Abstract
Genus Ocimum is known to have species possessing important therapeutic essential oil. The major phytoconstituents of essential oil in Ocimum species are phenylpropanoids and terpenoids. The essential oil is accumulated in the trichomes; the specialized structures predominantly found on leaves and other tissues. The development of trichome is integrated with development of plant and leaf and also tightly coordinated with the primary and secondary metabolic pathways producing essential oil constituents. In continuation to our studies on elucidating/understanding the mechanism of biosynthesis of essential oil pathways in Ocimum species, we have performed comparative transcriptome analysis to investigate the role of trichome-related gene expression in the regulation of biosynthetic pathways of essential oil. The essential oil biogenesis is tightly integrated with primary metabolic activities, the analysis for the expression pattern of genes related to primary metabolism and its relationship with secondary metabolism was evaluated in comparative manner. Physiological parameters in relation to primary metabolism such as photosynthetic pigment content, soluble sugar content, and invertase enzymes along with morphological parameters were analysed in O. basilicum and O. sanctum. Differential expression profiling uncovered about 8116 and 2810 differentially expressed transcripts in O. basilicum and O. sanctum, respectively. Enrichment of differentially expressed genes were analysed in relation to metabolic pathways, primary metabolism and secondary metabolism. Trichome related genes identified from the Ocimum species vis-à-vis their expression profiles suggested higher expression in O. basilicum. The findings in this study provide interesting insights into the role of trichome-related transcripts in relation to essential oil content in Ocimum species. The study is valuable as this is the first study on revealing the transcripts and their role in trichome development and essential oil biogenesis in two major species of Ocimum.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Ocimum genus is highly diverse and comprises of more than 60 species [1]. The genus is distributed all over the Asian and African sub-continent, with intra and inter-specific genetic diversity [2]. Amongst these, Ocimum basilicum and Ocimum sanctum are extensively used for specialized secondary metabolites found in their essential oils in medicinal and aroma industries. Recent efforts on plant metabolites are majorly focused on defining the metabolic pathways for the synthesis of secondary metabolites in cells. Pathways are classified on the basis of origin and function as primary metabolic pathways and, secondary metabolic pathways. Primary metabolic processes and pathways are largely responsible for the survival of plants and provide “C” flux for the secondary metabolic pathways [3]. On the other hand, secondary metabolites are reported to have important ecological, developmental and defense related functions [4]. Metabolites derived from secondary metabolic pathways include fragrances, pesticides, flavours, food additives, dyes, pigments etc, with applications in pharmaceuticals, agriculture and industrial products for plant defense, pollination etc. [2, 5, 6]. Secondary metabolites are classified into major chemical classes of flavonoids, phenolics, alkaloids, terpenoids and phenylpropanoids [6].
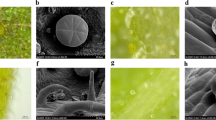
The secondary metabolites in Ocimum sp. are majorly documented from essential oil. The essential oil is sequestered in the characteristic anatomical structures known as glandular trichomes. Trichomes are epidermal outgrowths having specialized cells that have ability to secrete or store considerable quantities of specialized metabolites [7, 8]. Ocimum essential oils are reported to be widely used such as in treatment of stress, fever, cough relieving, arthritis, nervous disorders, and allergies, etc. [9, 10]. Due to these medicinal properties the market share of Ocimum oil is growing at an increasing pace. The essential oil of Ocimum species contains phenylpropanoids and terpenoids as major components of essential oils attributing specific functions for medicinal and aroma purposes. The diversity of Ocimum species studied by various groups at molecular and morphological levels [10, 11] however the mechanism and the factors responsible for variation in essential oil content and components are limited. Though some recent reports from our lab as well as from others have attempted to understand the regulation of essential oil accumulation [6, 11, 12, 33]. The Ocimum species differ distinctly in their composition such as in sesquiterpenoids, monoterpenoids as well as in phenylpropanoids based constituents in O. basilicum and in O. sanctum [6]. This observation prompted us to investigate transcriptomes of both the Ocimum species for transcripts responsible for essential oil sequestration in trichomes, the secretory glands. Transcription factor families such as R2R3MYB, bHLH, and C2H2 etc. have been shown to act as major regulators in the regulation of genes responsible for trichome development [13,14,15]. The transcription factors such as WRKY and AP2 have been associated with the secondary metabolite pathways in medicinal plants [16]. Expression of trichome related genes such as glabrous, caprice, enhancer of triptychon and transparent testa in coherence to each other influences the development, clustering, and density of the trichomes [17, 18].
Recent developments in the NGS technology platforms have facilitated the generation of huge transcriptomic resources. Platforms related to NGS have also created the opportunity to draw comparative profiles using the transcriptome data. The increased number of publications related to the comparative transcriptome studies, manifest its utilization in the investigation of pathways, gene mining, pathway networks and interaction studies [19,20,21,22,23].
Transcriptome analysis provides useful information regarding all major pathways in Ocimum. In the present study, transcriptome analysis is conducted to understand the variation in relation to trichome development related genes, which may be associated with the development of trichomes, sequestration of essential oil content and regulation of composition. To the best of our knowledge, the present work is first effort in the identification and investigation of trichomes related gene profiling in Ocimum species in relation to metabolic pathways. The findings of the present work will provide guidelines for future research directions in the understanding of the molecular regulation of trichome development and essential oil sequestration.
Results
Assembly of reads
The data of Ocimum sanctum and Ocimum basilicum transcriptomes was analysed from the SRA (https://www.ncbi.nlm.nih.gov/sra/). O. basilicum data with > 18 million reads (13.5 Gb) and O. sanctum data with > 16 million reads (12.2 Gb) was analysed and presented in Table 1. After initial adapter removal and filtering of raw reads, data was re-assembled using CLC genomics workbench 8.2 (https://digitalinsights.qiagen.com). The distribution of assembled transcripts was assessed in between 200 and 5000 bases. After proper assembling of the transcripts by taking optimum L50 values, average transcript lengths ranged from 1363 to 1964 bp for O. basilicum and O. sanctum, respectively.
