Abstract
The cytoplasmic male sterility (CMS) system is a useful tool for commercial hybrid cotton seed production. Two main CMS systems, CMS-D8 and CMS-D2, have been recognized with Rf2 and Rf1 as the restorer genes, respectively. The development of molecular markers tightly linked with restorer genes can facilitate the breeding of restorer lines. In this study, the InDel-1892 marker was developed to distinguish Rf2 and Rf1 simultaneously. Sequence alignment implied that CMS-D8-Rf2 has a 32 bp insertion and that CMS-D2-Rf1 has a 186 bp insertion at the InDel-1892 locus. The codominant marker was co-segregated with Rf1 and Rf2. Hence, this marker can be used for tracing Rf1 and Rf2 simultaneously and identifying the allele status at the restorer gene locus. The results of this study will facilitate efficient marker-assisted selection for restorer lines and hybrids of CMS systems.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Cotton (Gossypium hirsutum L.) is one of the most important economic crops and is a key source of fiber and oil worldwide. Hybrid breeding is an important way to improve cotton yield. At present, there are two ways to produce cotton hybrids, i.e. artificial emasculation and pollination (AEP) and cytoplasmic male sterility (CMS) systems. However, it is difficult to ensure the purity of hybrid seeds with emasculation by hand. CMS is a maternally inherited trait that can produce nonfunctional pollen grains or sterile male gametes in flowering plants [1]. The CMS system is an efficient and economical way of producing hybrids without the need for anther removal and has been widely utilized for commercial F1 hybrid seed production in many crops [2,3,4]. In upland cotton, the CMS-D2 line and its restorer line have been developed by introducing the cytoplasmic and nuclear gene Rf1 from wild diploid cotton G. harknessii Brandegee (D2-2 genome) into cultivated tetraploid upland cotton [5, 6], whereas the CMS-D8 line and its restorer line have an exotic cytoplasmic and nuclear gene Rf2 from the wild diploid species G. trilobum (DC.) Skovst (D8 genome) [7, 8]. According to previous reports, genetic mutations in the mitochondrial gene atpA are associated with CMS in cotton [9, 10]. Previous studies showed that Rf1 gene loci and Rf2 gene loci are not allelic, but these two genes are tightly linked at a genetic distance of 0.93 cM on chromosome D05, and Rf1 could restore the fertility of both CMS-D2 and CMS-D8 lines, while Rf2 could only restore the fertility of CMS-D8 sterile line [11]. The Rf1 gene functions in the sporophyte, while the Rf2 gene has a gametophytic restoration system [8].
Several studies have developed many markers linked with the Rf1 gene, including random amplification of polymorphic DNA (RAPD) markers (OPV_15, OPJ_11, R6952 and R6861) [12, 13], cleaved amplified polymorphic sequence (CAPS) markers (CAPS-R) [14], sequence tagged site (STS) markers (UBC1471400, UBC607500 and UBC679700) [15], and simple sequence repeat (SSR) markers [16]. Among them, the SSR marker NAU211 was identified as being near to the Rf1 gene with a genetic distance of 0.163 cM [17]; another SSR marker NAU4047 has been verified to be linked with the Rf1 gene at a genetic distance within 1 cM [18]. Our previous study found that the nearest SSR markers on both sides to Rf1 are BNL3535 and NAU3652 with genetic distances of 0.049 cM and 0.078 cM, respectively [14]. In comparison, only a few molecular markers closely linked to the Rf2 gene have been reported in cotton. The STS marker UBC188, linked with Rf2, was developed with a genetic distance of less than 2.9 cM [19]. Three RAPD markers, the CAPS-UBC722-HpaII marker, two AFLP markers, and the SSR CIR179-250 marker, were developed, but these markers are not tightly linked with Rf2 [20]. In addition, molecular markers can also be used to genotype different cytoplasmic types. In cotton, the atpA sequence characterized amplified region (SCAR) [9] and SSR160 markers [10] have been developed to identify sterile and fertile cytoplasmic types in individual plants.
