of the Rf1 fertility-restoring gene to a 100 kb region in cotton. Theor Appl Genet 112:1318–1325
Wang F, Stewart JM, Zhang J (2007) Molecular markers linked to the Rf2 fertility restorer gene in cotton. Genome 50(9):818–824
Feng J et al (2021) Physical mapping and InDel marker development for the restorer gene Rf2 in cytoplasmic male sterile CMS-D8 cotton. BMC Genomics 22:1–12
Feng Y et al (2015) Multiple loci not only Rf3 involved in the restoration ability of pollen fertility, anther exertion and pollen shedding to S type cytoplasmic male sterile in maize. Theor Appl Genet 128:2341–2350
Wang Z, Gerstein M, Snyder M (2009) RNA-Seq: a revolutionary tool for transcriptomics. Nat Rev Genet 10(1):57–63
Sheoran IS, Ross AR, Olson DJ, Sawhney VK (2009) Differential expression of proteins in the wild type and 7B-1 male-sterile mutant anthers of tomato (Solanum lycopersicum): a proteomic analysis. J Proteom 71(6):624–636
Sheoran IS, Sawhney VK (2010) Proteome analysis of the normal and Ogura (ogu) CMS anthers of Brassica napus to identify proteins associated with male sterility. Botany 88(3):217–230
Yue J, Ren Y, Wu S, Zhang X, Wang H, Tang C (2014) Differential proteomic studies of the genic male-sterile line and fertile line anthers of upland cotton (Gossypium hirsutum L). Genes Genomics 36:415–426
Hamid R, Tomar RS, Marashi H, Shafaroudi SM, Golakiya BA, Mohsenpour M (2018) Transcriptome profiling and cataloging differential gene expression in floral buds of fertile and sterile lines of cotton (Gossypium hirsutum L). Gene 660:80–91
Cheng C et al (2023) Identification of fertility restoration candidate genes from a restorer line R186 for Gossypium Harknessii cytoplasmic male sterile cotton. BMC Plant Biol 23(1):175
Zhang M et al (2020) Deficiencies in the formation and regulation of anther cuticle and tryphine contribute to male sterility in cotton PGMS line. BMC Genomics 21:1–18
Liu F et al (Dec. 2019) GhFAD2–3 is required for anther development in Gossypium hirsutum. BMC Plant Biol 19(1):393. https://doi.org/10.1186/s12870-019-2010-9
Chen L et al (May 2024) GhTKPR1_8 functions to inhibit anther dehiscence and reduce pollen viability in cotton. Physiol Plant 176(3):e14331. https://doi.org/10.1111/ppl.14331
Gong J et al (2024) Gh4CL20/20A involved in flavonoid biosynthesis is essential for male fertility in cotton (Gossypium hirsutum L). Plant Physiol Biochem, p. 108484
Zhang M et al (2021) GhGPAT12/25 are essential for the formation of anther cuticle and pollen exine in cotton (Gossypium hirsutum L). Front Plant Sci 12:667739
Dharmalingam Raja DR et al (2018) Identification of molecular markers associated with genic male sterility in tetraploid cotton (Gossypium hirsutum L.) through bulk segregant analysis using a cotton SNP 63K array., Accessed: Mar. 13, 2024. [Online]. Available: https://www.cabidigitallibrary.org/doi/full/10.5555/20193060278
Guo W, Zhang T, Pan J, Kohel RJ (1998) Identification of RAPD marker linked with fertility-restoring gene of cytoplasmic male sterile lines in upland cotton. Chin Sci Bull 43:52–54
Lan T-H, Cook CG, Paterson AH (1999) Identification of a RAPD marker linked to a male fertility restoration gene in cotton (Gossypium hirsutum L). J Agric Genomics 4:1–5
Liu L, Guo W, Zhu X, Zhang T (2003) Inheritance and fine mapping of fertility restoration for cytoplasmic male sterility in Gossypium hirsutum L. Theor Appl Genet 106:461–469
Zhang J, Stewart JM (2004) Identification of molecular markers linked to the fertility restorer genes for CMS-D8 in Cotton. Crop Sci 44(4):1209–1217
Feng J et al (2020) Development and utilization of an InDel marker linked to the fertility restorer genes of CMS-D8 and CMS-D2 in cotton. Mol Biol Rep 47:1275–1282
Wu J et al (Nov. 2017) Development of InDel markers for the restorer gene Rf1 and assessment of their utility for marker-assisted selection in cotton. Euphytica 213(11):251. https://doi.org/10.1007/s10681-017-2043-9
Toppo RR, Katageri IS (2016) Molecular markers for fertility restoration in cotton CGMS system. J Farm Sci 29(3):307–309
