Abstract
Background
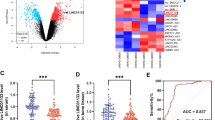
Osteogenic differentiation of human mesenchymal stem cells (hMSCs) holds significant clinical implications for patients with bone diseases. LncRNAs are an emerging group of epigenetic modulators involved in the osteogenesis of hMSCs. In this study, we explored lncRNA profiles that are upstream to the hsa-miR-214-3p/BMP2 axis in osteogenic differentiation of hMSCs.
Method
HMSCs were induced toward osteogenesis for 14 days. Between day 1 and day 14, qRT-PCR was conducted to compare the expressions of BMP2, Runx2, hsa-miR-214-3p, and biochemical assays to compare alkaline phosphatase and Alizarin Red S activities. 145 lncRNAs, which were experimentally confirmed upstream to hsa-miR-214-3p were examined. Five significantly upregulated lncRNAs, MEG3, SNHG16, FAM83H-AS1, MALAT1 and LINC00657 were downregulated in differentiated hMSCs and their impact on osteogenic differentiation were examined. Hsa-miR-214-3p was silenced in lncRNAs-downregulated hMSCs to further examine the association between lncRNAs and hsa-miR-214-3p/BMP2 axis.
Results
From day 1 to day 14, hMSCs underwent significant osteogenic differentiation, and KCNQ1OT1, MEG3, SNHG16, FAM83H-AS1, MALAT1 and LINC00657 were significantly upregulated. Downregulations of MEG3, SNHG16, FAM83H-AS1, MALAT1 and LINC00657 all suppressed osteogenic differentiation. However, qRT-PCR and RIP assay demonstrated that only MALAT1 and LINC00657 acted through hsa-miR-214-3p/BMP2 to regulate osteogenic differentiation. Furthermore, silencing hsa-miR-214-3p only rescued osteogenic differentiation in MALAT1- or LINC00657- downregulated hMSCs.
Conclusions
Our data strongly indicated that lncRNAs MALAT1 and LINC00657 acted through miR-214-3p/BMP2 axis to regulate osteogenic differentiation of hMSCs.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Human bone development starts in the early stages of embryonic life and consists of two osteogenic pathways, intramembranous ossification and endochondral ossification [1, 2]. Human mesenchymal stem cells (hMSCs), with tremendous self-renew capabilities and significant potential of differentiating into multiple cell lineages under various biological conditions [3, 4], can also act as osteoblastic precursors mainly through the activation of intramembranous ossification pathway [5, 6]. In clinical medicine, hMSCs had been extensively utilized as a reliably tissue-engineering tool for a variety of bone repair and regenerating applications [4, 7]. Thus, understanding the genetic mechanisms underlying osteogenic differentiation of hMSCs bears significant benefits toward curing bone-related deficits and diseases among human patients.
Long noncoding RNAs (lncRNAs) are groups of long (> 200 nucleotides) and non-protein-coding RNA transcripts that had been identified to play important roles in human health and diseases [8, 9]. In a recent study investigating the roles of lncRNAs in osteogenic differentiation of bone marrow‐derived mesenchymal stem cells, Tang and colleagues revealed that there were more than 1,000 lncRNAs which were either upregulated or downregulated from day 0 to day 10 of osteogenic differentiation [10]. In addition, functional regulations of lncRNAs during the process of mesenchymal stem cells induced osteogenic differentiation have been widely reported [10,11,12,13,14,15].
MicroRNAs (miRNAs) are another group of non-protein-coding but with short-length (17–25 nucleotides) RNA transcripts, that can transcriptionally suppress downstream messenger RNA (mRNA) through the binding on target genes’ three prime untranslated regions (3′-UTRs) [16]. Strong evidence has shown that miRNAs, through inhibitory effects on downstream mRNA/protein modulations, can actively regulate human cell development, such as proliferation, differentiation or apoptosis [17]. In mesenchymal stem cells-derived osteogenic differentiation, miRNAs have also been widely implicated to play functional roles by either inducing or suppressing key osteogenic genes and proteins [18,19,20,21].
A recent study by Wang and colleagues demonstrated that, lncRNA of KCNQ1 Opposite Strand/Antisense Transcript 1 (KCNQ1OT1) modulated bone marrow mesenchymal stem cell osteogenic differentiation through the interaction of an epigenetic axis of human mature microRNA-214 (hsa-miR-214-3p) and human bone morphogenetic protein 2 (BMP2) gene [14]. As BMP2 functions as one of the key components of osteogenic induction in mesenchymal stem cells [22, 23], it is important to explore the full scope of the upstream initiators of hsa-miR-214-3p / BMP2 axis, thus to further understanding the epigenetic signaling pathway responsible for osteogenic differentiation.