Marker-assisted selection (MAS) is one of the most effective methods for accurate breeding, which can use molecular markers to track functional genes associated with excellent agronomic traits [21, 22]. In addition, MAS helps breeders obtain plants with a recurrent parent genome that only requires two or three generations in the breeding process [23]. However, the application of the reported markers shows limited efficiency in MAS breeding because of inappropriate genetic distances and tedious application procedures, which require extensive time and labor. Insertion-deletion (InDel) markers are highly stable and accurate and occur frequently based on the InDel of DNA fragments at the same genomic locus [24]. InDel markers have proven convenient and efficient in molecular breeding because they are easy to amplify with polymerase chain reaction (PCR) and can be detected through agarose gel electrophoresis. More recently, many studies have attempted to develop InDel markers with different approaches. InDel markers have been successfully used in many domains including marker-assisted breeding [25, 26], genetic fine mapping, map-based cloning [27,28,29,30,31], and analyses of germplasm resources [32, 33]. The release of the upland cotton genomic sequence [34,35,36] has made it possible to explore InDel markers in upland cotton. Thus, InDel makers highly linked with target genes could be developed at a low cost to facilitate MAS in cotton breeding. Currently, four InDel markers tightly linked with the Rf1 gene have been successfully developed in cotton [37].
Unfortunately, no suitable PCR-based InDel markers were identified to distinguish both restorer genes (Rf1 and Rf2). In the present study, we developed a PCR-based codominant marker (InDel-1892) that could distinguish both Rf1 and Rf2 in cotton. Furthermore, sequence analysis was conducted at the InDel-1892 marker locus, and sequence variation was validated between the CMS-D2 and CMS-D8 systems. Finally, we used the InDel marker for restorer lines and commercial hybrid cotton in accurate MAS breeding.
Materials and methods
Plant materials and DNA extraction
In our study, three lines (i.e., male sterile, maintainer, and restorer lines) of two sets of cotton CMS systems, including CMS-D8-Rf2 BC1F1 and CMS-D2-Rf1 F2 populations, were used, which were provided by the Institute of Cotton Research (ICR), Chinese Academy of Agricultural Science, Anyang, Henan, China. Plants were grown at the Cotton Research Farm at the ICR.
Young leaves from each plant were collected and immediately frozen in liquid nitrogen. The leaf samples were stored at -80 °C until use. The total genomic DNA of each plant was isolated from cotton leaves using a modified cetyltrimethylammonium bromide (CTAB) method [38]. The concentration of DNA was calculated using a NanoDrop 2000C spectrophotometer (Thermo Scientific, Wilmington, DE, USA), whereas the quality of DNA was estimated by 1.2% agarose gel electrophoresis. The DNA extraction and fertility survey of the CMS-D2-Rf1 BC4F2 population had been completed by our team previously [14].
Development of InDel markers for restorer genes
InDel markers linked to fertility restorer genes were selected based on our earlier report [37]. Here, the InDel markers that are closely linked to the CMS-D2 restorer gene (Rf1) were used to screen the male sterile, maintainer and restorer lines of CMS-D8 system, including InDel-3434, InDel-7525, InDel-9356, InDel-7138, and InDel-1892 (Table 1, InDel-3434, InDel-7525 and InDel-9356 have been reported [37]). These primers were designed using Oligo7 software [39] and synthesized commercially (TSINGKE Biological Technology, Zhengzhou, China). A total of 20 µL of PCR mixture was prepared that contained 1× reaction buffer, 2.0 mM MgCl2, 0.2 mM dNTPs, 0.5 mM each primer, 1 U Taq DNA polymerase (Takara, Japan), and 50 ng DNA template. The PCR amplification conditions were as follows: 35 cycles of denaturation at 94 °C for 30 s, annealing at 58 °C for 30 s, and extension at 72 °C for 30 s. Then, the reaction was held at 4 °C. The genetic analyses were conducted in the BC1F1 population (500 progenies) of CMS-D8 and the F2 population (500 progenies) of CMS-D2 that were produced in our previous study.
