Abstract
Background
Soil salinity drastically reduced wheat growth and production in Pakistan. It is a need of an hour to identify the best suitable salt tolerance or resistant wheat varieties which shows good growth under salinity affected areas. In presented study, two wheat varieties Johar (salt tolerant) and Sarsabaz (salt sensitive) were examined under NaCl stress conditions.
Methods
Antioxidant enzyme activities were investigated in 10-days old wheat seedlings under 200 mM NaCl stress in hydroponic conditions. To investigate the various growth parameters, antioxidant enzyme activities such as superoxide dismutase (SOD: EC 1.15.1.1), catalase (CAT: EC 1.11.1.6) and ascorbate peroxidase (APX: EC 1.11.1.11) were monitored and studied. Besides this various growth parameters such as length of the roots, shoots, as well as Physiological parameters likes lipid peroxidation by malondialdehyde (MDA), hydrogen peroxide (H2O2), and proline contents and antioxidant enzyme activities were estimated. The effect of salinity was also observed on gene transcription level and eventually expression level.
Results
Shoot and root length were decreased in Sarsabaz variety while it showed opposite trend in johar at 200 mM salt concentration. The concentration of proline showed a noticeable rise in salt dependency. Higher concentrations of Proline in Johar were observed as compared to Sarsabaz. SOD showed the increase in activity for antioxidant enzymes. Significant increase of SOD levels were observed in shoot tissues as compared to root tissues. The results indicated that the shoots were more susceptible to salt stress. Activity of APX showed similar affects in both varieties. The production of CAT enzyme in the shoot and root tissues of both varieties showed substantial growth under increased salt stress. Furthermore, NaCl stress has increased the expression of certain genes coding for antioxidant enzymes such as catalase, superoxide dismutase, and peroxidase. Maximum expression of all the antioxidant enzyme coding genes were observed in Johar (tolerant) at 48 h exposure to salt. In contrast the expression of the all mentioned genes in Sarsabaz variety were found maximum at early hours (24 h) and gradually decreased at 48 h.
Conclusion
The study showed that the selected salt tolerant wheat variety Johar is significantly resistant to 200 mM NaCl salt level as compared to Sarsabaz.
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
Introduction
Wheat is the third most leading cereal crop in the world. Around 85 percent of the world’s population depend on wheat [1]. The rapidly increasing population and decreasing agricultural lands in Pakistan demand higher productivity of wheat [2].
Salt stress caused oxidative damage by creating ionic imbalance and producing reactive oxygen species (ROS) like radical superoxide, hydrogen peroxide (H2O2) and radical hydroxyl [3, 4]. These are highly cytotoxic and can interfere with important biosynthetic pathways, resulting in peroxidation, protein denaturation, and DNA mutation [5,6,7,8]. The ROS in plant cells is also responsible for aggregation of membrane lipids, proteins and nucleic acids [9, 10].
Plants formed an efficient scavenging mechanism consisting of enzyme antioxidants such as peroxidase, catalase, ascorabate Xia [11]. These enzymes are also very effective metabolic stress indicators and their elevated expression which indicate a tendency for remediation. In an antioxidant system,antioxidant compounds (ascorbate, glutathione, tochopherols, salicylate) and antioxidant enzymes control the production of reactive oxygen species (ROS) [12, 13]Several studies showed that, in an attempt to defend itself against oxidative damage, plants under different abiotic stresses alter the activity of antioxidant enzymes [4, 13,14,15,17].
SOD constitutes the first line of defense which resist oxidative damage in plants and catalyzes the disputation of two superoxide radicals to hydrogen peroxide and oxygen [11, 18]. There are three groups of SOD categorized by the metal cofactor, namely Cu/Zn-SOD, Fe-SOD and Mn-SOD [19]. CAT is a tetrameric heme protein and mostly located in peroxisomes [20]. CAT has the highest conversion efficiency of all antioxidant enzymes and one molecule of CAT can remove about six million molecules of H2O2 per minute [9, 21]. In stressed plants, the SOD genes are regulated by producing, tissue-specific and environmental signals [14, 22, 23]. The CATs genes respond differentially to various stresses conditions [24, 25]. The APX genes show differential modulation by several abiotic stresses in plants [26, 27]. The expression of wheat GPX genes were altered when wheat plants were submitted to salt, H2O2 and abscisic acid treatment. Moreover, other findings suggest that GPX genes not only act as scavenger of H2O2 to control abiotic stress responses but also play important roles in salt and ABA-signaling cascades [28].
The objectives of this study was to investigate the changes in antioxidant enzyme activities, osmotic adjustment and transcript levels of antioxidant enzyme-related genes against salt stress in two wheat varieties Johar and Sarsabaz at seedling stage.
Material and methods
The experiments were held at the Department of Botany, Islamia University of Bahawalpur. Seeds of 12 wheat verities were obtained from Nuclear Institute of Agriculture (NIA) Tandojam Sindh, Pakistan.
Seed sterilization & germination
The seeds were surface sterilized for 2 min with 10% ethanol, then washed thrice with sterile H2O and submerged into sterile H2O for 24 h. Then seeds were moved to plastic pots lined with cheese cloth. Each pot contains eight seeds and 400 mL of half strength Hoagland’s solution [29]. Pots were kept at 16 h light and 8 h dark cycle.
200 mM NaCl is used for Salt treatment. Among 12 varieties, two varieties of wheat Johar and Sarsabz were selected on the bases of preliminary salt tolerance tests. These selected varieties were then sprouted in petri dishes [30]. Three-day old seedlings (5 replicates) were moved to Hoagland solution. One week old seedlings (3 replicates) were moved to the same medium with 200 mM of NaCl and without NaCl as control for next 3 days. Ten days old seedlings were taken to perform the enzyme assays and mRNA extraction. These plantlets were kept in liquid nitrogen immediately for further studies.
Estimation of growth
Growth analysis of wheat seedlings were carried out after 10 days. The uprooted plants were washed with distilled water. Growth attributes such as root length, shoot length, were measured and various plant parts were separated for further experimentation.