In this study, by inducing osteogenic differentiation among hMSCs, we studied the lncRNAs potentially upstream to hsa-miR-214-3p / BMP2 axis. Using qRT-PCR analysis, we identified several lncRNAs which were significantly upregulated through the course of hMSC osteogenic differentiation and were experimentally confirmed to bind hsa-miR-124-3p. Then, through quantitative biochemical assays and gene knockdown approaches, we demonstrated that Metastasis Associated Lung Adenocarcinoma Transcript 1 (MALAT1) and long intergenic non-protein coding RNA 657 (LINC00657) were upstream to miR-214-3p / BMP2 axis and regulated osteogenic differentiation of hMSCs.
Materials and methods
Human mesenchymal stem cells and osteogenic differentiation
Human mesenchymal stem cells (hMSCs) were purchased from Lonza (Lonza, Shanghai, China) and maintained in mesenchymal stem cell growth medium (BulletKit, Lonza, Shanghai, China) according to the manufacturer’s recommendation in 6-well plates (VWR. Shanghai, China) in a humidified environment of 5% CO2 at 37 °C. Cells were sub-cultured once reaching ~ 85% confluency. To induce osteogenesis, hMSCs were lifted from 6-well plates and re-cultured in 12-well plates (VWR, Shanghai, China) at a density of 5,000 cells/cm2, also in a mesenchymal stem cell growth medium for 24 h. Then, the culture medium was changed to hMSC Osteogenic Differentiation Medium (BulletKit, Lonza, Shanghai, China) and osteogenic differentiation was induced for 14 days according to the manufacturer’s recommendation.
RNA isolation and quantitative real-time reverse transcription-PCR (qRT-PCR)
Total RNA was extracted from differentiated hMSCs using an RNAqueous-4PCR Total RNA Isolation Kit (Thermo Fisher Scientific, Shanghai, China) and converted to first-strand cDNA using a SuperScript VILO cDNA Synthesis Kit (Thermo Fisher Scientific, Shanghai, China), according to the manufacturer’s recommendations. Quantitative real-time reverse transcription-PCR (qRT-PCR) was conducted on a QuantStudio 5 Real-Time PCR Systems (Thermo Fisher Scientific, Shanghai, China) using an SYBR Premix Ex Taq Kit (TaKaRa, China) according to the manufacturer’s recommendations and U6 snRNA was used for internal control. For lncRNA detections, all forward and reverse primers were designed and purchased from CapitalBio (CapitalBio, Beijing, China). The primers for hsa-miR-214-3p were forward, F 5′-TGCCTGTCTACACTTGCTGTGC-3′ & reverse, 5′-GCGAGCACAGAATTAATACGAC-3′, for BMP2 were forward, 5′-ACTCGAAATTCCCCGTGACC-3′ & reverse, 5′-CCACTTCCACCACGAATCCA-3′; for Runx2 were forward, 5′-CGAATAACAGCACGCTATTAA-3′ & reverse, 5′-GTCGCCAAACAGATTCATCCA-3′, and for U6 snRNA were forward 5′-CTCGCTTCGGCAGCACA-3′ & reverse 5′-AACGCTTCACGAATTTGCGT-3′. Finally, relative expression levels were characterized using the 2−∆∆Ct method.
Alkaline phosphatase (ALP) assay
Differentiated hMSCs were lysed using a Protein Extraction Kit (Abcam, Shanghai, China), and the total protein contents were determined using a NanoOrange™ Protein Quantitation Kit (Thermo Fisher Scientific, Shanghai, China), according to the manufacturers’ recommendations. Then, alkaline phosphatase (ALP) Assay was conducted using an Alkaline Phosphatase Assay Kit (Fluorometric, Abcam, Shanghai, China) according to the manufacturer’s recommendation. Briefly, an equal amount of protein lysate (10 uL) was collected from each sample and incubated with 20 μl non-fluorescent 4-Methylumbelliferyl phosphate disodium salt (MUP) medium in a 96-well plate (VWR, Shanghai, China) for 30 min at room temperature (RT). Then, the plate was mounted on a Varioskan Lux Microplate Reader (Thermo Fisher Scientific, Shanghai, China) and fluorescence was measured at extension/emission of 360/440 nm. The relative ALP activities were normalized to the activities under control conditions.
Alizarin red S (ARS) quantification assay
The method of quantitatively measuring Alizarin Red S (ARS) activity among differentiated hMSCs was described previously [24], and conducted using an ARS staining quantification assay (ScienCell Research Laboratories, San Diego, CA, USA) according to the manufacturer’s recommendation. Briefly, differentiated hMSCs were washed 3 times with 1X PBS (Thermo Fisher Scientific, Shanghai, China), fixed with 4% formaldehyde (MilliporeSigma, Shanghai, China) for 15 min at RT, and then stained with ARS dye for 30 min at RT. After washing 5 times with diH2O (MilliporeSigma, Shanghai, China), 400 µL of 10% acetic acid (MilliporeSigma, Shanghai, China) was added into wells for 30 min at RT. Then, cells were collected, centrifuged and heated at 85 °C for 10 min. After centrifugation, 200 uL supernatant of each sample was loaded in a 96-well plate and neutralized with 75 uL ammonium hydroxide (MilliporeSigma, Shanghai, China). Then, the plate was mounted on a Varioskan Lux Microplate Reader (Thermo Fisher Scientific, Shanghai, China) and absorbance was measured at 405 nm. The relative ARS activities were normalized to the activities under control conditions.