Re-annotation of the transcriptomes
Comprehensive analyses of the O. basilicum and O. sanctum transcriptomes were carried out and assembled transcripts were annotated by using different bioinformatics tools, softwares and online web servers. On annotation by Blast2Go (B2G), it became clear that approximately 121,797 (85.06%) of transcripts were significantly annotated in O. basilicum while 65,851 (87.66%) transcripts were annotated in case of O. sanctum in comparison to previously published report [33]. Distributions of data were depicted for transcripts at various steps of assignment in O. basilicum and O. sanctum transcriptomes (Fig. 1a). Mapping of sequences, validated the evidence code which provides functional terms and information about the annotations either derived from automatic/computational or manually curated ones. O. basilicum and O. sanctum transcriptomes showed maximum curation with IEA (Inferred from electronic annotation) while second annotation were to IBA (Inferred from biological aspects of ancestor) and likewise from IDA (Inferred from direct assay), ISS (Inferred from sequence or structural similarity), ISM (Inferred from sequence model) etc. Mapping with the database source Ensembl Plants (63,942 & 85,434), ensembl (1867 & 3833), and Uniprot (1,080,118 & 1,541,565) etc, GO annotations were retreived, for O. basilicum and O. sanctum, respectively. 51,604 transcripts of O. basilicum and 52,152 transcripts of O. sanctum were mapped for the GO terms (Table 1).
On blastx with Uniprot and non-redundant database, transcripts from O. basilicum showed maximum hits with Erythranthe guttata (> 90,000 hits), followed by Sesamum indicum (> 70,000 hits), Nicotiana tabacum (> 60,000 hits) and Salvia militiorrhiza etc. Likewise, transcripts from O. sanctum showed maximum hits against the Sesamum indicum (> 90,000 hits), followed by Erythranthe guttata (> 70,000 hits), Nicotiana tabacum (> 60,000 hits) and Salvia militiorrhiza etc. maximum hits were with the order lamiales (Fig. 1b).
Global and secondary metabolism pathway analysis
To get the details of the genes with respect to the primary metabolism of the plant, standalone version of blast is used for identification of mined transcripts from the transcriptome. To establish the relation amongst global pathways (primary metabolism) and specialized pathways (secondary metabolism), in silico transcriptome screening was performed by isolating and mapping the transcripts for global and secondary metabolic pathways. Transcripts related to metabolism, degradation, synthesis, translation etc. were identified. Abundance of transcripts was more for O. sanctum as compared to O. basilicum for the pathways taken into consideration (Fig. 1c). In O. basilicum 1177 transcripts were assigned to the primary metabolic pathways and 3653 transcripts assigned from O. sanctum were assigned to the primary metabolic pathways. Total 89 transcripts related to photosynthesis were found in O. basilicum and 92 in O. sanctum. Likewise, Kreb's cycle, sucrose metabolism, lipid metabolism and signal transduction related transcripts were found to be abundant in O. sanctum. FPKM values were calculated to get an idea of the up/down regulation of primary and secondary metabolic pathways in both the Ocimum species (Fig. 2a, b). There were considerable differences in expression of transcripts, pathway genes related to the glycolysis & glyconeogenesis, transport and signal transduction were up regulated in O. basilicum. While genes related to photosynthesis, starch and sucrose metabolism were relatively upregulated in O. sanctum (Fig. 2a). Similarly, FPKM values indicate transcripts related to pathway of monoterpenoids, diterpenoids, flavonoids, and isoquinoline alkaloids were upregulated in O. basilicum, and sesquiterpenoids/triterpenoids were upregulated in O. sanctum transcriptome, respectively (Fig. 2b).
KEGG based classification showed transcripts mapping with metabolic pathways (KO1100), biosynthesis of secondary metabolites (KO1110), carbon metabolism (KO1200), fatty acid metabolism (KO01212), biosynthesis of amino acids (01230), 2-oxocarboxylic acid metabolism (01210) etc. Mapping with the KEGG pathway database retrieved 154 and 152 pathways for O. sanctum and O. basilicum respectively (Table 1). Major global pathways showed 8025 & 6038 transcripts in KEGG pathway with only 851 (10.95%) and 661 (10.60%) enzymes identified for the O. basilicum and O. sanctum. Likewise, biosynthesis of secondary metabolism retrieved 1524 and 1357 transcripts with 340 and 289 EC assigned for each pathway (Table 2).
Enzymatic classification of the transcripts in Ocimum species
In order to understand the enzyme distribution in Ocimum species, a comprehensive analysis was conducted for the major classes of the enzymes. Transcript's distribution against six categories of enzymes such as transferases, oxido-reductases, hydrolases, lyases, ligases and isomerases were retrieved. Results revealed that more than > 16,000 transcripts were assigned to the different enzyme classes with maximum being the 6295 for hydrolases and 5513 for transferases in O. sanctum, whereas relatively less number of transcripts were observed for these two classes in O. basilicum. Both O. basilicum and O. sanctum showed maximum hits with the hydrolases followed by the transferases (Fig. 2c). In hydrolases group, enzymes acting on ester bonds accounted for 348 transcripts in O. basilicum and 320 transcripts in O. sanctum, respectively. Esters considered as the phytochemicals responsible for the mild aroma for which maximum number of transcripts were assigned to the O. basilicum and relatively less to the O. sanctum. Within transferases maximum numbers of transcripts were assigned to transferring phosphorus groups approximately 733 sequences, followed by glycosyltransferases and acyltransferases. Oxidoreductases account for approximately 2500 transcripts followed by lyases, isomerases and ligases, in both the Ocimum species. Total 865 enzyme Ids were assigned to O. basilicum sequences and 922 enzyme Ids to O. sanctum after KEGG classification (Table 1).