Estimation of chlorophyll content
For spectrophotometric estimation of Chlorophyll a, Chlorophyll b, and the carotenoids, a 50 mg sample in mortar was crushed and homogenized with 4 ml of 80% (v/v) acetone. The solution is filtered through a paper filter. The filtrate volume was made up to 5 mL by adding cold acetone. The absorption of solution was detected at 663.3, 646.8, and 470 nm, and the pigment concentrations were calculated as defined by Lichtenthaler [31].
Estimation of electrolyte leakage (EL) in leaves
The completely formed 1–2 leaves were cut in 0.5 cm pieces and dipped into 7 ml sterilize H2O in glass tubes and incubated for 24 h in rotatory shaker at 250 rpm. After the initial reading of the primary leaf conductivity (Ec–i) was noted then samples were heat at 120 °C for thirty minutes and cooled to room temperature for final conductivity (Ec–f) recording by EC meter. The leakage of electrolytes was estimated by the formula given [32],
Estimation of proline
Proline content was determined taking 0.5 g of fresh leaves and 5 ml of 3% sulfosalicylic acid were added into a test tube and then kept into boiling water for 10 min. The extract was filtered into a clean test tube. 2 ml of extract was put into a test tube with a stopper, and then 2 ml of glacial acetic acid and 2 ml of acidic ninhydrin were added. The mixture was incubated at boiling water for 30 min. After cooling the mixture 4 ml of toluene was added into the test tube. The supernatant was collected into a 10 ml centrifuge tube after centrifuging for 5 min at 3000 rpm. The absorbance of the supernatant was recorded at 520 nm using toluene as a blank [33].
Estimation of H2O2 and MDA content
Patterson [34] method was employed for the detection of hydrogen per oxide content. The content of MDA was calculated using the Hodges et al. [35] test.
Estimation of potassium and sodium ions
The concentrations of Na+ and K+ in leaf and root were measured according to the methods of Weimberg [36]. Briefly, the samples were washed with distilled water to remove any traces of the Na+ from the leaf and root tissues, then dried at 50 °C for 4 days. Thereafter, the dried leaf and root tissues were ground into a fine powder in liquid nitrogen. The powder was then placed in 5 mL nitric acid overnight. Thereafter, the digested solution was diluted to 25 mL with double-distilled water. The concentration of Na+ and K+ in the acid-digested tissue was measured using a flame photometer.
Determination of osmotic adjustment substances
Protein contents were measured using the [37]. Bovine serum was used as standard. The level of total free amino acids in wheat leaf was estimated by the method described by Misra et al. [38] and Moore & Stein [39].
Evaluation of antioxidant enzymes
SOD activity was determined according to Beyer and Fridovich [40]. Twenty microlitres (20 ul) of enzyme solution was mixed with 3 ml SOD reaction solution (pH 7.8 phosphate buffer 1.5 ml, 750 mol−1 NBT0.3 ml, 130 mmol−1 Met 0.3 ml, 20 mol−1 FD 0.3 ml, 100 mol−1 EDTA-Na2 0.3 ml, distilled water 0.3 ml). The control and enzyme solution were placed for 30 min in 4000 l × light. The blank was placed in the dark and read at 560 nm.
Peroxidase (POD) activity was determined using Hernández et al. [41] method. Twenty microliters (20ul) of enzyme solution was mixed with 3 ml POD reaction solution (1.4 μ l guaiacol, 0.85 μ l 30% H2O2 and 0.1 mol−1 pH 6.0 phosphate buffer). The absorbance values was recorded every 30 s at 470 nm.
Catalase (CAT) activity was assayed as a decrease in absorbance at 240 nm for 1 min following the decomposition of H2O2 according to Aebi [42]. The reaction mixture contained 50 mM phosphate buffer (pH 7.0) and 15 mM H2O2.
Ascorbate peroxidase (APX) activity was determined according to Nakano and Asada [43]. The assay mixture consisted of 0.5 mM ASA, 0.1 mM H2O2, 0.1 mM EDTA, 50 mM sodium phosphate buffer (pH 7.0), and 0.15 ml enzyme extract.
RNA extraction from plant tissues
200 mg lypholysed plant was weighed and finely ground with liquid nitrogen. Homogenized tissues were allowed to thaw at room temperature. 1 ml of Trizol was added in homogenized tissue in 1.5 ml microfuge tubes. Tubes were invert mix for 4–5 times. Incubate it at room temperature for 5 min. 400 μL of chloroform was added and incubated at RT for 3 min. Mixture was allowed to centrifuge at 12,000 rpm for 10 min at four degree centigrade for phase dispersion. (During this centrifugation step, new microfuge tubes were labeled and placed in ice box to cool down). Supernatant was transferred in new 1.5 ml tube placed on ice and isopropanol was added in equal ratio. Tubes were incubated on ice (− 20 °C) for 10 min in horizontal form to precipitate the RNA. Samples were centrifuged at 4 °C centigrade with 12,000 rpm for 10 min. Supernatant was discarded. Pellet was washed 2 times with 1 ml of 70% ethanol at 7500 rpm for 5 min at 4 °C. Pellet was allowed to dry fully. 40 μL of RNase free DEPC treated water is used to dissolve the RNA pellet Store at − 80 °C.
RNA evaluation and quantification
RNA standard and quantity were evaluated by using Nano drop. Absorbance at 260, 280 and 320 nm were calculated. The 260/280 ratio was from 1.9 to 2.2 denoting good quality of RNA whereas RNA quantity was about 800–1200 ng/μL.
cDNA synthesis
cDNA was synthesized from isolated RNA using cDNA kit (Vivantis cDSK01-050). For primer preparation mix the reagents i.e., RNA template (7 μL), primer oligos (T)8 (1 μL) dNTPS (1 μL), Nuclease free water (1 μL) and centrifuge the tubes. Incubate the mixture at 65 °C for 5 min and chill on ice for 2 min. Briefly spin down the mixture.