LncRNA knockdown assay
The small interfering RNA (shRNAs) specifically against human MEG3 (sh_MEG3), SNHG16 (sh_SNHG16), FAM83H-AS1 (sh_FAM83HAS1), MALAT1 (sh_MALAT1) and LINC00657 (sh_657) were designed and manufactured by Genepharma (Genepharma, Shanghai, China). In addition, a non-specific control snRNA (sh_C) was also purchased from Genepharma. The method of shRNA transfection was described in previous publications [25,26,27]. Briefly, The differentiated hMSCs were transfected twice with shRNAs, one on day 1 and the other on day 7, using Lipofectamine 3000 reagents (Thermo Fisher Scientific, Shanghai, China). qRT-PCR was used to verify the knockdown efficiency.
RNA immunoprecipitation (RIP) assay
RNA immunoprecipitation (RIP) assay was conducted using a Magna RIP kit (MilliporeSigma, Shanghai, China) according to the manufacturer's recommendation. Briefly, on day 14, differentiated hMSCs were collected and lysed using the RIP lysis buffer. Then, cell lysates were treated with a rabbit polyclonal anti-human BMP2 antibody (1:500, Abcam, Shanghai, China) or a control IgG (1:2000, MilliporeSigma, Shanghai, China) in RIP buffer for 4 h at 4 °C, followed by another treatment of Proteinase K (MilliporeSigma, Shanghai, China) for 30 min at 55 °C. Finally, total RNA was extracted from immunoprecipitated precipitates using an RNAqueous-4PCR Total RNA Isolation Kit (Thermo Fisher Scientific, Shanghai, China), followed by qRT-PCR assays to probe MEG3, SNHG16, FAM83H-AS1, MALAT1 and LINC00657.
Hsa-miR-214-3p knockdown assay
The synthetic human miRNA inhibitor specifically targeting hsa-miR-214-3p (miR214_I) was designed and manufactured by Genepharma (Genepharma, Shanghai, China). In addition, a non-specific human miRNA inhibitor (miRNC_I) was also purchased from Genepharma. The method of shRNA transfection was described in previous publications [28, 29]. The differentiated hMSCs were transfected twice with miRNA inhibitor, one at day 2 and the other at day 8, using Lipofectamine 3000 reagents (Thermo Fisher Scientific, Shanghai, China). qRT-PCR was used to verify the knockdown efficiency.
Statistical analysis
Statistical analysis was conducted using a two-tailed Student's t-test on SPSS software (Version 21.0, SPSS, Chicago, IL, USA). All experiments were conducted in three or more independent repeats. All data were demonstrated as the mean ± SEM p values < 0.05 were considered statistically significant.
Results
Characterizations of hsa-miR-214-3p/BMP2 epigenetic axis and osteogenic differentiation in hMSCs
Human mesenchymal stem cells (hMSCs) were induced with osteogenic differentiation for 14 days (Fig. 1A). On day 1 and day 14, cells were collected and compared by qRT-PCR. It showed that, hsa-miR-214-3p was downregulated, whereas BMP2 and Runx2 were upregulated in hMSCs from day 1 to day 14 (Fig. 1B–D, *p < 0.05). It also showed that, ALP and Alizarin Red S activities were significantly upregulated in differentiated hMSCs from day 1 to day 14 (Fig. 1E–G, *p < 0.05).