Differential gene expression
Mapping of O. basilicum and O. sanctum transcriptomes examine the expression differences in the form of transcriptome abundance estimation. Satisfying logFC ≥ 2 and FDR ≤ 0.001, corrected P-value calculated by Benjamin-Hochberg procedure, a total of 8116 and 2810 DE (Differentially expressed) transcripts were observed (Table 1). In both the species, differential expression was high in O. basilicum.
Functional enrichment analysis of DE transcripts
Enrichment of GO terms and KEGG was conducted using KOBAS server to explore the biological functions of the DE transcripts. In the two-way comparison 77.9% (6330 of 8116) and 74.9% (2107 of 2810) in O. basilicum and O. sanctum respectively, transcripts were functionally annotated. Significant correlations were found for the 10 and 16 KEGG terms and 69 and 40 GO terms, for the O. basilicum and O. sanctum respectively (Table 1). For O. basilicum enriched KEGG terms were for metabolic pathways, biosynthesis of secondary metabolites, biosynthesis of amino acids, amino sugar and nucleotide sugar metabolism, biosynthesis of antibiotics, carbon metabolism, pentose phosphate pathway, porphyrin and chlorophyll metabolism, and in O. sanctum they were metabolic pathways, biosynthesis of antibiotics, biosynthesis of secondary metabolites, biosynthesis of amino acids, carbon metabolism, citrate cycle (TCA cycle), cysteine and methionine metabolism, fatty acid metabolism, biosynthesis of unsaturated fatty acids, ubiquitin mediated proteolysis, 2-oxocarboxylic acid metabolism, fatty acid biosynthesis, amino sugar and nucleotide sugar metabolism, lipoic acid metabolism, steroid biosynthesis and glyoxylate and dicarboxylate metabolism. For GO terms enrichment, maximum terms were placed in biological process category (32 & 18), followed by molecular function (20 & 15) and cellular component (17 & 7) for O. sanctum and O. basilicum respectively (Supplementary table 1). Network of the enriched terms on the basis of biological process predicted for the Ocimum species (Supplementary Fig. 1). Statistical validation of enrichment analysis of DE transcripts on the basis of P-value was also carried out.
Physiological studies in different developmental stages of O. basilicum and O. sanctum
Key physiological parameters (such as plant height, leaf length/width, internodal space, and inflorescence length of the mature plants were measured to study the basic physiological nature of the species. Plant height was found to be more in O. sanctum as compared to O. basilicum plants by 1.5 folds (Fig. 3a), remaining parameters such as inflorescence length and leaf length were recorded higher by 1.5–6.0 folds in O. basilicum as compared to O. sanctum. Total photosynthetic pigments content analysis in O. basilicum and O. sanctum at different developmental stages showed higher Chl a, Chl b, Chl (a + b) and carotenoid in mature plants of both species by 1.20–3.14 folds relative to seedlings. Further, these components were higher in O. sanctum in comparison to O. basilicum (Fig. 3c).
Physiological studies during developmental stages of Ocimum basilicum and O. sanctum. a Physiological parameters of the glass house-grown seedling plants of Ocimum species. b Total sugar and reducing sugar content in seedling and mature tissues of Ocimum species. c Total photosynthetic pigment content in Ocimum developmental stages. d Estimation of acid (grey), alkaline (black) and neutral (white) Invertase enzyme activity of Ocimum species; Ob = O. basilicum; Os = O. sanctum, O. basilicum seedling (ObS), O. basilicum mature (ObM), O. sanctum seedling (OsS) and O. sanctum mature (OsM)
Comparative analysis of sugar metabolism in different developmental stages of both Ocimum species showed significant variance in total sugars, reducing sugars and invertase enzyme activity. Total sugar content was higher by 1.25 folds in O. basilicum seedling as compared to O. sanctum seedling while in case of mature plants, O. sanctum showed higher sugar content by 2.12 folds than O. basilicum. O. basilicum seedling showed comparatively higher total reducing sugars (0.37 folds) than O. sanctum seedling. On comparison of O. basilicum mature with the O. sanctum mature, for reducing sugar levels latter exhibited 2.18 folds increased levels (Fig. 3b). Invertase enzyme activity was found maximum in acidic medium followed by neutral and alkali medium. Comparatively invertase activity was higher (approximately 1.5 folds) in O. basilicum seedlings as well as mature plants in comparison to O. sanctum (Fig. 3d).
Comparative analysis of essential oil yield showed distinct variation (0.10%–0.54%) in O. basilicum and O. sanctum at seedling and mature stage of plants (Supplementary Fig. 2).
Genes related to trichome development
Trichomes on the leaf surfaces are thought to be morphological characteristic that are involved in essential oil sequestration in Ocimum species. Trichomes play crucial role for the differential essential oil components in O. basilicum and O. sanctum. Transcriptomes of O. basilicum and O. sanctum were searched for the list of descriptions as for A. thaliana genes, approximately 120 genes are responsible for the trichome development [23]. BLASTx analysis of these genes showed 2087 transcripts matching in O. basilicum and 1333 in O. sanctum transcriptomes (Fig. 4a). 16 genes related to trichome development were identified, maximum number of transcripts were annotated for GLABROUS1 (GL1), and less transcripts annotated for Zinc finger protein 5 (ZFP5) in both the Ocimum species.
Real time quantification analysis of genes related to trichome development
Six genes out of 16 were chosen for the relative quantification analysis; GIS2 was only gene which showed higher expression in seedlings stage of O. sanctum (Fig. 4b). CPC, GL1, ETC3, GL3 and, TT8 were relatively higher expressed in O. basilicum seedling as well as mature stages of plant development. CPC gene showed higher expression in O. basilicum seedlings (Fig. 4b), while it showed higher expression in case of O. sanctum mature plants (Fig. 4c). Mapping of genes related to trichome against the KEGG database was also conducted (Supplementary Table 2). Majority of the genes isolated were unregulated according to FPKM values in O. basilicum and O. sanctum.