For cDNA Synthesis prepare the Mix according to instructions, Reverse transcriptase (0.2 μL), Buffer 2X (M-MULV, 2 μL, nuclease free H2O (7.8 μL) and make 10 μL final volume. Add 10 μl of the cDNA Synthesis Mix into each RNA-primer mixture. Mix gently and centrifuge briefly. Incubate at 42 °C for 60 min. Terminate the reaction by incubate the tubes at 85 °C for 5 min. Chill the tubes on ice and centrifuge the tube briefly. The synthesized cDNA can be directly used for downstream application or stored at − 20 °C. The list of primers is given in Table 1.
Real time—PCR
Real time polymerase chain reactions (40 cycles) were performed on Mic PCR (Bio Molecular System). To illuminate each sample with a beam of light of at least one specified wavelength and detect the fluorescence emitted by the excited fluorophore. The relative level of gene expression was calculated using the Klein method [44]. The gene transcription levels in control plants were normalized as 1. Following primers were used in Table 1.
Statistical analysis
All of the data was analyzed using IBM SPSS Statistics 2.3 software and a One-way analysis of variance and honest significant difference (HSD) test was done. Principal component analysis was also measured and constructed by Origin 2b software.
Results
Effect of NaCl on growth
It was observed that increased salt concentration decreased the shoot development duration of both verities. The 200 mM NaCl salt concentration increased the shoot length of Johar by 15.2 cm while the shoot length of susceptible variety Sarsabaz decrease (10.6 cm). Root length of tolerant Johar increased (15.32 cm) under 200 mM NaCl stress whereas root length of susceptible Sarsabaz showed decrease. The physiological changes due to the salinity stress were assessed through calculating the ratio of root and shoot length (Fig. 1).
Effect of NaCl stress (200 mM) on plant growth parameters (upper row), Chlorophyll a, Chlorophyll b and Carotenoid (lower row), of tolerant (Johar) and sensitive (Sarsabz) varieties of Triticum aestivum. Period of stress application (48 h):T1 = control Johar plant, T2 = control Sarsabz plant, T3 = Nacl treated Johar plant and T4 = NaCl treated Sarsabz plant. Data are means ± SE at P ≤ 0.01
Effect of NaCl on chlorophyll content
Triticum avestium varieties grown under high salt stress; Sarsabaz variety showed lower values of chlorophyll a, chlorophyll b compared to the Johar variety (Fig. 1) The Johar genotype was proved most tolerant variety by enhanced chlorophyll “a” and chlorophyll “b” under 200 mM salt stress. S arsabaz variety showed lower carotenoid levels as compared to the Johar variety (Fig. 1). The Johar proved to be the most tolerant variety by enhanced carotenoids percentage under 200 mM salt stress.
Effect of NaCl on electrolyte leakage
The impact of salt stress on plants was estimated by the measurement of the electrolyte leakage test. Significant increase of EL percent was observed in Sarsabaz sensitive plant under 200 mM NaCl. However, resistant variety (Johar) cultivar produced decreased electrolyte leakage (Fig. 2).
Effect of NaCl stress (200 mM) on oxidative stress parameters in shoot and root of tolerant (Johar) and sensitive (Sarsabz) varieties of Triticum aestivum. Period of stress application (48 h): T1 = control Johar plant, T2 = control Sarsabz plant, T3 = Nacl treated Johar plant and T4 = NaCl treated Sarsabz plant. Data are means ± SE at P ≤ 0.01
Effect of NaCl on H2O2 and MDA content
Determining the MDA content, that is the result of lipid peroxidation, has established the impact of salt stress on plant tissues. MDA content in the roots of both of verities showed significant changes under increased salt concentrations. Concentrations of MDA in shoot and roots increased under 200 mM NaCl stress in the Sarsabaz variety (Fig. 2).
Measuring the content of H2O2 has established the impact of salt stress on plant tissues. H2O2 concentrations decreased in Sarsabaz variety under NaCl stress of 200 mM (Fig. 2). Results revealed that regulation of increasing variety indicates limited oxidative stress that has been decreased slowly and dramatically (p > 0.01).
Osmotic adjustment substance contents in response to NaCl stress
Effect of NaCl on proline content
Proline is the remarkable organic osmolyte which accumulates in plants as a result of response to environmental stresses i.e., severe temperature, H2O deficiency conditions, salt stress and heavy metals. The proline estimation serves as a physiological marker for salt stress. Proline content varies from 2.3 to 3.5 in the shoot and 2.6–4 in the root under control conditions. Maximum increased proline output was observed in Johar (3.2 gμ g1 and 4.2 gμ g1) in the shoot and root respectively. Minimal proline production was observed in Sarsabaz variety (Fig. 3).
Effect of NaCl stress (200 mM) on osmoprotectants in shoot and root of tolerant Johar) and sensitive (Sarsabz) varieties of Triticum aestivum. Period of stress application (48 h):T1 = control Johar plant, T2 = control Sarsabz plant, T3 = Nacl treated Johar plant and T4 = NaCl treated Sarsabz plant. Data are means ± SE at P ≤ 0.01
Total soluble sugars
Figure 3 shows that wheat variety, salinity and their association had major effects on overall soluble sugar (p > 0.01) The two salinity-stressed wheat varieties showed a significantly lower accumulation of sugars in roots relative to shoot tissues. The reducing sugars of shoot decreased in Sarsabaz.
Total soluble amino acids
Varieties, salinity and association of varieties had major effects on total soluble amino acids (p > 0.01) among the two salinity stressed wheat varieties. Root tissues displayed significantly more amino acid accumulation than shoot. The amino acids of shoot decreased as salinity increased in only Sarsabaz (Fig. 3).
Determination of potassium and sodium ions
After 48 h of stress measurement of sodium aggregation revealed two significant variations in Na transport between Johar and Sarsabaz. Sarsabaz sequestered more root sodium than shoot under 200 mM NaCl stress. In comparison, more potassium was stored in the leaf of Johar than Sarsabaz. In both tested varieties, the roots exhibited higher sodium compared to shoots. The highest sodium content (6.82 mg g1 DW) was observed in roots of Sarsabaz plants under 200 mM NaCl (Fig. 4).