Osteogenic differentiation of hMSCs. A Human mesenchymal stem cells (hMSCs) were self-renewed by incubating with the mesenchymal stem cell growth medium. hMSCs were also replated, and induced with hMSC osteogenic differentiation medium for 14 days. One day 1 and day 14, cells were collected. qRT-PCR was conducted to examine expression levels of hsa-miR-214-3p, BMP2, Runx2 and lncRNAs. Biochemical assays were also performed to measure alkaline phosphatase (ALP) and Alizarin Red S activities. B, C, D qRT-PCR was conducted to compare the expression levels of miR-214-3p (B), BMP2 (C) and Runx2 (D) between day 1 and day 14 differentiated hMSCs (*p < 0.05). E alkaline phosphatase (ALP) activities were compared between day 1 and day 14 differentiated hMSCs (*p < 0.05). F Alizarin Red S activities were compared between day 1 and day 14 differentiated hMSCs (*p < 0.05). F Alizarin Red staining was compared between day 1 and day 14 differentiated hMSCs
Characterizations of upregulated lncRNAs upstream to hsa-miR-214-3p during osteogenic differentiation in hMSCs
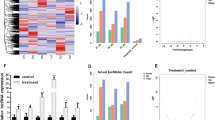
In a previous study, lncRNA of KCNQ1OT1 was demonstrated to be upstream to hsa-miR-214-3p / BMP2 epigenetic axis [14]. In this study, we used the bioinformatic search tool LncBase Experimental V.2 [30], to examine 145 lncRNAs which were experimentally characterized to be upstream to hsa-miR-214-3p (the list of 145 lncRNAs was included in the supplement document of “supplement01”). qRT-PCR analysis was conducted and we focused on those lncRNAs which were upregulated at least 2 times from day 1 to day 14 in differentiated hMSCs. As the result, we found six candidate lncRNAs, KCNQ1OT1, maternally expressed 3 (MEG3), small Nucleolar RNA Host Gene 16 (SNHG16), FAM83H antisense RNA 1 (FAM83H-AS1), Metastasis Associated Lung Adenocarcinoma Transcript 1( MALAT1) and long intergenic non-protein coding RNA 657 (LINC00657), who could bind hsa-miR-214-3p (Fig. 2A) and were markedly upregulated in differentiated hMSCs from day 1 to day 14 (Fig. 2B, *p < 0.05).
LncRNAs upstream to hsa-miR-214-3p are significantly upregulated in osteogenic differentiation of hMSCs. A Diagrams were shown for the bindings of KCNQ1 Opposite Strand/Antisense Transcript 1 (KCNQ1OT1), maternally expressed 3 (MEG3), small Nucleolar RNA Host Gene 16 (SNHG16), FAM83H antisense RNA 1 (FAM83H-AS1), long intergenic non-protein coding RNA 657 (LINC00657) on hsa-miR-214-3p. B Between day 1 and day 14 differentiated hMSCs, qRT-PCR was conducted to compare expression levels of KCNQ1OT1, MEG3, SNHG16, FAM83H-AS1 and LINC00657 (*p < 0.05)
Downregulations of lncRNAs upstream to hsa-miR-214-3p suppressed osteogenic differentiation in hMSCs
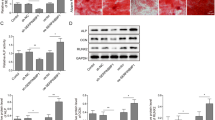
We used genetic engineering methods to downregulate MEG3, SNHG16, FAM83H-AS1 and LINC00657 in differentiated hMSCs. On day 14 of osteogenic differentiation, for each lncRNA, qRT-PCR confirmed that its endogenous expression level was specifically and significantly downregulated by transfection of corresponding shRNAs (Fig. 3A, *p < 0.05; ∆p > 0.05). Then, quantitative assessments demonstrated that characteristic features of osteogenic differentiation, including BMP2 expression level (Fig. 3B), Runx2 expression level (Fig. 3C), ALP activities (Fig. 3D) and Alizarin Red S activities (Fig. 3E) were all substantially suppressed by lncRNA downregulations (*p < 0.05; ∆p > 0.05).
The effects of LncRNAs downregulation on osteogenic differentiation of hMSCs. A Differentiated hMSCs were transfected with lncRNA-specific shRNAs, including sh_MEG3, sh_SNHG16, sh_MALAT1, sh_FAM83HAS1 and sh_657, and a non-specific shRNA, sh_C. At day 14 of differentiation, qRT-PCR was conducted to compare the expression levels of MEG3, SNHG16, AMALT1, FAM83H-AS1 and LINC00657 among un-transfected hMSCs, hMSCs transfected with sh_C and hMSCs transfected with lncRNA-specific shRNAs (*p < 0.05; ∆p < 0.05). B and C At day 14, qRT-PCR was conducted in differentiated hMSCs to compare the expression levels of BMP2 (B) and Runx2 (C) among un-transfected hMSCs, hMSCs transfected with sh_C and hMSCs transfected with lncRNA-specific shRNAs (*p < 0.05; ∆p < 0.05). D At day 14, ALP activities were compare among un-transfected hMSCs, hMSCs transfected with sh_C and hMSCs transfected with lncRNA-specific shRNAs (*p < 0.05; ∆p < 0.05). E At day 14, Alizarin Red S activities were compare among un-transfected hMSCs, hMSCs transfected with sh_C and hMSCs transfected with lncRNA-specific shRNAs (*p < 0.05; ∆p < 0.05)
MALAT1 and LINC00657 were directly associated with hsa-miR-214-3p / BMP2 axis in osteogenic differentiation of hMSCs
We further investigated the direct association between lncRNAs and hsa-miR-214-3p / BMP2 axis in osteogenic differentiation of hMSCs.