Analysis of gene sequences of the InDel-1892 marker locus
To determine the sequence difference between the two restorer lines on the InDel-1892 marker locus, target segments were cloned. In the cloning process, we used the TaKaRa MiniBEST DNA Fragment Purification Kit Ver. 4.0, pEASY-T5 Zero Cloning Kit (TRANSGEN BIOTECH, Beijing, China) and DH5α Chemically Competent Cell kit (TRANSGEN BIOTECH, Beijing, China). In the end, four centrifuge tubes of DH5α containing the target fragment were sent to Sunya Company (www.biosunya.com) for Sanger sequencing. Multiple sequence alignments were analyzed by Mega7 and Genedoc (http://www.nrbsc.org/gfx/genedoc).
Marker-assisted selection of restorer lines
Restorer lines were used as the male parent to cross with the female parent line, which has excellent agronomic traits no containing restorer genes. Subsequently, four successive backcrosses were performed using that female parent as the recurrent parent. During the backcrossing process, marker InDel-1892 was used in each generation for marker-assisted selection of those plants carrying the restorer gene with improved agronomic traits for further backcrossing. After the fourth backcross generation, we obtained the BC4F1 population of CMS-D8 and the BC4F2 population of CMS-D2. During anthesis, visual fertility investigations were performed three times on each individual of these two populations. Specifically, fertile flowers exhibit plenty of pollen grains, while sterile flowers have no visible pollen grains. In addition, these two populations were all screened with the InDel-1892 marker.
Identification of the CMS hybrids
The InDel-1892 marker is used for genotyping the CMS-D8 hybrid, CMS-D2 hybrids (CCRI83, CCRI99), and conventional commercial hybrid varieties (CCRI29, CCRI52, CCRI54). The atpA SCAR marker we identified previously [9] is used to distinguish the CMS cytoplasm from other types of cytoplasm.
Results
Identification of the InDel-1892 marker and validation of InDel markers
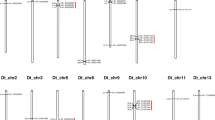
In this study, five previously developed InDel markers (Table 1), closely linked to the Rf1 gene, were used to screen the male sterile, maintainer and restorer lines of CMS-D8 system, including InDel-3434, InDel-7138, InDel-7525, InDel-9356, and InDel-1892. Interestingly, among the five InDel markers, only InDel-7138 and InDel-1892 markers were also polymorphic among the corresponding male sterile, maintainer, and restorer lines of CMS-D8 system (Fig. 1a). Obviously, for these two InDel markers, there was no difference in the size of the PCR product between the male sterile line and the maintainer line, whereas the restorer line showed another larger band (Fig. 1a). However, our further analysis of the InDel-7138 marker based on PCR amplification and agarose gel electrophoresis found that the InDel-7138 marker could not simultaneously distinguish these two fertility restoration loci (Rf1 and Rf2) in cotton (Supplementary Fig. S1). Therefore, we mainly focus on the analysis of another marker InDel-1892 and its application in two cotton CMS systems. The results of genetic analyses revealed that the InDel-1892 marker co-segregated with both Rf2 and Rf1 genes based on the marker-trait co-segregation analysis. Moreover, we also provided a rough physical map to show the distribution of the InDel markers used in this study on the D05 chromosome and their relationship with the Rf1 and Rf2 loci (Fig. 1b).
a Screening and identification of polymorphic InDel markers based on PCR amplification in cotton CMS-D8 system. M: DL2000 DNA marker; A: CMS-D8-sterile line; B: CMS-D8-maintainer line; R: CMS-D8-restorer line. b A ough physical map showing the distribution of InDel markers on the D05 chromosome and their relationship to the Rf1 and Rf2 loci. The green and red bar segments highlight the locations of the Rf1 and Rf2 genes, respectively. The purple and bold InDel-1892 is a newly developed InDel marker co-isolated with both Rf1 and Rf2 genes. (Color figure online)
Sequence analysis of the InDel-1892 marker locus
The sequences of the target regions amplified by the InDel-1892 marker among the two cotton CMS systems were subsequently selected for multiple sequence alignment with the cotton TM-1 reference genome [36]. Compared with the genomic sequence of cotton TM-1, the male sterile lines and maintainer lines of the two cotton CMS systems were found to have the same sequence length at the target region (376 bp), except for a few base differences (Fig. 2). Interestingly, the sequences of the restorer lines of CMS-D8 and CMS-D2 showed different lengths of insertions and/or mutations at the target region, including a 32 bp insertion in the restorer line of CMS-D8 (408 bp) and a 186 bp insertion in the restorer line of CMS-D2 (562 bp) (Fig. 2).