Effect of NaCl stress (200 mM) on ion contents in root and shoot of tolerant (Johar) and sensitive (Sarsabz) varieties of Triticum aestivum. Period of stress application (48 h): T1 = control Johar plant, T2 = control Sarsabz plant, T3 = Nacl treated Johar plant and T4 = NaCl treated Sarsabz plant. Data are means ± SE at P ≤ 0.01
Determination of Na/k Ratio
Higher ratio of Na/K in the salt sensitive cultivar Sarsabaz was observed. On the contrary, Johar had less Na/K ratio than Sarsabaz (Fig. 4). The highest ratio of Na/K was observed in Sarsabaz roots under 200 mM NaCl.
Antioxidant enzymes activities
Effect of NaCl on catalase activity
Data in the Fig. 5 showed that salinity had a significant impact on the antioxidant enzyme activities. The salt stress produced a significant increase in catalase activity in the Johar plant at 200 mM NaCl concentration. Catalase activity was slightly higher in root (6.0 min−1 g−1 FW) and (5.33 min−1 g−1 FW) in shoot of Johar variety relative to Sursabaz variety showing limited catalase activity in both tissues of plant.
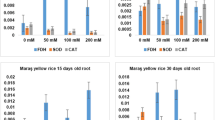
Effect of NaCl stress (200 mM) on antioxidant enzymes in shoot and root of tolerant (Johar) and sensitive (Sarsabz) varieties of Triticum aestivum. Period of stress application (48 h):T1 = control Johar plant, T2 = control Sarsabz plant, T3 = Nacl treated Johar plant and T4 = NaCl treated Sarsabz plant. Data are means ± SE at P ≤ 0.01
Effect of NaCl on superoxide dismutase activity
It indicates that SOD is inducible under higher salt concentration. SOD behavior increased dramatically under 200 mM NaCl concentration for Johar. Higher percent increase was observed in the shoot (105.2 min−1 g−1 FW). In the Johar, the root SOD activity was 115.5 min−1 g−1 FW (Fig. 5).
Effect of NaCl on ascorbate peroxidase activity
APX activity indicates a rise under 200 mM NaCl concentration in the Johar variety. A higher percentage increase was observed in the shoots tissues of the Johar (2.77 min−1 g−1 FW) (Fig. 5).
Effect of NaCl on peroxide dismutase activity
POD tends to induce under higher salt concentration. POD behavior dramatically increased under 200 mM NaCl concentration for Johar variety. Higher percent rise was observed in Johar shoot tissue (2.83 min−1 g−1 FW). Whereas in root tissues POD activity was 3.33 min−1 g−1 FW (Fig. 5).
Gene expression of antioxidant enzymes in response to salt stress
Expression of CAT gene
According to the analysis of variance, it was observed that the level of transcripts of the genes of CAT, SOD, and APX enzymes in wheat seedlings subjected to 200 mM NaCl salt stress significantly change in period of stress (Fig. 6) In this study, maximum expression of CAT gene was observed in Johar at 48 h compared to sensitive variety (Sarsabaz) which shows highest gene expression only at early hours (24 h) and gradually decrease at 48 h (Fig. 6).
Expression of MnSOD gene
The expression profile of the MnSOD gene under salt stress (200 mM NaCl) was observed. In this study minimum expression of MnSOD gene was observed at early hours (24 h) in both tolerant (Johar) and sensitive variety (Sarsabaz) and it gradually increased after 24 h which shows highest expression at 48 h (Fig. 6).
Expression of APX gene
In this study, we determine a differential response of APX activity in two wheat varieties under 200 mM NaCl stress. APX activity was observed highest at early hours (24 h) in both salt tolerant variety (Johar) and sensitive variety (sarsabaz) and it gradually decrease to show minimum expression at 48 h (Fig. 6).
Correlation between antioxidant enzyme activity and relevant gene expressions
According to CAT enzyme activity and gene expression profile of CAT gene (Fig. 6), we observed that both CAT enzyme activity and expression was greater in Johar variety compared to Sarsabaz. SOD enzyme efficiency of the salt–tolerant varieties (Johar) increases in both root and shoot tissues. In contrast, Super dismutase activity in variety Sarsabaz slightly increased relative to the control plants. According to gene expression we observed that MnSOD gene expression highest at 48 h in Johar followed by Sarsabaz.
In the Johar cultivar APX activity increased under salt stress with respect to control plants. Moreover, this increase was more in Johar than in Sarsabaz. In the present study we observed maximum APX enzyme activity in shoot and roots of Johar compared to Sarsabaz variety. In the presence of high NaCl concentration (200 mM), APX gene expression of the salt -tolerant variety was higher relative to untreated plants and higher in Sarsabaz variety.
Discussion
In presented study, salt tolerance of two wheat varieties were examined. At germination stage, Johar cultivar was more tolerant to salinity than Sarsabaz. In Sarsabaz, the higher concentration of sodium chloride reduced the plant growth is higher in particular relative water content.
Plant growth particularly plant biomass and root and shoot length were reduced under saline conditions (Fig. 1). The reduction in growth at high salt stress resulted from a reduction in osmotic reliability might be due to saturated ion uptake systems, [45, 46]. It was observed that salt stress affects plant development and growth by causing nutrient deficiency [potassium (K+), calcium (Ca) and Magnesium (Mg+)]. Excessive growth decrease was seen in leaf tissues as compared to roots under 200 mM of salinity stress [47]. Development of the shoot in most crop species is highly susceptible to salinity as compared to root growth [48]. Hernandez et al. [41] found that salt stress contributed to a decrease in the area of the leaves as well as a decrease in dry and fresh weight in both the pea roots and leaves. Salt stress causes decreased wet and dry weight in Maize genotypes [49].
Two varieties of wheat that we observed seemed to have same sodium storage ability in roots and shoots, Johar showed different Na + accumulation efficiency in both tissues (Fig. 4). Sana et al. [50] reported that the storage capacity of Na + in roots was comparable in tolerant and sensitive genotypes of wheat but higher in leaf blades of sensitive varieties.