First, on day 14 of osteogenic differentiation, qRT-PCR showed that, downregulations of MALAT1 and LINC00657 upregulated hsa-miR-214-3p whereas downregulation of MEG3, SNHG16 or FAM83HAS1 had no effect on hsa-miR-214-3p expression in hMSCs (Fig. 4A, *p < 0.05; ∆p > 0.05).
The association between LncRNAs and hsa-miR-214-3p / BMP2 axis in osteogenic differentiation of hMSCs. A Differentiated hMSCs were transfected with lncRNA-specific shRNAs, including sh_MEG3, sh_SNHG16, sh_MALAT1, sh_FAM83HAS1 and sh_657, and a non-specific shRNA, sh_C. At day 14 of differentiation, qRT-PCR was conducted to compare their expression levels of hsa-miR-214-3p (*p < 0.05; ∆p < 0.05). B Diagrams were shown for the DNA bindings of hsa-miR-214-3p on BMP2. C At day 14 of differentiation, a RIP assay was conducted using BMP2 and IgG antibodies, followed by qRT-PCR to examine the expression levels of MEG3, SNHG16, AMALT1, FAM83H-AS1 and LINC00657 (*p < 0.05; ∆p < 0.05)
Second, the correlation between lncRNAs and BMP2 in osteogenic differentiation of hMSCs was investigated. In a previous study, it was demonstrated that hsa-miR-214-3p could directly bind BMP2 to form hsa-miR-214-3p / BMP2 axis [14] (Fig. 4B). Through an RNA immunoprecipitation (RIP) assay, we demonstrated that MALAT1 and LINC00657 were significantly enriched in the samples pulled down by BMP2 antibody, whereas MEG3, SNHG16 or FAM83HAS1 were not specifically associated with BMP2 (Fig. 4C, *p < 0.05; ∆p > 0.05).
MALAT1 and LINC00657 regulated osteogenic differentiation of hMSCs through hsa-miR-214-3p
Finally, we silenced hsa-miR-214-3p in differentiated hMSCs which were pre-transfected with lncRNAs-specific shRNAs. qRT-PCR confirmed that, at day 14 of differentiation, hsa-miR-214-3p was significantly downregulated in cells transfected with miR214_I, as compared to cells transfected with miRNC_I (Fig. 5A, *p < 0.05).
The effects of silencing hsa-miR-214-3p on lncRNA-associated osteogenic differentiation of hMSCs. A Differentiated hMSCs, which were transfected with lncRNA-specific shRNAs, were further transfected with a hsa-miR-214-3p specific inhibitor (miR214_I), or a non-specific control miRNA inhibitor (miRNC_I). qRT-PCR analysis showed that hsa-miR-214-3p was significantly silenced in cells transfected with miR214_I, than in cells transfected with miRNC_I (*p < 0.05). B and C At day 14 of differentiation, qRT-PCR was conducted to compare gene expression levels of BMP2 (B) and Runx2 (C) between differentiated hMSCs which were transfected with lncRNA-specific shRNAs only (shRNA_only), and those transfected with lncRNA-specific shRNAs and then double-transfected with miRNC_I or miR214_I (*p < 0.05; ∆p < 0.05). D At day 14 of differentiation, ALP activities were compared between differentiated hMSCs which were transfected with lncRNA-specific shRNAs only (shRNA_only), and those transfected with lncRNA-specific shRNAs and then double-transfected with miRNC_I or miR214_I (*p < 0.05; ∆p < 0.05). E At day 14 of differentiation, Alizarin Red S activities were compare between differentiated hMSCs which were transfected with lncRNA-specific shRNAs only (shRNA_only), and those transfected with lncRNA-specific shRNAs and then double-transfected with miRNC_I or miR214_I (*p < 0.05; ∆p < 0.05)
Then, double-transfected hMSCs were examined with characteristic features of osteogenic differentiation at day 14. qRT-PCR showed that, hsa-miR-214-3p knockdown only induced BMP2 and Runx2 upregulations in cells with MALAT1 or LINC00657 downregulations (Fig. 5B, C, *p < 0.05; ∆p > 0.05). In addition, hsa-miR-214-3p knockdown only induced ALP and Alizarin Red S activity upregulations in cells with MALAT1 or LINC00657 downregulations (Fig. 5B, C, *p < 0.05; ∆p > 0.05).
Thus, our data indicated that MALAT1 and LINC00657, not MEG3, SNHG16 or FAM83H-AS1, regulated osteogenic differentiation of hMSCs through hsa-miR-214-3p.
Discussion
In this study, we investigated the lncRNA profiles upstream to hsa-miR-214-3p / BMP2 axis during the osteogenic differentiation of hMSCs.
First, we induced hMSCs toward osteogenic differentiation for 14 days in vitro, and used multiple quantitative assays to confirm this process. Notably, key genetic transcripts of osteogenic differentiation, BMP2 and Runx2 [23, 31], were confirmed by qRT-PCR to be significantly upregulated. In addition, we used biochemical methods to confirm that ALP and Alizarin Red S activities, which were standard cellular indicators of mineralization and osteogenesis [32, 33], were also markedly upregulated.