Discussion
A co-ordination of a number of processes controlling the partitioning of plant primary and secondary metabolisms, such as photosynthesis, TCA cycle, glycolysis, transport and terpenoid (mono/sesqui/tri), phenylpropanoid metabolism remains unclear. Release of CO2 is closely connected to the metabolic branching of the primary and secondary metabolism. Pyruvate acts as a substrate for various secondary metabolism pathways, despite being the major metabolite of the primary pathways [24,25,26,27]. Here, analysis of O. basilicum and O. sanctum transcriptome data and enrichment analysis of differential genes provided a broad distribution pattern of various transcripts belonging to different metabolic pathways either be primary or secondary metabolic pathways [6, 27].
Re-assembly and Re-annotation of transcriptomes for intra-species comparison
Previous studies have compared the transcriptomes of plants to date for the identification of expression pattern of genes, either across treatments [22, 23, 28, 29] or across single species ecotypes [27]. Due to the advent of the genomic resources and high throughput bioinformatics tools, comparative analyses of distant species transcriptome became possible [28]. Comparisons of species of same genus are rare, and they either use transcriptomic or genomic data of one of the species as a reference or of a related species [30, 31]. The following study re-analysed, re-assembled and re-annotated the transcriptomes of medicinally important species O. basilicum and O. sanctum. Earlier attempts for gene mining from the already reported transcriptomes have provided deeper insights into the pathway assignment, mapping, and gene discovery [16, 19, 21, 32]. Thus, this approach was useful in profiling novel aspects of secondary metabolism in MAPs [33, 16, 19, 23]. As genome of O. sanctum is available in the public domain and utilized in the study as the reference for making the genes full length. The genome of O. sanctum was merely used in this study as substrate to overlap the reads and assemble i.e. genome guided assembly [34]. The study uses the transcriptomes from the public domain and utilizes them as base for the mining of the genes related to the trichome development. A comprehensive comparative profile of the transcriptome of O. basilicum and O. sanctum were made, which provided high depth, quality and coverage. O. basilicum transcriptome accounted for more number of transcripts relative to O. sanctum. Only 12 and 9 sequences were left without analysis in case of O. basilicum and O. sanctum respectively. Assembly and annotated transcripts for nearly all the pathway genes of primary metabolic pathways and secondary metabolic pathways were obtained. The assembled transcripts were aligned with the protein sequences from the UniProt KB virdiplantae and mRNA sequences available for the different plant species at the nr database, which annotated the maximum transcripts and helped in isoform prediction. Maximum blast hits were obtained for the O. sanctum relative to O. basilicum when subjected to homology search with the plant databases. Annotation with uniprot database provided maximum hits with order lamiales, while significant hits were also observed with the member plants of family solanaceae, which may be due to the presence of large amount of data for the plants of this family in the databases. Evidence code distribution gives an idea about how many annotations were derived from automatic/computational annotations or from manually curated ones. Data curation showed that the transcripts from O. basilicum and O. sanctum were maximal aligned to the electronic annotation followed by the biological aspects. Ocimum species are known for their remarkable medicinal and aromatic properties [6, 10, 12], due to the presence of a varied mix of different metabolites. As an expansion of the previous study, In current work the global metabolism and secondary metabolism related pathways were analysed for O. basilicum and O. sanctum in comparative manner. Primary metabolites are linked to the secondary metabolites or it can be illustrated as majority of secondary metabolic pathways uses the product of primary metabolic pathways as their substrates [30, 34]. It was observed that photosynthesis, starch and signal transduction genes were showing more expression in O. sanctum relative to O. basilicum may be accounting for more metabolic activity in the former. High level of expression by glycolysis in O. basilicum may also imply to stress management. These results were in accordance to the previous studies on plants for regulation of genes involved in photosynthesis and chlorophyll biosynthesis [35, 36]. Further, more expression of pathway genes related to monoterpenoid, diterpenoid and flavonoid in O. basilicum relative to O. sanctum, may be leading to products of these pathways in former plant species. As revealed by our group previously that O. basilicum is phenylpropanoid rich species while sesquiterpenoids were found more in O. sanctum [6, 12, 13].
On enzyme classification basis, maximum numbers of transcripts were coded for hydrolases followed by transferases, and oxidoredutases in both the Ocimum species. More number of transferases in O. sanctum may be responsible for the formation of diverse and derivatives of metabolites including volatile esters [37,38,39]. Transferases are class of enzymes that enact the transfer of functional group from one molecule to another [38]. It has been shown in earlier studies that O. sanctum essential oil pool comprises higher diversity of metabolites as compared to O. basilicum [6, 12]. Likewise, transcripts for hydrolases, oxidoreductases, isomerases, and ligases were more abundant in O. sanctum. This may be due to some specific functions, which further require in-depth investigation to explore their specific roles. The GO ontology repository was used for assigning the molecular function, biological role, and cellular component of the large number of genes [31]. Significant enhancement in annotated data was done by merging the annotations with interproscan analysis results. Maximum numbers of transcripts were assigned to the molecular function as shown by the GO assignments followed by the biological process and cellular component.
In the functional analysis of DE transcripts between the two parental species, up-regulated transcripts in the transcriptome of O. sanctum were enriched with more number of pathway functions associated with biosynthesis and metabolism (i.e. ‘Citrate cycle’, ‘cysteine and methionine’, ‘fatty acid’, ‘unsaturated fatty acids’, ‘proteolysis’ and ‘2-oxocarboxylic acid’) than O. basilicum. This may explain the better production of secondary metabolites, and better divergence of primary carbon flux to the secondary metabolism pathways. Network linking the primary metabolite pathway with the secondary metabolite were predicted with the help of enriched genes, which showed the interdependence of these pathways. We note, however, that up-regulated genes do not necessarily confer better performance or better production of secondary metabolites, and the true implications could only be established through future functional studies. Similar kind of differential expression studies on transcriptome are also conducted in the recent past [21, 40].