Bharati M, Khadi B, Katageri I, Vamadevaiah H, Fakrudin B (2010) Molecular characterization of genetic male sterile genotypes in cotton (Gossypium spp). Indian J Genet Plant Breed 70(01):94–96
Andersen JR, Lübberstedt T (2003) Functional markers in plants. Trends Plant Sci 8(11):554–560
Varshney RK, Graner A, Sorrells ME (2005) Genomics-assisted breeding for crop improvement. Trends Plant Sci 10(12):621–630
Salgotra RK, Gupta BB, Stewart CN (2014) From genomics to functional markers in the era of next-generation sequencing, Biotechnol Lett, vol. 36, no. 3, pp. 417–426, Mar. https://doi.org/10.1007/s10529-013-1377-1
Ingvardsen CR, Schejbel B, Lübberstedt T (2008) Functional Markers in Resistance Breeding, in Progress in Botany, vol. 69, U. Lüttge, W. Beyschlag, and J. Murata, Eds., in Progress in Botany, vol. 69., Berlin, Heidelberg: Springer Berlin Heidelberg, pp. 61–87. https://doi.org/10.1007/978-3-540-72954-9_3
Salgotra RK, Stewart CN Jr (2020) Functional markers for precision plant breeding. Int J Mol Sci 21(13):4792
Lin Z, Wang Y, Zhang X, Zhang J (Oct. 2012) Functional markers for cellulose synthase and their comparison to SSRs in cotton. Plant Mol Biol Rep 30(5):1270–1275. https://doi.org/10.1007/s11105-012-0432-8
Pranathi K et al (2016) Comparative analysis of sequences of mitochondrial genomes of wild abortive male sterile (WA-CMS) and male fertile lines of rice, development of functional markers for WA-CMS trait and their use in assessment of genetic purity of seeds of WA-CMS lines. Mol Breeding 36:1–12
Chen C et al (2013) A co-dominant marker BoE332 applied to marker-assisted selection of homozygous male-sterile plants in cabbage (Brassica oleracea var. capitata L). J Integr Agric 12(4):596–602
Abuyusuf M, Nath UK, Kim H-T, Islam MR, Park J-I, Nou I-S (2019) Molecular markers based on sequence variation in BoFLC1. C9 for characterizing early-and late-flowering cabbage genotypes. BMC Genet 20:1–11
Naresh P, Lin S, Wang Y, Kumar S (2018) Molecular markers associated to two non-allelic genic male sterility genes in peppers (Capsicum annuum L). Front Plant Sci 9:354475
Kim S-C et al (2014) Development of a SCAR marker for sex identification in asparagus. Korean J Plant Resour 27(3):236–241
Li J et al (2012) Map-based cloning of a recessive genic male sterility locus in Brassica napus L. and development of its functional marker. Theor Appl Genet 125:223–234
Andeden E et al (2011) Distribution of vernalization and photoperiod genes (Vrn-A1, Vrn-B1, Vrn-D1, Vrn-B3, Ppd-D1) in Turkish bread wheat cultivars and landraces. Cereal Res Commun 39(3):352–364
Dunford RP et al (2002) Comparative mapping of the barley Ppd-H1 photoperiod response gene region, which lies close to a junction between two rice linkage segments. Genetics 161(2):825–834
Qi Y, Wang L, Gui J, Zhang L, Liu Q, Wang J (2017) Development and validation of a functional co-dominant SNP marker for the photoperiod thermo-sensitive genic male sterility pms3 (p/tms12-1) gene in rice, Breeding Science, vol. 67, no. 5, pp. 535–539
Hecht V et al (2005) Conservation of Arabidopsis flowering genes in model legumes. Plant Physiol 137(4):1420–1434
Funding
No funding was received.
Author information
Authors and Affiliations
Contributions
All authors contributed to the paper as follows; idea of the article: A. Subramanian; literature search: V. Deepa Dharsini, N. Premalatha, N. Manikanda Boopathi, M. Djanaguiraman, V. P. Santhanakrishnan; draft manuscript preparation: V. Deepa Dharsini, A. Subramanian; critical revision of the work: A. Subramanian. All authors reviewed and approved the final version of the manuscript.
Corresponding author
Ethics declarations
Ethical approval
Not applicable.
Consent to participate
Not applicable.
Consent to publish
Not applicable.
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Dharsini, V.D., Subramanian, A., Premalatha, N. et al. Fertile grounds: exploring male sterility in cotton and its marker development. Mol Biol Rep 51, 961 (2024). https://doi.org/10.1007/s11033-024-09893-9
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s11033-024-09893-9