In a previous study, it was revealed that lncRNA of KCNQ1OT1 was upstream to hsa-miR-214-3p / BMP2 axis and functionally regulated osteogenic induction of bone marrow mesenchymal stem cells [14]. In addition, it was demonstrated that increased osteoclastic miR-214-3p was associated with both elevated serum exosomal miR-214-3p and reduced bone formation in elderly women with fractures and in ovariectomized (OVX) mice [34]. Also, a very recent study showed that miR-214-3p was downregulated in periodontal ligament stem cells after osteogenic induction [35]. Based on these information, the strategy of this study was to further explore the lncRNA candidates upstream to hsa-miR-214-3p / BMP2 axis, thus broadening our knowledge on epigenetic regulations in osteogenesis. Using qRT-PCR analysis, we examined 145 lncRNAs, which were experimentally confirmed to be able to bind hsa-miR-214-3p [30] (the list of 145 lncRNAs was included in the supplement document of “supplement01”). Among them, we identified six lncRNAs, KCNQ1OT1, MEG3, SNHG16, FAM83H-AS1, MALAT1 and LINC00657 to be upregulated at least two folders in hMSCs from day 1 to day 14 of osteogenic differentiation. The result regarding KCNQ1OT1 was the confirmation to the previous study [14], whereas the findings on the other five lncRNAs were novel and we decided to further investigate them.
Next, through qRT-PCR and quantitative biochemical assays, we found that, inhibiting MEG3, SNHG16, FAM83H-AS1, MALAT1 and LINC00657 could all functionally suppress osteogenic differentiation in hMSCs, demonstrated by downregulated BMP2 and Runx2 expressions, as well as decreased ALP and Alizarin Red S activities. Interestingly, while we further looked into hsa-miR-214-3p expressions in lncRNAs-downregulated hMSCs, we discovered that only MALAT1 and LINC00657 upregulated hsa-miR-214-3p, whereas MEG3, SNHG16 and FAM83H-AS1 did not. In addition, qRT-PCR analysis following RIP assay with BMP2-specific antibody showed that only MALAT1 and LINC00657 were enriched in pull-down precipitates, whereas MEG3, SNHG16 and FAM83H-AS1 were not. In previous studies, it was demonstrated MEG3 correlated with miR-133a-3p and miR-140-5p [13, 36], SNHG16 correlated with miR-485 [11], and FAM83H-AS1 correlated miR-541-3p [15] in osteogenic development of mesenchymal stem cells. Thus, these evidences strongly indicated that in hMSCs, MALAT1 and LINC00657 regulated osteogenic differentiation through hsa-miR-214-3p / BMP2 axis, whereas MEG3, SNHG16 and FAM83H-AS1 may act through other downstream miRNA-mRNA signaling pathways.
This notion was further supported in our loss-of-function probe while we further silenced hsa-miR-214-3p in lncRNAs-downregulated hMSCs. Our data demonstrated that, silencing hsa-miR-214-3p could augment BMP2 and Runx2 expressions, as well as ALP and Alizarin Red S activities in MALAT1- or LINC00657- downregulated hMSCs, but not in MEG3-, SNHG16- or FAM83H-AS1- downregulated hMSCs, thus confirming that MALAT1 and LINC00657 were upstream of hsa-miR-214-3p / BMP2, whereas MEG3, SNHG16 or FAM83H-AS1 was not.
In summary, we demonstrated five lncRNAs (in addition to KCNQ1OT1) to be significantly upregulated in osteogenic induction of hMSCs, and further identified MALAT1 and LINC00657 to be upstream of hsa-miR-214-3p / BMP2 to functionally regulate osteogenic differentiation. Future studies are needed to fully explore the functional roles of epigenetic regulations, as well as their downstream effects on gene transcriptions and protein translations in osteogenic differentiation.
Data availability
All data were released in this study.