Proline is an important osmolyte [51, 52] that plays a special role in exposure to various stress conditions as an antioxidant protection compounds and a signaling molecule in the plant system [6]. Although Proline action mechanism in response to stress is not well understood, this may be involved in water stress membrane stabilization [5]. The excessive presence of proline in plants is known to retard development and adversely impacts many metabolisms of tissues [53]. Build up of well suited solutes such as Proline and GB under salinity is hypothesized to raise plant salt resistance [7, 48].
The proline content under osmotic stress save plants by controlling osmotic equilibrium, Preventing EL and ROS concentrations to control oxidative bursts in plants [52]. Similar results were observed in this research, higher Proline values in roots and shoots of tolerant variety Johar (Fig. 3). Environmental stresses increase superoxide development efficiency depending on the species, stress duration and severity, and plant age. There is growing evidences that membrane wound under salinity is associated with high production of AOS [52,53,56], Hernandez et al. 1998. In accordance with these studies, our findings demonstrate that salt stress produced oxidative stress in leaves of two wheat varieties shown by a collection of oxygen ions, rise in MDA and hydrogen peroxide content. However, the amount of lipid peroxidation and the content of hydrogen peroxide in the Sarsabaz were higher than in the other varieties under NaCl stress. These data indicate that the Johar under NaCl stress was better protected against oxidative damage. According to our findings, low MDA is significant in respect of salt resistance as shown in various observations. Salt resistant tobacco 58 and wheat varieties [56, 57] also showed low MDA levels, that is a significant sign of increasing oxidative retardation limiting potential salt stress.
For the estimation of lipid peroxidation level the MDA content is used as an indicator. The determination of the MDA level and the membrane lipid peroxidation extent, which has been used as a method to evaluate magnitude of the oxidative stress and the extent of non-tolerance of the plant [5, 59]. Many studies have recorded an increase in H2O2 concentration after exposure to stress, it is related to stress intensity and duration. In addition, the amount of H2O2 varies between different cellular compartments and is correlated with stress form [60].
Tolerance to the stress of salinity in higher plants is associated with antioxidant systems and substrates [61]. The increases in concentration of antioxidant compounds are indicators of plant resistance/adaptation to salinity condition. Variation in the efficiency of the enzymes is therefore associated with plant oxidative stress resistance [13]. Changes in antioxidant concentration may act as a signal for modulating reactive oxygen species scavenging mechanisms and transduction of reactive oxygen species signals [17].
Comparison of the enzymatic efficiency for two wheat verities; the salt tolerant Johar showed the higher activity as compared to Sarsabaz. This result indicated that the Johar is more able to scavenge ROS than the Sarsabaz. The decrease in Johar’s H2O2 content is thus the product of superoxide dismutase reaction; that is followed by an increased enzymatic capability to decompose it. This was especially evident in Johar, where under low or high salinity a significant increase in SOD, CAT, POD and APX activities were observed (Fig. 5).
SOD level raised under 200 mM NaCl in the shoot tissues of Johar (Fig. 5). This link between salinity resistant and increased Superoxide dismutase activity was demonstrated in several studies. [14, 18, 62, 63]. High production of superoxide dismutase in Johar at higher salt level was able to cope with injurious effects of reactive oxygen. Ascorbate peroxidase uses AsA as an electron donor for reduction of hydrogen peroxide to H2O with monodehydro ascorbate generation, [23] and plays a major role in controlling reactive oxygen species during stress [61].
It is observed in several salt resistant varieties that salinity increase ascorbate peroxidase activity [26, 27, 64]. The Sarsabaz’s vulnerability to salinity may be due to ascorbate peroxidase efficiency inhibition at a high salt stress level.
In wheat, the superoxide dismutase transcript was high in response to differential heat sensitive experimental treatments. [51], suggested an increased resistance to environmental stresses. In plants, the superoxide dismutase genes are regulated by growth, tissue and environmental stresses [25, 65, 66].
In Johar, higher CAT activity was observed in the presence of a high level of salt stress (200 mM NaCl) (Fig. 5), resulting in good coping with ROS. Higher CAT activity, close to our findings, has been observed in many salt resistant varieties i.e., sesame, maize, and wheat [67, 68]. The maximum expression of the CAT gene was observed in Johar in at 48 h relative to the susceptible variety (Sarsabaz) showing the highest gene expression only at the early hours (24 h) and steadily decreasing with increase in time till 48 h Fig. 6. Similar results were observed when the wheat CAT gene expressed in transgenic rice increases resistance compared to non-transgenic plants to low-temperature stress [69]. The catalase genes react to many stress conditions differently [27, 70].
In this analysis, APX activity was observed to be highest in early hours (24 h) and steadily decreased to show minimum expression at 48 h. Ascorbate peroxidase proteins are dispersed in chloroplasts, mitochondria, cytosol and peroxisomes The ascorbate peroxidase genes show differential modulation in plants through various environmental stresses [26, 64, 71, 72].
Conclusion
The findings of this study indicate that the ability of Tritium aestivum to cope with salt stress depends on antioxidant stress defense mechanisms. An increase in antioxidant enzyme activity and the resulting increase in ROS-scavenging capacity can improve the salt tolerance of plants. Our results show that there were increases in antioxidant enzyme activity and in the transcript levels of genes encoding these enzymes under salt stress. These findings suggest that the response of salt tolerant variety Johar to salt stress involves CAT, SOD, APX, POD, and GPX. These results increase our understanding of how T. aestivum responds to salt stress and provide evidence for the effectiveness of its detoxification mechanisms. Thus, we concluded that insight into any plant’s antioxidant enzyme mechanism implies tolerance efficiency.
Data availability
All data are available when need.