Marker-assisted selection of restorer lines of two cotton CMS systems
Considering the different sequence sizes at InDel-1892 marker locus among the three lines (i.e., male sterile, maintainer, and restorer lines) of two sets of cotton CMS systems (Fig. 2), the co-dominant marker InDel-1892 can be used for simultaneous marker-assisted selection breeding of two types of CMS restorer lines with improved agronomic traits. The BC1F1 individuals that produced two PCR products, that is, a 376 bp product and a 408 bp product for CMS-D8 and a 376 bp product and a 562 bp product for CMS-D2 (Fig. 3), were considered heterozygous for the restorer genes Rf2 and Rf1, respectively, and were chosen for successive backcrossing in each generation.
Subsequently, we randomly selected 96 individual plants from the BC4F1 population of CMS-D8 to analyze the genotype with the InDel-1892 marker. Based on visible fertility investigations, individual phenotypes are distinguished into fertile and sterile types. Clearly, the results of agarose gel electrophoresis showed two different banding patterns. The sterile plants that produced a single small PCR product were considered homozygous and lacked the restorer gene allele [S (rf2rf2)], whereas the fertile plants that produced two fragments were considered heterozygous at the restorer gene locus [S (Rf2rf2)]. Furthermore, the segregating ratio followed a 1 (Rf2rf2):1 (rf2rf2) (50 Rf2rf2: 46 rf2rf2, χ20.05 = 0.1667 < 3.841) ratio, and this result was consistent with the results of fertility investigations in the field (see Fig. 4a as an example). Similarly, 96 individual plants were taken from the BC4F2 population of CMS-D2 at random and analyzed by PCR amplification and agarose gel electrophoresis using the InDel-1892 marker. The segregation ratio of three different banding types followed a 1 (Rf1Rf1):2 (Rf1rf1):1 (rf1rf1) (25 Rf1Rf1: 47 Rf1rf1: 24 rf1rf1, χ20.05 = 0.9692 < 5.991) ratio (see Fig. 4b as an example), which was in accordance with the results of field fertility surveys in our previous study [14]. Taken together, we propose that the newly developed InDel marker InDel-1892 can be effectively used for simultaneous marker-assisted selection breeding of the restorer lines of two cotton CMS systems in the future.
The utility of the InDel-1892 marker for CMS hybrid identification
InDel-1892 was further used to distinguish the CMS hybrids from other conventional commercial hybrid varieties. The hybrids of the CMS-D8 system have two bands at 376 bp and 408 bp, and the hybrids of the CMS-D2 system, CCRI 83 and CCRI 99, have two bands at 376 bp and 562 bp, respectively. However, the conventional commercial hybrid varieties produced by artificial emasculation and pollination, CCRI 29, CCRI 52 and CCRI 54, have only one band at 376 bp and lack the two restorer genes (Fig. 5a). The InDel-1892 marker could not only identify whether the plants contained a restorer gene but also distinguish the Rf1 or Rf2 restorer genes at the same time. Next, the plants were scanned with an atpA SCAR marker [9], as the hybrids of the CMS-D8 system and CMS-D2 system (CCRI 83 and CCRI 99) have sterile cytoplasm, and a 611 bp fragment was amplified (Fig. 5b). Therefore, the CMS hybrids with heterozygous restorer gene sites and sterile cytoplasm were differentiated by the genotyping of restorer genes and the identification of cytoplasm type.