Abbreviations
- hMSC:
-
Human mesenchymal stem cell
- ALP:
-
Alkaline phosphatase
- lncRNAs:
-
Long noncoding RNAs
- 3′-UTR:
-
Three prime untranslated region
- KCNQ1OT1:
-
KCNQ1 Opposite Strand/Antisense Transcript 1
- BMP2:
-
Bone morphogenetic protein 2
- MALAT1:
-
Metastasis Associated Lung Adenocarcinoma Transcript 1
- LINC00657:
-
Long intergenic non-protein coding RNA 657
References
Olsen BR, Reginato AM, Wang W (2000) Bone development. Annu Rev Cell Dev Biol 16:191–220
Berendsen AD, Olsen BR (2015) Bone development. Bone 80:14–18
Pittenger MF, Mackay AM, Beck SC, Jaiswal RK, Douglas R, Mosca JD, Moorman MA, Simonetti DW, Craig S, Marshak DR (1999) Multilineage potential of adult human mesenchymal stem cells. Science 284(5411):143–147
Abdallah BM, Kassem M (2008) Human mesenchymal stem cells: from basic biology to clinical applications. Gene Ther 15(2):109–116
Kronenberg HM (2003) Developmental regulation of the growth plate. Nature 423(6937):332–336
Scotti C, Tonnarelli B, Papadimitropoulos A, Scherberich A, Schaeren S, Schauerte A, Lopez-Rios J, Zeller R, Barbero A, Martin I (2010) Recapitulation of endochondral bone formation using human adult mesenchymal stem cells as a paradigm for developmental engineering. Proc Natl Acad Sci USA 107(16):7251–7256
Pountos I, Corscadden D, Emery P, Giannoudis PV (2007) Mesenchymal stem cell tissue engineering: techniques for isolation, expansion and application. Injury 38(Suppl 4):S23-33
Kwok ZH, Tay Y (2017) Long noncoding RNAs: lincs between human health and disease. Biochem Soc Trans 45(3):805–812
Maass PG, Luft FC, Bahring S (2014) Long non-coding RNA in health and disease. J Mol Med (Berl) 92(4):337–346
Tang S, Xie Z, Wang P, Li J, Wang S, Liu W, Li M, Wu X, Su H, Cen S et al (2019) LncRNA-OG promotes the osteogenic differentiation of bone marrow-derived mesenchymal stem cells under the regulation of hnRNPK. Stem Cells 37(2):270–283
Asila A, Yang X, Kaisaer Y, Ma L (2021) SNHG16/miR-485–5p/BMP7 axis modulates osteogenic differentiation of human bone marrow-derived mesenchymal stem cells. J Gene Med 23(3):e3296
Huang XZ, Huang J, Li WZ, Wang JJ, Song DY, Ni JD (2020) LncRNA-MALAT1 promotes osteogenic differentiation through regulating ATF4 by sponging miR-214: Implication of steroid-induced avascular necrosis of the femoral head. Steroids 154:108533
Li Z, Jin C, Chen S, Zheng Y, Huang Y, Jia L, Ge W, Zhou Y (2017) Long non-coding RNA MEG3 inhibits adipogenesis and promotes osteogenesis of human adipose-derived mesenchymal stem cells via miR-140-5p. Mol Cell Biochem 433(1–2):51–60
Wang CG, Liao Z, Xiao H, Liu H, Hu YH, Liao QD, Zhong D (2019) LncRNA KCNQ1OT1 promoted BMP2 expression to regulate osteogenic differentiation by sponging miRNA-214. Exp Mol Pathol 107:77–84
Wu H, Cao F, Zhou W, Wang G, Liu G, Xia T, Liu M, Mi B, Liu Y (2020) Long Noncoding RNA FAM83H-AS1 Modulates SpA-inhibited osteogenic differentiation in human bone mesenchymal stem cells. Mol Cell Biol 40(5).
Dalmay T (2013) Mechanism of miRNA-mediated repression of mRNA translation. Essays Biochem 54:29–38
Chekulaeva M, Filipowicz W (2009) Mechanisms of miRNA-mediated post-transcriptional regulation in animal cells. Curr Opin Cell Biol 21(3):452–460
Kim YJ, Bae SW, Yu SS, Bae YC, Jung JS (2009) miR-196a regulates proliferation and osteogenic differentiation in mesenchymal stem cells derived from human adipose tissue. J Bone Miner Res 24(5):816–825
Zhang JF, Fu WM, He ML, Xie WD, Lv Q, Wan G, Li G, Wang H, Lu G, Hu X et al (2011) MiRNA-20a promotes osteogenic differentiation of human mesenchymal stem cells by co-regulating BMP signaling. RNA Biol 8(5):829–838
Eskildsen T, Taipaleenmaki H, Stenvang J, Abdallah BM, Ditzel N, Nossent AY, Bak M, Kauppinen S, Kassem M (2011) MicroRNA-138 regulates osteogenic differentiation of human stromal (mesenchymal) stem cells in vivo. Proc Natl Acad Sci USA 108(15):6139–6144
Guo L, Zhao RC, Wu Y (2011) The role of microRNAs in self-renewal and differentiation of mesenchymal stem cells. Exp Hematol 39(6):608–616
Ryoo HM, Lee MH, Kim YJ (2006) Critical molecular switches involved in BMP-2-induced osteogenic differentiation of mesenchymal cells. Gene 366(1):51–57
Chen Z, Wu C, Gu W, Klein T, Crawford R, Xiao Y (2014) Osteogenic differentiation of bone marrow MSCs by beta-tricalcium phosphate stimulating macrophages via BMP2 signalling pathway. Biomaterials 35(5):1507–1518
Gregory CA, Gunn WG, Peister A, Prockop DJ (2004) An Alizarin red-based assay of mineralization by adherent cells in culture: comparison with cetylpyridinium chloride extraction. Anal Biochem 329(1):77–84
Xie P, Cao H, Li Y, Wang J, Cui Z (2017) Knockdown of lncRNA CCAT2 inhibits endometrial cancer cells growth and metastasis via sponging miR-216b. Cancer Biomark 21(1):123–133
Hu X, Ding D, Zhang J, Cui J (2019) Knockdown of lncRNA HOTAIR sensitizes breast cancer cells to ionizing radiation through activating miR-218. Biosci Rep 39(4).