References
Chaves MM, Pereira JS, Maroco J, Rodrigues ML (2002) How plants cope with water stress in the field: photosynthesis and growth. Ann Bot 89:907–916
Iqbal MA, Rahim J, Naeem W, Hassan S, Khattab Y, Sabagh A (2021) Rainfed winter wheat (Triticum aestivum L.) cultivars respond differently to integrated fertilization in Pakistan. Fresenius Environ Bull 30(4):3115–3121
Naheed R, Zahid M, Aqeel M et al (2022) Mediation of growth and metabolism of Pisum sativum in salt stress potentially be credited to thiamine. J Soil Sci Plant Nutr. https://doi.org/10.1007/s42729-022-00854-4
Naheed R, Zahid M, Aqeel M, Maqsood MF, Kanwal H, Khalid N, Hashem M, Alamri S, Noman A (2022) Mediation of growth and metabolism of Pisum sativumin salt stress potentially be credited to thiamine. J Soil Sci Plant Nutr. https://doi.org/10.1007/s42729-022-00854-4
Banik N, Bhattacharjee S (2020) Complementation of ROS scavenging metabolites with enzymatic antioxidant defense system augments redox-regulation property under salinity stress in rice. Physiol Mol Biol Plants 26:1623–1633
García-Caparrós P, De Filippis L, Gul A, Hasanuzzaman M, Ozturk M, Altay V, Lao MT (2020) Oxidative stress and antioxidant metabolism under adverse environmental conditions: a review. Bot Rev 87:421
Hossain MS (2019) Present scenario of global salt affected soils, its management and importance of salinity research. Int Res J Biol Sci 1:1–3
Tufail A, Aqeel M, Khalid N, Ahsan M, Khilji SA, Ahmad F, Hameed M, Noman A, Alamri S, Hashem M (2020) Salt toxicity in a natural habitat induces structural and functional modifications and modulate metabolism in Bermuda grass (Cynodon dactylon pers.) ecotypes. Appl Ecol Environ Res 18(5):6569–6588
Gill SS, Tuteja N (2010) Reactive oxygen species and antioxidant machinery in abiotic stress tolerance in crop plants. Plant Physiol Biochem 48:909–930
Pérez-López U, Robredo A, Lacuesta M, Sgherri C, Muñoz-Rueda A, Navari-Izzo F (2009) The oxidative stress caused by salinity in two barley cultivars is mitigated by elevated CO2. Physiol Plant 135:29–42
Zhang X, Yin HaiBo, Chen SH, He J, Guo SL (2014) Changes in antioxidant enzyme activity and transcript levels of related genes in Limonium sinense Kuntze seedlings under NaCl stress. J Chem 5(1):1–6
Rashda N, Humaira A, Hina K, Fozia F, Mohammad I, Abo G, Al-Mushhin AAM, Dilfuza J, Mohammad JA, Sehar S, Muhammad A, Ali N, Kamel H (2021) Growth attributes, biochemical modulations, antioxidant enzymatic metabolism and yield in Brassica napus varieties for salinity tolerance. Saudi J Biol Sci 28(10):5469–5479
Wani PA, Garba SH, Wahid S, Hussaini NA, Mashood KA (2019) Prevention of oxidative damage and phytoremediation of Cr (VI) by chromium (VI) reducing Bacillus subtilus PAW3 in cowpea plants. Bull Environ Contam Toxicol 103(3):476–483
Anton E, Aleksandr B, Ulyana Z, Dmitrii K, Alexey D (2019) Stress induced changes in the expression of antioxidant system genes for rice (Oriza sativa L.) and bread wheat (Triticum aestivum L.). Peer J. 29(7):1–10
Moradbeygi H, Jamei R, Heidari R, Darvishzadeh R (2020) Investigating the enzymatic and non-enzymatic antioxidant defense by applying iron oxide nanoparticles in Dracocephalum moldavica L. plant under salinity stress. Sci Hortic 272:109537
Moradbeygi H, Jamei R, Heidari R, Darvishzadeh R (2020) Fe2O3 nanoparticles induced biochemical responses and expression of genes involved in rosmarinic acid biosynthesis pathway in Moldavian balm under salinity stress. Physiol Plant 169:555–570
Mushtaq T, Shah AA, Akram W, Yasin NA (2020) Synergistic ameliorative effect of iron oxide nanoparticles and Bacillus subtilis S4 against arsenic toxicity in Cucurbita moschata: polyamines, anti-oxidants, and physiochemical studies. Int J Phytoremediat 22:1408–1419
Eteshami H, Hamideh F, Muhammad R (2021) Interaction of nanoparticles and salinity stress at physiological, biochemical and molecular levels in plants. Ecotoxicol Environ Safety 5(1):6654
Abdoli S, Ghassemi-Golezani K, Alizadeh-Salteh S (2020) Responses of ajowan (Trachyspermum ammi L.) to exogenous salicylic acid and iron oxide nanoparticles under salt stress. Environ Sci Pollut Res 27:36939–36953. https://doi.org/10.1007/s11356-020-09453-1
Mhamdi A, Queval G, Chaouch S, Vanderauwera S, Van Breusegem F, Noctor G (2010) Catalase function in plants: a focus on Arabidopsis mutants as stress-mimic models. J Exp Bot 61:4197–4220
González A, Vidal C, Espinoza D, Moenne A (2021) Anthracene induces oxidative stress and activation of antioxidant and detoxification enzymes in Ulvalactuca (Chlorophyta). Sci Rep 11:7448
Alscher RG, Erturk N, Heath LS (2002) Role of superoxide dismutase in controlling oxidative stress in plants. J Exp Bot 53:1331–1341
Tatiana R, Marcelo NA, Leticia CB, Isabel LV, Eugenia JBB, Ariano MMJ, Mara AC (2017) Gene expression and activity of antioxidant enzymes in rice plants, cv. BRS AG, under saline stress. Physiol Mol Biol Plants 23(4):856–875
Scandalios JG (2002) The rise of ROS. Trends Biochem Sci 27(9):483–486
Scandalias JG (2005) Responses of plant antioxidant defense genes to environmental stress. Adv Genet 28:1–41
Rosa SB, Caverzan A, Teixeira FK, Lazzarotto F, Silveira JA, Ferreira-Silva SL, Abreu-Neto J, Margis R, MargisPinheiro M (2010) Cytosolic APx knockdown indicates an ambiguous redox responses in rice. Phytochemistry 71:548–558