Discussion
Hybrid breeding could greatly increase the productivity of major crops by utilizing the phenomenon of heterosis. CMS systems serve as an important pollination control tool in hybrid cotton seed production for saving labor and time compared with artificial emasculation and pollination (AEP). The efficiency of CMS systems largely depends on restorer lines [40]. Maintaining the genetic purity of the parental lines (male sterile lines, maintainer lines, and restorer lines) and the F1 seeds is important because it directly affects the yield of “three-line” hybrid cotton. Many studies with different types of molecular markers have been performed to map restorer genes in cotton [14,15,16, 41]. However, these markers are difficult to apply to marker-assisted selection (MAS) breeding and cannot ensure accurate breeding. The percent heritability of the markers and their lower cost are the main factors supporting the utilization of these markers in cotton breeding programs [42]. For example, simple sequence repeat (SSR) markers provide low polymorphic levels and require complicated analytical procedures. High-throughput analysis of single nucleotide polymorphism (SNP) markers requires costly procedures [43], e.g., genotyping-by-sequencing, because cotton has a relatively large genome. Furthermore, there is still an urgent need for closer, more versatile, and codominant markers linked to Rf1 and Rf2.
Recently, the release of the cotton genome sequence [44, 34,35,36, 45] has promoted the discovery of more polymorphisms within cotton cultivars. Highly reproducible and abundant InDel markers were exploited based on the BLAST analysis of cotton genomic sequences. High-throughput sequencing technologies have reduced the cost of developing InDel markers in recent years [46]. In addition, InDel markers, especially those with large insertions/deletions, are more stable and can be easily amplified and analyzed on agarose gels compared with other kinds of molecular markers. Therefore, InDel markers present a feasible option for efficient and accurate MAS breeding and tracking restorer genes in CMS cotton. Accurate MAS breeding can select restorer genes using closely linked markers at the genome level and shorten the breeding period compared with the conventional method of “grow-out testing” [47, 48].
The newly developed InDel-1892 marker is the first InDel marker that can be used for tracing Rf1 and Rf2 simultaneously and identifying the allele status at the restorer gene locus in two sets of cotton CMS systems. Therefore, the application of InDel-1892 marker can improve the accuracy of MAS breeding for cotton restorer lines. Furthermore, the InDel-1892 marker is more reliable, efficient, and economical compared with SSR, CAPS and STS markers in tracing Rf1 and Rf2 genes. The developed InDel-1892 marker would empower effective selection in CMS system breeding programs and can be used (I) to genotype each plant from any CMS segregation population at any stage of cotton growth, if necessary; (II) to identify the selfing progenies of heterozygous plants as an alternative to test crossing, which requires at least two years and additional labor for field work; (III) to identify CMS hybrids in combination with cytoplasmic genotyping; and (IV) as a foreground marker to create new restorer lines containing Rf1 or Rf2. Fine-mapping at the molecular and physical levels using InDel-1892 will facilitate the isolation of the two restorer genes. Our study provides valuable information for MAS and promotes the application of a simplified hybrid seed production technique in cotton.
References