The primary metabolic pathways being basic pathways provided the ‘C’ source and energy to the plants for plant growth and development. Validation of the findings with the estimation of total photosynthetic content of Ocimum at different developmental stages showed that the O. basilicum exhibited lower chlorophyll content and carotenoids. In fact analyses at seedling stage of O. sanctum showed more amounts of photosynthetic pigments as to that of mature O. basilicum. Further to confirm, photosynthetic rate was also estimated of the leaf which was found decreased in case of O. basilicum seedlings and mature stage plants. Thus, O. basilicum exhibited elevated levels of primary metabolic and expectedly secondary metabolic expression as compared to O. sanctum. Such correlations were previously shown for plants possessing secondary metabolites [41, 42]. Various studies are conducted in medicinally important plants as well as Arabidopsis to show the effect of chemical stimulants on primary and secondary metabolic pathways [30, 34].
Trichome development in O. basilicum and O. sanctum
Glandular trichomes are specialized structures or epidermal outgrowths with secretary head made of cells to store and sequester large quantities of volatile metabolites [6, 43]. To understand the relationship between essential oil content and trichomes; the morphology, density, and size of trichomes were discussed earlier [6]. Glandular trichomes classified as peltate and capitate trichomes as two major types are implicated in the biogenesis and sequestering of essential oil though mechanism is yet to be established [6]. In the present study, the genes involved in trichome development were studied to get deeper understanding of essential oil biosynthesis in glandular trichomes. In Ocimum species, transcripts for R2R3MYB, R3MYB, bHLH, and C2H2 were predominantly detected and expression pattern suggested that many of the transcripts related to trichome development were expressed higher in O. basilicum. Recent studies on A. thaliana suggested that trichome development processes were controlled by transcription factors majorly including basic helix-loop helix (bHLH) one of the largest family with 160 members in Arabidopsis consisting of 12 sub-groups, and R2R3MYB with 25 sub-groups consisting of 126 members in Arabidopsis [44]. C2H2 zinc finger type transcription factors with 176 members in Arabidopsis were also involved in the trichome formation on the stems, flower, and inflorescence tissues [13, 14]. Genes controlled by R3MYB TFs were CPC, ETC1, and ETC3 [17], while GL3, EGL3, TT8, GL2, and GIS were controlled by bHLH TFs and GIS2 by C2H2 TFs [13, 45].
Higher number of transcripts in O. basilicum may be responsible for better trichome development in the particular species. Studies were further confirmed by the qRT-expression analysis of transcripts which exhibited relatively less expression of these genes in O. sanctum. Relative higher expression of trichome related genes may be responsible for the higher levels of essential oil in the O. basilicum relative to that of O. sanctum. Hence, the study suggested that the trichomes related genes may be responsible for the overall essential oil content in plants while the specific component in the essential oil depended on the secondary metabolic pathway genes as studied earlier [6, 10].
Materials and methods
Sampling and RNA extraction
O. basilicum and O. sanctum plants were grown in the greenhouse at Central Institute of Medicinal and Aromatic Plants CSIR, Lucknow, India, during which all plants showed normal growth. Seedlings were collected for analysis from each plant after one month and mature plants after three months of sowing. Sample leaf tissue were collected and frozen in liquid N2 and stored at − 80 °C until RNA extraction. Total RNA was extracted using the CTAB method as describe earlier [6]. Intergrity of isolated RNA was assessed using the agarose gel electrophoresis. Quality and quantity assessment of RNA was done using the nanodrop, before proceeding for the cDNA synthesis. All the reactions were carried out in triplicate consisting of cDNA (~ 100 ng), syber green master mix and gene specific primer (5 picomole). Following PCR conditions: initial denaturation at 95 °C for 2 min, followed by 40 cycles of denaturation at 95 °C for 15 s, and annealing at 60 °C for 30 s was utilized. The relative expression level of each selected gene was normalized with the actin gene and detected by the 2−ΔΔCt method.
Assembly and redundancy reduction
O. basilicum project code (PRJNA240352), and O. sanctum project code (PRJNA238293), data downloaded from NCBI repositories using the FTP client. CLC genomics work bench [46] was utilized for assembly of the data. For conducting the differential expression of the two species, O. sanctum was considered as the parent species. In order to carry out the comparative transcriptome study, the individual transcriptome of both species were assembled. Quality of the assembled transcripts was assessed using the full-length coding sequences with taking O. sanctum genome as the reference. To make full length BLASTx search made with the virdiplantae protein database, of Uniprot, Ensemble/Swissprot criteria was set as [e-value ≤ 1e-30, max_target_seqs = 1].
Functional annotation and similarity search
Sequences assembled were subjected to the BLAST search with mRNA and protein sequences of different plant species (https://www.ncbi.nlm.nih.gov/blast/cgi/), of both the transcriptomes. These CLC assembled transcripts were analysed by various bioinformatics tools, for their effective assignment and functional analysis. Different modules of Blast2GO were followed to functionally annotate both transcriptomes. Annotated files of transcriptome of O. basilicum and O. sanctum were analysed separately for the secondary metabolic and global pathway analysis. KEGG analysis was performed in order to retrieve pathway maps, for each species. Parallel to this, the transcriptomes were also subjected to a local BLAST search against the previously isolated genes from the public databases for various pathways to make full length. Full-length pathway gene transcripts were assembled and mapped to respective genes submitted in non-redundant databases. Enzyme mapping was performed by the blast2go module for the distribution of enzyme classes in the Ocimum species. The expressions of transcripts belonging to metabolic pathways were evaluated using the FPKM values for each transcript coding for specific gene as described earlier [33, 31].