Soraya MPB, Sedigheh FO, Ali A (2019) Expression of dehydrin and antioxidant genes and enzymatic antioxidant defense under drought stress in wild relatives of wheat. Biotech Biotec Equip 33(1):1063–1073
Zhai CZ, Zhao L, Yin LJ, Chen M, Wang QY, Li LC, Xu ZS, Ma YZ (2013) Two wheat glutathione peroxidase genes whose products are located in chloroplasts improve salt and H2O2 tolerances in Arabidopsis. PLoS ONE 8(10):e73989
Hoagland DR, Arnon DI (1950) The water-culture method for growing plants without soil, 2nd edn. University of California Agricultural Experiment Station, Berkeley, Circular
Brini F, Amara I, Feki K, Hanin M, Khoudi H, Masmoudi K (2009) Physiological and molecular analyses of seedlings of two Tunisian durum wheat (Triticum turgidum L. subsp. Durum [Desf.]) varieties showing contrasting tolerance to salt stress. Acta Physiol Plant 31:145–154
Lichtenthaler HK, Wellburn AR (1983) Determinations of total carotenoids and chlorophylls a and b of leaf extracts in different solvents. Biochem Soc Trans 11:591–592
Li W, Li Q (2017) Effect of environmental salt stress on plants and the molecular mechanism of salt stress tolerance. Int J Environ Sci Nat Res. https://doi.org/10.19080/IJESNR.2017.07.555714
Bates LS, Waldren RP, Teare ID (1973) Rapid determination of free proline for water-stress studies. Plant Soil 39:205–207
Peterson RKD, Higley LG (2000) Biotic stress and yield loss, 1st edn. CRC Press, Washington, p 261
Hodges DM, DeLong JM, Forney CF, Prange RK (1999) Improving the thiobarbituric acid-reactive-substances assay for estimating lipid peroxidation in plant tissues containing anthocyanin and other interfering compounds. Planta 207:604–611
Weimberg R (1987) Solute adjustments in leaves of two species of wheat at two different stages of growth in response to salinity. Physiol Plant 70:381–388
Gil R, Bautista I, Boscaiu M, LidoÂn A, Wankhade S, SaÂnchez H (2014) Responses of five Mediterranean halophytes to seasonal changes in environmental conditions. AoB Plants 6:1–18
Misra PS, Mertz ET, Glower DV (1975) Cereal Chem 52: 844
Moore S, Stein WH (1984). In: Colowick SP, Kaplan ND (eds) Methods in enzymol. Academic Press, New York, p 468
Beyer W, Fridovich I (1987) Assaying for superoxide dismutase activity: some large consequences of minor changes in conditions. Anal Biochem 161:559–566
Hernandez JA, Campillo A, Jimenez A, Alarcon JJ, Sevilla F (1999) Response of antioxidant systems and leaf water relations to NaCl stress in pea. New Phytol 141:241–251
Aebi H (1984) Catalase in vitro. Methods Enzymol 105:121–126
Nakano Y, Asada K (1981) Hydrogen peroxide is scavenged by ascorbate-specific peroxidase in spinach chloroplasts. Plant Cell Physiol 22:867–880
Klein D (2002) Quantification using real time PCR technology: applications and limitations. Trends Mol Med 8:257–260
Molotoks A, Smith P, Dawson TP (2021) Impacts of land use, population, and climate change on global food security. Food Energy Secur 10:e261
Romano-Armada N, Yanez-Yazlle MF, Irazusta VP, Rajal VB, Moraga NB (2020) Potential of bioremediation and PGP traits in Streptomyces as strategies for bio-reclamation of salt-afected soils for agricultural. Pathogens 9:117
Peng Z, He S, Sun J, Pan Z, Gong W, Lu Y, Du X (2016) Na+ compartmentalization related to salinity stress tolerance in upland cotton (Gossypium hirsutum) seedlings. Sci Rep 4(6):34548. https://doi.org/10.1038/srep34548
Tanveer M, Ahmed HAI (2020) ROS signaling in modulating salinity stress tolerance in plants. In: Hasanuzzaman M, Tanveer M (eds) Salt and drought stress tolerance in plants. Springer, Cham, pp 299–314
Noureen Z, Zulfiqar AR, Saqib M (2020) Effect of salinity stress on various growt and physioloical attributes of two contrasting Maize genotypes. Agri Agribusi Biotech 63:e20200072
Tounsi S, Feki K, Hmidi D, Masmoudi K, Brini F (2017) Salt stress reveals differential physiological, biochemical and molecular responses in T. monococcum and T. durum wheat genotypes. Physiol Mol Biol Plants 23(3):517–528
Kumar RR, Sharma SK, Goswami S, Singh K, Gadpayle KA, Singh GP, Pathak H, Rai RD (2013) Transcript profiling and biochemical characterization of mitochondrial superoxide dismutase (mtSOD) in wheat (Triticum aestivum) under different exogenous stresses. Aust J Crop Sci 7:414–424
Zelm EV, Zhang Y, Testerink C (2020) Salt tolerance mechanism in plants. Ann Rev Plant Biol 71:403–433
Tokihiko N, Miki F, Motoaki S, Tomohiko K, Satoshi T, Kazuo S (2003) Toxicity of free proline revealed in an Arabidiopsis T-DNA tagged mutant deficient in proline deydroenase. Plant Cell Physiol 44(5):541–548
Hernandez JA, Del Rio LA, Sevilla F (1994) Salt stress induced changes in superoxide dismutase isozyme in leaves and mesophyll protoplast from Vigna unguiculata (L) Walp. New Phytol 126:37–44
Hernandez JA, Olmos E, Corpas FJ, Sevilla F, Del Rio LA (1995) Salt induced oxidative stress in chloroplast of pea plants. Plant Sci 105:151–167
Sairam RK, Rao KV, Srivastava GC (2002) Differential response of wheat genotypes to long term salinity stress in relation to oxidative stress, antioxidant activity and osmolyte concentration. Plant Sci 163:1037–1046
Mandhania S, Madan S, Sawhney V (2006) Antioxidant defense mechanism under salt stress in wheat seedling. Biol Plant 227:227–231
Ruiz JM, Blasco B, Rivero RM, Romero L (2005) Nicotine‐free and salt‐tolerant tobacco plants obtained by grafting to salinity‐resistant rootstocks of tomato. Physiol Plant 124(4):465–475.