Schnable PS, Wise RP (1998) The molecular basis of cytoplasmic male sterility and fertility restoration. Trends Plant Sci 3(5):175–180
Sun CQ, Huang ZZ, Wang YL, Chen FD, Teng NJ, Fang WM, Liu ZL (2011) Overcoming pre-fertilization barriers in the wide cross between Chrysanthemum grandiflorum (Ramat.) Kitamura and C. nankingense (Nakai) Tzvel. by using special pollination techniques. Euphytica 178(2):195–202
Bohra A, Jha UC, Adhimoolam P, Bisht D, Singh NPJPCR (2016) Cytoplasmic male sterility (CMS) in hybrid breeding in field crops. Plant Cell Rep 35(5):967–993
Budar F, Pelletier G (2001) Male sterility in plants: occurrence, determinism, significance and use. C R Acad Sci III 324(6):543–550. doi:https://doi.org/10.1016/s0764-4469(01)01324-5
Meyer VG (1975) Male sterility from Gossypium harknessii J Hered.https://doi.org/10.1186/s42397-018-0013-y
Weaver DB, Weaver JB (1977) Inheritance of pollen fertility restoration in cytoplasmic male-sterile upland cotton 1. Crop Sci 17(4):497–499
Zhang JF, Stewart JMD (2001) CMS-D8 restoration in cotton is conditioned by one dominant gene. Crop Sci 41(2):283–288
Stewart JA (1992) New cytoplasmic male sterile and restorer for cotton. In: Proc. Beltwide Cotton Conf. National Cotton Council, Memphis
Wu J, Gong Y, Cui M, Qi T, Guo L, Zhang J, Xing CJE (2011) Molecular characterization of cytoplasmic male sterility conditioned by Gossypium harknessii cytoplasm (CMS-D2) in upland cotton. Euphytica 181(1):17–29
Zhang X, Meng Z, Zhou T, Sun G, Shi J, Yu Y, Zhang R, Guo S (2012) Mitochondrial SCAR and SSR markers for distinguishing cytoplasmic male sterile lines from their isogenic maintainer lines in cotton. Plant Breed 131(4):563–570. https://doi.org/10.1111/j.1439-0523.2012.01971.x
Zhang JF, Stewart JM (2001) Inheritance and genetic relationships of the D8 and D2–2 restorer genes for cotton cytoplasmic male sterility. Crop Sci.https://doi.org/10.2135/cropsci2001.412289x
Guo W, Zhang T, Pan J, Kohel RJ (1998) Identification of RAPD marker linked with fertility-restoring gene of cytoplasmic male sterile lines in upland cotton. Chin Sci Bull 43(1):52–54
Lan TH, Cook CG, Paterson AH (1999) Identification of a RAPD marker linked to a male fertility restoration gene in cotton (Gossypium hirsutum L.). J Agric Genomics 4:1–5
Wu J, Cao X, Guo L, Qi T, Wang H, Tang H, Zhang J, Xing C (2014) Development of a candidate gene marker for Rf 1 based on a PPR gene in cytoplasmic male sterile CMS-D2 Upland cotton. Mol Breed 34(1):231–240
Feng C, Mc D, Stewart J, Zhang J (2005) STS markers linked to the Rf1 fertility restorer gene of cotton. Ther Appl Genet.https://doi.org/10.1007/s00122-004-1817-3
Liu L, Guo W, Zhu X, Zhang T (2003) Inheritance and fine mapping of fertility restoration for cytoplasmic male sterility in Gossypium hirsutum L. Theor Appl Genet 106(3):461–469
Yang L (2009) Map-based cloning of fertility restoring gene of CMS and analysis of PPR gene family in cotton. Ph.D. Dissertation, Nanjing Agric Univ
Yin J, Guo W, Yang L, Liu L, Zhang T (2006) Physical mapping of the Rf1 fertility-restoring gene to a 100 kb region in cotton. Theor Appl Genet 112(7):1318–1325. doi:https://doi.org/10.1007/s00122-006-0234-1
Zhang J, Stewart JM (2004) Identification of molecular markers linked to the fertility restorer genes for CMS-D8 in cotton. Crop Sci.https://doi.org/10.2135/cropsci2004.1209
Wang F, Stewart JM, Zhang J (2007) Molecular markers linked to the Rf2 fertility restorer gene in cotton. Genome 50(9):818–824. doi:https://doi.org/10.1139/g07-061
Collard BC, Mackill DJ (2007) Marker-assisted selection: an approach for precision plant breeding in the twenty-first century. Philos Trans R Soc Lond B Biol Sci 363(1491):557–572