Differential expression and functional enrichment
To quantify the expression abundance of each transcript, the Blast2Go module was used, employing Kallisto [47] to generate normalized abundance estimates across transcripts of the both transcriptomes. To identify the differentially expressed (DE) transcripts, EdgeR was used, following the false discovery rate (FDR) of ≤ 0.001 and a log2 fold-change (logFC) of ≥ 2 [48]. Microsoft excel was used to generate the heat maps for the DE transcripts and clustering was done using the heat mapper [49]. Differentially expressed genes were subjected to the A. thaliana and O. sativa to get the enriched Kyoto Encyclopedia of Genes and Genomes (KEGG) and Gene Ontology (GO) terms [e-value ≤ 1e-5], using the KOBAS server (http://kobas.cbi.pku.edu.cn). AmiGO (amigo.geneontology.org) was used getting the GO terms definitions for the differentially expressed transcripts. Cytoscape (https://cytoscape.org/) was used for making the network plot of the DE transcripts.
Identification of trichome related genes
To determine the molecular factors behind the variations in the trichomes of the Ocimum species, genes related to the trichome were isolated namely caprice (CPC), enhancer of triptychon caprice (ETC), glabrous inflorescence stems (GIS), glabrous (GL), squamosa promoter binding like (SPL), trichomeless (TCL), triptychon (TRY), transparent testa (TT), ubiquitin-protein ligase (UPL) and zinc finger protein (ZFP). First, A. thaliana TAIR10 proteins with functional descriptions were extracted from the database. Trichome related genes identified from the both the transcriptomes were then queried against these trichome-related protein sequences using BLASTx [e-value ≤ 1e-3], and significant matches were recorded.
Estimation of total photosynthetic pigments content
Total photosynthetic pigments content such as total chlorophyll (Chl a & b), chlorophyll a (Chl a), chlorophyll b (Chl b), and carotenoids were estimated from fresh leaves of seedlings and mature plants of the both Ocimum species using earlier reported method [26].
Estimation of total sugar, reducing sugar, and invertase enzyme activity
Sugar level and invertase enzyme activity were estimated in seedlings and mature plants of both Ocimum species as reported earlier [39].
Conclusions
In the present work, comparative study of transcriptomes of O. basilicum and O. sanctum was performed with respect to overall global metabolic pathways as well as secondary metabolic pathways in relation to secondary metabolite phytochemicals and trichome development. Genes for the metabolism and for phytoconstituents were differentially regulated in O. basilicum and O. sanctum. Trichome related genes were putatively involved in the regulation of essential oil in the trichomes. Further studies in continuation will help in delineating the exact mechanism of trichome development and essential oil regulation in integration with overall metabolism in Ocimum species.
Abbreviations
- Ob:
-
O. basilicum
- Os:
-
O. sanctum
- CPC:
-
Caprice
- ETC1:
-
Enhancer of TRY and CPC1
- ETC2:
-
Enhancer of TRY and CPC2
- ETC3:
-
Enhancer of TRY and CPC3
- GIS:
-
Glabrous inflorescence stems
- GIS2:
-
Glabrous inflorescence stems2
- GL1:
-
Glabrous1
- GL2:
-
Glabra2
- GL3:
-
Glabra3
- SPL3:
-
Squamosa promoter binding protein Like3
- TCL1:
-
Trichomeless1
- TCL2:
-
Trichomeless2
- TRY:
-
Triptychon
- TT8:
-
Transparent testa8
- UPL3:
-
Ubiquitin protein ligase3
- ZFP:
-
Zinc finger protein
- KEGG:
-
Kyoto encyclopedia of genes and genomes
- B2G:
-
Blast2Go
- BLAST:
-
Basic local alignment search tool
References
Paton A, Harley MR, Harley MM (1999) In: Hiltunen R, Holm Y (eds). Basil: the genus Ocimum. Harwood Academic Publishers, Reading, pp 1–38
da Silveira Agostini-Costa T, Vieira R, Bizzo RH, Silveira D, Gimenes MA (2012) Secondary metabolites. In: Dhanarasu E (ed) Chromatography and its applications. InTech, London, pp 131–164
Shih ML, Morgan JA (2020) Metabolic flux analysis of secondary metabolism in plants. Metab Eng Commun 1:10:e00123. https://doi.org/10.1016/j.mec.2020.e00123
Bartwal A, Mall R, Lohani P, Guru SK, Arora S (2013) Role of secondary metabolites and brassinosteroids in plant defense against environmental stresses. J Plant Growth Regul 32(1):216–232. https://doi.org/10.1007/s00344-012-9272-x
Pagare S, Bhatia M, Tripathi N, Pagare S, Bansal YK (2015) Secondary metabolites of plants and their role: overview. Curr Trends Biotechnol Pharm 9(3):293–304
Maurya S, Chandra M, Yadav RK, Narnoliya LK, Sangwan RS, Bansal S, Sandhu P, Singh U, Kumar D, Sangwan NS (2019) Interspecies comparative features of trichomes in Ocimum reveal insights for biosynthesis of specialized essential oil metabolites. Protoplasma. https://doi.org/10.1007/s00709-018-01338-y
Huchelmann A, Boutry M, Hachez C (2017) Plant glandular trichomes: natural cell factories of high biotechnological interest. Plant Physiol 175:00727. https://doi.org/10.1104/pp.17.00727
Oksanen E (2018) Trichomes form an important first line of defence against adverse environment—New evidence for ozone stress mitigation. Plant Cell Environ 41:1497–1499. https://doi.org/10.1111/pce.13187
Prakash P, Gupta N (2005) Therapeutic uses of Ocimum sanctum linn (Tulsi) with a note on eugenol and its pharmacological actions: a short review. Indian J Physiol Pharmacol 49:125–131. https://doi.org/10.7860/JCDR/2014/9122.4629