Ashraf M (2009) Biotechnological approach of improving plant salt tolerance using antioxidants as markers. Biotechnol Adv 27:84–93
Slesak I, Libik M, Karpinska B, Karpinski S, Miszalski Z (2007) The role of hydrogen peroxide in regulation of plant metabolism and cellular signalling in response to environmental stresses. Acta Biochim Pol 5:39–50
Ahmad P, Jaleel CA, Sharma S (2010) Antioxidant defence system, lipid peroxidation, proline-metabolizing enzymes, and biochemical activities in two Morus alba genotypes subjected to NaCl stress. Russ J Plant Physiol 57:509–517
Karolina D, Magdalena Z, Andreas B, Hubert S, Krzysztof K, Michal N (2019) Analysis of wheat gene expression related to the oxidative stress response and signal transduction under short term osmotic stress. Sci Rep 9:2743
Xu FJ, Jin CW, Liu WJ, Zhang YS, Lin XY (2011) Pretreatment with H2O2 alleviates aluminum-induced oxidative stress in wheat seedlings. J Integr Plant Biol 53:44–53
Caverzan C, Bonifacio A, Carvalho FEL, Andrade CMB, Passaia G, Schünemann M, Maraschin FS, Martins MO, Teixeira FK, Rauber R et al (2014) The knockdown of chloroplastic ascorbate peroxidases reveals its regulatory role in the photosynthesis and protection under photo-oxidative stress in rice. Plant Sci 214:74–87
Scandalios JG, Guan L, Polidoros AN (1997) Catalases in plants: gene structure, properties, regulation, and expression. Cold Spring Harb Monogr Ser 34:343–406
Menezes BL, Teixeira FK, Kamei CLA, Margis PM (2004) Salt stress induces altered expression of genes encoding antioxidant enzymes in seedlings of a Brazilian indica rice (Oryza sativa L.). Plant Sci 166(2):323–331
Azevedo NA, Prisco JT, Filho JE, Lacerda CF, Silva JV, Costa PH, Filho EG (2004) Effects of salt stress on plant growth, stomatal response and solute accumulation of different maize genotypes. Braz J Plant Physiol 16:31–38
Koca H, Bor M, Ozdemir F, Turkan I (2007) Effect of salt stress on lipid peroxidation, antioxidative enzymes and proline content of sesame cultivars. Environ Exp Bot 2007(60):344–351
Mehdi R, Mojtaba K, Fereshteh MH, Sanam SC (2021) Antioxidant gene expression analysis and evaluation of total phenol content and oxygen scavenging system in tea accessions under normal and drought stress conditions. BMC Plant Biol 21:494
Doroskov AV, Bobroviskikah AV (2018) Using the method of systems biology for predicting perspective target genes to select C3 and C4 cereals for oxidative stress resistance. Vavilovskii Zhurnal Genetiki I Selekitsii 22(1):122–131
Raza A, Sharif Y, Chen K, Wang L, Fu H, Zhuang Y, Zhuang W (2022) Genome-wide characterization of ascorbate peroxidase (APX) gene family in peanut (Arachis hypogea L.) revealed their crucial role in growth and multiple stress tolerance. Front Plant Sci 2466
Caverzan A, Passaia G, Rosa SB, Ribeiro CW, Lazzarotto F, Margis-Pinheiro M (2012) Plant responses to stresses: role of ascorbate peroxidase in the antioxidant protection. Genet Mol Biol 35:1011–1019
Choudhury S, Panda P, Sahoo L, Panda SK (2013) Reactive oxygen species signaling in plants under abiotic stress. Plant Signal Behav 8:23681
Epstein E, Norlyn JD, Rush DW, Kingburry RW, Kelley DB, Cunnigham GA, Wrona AF (1980) Saline culture of crops: a genetic approach. Science 210:399–404
Liang X, Zhang L, Natarajan SK, Becker DF (2013) Proline mechanism of stress survival. Antioxid Redox Signal 19(9):998–1011
Pitman MG, Lauchli A (2002) Global impact of salinity and agricultural ecosystems. In: Lauchli A, Luttge V (eds) Salinity: environment-plants molecules. Kluwer, Dordrecht, pp 375–378
Rengasamy P (2010) Osmotic and ionic effects of various electrolytes on the growth of wheat. Aust J Soil Res 48:120–124
Sairam RKK, Chandrasekhar V, Srivastava GCC (2001) Comparison of hexaploid and tetraploid wheat cultivars in their responses to water stress. Biol Plant 44:89–94
Acknowledgements
The authors would like to acknowledge the Deanship of Scientific Research at King Khalid University, Saudi Arabia, for their technical support RGP.2/169/42.
Funding
Funding for research was provided from project number RGP.2/169/42, King Saud University, Saudi Arabia.
Author information
Authors and Affiliations
Contributions
MR, Conceptualization and Validation; MG, Performed experiments; AAS, Statistical analysis, review and drafting; RTA, Review; MJ, Writing; SU, Drafting and funding support.
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare no conflict of interest.
Ethical approval
This article does not contain any studies with human participants or animals performed by any of the authors.
Consent to participate
Not applicable.
Consent to publication
Not applicable.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Ramzan, M., Gillani, M., Shah, A.A. et al. Triticum aestivum: antioxidant gene profiling and morpho-physiological studies under salt stress. Mol Biol Rep 50, 2569–2580 (2023). https://doi.org/10.1007/s11033-022-07990-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11033-022-07990-1