Zhou YL, Uzokwe VNE, Zhang CH, Cheng LR, Wang L, Chen K, Gao XQ, Sun Y, Chen JJ, Zhu LH (2011) Improvement of bacterial blight resistance of hybrid rice in China using the gene derived from wild rice. Crop Prot 30(6):637–644
Watson A, Ghosh S, Williams MJ, Cuddy WS, Simmonds J, Rey M, Hatta MAM, Hinchliffe A, Steed A, Reynolds DJNP (2017) Speed breeding is a powerful tool to accelerate crop research and breeding. Nat Plans 4(1):23–29
Weber JL, David D, Heil J, Fan Y, Zhao C, Marth G (2002) Human diallelic insertion/deletion polymorphisms. Am J Hum Genet 71(4):854–862
Hayashi K, Yoshida H, Ashikawa (2006) Development of PCR-based allele-specific and InDel marker sets for nine rice blast resistance genes. Tagtheor Appl Geneticstheor Angewandte Genetik 113(2):251–260
Zhao J, Zhang S, Dong J, Yang T, Mao X, Liu Q, Wang X, Liu B (2017) A novel functional gene associated with cold tolerance at the seedling stage in rice. Plant Biotechnol J 15(9):1141
Ye XL, Hu FY, Ren J, Huang SN, Liu WJ, Feng H, Liu ZY (2016) Fine mapping and candidate gene analysis of Brtri1 agene controlling trichome development in Chinese cabbage (Brassica rapa L. ssp pekinensis). Genet Mol Res. doi:https://doi.org/10.4238/gmr15048924
Srivastava R, Singh M, Bajaj D, Parida SK (2016) A high-resolution InDel (insertion-deletion) markers-anchored consensus genetic map identifies major QTLs Governing Pod Number and Seed Yield in Chickpea. Front Plant Sci 7:1362. https://doi.org/10.3389/fpls.2016.01362
Singh VK, Khan AW, Saxena RK, Sinha P, Kale SM, Parupalli S, Kumar V, Chitikineni A, Suryanarayana V, Sameer Kumar CV (2017) Indel-seq: a fast forward genetics approach for identification of trait associated putative candidate genomic regions and its application in pigeonpea (Cajanus cajan). Plant Biotechnol J 15(7):906–914
Zhang H, Wu J, Dai Z, Qin M, Hao L, Ren Y, Li Q, Zhang L (2016) Allelism analysis of BrRfp locus in different restorer lines and map-based cloning of a fertility restorer gene, BrRfp1, for pol CMS in Chinese cabbage (Brassica rapa L.). Tagtheor Appl Genet Angewandte Genetik 130(3):539–547
Liu XP, Gao BZ, Han FQ, Fang ZY, Yang LM, Zhuang M, Lv HH, Liu YM, Li ZS, Cai CC (2017) Genetics and fine mapping of a purple leaf gene, BoPr, in ornamental kale (Brassica oleracea L. var. acephala). Bmc Genomics 18(1):230
Lv H, Wang Q, Han F, Liu X, Fang Z, Yang L, Zhuang M, Liu Y, Li Z, Zhang Y (2017) Genome-wide indel/SSR scanning reveals significant loci associated with excellent agronomic traits of a cabbage (Brassica oleracea) elite parental line '01–20'. Sci Rep 7:41696. doi:https://doi.org/10.1038/srep41696
Lu BR, Cai X, Xin J (2009) Efficient indica and japonica rice identification based on the InDel molecular method: Its implication in rice breeding and evolutionary research. Prog Nat Sci 19(10):1241–1252
Li F, Fan G, Lu C, Xiao G, Zou C, Kohel RJ, Ma Z, Shang H, Ma X, Wu J (2015) Genome sequence of cultivated Upland cotton (Gossypium hirsutum TM-1) provides insights into genome evolution. Nat Biotechnol 33(5):524–530
Wang M, Tu L, Yuan D, Zhu, Shen C, Li J, Liu F, Pei L, Wang P, Zhao G, Ye Z, Huang H, Yan F, Ma Y, Zhang L, Liu M, You J, Yang Y, Liu Z, Huang F, Li B, Qiu P, Zhang Q, Zhu L, Jin S, Yang X, Min L, Li G, Chen LL, Zheng H, Lindsey K, Lin Z, Udall JA, Zhang X (2019) Reference genome sequences of two cultivated allotetraploid cottons, Gossypium hirsutum and Gossypium barbadense Nat Genet 51(2):224–229. https://doi.org/10.1038/s41588-018-0282-x