Pattanayak P, Behera P, Das D, Panda S (2010) Ocimum sanctum Linn. A reservoir plant for therapeutic applications: an overview. Pharmacogn Rev 4:95. https://doi.org/10.4103/0973-7847.65323
Bansal S, Narnoliya LK, Mishra B, Chandra M, Yadav RK, Sangwan NS (2018) HMG-CoA reductase from Camphor Tulsi (Ocimum kilimandscharicum) regulated MVA dependent biosynthesis of diverse terpenoids in homologous and heterologous plant systems. Sci Rep 8:3547–3561. https://doi.org/10.1038/s41598-017-17153-z
Maurya S, Sangwan NS (2019) Profiling of essential oil constituents in Ocimum species. Proc Natl Acad Sci India Sect B Biol Sci 26:1–7. https://doi.org/10.1007/s40011-019-01123-8
Zhao H, Wang X, Zhu D, Cui S, Li X, Cao Y, Ma L (2012) A single amino acid substitution in IIIf subfamily of basic helix-loop-helix transcription factor AtMYC1 leads to trichome and root hair patterning defects by abolishing its interaction with partner proteins in Arabidopsis. J Biol Chem 287:14109–14121. https://doi.org/10.1074/jbc.M111.280735
Gan L, Xia K, Chen J-G, Wang S (2011) Functional characterization of TRICHOMELESS2, a new single-repeat R3 MYB transcription factor in the regulation of trichome patterning in Arabidopsis. BMC Plant Biol 11:176–188. https://doi.org/10.1186/1471-2229-11-176
Liang G, He H, Li Y, Ai Q, Yu D (2014) MYB82 functions in regulation of trichome development in Arabidopsis. J Exp Bot 65:3215–3223. https://doi.org/10.1093/jxb/eru179
Tripathi S, Srivastava Y, Sangwan RS, Sangwan NS (2020) In silico mining and functional analysis of AP2/ERF gene in Withania somnifera. Sci Rep 10:1–12. https://doi.org/10.1038/s41598-020-60090-7
Kirik V, Simon M, Wester K, Schiefelbein J, Hulskamp M (2004) Enhancer of try and CPC 2 (ETC2) reveals redundancy in the region-specific control of trichome development of Arabidopsis. Plant Mol Biol 55:389–398. https://doi.org/10.1007/s11103-004-0893-8
Hung FY, Chen JH, Feng YR, Lai YC, Yang S, Wu K (2020) Arabidopsis JMJ29 is involved in trichome development by regulating the core trichome initiation gene GLABRA3. Plant J. https://doi.org/10.1111/tpj.14858
Sangwan RS, Tripathi S, Singh J, Narnoliya LK, Sangwan NS (2013) De novo sequencing and assembly of Centella asiatica leaf transcriptome for mapping of structural, functional and regulatory genes with special reference to secondary metabolism. Gene 525:58–76. https://doi.org/10.1016/j.gene.2013.04.057
Tripathi S, Sangwan RS, Mishra B, Jadaun JS, Sangwan NS (2019) Berry transcriptome: insights into a novel resource to understand development dependent secondary metabolism in Withania somnifera (Ashwagandha). Physiol Plant. https://doi.org/10.1111/ppl.12943
Marks MD, Wenger JP, Gilding E, Jilk R, Dixon RA (2009) Transcriptome analysis of Arabidopsis wild-type and gl3-sst sim trichomes identifies four additional genes required for trichome development. Mol Plant 2:803–822. https://doi.org/10.1093/mp/ssp037
Tripathi S, Sangwan RS, Narnoliya LK, Srivastava Y, Mishra B, Sangwan NS (2017) Transcription factor repertoire in Ashwagandha (Withania somnifera) through analytics of transcriptomic resources: insights into regulation of development and withanolide metabolism. Sci Rep 7:16649–16666. https://doi.org/10.1038/s41598-017-14657-6
Narnoliya LK, Sangwan RS, Singh SP (2018) Transcriptome mining and in silico structural and functional analysis of ascorbic acid and tartaric acid biosynthesis pathway enzymes in rose-scanted Geranium. Mol Biol Rep 45:315–326. https://doi.org/10.1007/s11033-018-4164-1
Singh N, Luthra R (1988) Sucrose metabolism and essential oil accumulation during lemongrass (Cymbopogon flexuosus Stapf) leaf development. Plant Sci 57:127–133
Farooqi AHA, Bansal RP, Luthra R, Sangwan Neelam S, Sangwan RS (2000) Physiological, biochemical and environmental aspects of essential oil biosynthesis in aromatic grasses (Cymbopogon spp). In: Kumar S et al (eds) Aromatic grass monograph. CIMAP, Lucknow, pp 199–222
Bose SK, Yadav RK, Mishra S, Sangwan RS, Singh AK, Mishra B, Srivastava AK, Sangwan NS (2013) Effect of gibberellic acid and calliterpenone on plant growth attributes, trichomes, essential oil biosynthesis and pathway gene expression in differential manner in Mentha arvensis L. Plant Physiol Biochem 66:150–158
Yadav RK, Sangwan RS, Sabir F, Srivastava AK, Sangwan NS (2014) Effect of prolonged water stress on specialized secondary metabolites, peltate glandular trichomes, and pathway gene expression in Artemisia annua L. Plant Physiol Biochem 74:70–83. https://doi.org/10.1016/j.plaphy.2013.10.02
Yu L, Liu Y, Xu F (2019) Comparative transcriptome analysis reveals significant differences in the regulation of gene expression between hydrogen cyanide- and ethylene-treated Arabidopsis thaliana. BMC Plant Biol 19:1–19. https://doi.org/10.1186/s12870-019-1690-5
Gerstein MB, Rozowsky J, Yan KK, Wang D, Cheng C, Brown JB, Davis CA, Hillier L, Sisu C, Li JJ, Pei B (2014) Comparative analysis of the transcriptome across distant species. Nature 512:445–448. https://doi.org/10.1038/nature13424
Scheible W-R (2004) Genome-wide reprogramming of primary and secondary metabolism, protein synthesis, cellular growth processes, and the regulatory infrastructure of Arabidopsis in response to nitrogen. Plant Physiol 136:2483–2499. https://doi.org/10.1104/pp.104.047019