Zhang C, Bao C, Zhang X, Lin X, Pan D, Chen Y (2019) Knockdown of lncRNA LEF1-AS1 inhibited the progression of oral squamous cell carcinoma (OSCC) via Hippo signaling pathway. Cancer Biol Ther 20(9):1213–1222
Schoolmeesters A, Eklund T, Leake D, Vermeulen A, Smith Q, Force Aldred S, Fedorov Y (2009) Functional profiling reveals critical role for miRNA in differentiation of human mesenchymal stem cells. PLoS ONE 4(5):e5605
Tome M, Lopez-Romero P, Albo C, Sepulveda JC, Fernandez-Gutierrez B, Dopazo A, Bernad A, Gonzalez MA (2011) miR-335 orchestrates cell proliferation, migration and differentiation in human mesenchymal stem cells. Cell Death Differ 18(6):985–995
Paraskevopoulou MD, Vlachos IS, Karagkouni D, Georgakilas G, Kanellos I, Vergoulis T, Zagganas K, Tsanakas P, Floros E, Dalamagas T et al (2016) DIANA-LncBase v2: indexing microRNA targets on non-coding transcripts. Nucleic Acids Res 44(D1):D231-238
Bruderer M, Richards RG, Alini M, Stoddart MJ (2014) Role and regulation of RUNX2 in osteogenesis. Eur Cell Mater 28:269–286
Siffert RS (1951) The role of alkaline phosphatase in osteogenesis. J Exp Med 93(5):415–426
Chen CH, Ho ML, Chang JK, Hung SH, Wang GJ (2005) Green tea catechin enhances osteogenesis in a bone marrow mesenchymal stem cell line. Osteoporos Int 16(12):2039–2045
Li D, Liu J, Guo B, Liang C, Dang L, Lu C, He X, Cheung HY, Xu L, Lu C et al (2016) Osteoclast-derived exosomal miR-214-3p inhibits osteoblastic bone formation. Nat Commun 7:10872
Feng Y, Wan P, Yin L (2020) Long noncoding RNA X-inactive specific transcript (XIST) promotes osteogenic differentiation of periodontal ligament stem cells by sponging microRNA-214–3p. Med Sci Monit 26:e918932
Wang Q, Li Y, Zhang Y, Ma L, Lin L, Meng J, Jiang L, Wang L, Zhou P, Zhang Y (2017) LncRNA MEG3 inhibited osteogenic differentiation of bone marrow mesenchymal stem cells from postmenopausal osteoporosis by targeting miR-133a-3p. Biomed Pharmacother 89:1178–1186
General Assembly of the World Medical A (2014) World Medical Association Declaration of Helsinki: ethical principles for medical research involving human subjects. J Am Coll Dent 81(3):14–18.
Acknowledgements
N/A.
Funding
None.
Author information
Authors and Affiliations
Contributions
Conceptualization: YT. Methodology: JL, HZ, ZW, YT. Validation: YT. Formal analysis and investigation: JL, HZ, ZW, ZC, XM, WC, XJ, DZ, WH. Writing—original draft preparation: JL. Writing—review and editing: PW. Resources: YT. Supervision: YT.
Corresponding author
Ethics declarations
Conflict of interest
None.
Ethical approval and consent to participate
The approval to conduct this study was reviewed and approved by the Human Research Ethic Committees at the Liaocheng Chinese Medical Hospital, Shandong University of Traditional Chinese Medicine and Yanggu People’s Hospital, in Shandong Province, China. In addition, all protocols were conducted in accordance with the principles of the Declaration of Helsinki (2013) [37]. In addition, since there was no data or samples of human patients in this study, there was no action to obtain informed consent, written or verbal, from all participants.
Consent for publication
All authors consent on the publication of this study.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Li, J., Zhuang, H., Wang, Z. et al. lncRNAs MALAT1 and LINC00657 upstream to miR-214-3p/BMP2 regulate osteogenic differentiation of human mesenchymal stem cells. Mol Biol Rep 49, 6847–6857 (2022). https://doi.org/10.1007/s11033-022-07136-3
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11033-022-07136-3