Zhang T, Hu Y, Jiang W, Fang L, Guan X, Chen J, Zhang J, Saski CA, Scheffler BE, Stelly DM, Hulse-Kemp AM, Wan Q, Liu B, Liu C, Wang S, Pan M, Wang Y, Wang D, Ye W, Chang L, Zhang W, Song Q, Kirkbride RC, Chen X, Dennis E, Llewellyn DJ, Peterson DG, Thaxton P, Jones DC, Wang Q, Xu X, Zhang H, Wu H, Zhou L, Mei G, Chen S, Tian Y, Xiang D, Li X, Ding J, Zuo Q, Tao L, Liu Y, Li J, Lin Y, Hui Y, Cao Z, Cai C, Zhu X, Jiang Z, Zhou B, Guo W, Li R, Chen ZJ (2015) Sequencing of allotetraploid cotton (Gossypium hirsutum L. acc. TM-1) provides a resource for fiber improvement. Nat Biotechnol 33(5):531–537. https://doi.org/10.1038/nbt.3207
Wu J, Meng Z, Zhang X, Guo L, Qi T, Wang H, Tang H, Zhang J, Xing C (2017) Development of InDel markers for the restorer gene Rf1 and assessment of their utility for marker-assisted selection in cotton. Euphytica 213(11):251
Zhang JF, Stewart JM (2000) Economical and rapid method for extracting cotton genomic DNA. J Cotton Sci 4(3):193–201
Rychlik W (2007) OLIGO 7 primer analysis software.Methods in molecular biology. PCR Primer Des 402:35–60. https://doi.org/10.1007/978-1-59745-528-2_2
Rai KN, Khairwal IS, Dangaria CJ, Singh AK, Rao ASJE (2009) Seed parent breeding efficiency of three diverse cytoplasmic-nuclear male-sterility systems in pearl millet. Euphytica 3:495
Wang F, Yue B, Hu JG, Stewart JM, Zhang JF (2009) A target region amplified polymorphism marker for fertility restorer gene Rf1 and chromosomal localization of rf1 and Rf2 in cotton. Crop Sci 49(5):1602–1608
Winter P, Kahl G (1995) Molecular marker technologies for plant improvement. World J Microbiol Biotechnol 11(4):438–448
Gut IG (2001) Automation in genotyping of single nucleotide polymorphisms. Hum Mutat 17(6):475–492. doi:https://doi.org/10.1002/humu.1131
Paterson AH, Wendel JF, Heidrun G, Hui G, Jerry J, Dianchuan J, Danny L, Showmaker KC, Shengqiang S, Joshua U (2012) Repeated polyploidization of Gossypium genomes and the evolution of spinnable cotton fibres. Nature 492(7429):423–427
Wang K, Wang Z, Li F, Ye W, Wang J, Song G, Yue Z, Cong L, Shang H, Zhu S, Zou C, Li Q, Yuan Y, Lu C, Wei H, Gou C, Zheng Z, Yin Y, Zhang X, Liu K, Wang B, Song C, Shi N, Kohel RJ, Percy RG, Yu JZ, Zhu YX, Wang J, Yu S (2012) The draft genome of a diploid cotton Gossypium raimondii Nat Genet 44(10):1098–1103. https://doi.org/10.1038/ng.2371
Varshney RK, Nayak SN, May GD, Jackson SA (2009) Next-generation sequencing technologies and their implications for crop genetics and breeding. Trends Biotechnol 27(9):522–530
Boopathi Manikanda N (2013) Genetic mapping and marker assisted selection. Springer, Berlin
Boopathi NM, Sathish S, Kavitha P, Dachinamoorthy P, Ravikesavan R (2015) Molecular breeding for genetic improvement of cotton (Gossypium spp.). In: Al-Khayri JM (ed) Advances in plant breeding strategies: breeding, biotechnology and molecular tools, pp 613–645. Springer, Berlin, https://doi.org/10.1007/978-3-319-22521-0_21
Funding
The research was supported by grants from the National Natural Science Foundation of China (31871679) and the Central Public-interest Scientific Institution Basal Research Fund (1610162019010303).
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare that no conflict of interest.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
11033_2019_5240_MOESM1_ESM.pdf
Fig. S1 Polymorphic analysis of the InDel-7138 marker in the CMS-D8 and the CMS-D2 systems. Supplementary Material 1 (PDF 346 kb)
Rights and permissions
About this article
Cite this article
Feng, J., Zhu, H., Zhang, M. et al. Development and utilization of an InDel marker linked to the fertility restorer genes of CMS-D8 and CMS-D2 in cotton. Mol Biol Rep 47, 1275–1282 (2020). https://doi.org/10.1007/s11033-019-05240-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11033-019-05240-5