Primmer CR, Papakostas S, Leder EH, Davis MJ, Ragan MA (2013) Annotated genes and nonannotated genomes: cross-species use of gene ontology in ecology and evolution research. Mol Ecol 22:3216–3241. https://doi.org/10.1111/mec.12309
Keeling CI, Weisshaar S, Ralph SG, Jancsik S, Hamberger B, Dullat HK, Bohlmann J (2011) Transcriptome mining, functional characterization, and phylogeny of a large terpene synthase gene family in spruce (Picea spp.). BMC Plant Biol 11:43. https://doi.org/10.1186/1471-2229-11-43
Rastogi S, Meena S, Bhattacharya A, Ghosh S, Shukla RK, Sangwan NS, Lal RK, Gupta MM, Lavania UC, Gupta V, Nagegowda DA, Shasany AK (2014) De novo sequencing and comparative analysis of holy and sweet basil transcriptomes. BMC Genom 15:588. https://doi.org/10.1186/1471-2164-15-588
Winkelblech J, Fan A, Li S-M (2015) Prenyltransferases as key enzymes in primary and secondary metabolism. Appl Microbiol Biotechnol 99:7379–7397. https://doi.org/10.1007/s00253-015-6811-y
Bilgin DD, Zavala JA, Zhu JI, Clough SJ, Ort DR, DeLUCIA EH (2010) Biotic stress globally downregulates photosynthesis genes. Plant Cell Environ 33:1597–1613. https://doi.org/10.1111/j.1365-3040.2010.02167.x
Rojas CM, Senthil-Kumar M, Tzin V, Mysore KS (2014) Regulation of primary plant metabolism during plant-pathogen interactions and its contribution to plant defense. Front Plant Sci 5:1–12. https://doi.org/10.3389/fpls.2014.00017
Sangwan RS, Sangwan NS (2000) Metabolic and molecular analysis of chemotypic diversity in aromatic grasses (Cymbopogon spp.). In: Kumar S et al (eds) Aromatic Grass Monograph. CIMAP, Lucknow, pp 223–247
Sharma PK, Sangwan Neelam S, Mishra BN, Sangwan RS (2009) Coherent ontogenic dynamics of geraniol acetyltransferase activity and geranyl acetate concentration in flowers and leaves of aroma grass Cymbopogon martini var Motia. Plant Growth Regul 57:103–10851
Sharma P, Sangwan Neelam S, Bose SK, Sangwan RS (2013) Biochemical characteristics of a novel vegetative tissue geraniol acetyltransferase from a monoterpene oil grass (Palmarosa, Cymbopogon martinii var. Motia) leaf. Plant Sci 203–204: 63–73
Ng WL, Wu W, Zou P, Zhou R (2019) Comparative transcriptomics sheds light on differential adaptation and species diversification between two Melastoma species and their F1 hybrid. AoB Plants. https://doi.org/10.1093/aobpla/plz019
Fasbender L, Yáñez-Serrano AM, Kreuzwieser J, Dubbert D, Werner C (2018) Real-time carbon allocation into biogenic volatile organic compounds (BVOCs) and respiratory carbon dioxide (CO2) traced by PTR-TOF-MS, 13 CO2 laser spectroscopy and 13 C-pyruvate labelling. PLoS ONE 13:1–22. https://doi.org/10.1371/journal.pone.0204398
Delfin JC, Watanabe M, Tohge T (2019) Understanding the function and regulation of plant secondary metabolism through metabolomics approaches. Theor Exp Plant Physiol 31:127–138. https://doi.org/10.1007/s40626-018-0126-1
Wagner GJ (1991) Secreting glandular trichomes: more than just hairs. Plant Physiol 96:675–679. https://doi.org/10.1104/pp.96.3.675
Pattanaik S, Patra B, Singh SK, Yuan L (2014) An overview of the gene regulatory network controlling trichome development in the model plant, Arabidopsis. Front Plant Sci 5:1–8. https://doi.org/10.3389/fpls.2014.00259
Gan Y (2006) Glaborous inflorescense stem modulates the regulation by gibberellins of epidermal differentiation and shoot maturation in Arabidopsis. Plant Cell Online 18:1383–1395. https://doi.org/10.1105/tpc.106.041533
Liu CH, Di YP (2020) Analysis of RNA sequencing data using CLC Genomics Workbench. In: Keohavong P, Grant SG (eds) Molecular toxicology protocols. Humana, New York, pp 61–113
Bray NL, Pimentel H, Melsted P, Pachter L (2016) Near-optimal probabilistic RNA-seq quantification. Nat Biotechnol 34:525–527. https://doi.org/10.1038/nbt.3519
Haas BJ, Papanicolaou A, Yassour M, Grabherr M, Blood PD, Bowden J, Couger MB, Eccles D, Li B, Lieber M, MacManes MD (2013) De novo transcript sequence reconstruction from RNA-seq using the Trinity platform for reference generation and analysis. Nat Protoc 8:1494–1512. https://doi.org/10.1038/nprot.2013.084
Babicki S, Arndt D, Marcu A et al (2016) Heatmapper: web-enabled heat mapping for all. Nucleic Acids Res 44:W147–W153. https://doi.org/10.1093/nar/gkw419
Acknowledgements
NSS is thankful to networking projects BSC-0203, and BSC0107 for financial grant from CSIR, N Delhi. MC is thankful to UGC for Junior and senior research fellowship and CIMAP-JNU-PhD program for registration.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
Authors declare no conflict of interest; all the co-authors are well aware of the publication of this work. Research not involved Human participants and/or Animals.
Research involving human and animal rights
Research not involved Human participants and/or Animals.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Rights and permissions
About this article
Cite this article
Chandra, M., Kushwaha, S. & Sangwan, N.S. Comparative transcriptome analysis to identify putative genes related to trichome development in Ocimum species. Mol Biol Rep 47, 6587–6598 (2020). https://doi.org/10.1007/s11033-020-05710-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11033-020-05710-1