Abstract
Attachment of isoprene units to various acceptors by prenylation plays an important role in primary and secondary metabolism of living organisms. Protein prenylation belongs to posttranslational modification and is involved in cellular regulation process. Prenylated secondary metabolites usually demonstrate promising biological and pharmacological activities. Prenyl transfer reactions catalyzed by prenyltransferases represent the key steps in the biosynthesis and contribute significantly to the structural and biological diversity of these compounds. In the last decade, remarkable progress has been achieved in the biochemical, molecular, and structural biological investigations of prenyltransferases, especially on those of the members of the dimethylallyltryptophan synthase (DMATS) superfamily. Until now, more than 40 of such soluble enzymes are identified and characterized biochemically. They catalyze usually regioselective and stereoselective prenylations of a series of aromatic substances including tryptophan, tryptophan-containing peptides, and other indole derivatives as well as tyrosine or even nitrogen-free substrates. Crystal structures of a number of prenyltransferases have been solved in the past 10 years and provide a solid basis for understanding the mechanism of prenyl transfer reactions.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Prenyltransferases catalyze the transfer reactions of prenyl moieties from different prenyl donors, e.g., dimethylallyl (DMAPP with a branched C5-chain), geranyl (GPP, C10), farnesyl (FPP, C15), or geranylgeranyl (GGPP, C20) diphosphate, to various aliphatic or aromatic acceptors of both low and high molecular substances including proteins and nucleic acids (Dumelin et al. 2012; Heide 2009a; Li 2009a; Oldfield and Lin 2012; Palsuledesai and Distefano 2015; Xie et al. 2007; Yazaki et al. 2009). The prenylation reactions can take place in regular manner (regular prenylation) by connection of the prenyl moieties via their C-1 to an acceptor or reverse manner (reverse prenylation) via their C-3 atoms (Heide 2009a; Yu and Li 2012).
Prenylated secondary metabolites including indole alkaloids, flavonoids, coumarins, xanthones, quinones, and naphthalenes are widely distributed in terrestrial and marine organisms (Fig. 1). They exhibit a wide range of biological activities such as cytotoxicity, antioxidant (Sunassee and Davies-Coleman 2012), and antimicrobial activities (Liu et al. 2013a; Oya et al. 2015), which are often distinct from their non-prenylated precursors. Prenyl transfer reactions usually represent the key steps in the biosynthesis of such compounds. Furthermore, the prenylated products can be further modified by cyclization, hydroxylation, oxidation, and so on (Raju et al. 2011; Tagami et al. 2013). Therefore, prenyltransferases contribute significantly to structural and biological diversity of natural products.
Based on their primary amino acid sequences, biochemical and structural characteristics, prenyltransferases are categorized into different subgroups (Table 1). Prenyl diphosphate synthases use prenyl diphosphate as donor and isopentenyl diphosphate (IPP) as acceptor (Oldfield and Lin 2012), while protein prenyltransferases catalyze the transfer reactions of C15 chain from FPP or C20 chain from GGPP to cysteine residues of proteins (Palsuledesai and Distefano 2015). Peptide prenyltransferases of the TruF family catalyze O-prenylations of tyrosyl residues of cyclic peptides in the presence of DMAPP. DMAPP and GPP serve as prenyl donors for tRNA prenyltransferases from bacteria (Dumelin et al. 2012; McIntosh et al. 2011, 2013; Xie et al. 2007). A large prenyltransferase group uses diverse nitrogen-containing and nitrogen-free aromatic compounds as substrates, and among them, the dimethylallyltryptophan synthase (DMATS) superfamily is one of the most investigated subgroup. The name of this superfamily was given due to their sequence similarity to DMATS involved in the biosynthesis of ergot alkaloids of Claviceps sp. (Tsai et al. 1995).
Since the first reviews on the DMATS enzymes in 2009 (Li 2009a, b; Steffan et al. 2009), significant progress has been achieved for this enzyme group. Thirty-two new enzymes have been characterized biochemically. The natural substrates of the DMATS enzymes were found to be much broader than just indole derivatives known at that time. Tyrosine, xanthones, tricyclic or tetracyclic aromatic or even nonaromatic compounds also serve as natural substrates of enzymes with sequence similarity to DMATS. In contrast, the spectrum of the natural prenyl donors of these enzymes is relatively narrow, and most of the members of the DMATS superfamily use DMAPP for prenylation. In addition, crystal structures of four DMATS enzymes were solved and used as basis for understanding of the reaction mechanism. Since 2009, several reviews on different aspects of the DMATS superfamily, mainly on indole prenyltransferases, have appeared (Tanner 2014; Walsh 2014; Yu and Li 2012). A systematic summary covering classification, occurrence, natural function, biochemical properties, crystal structures, and reaction mechanism of this intriguing enzyme group is now necessary. For a better understanding of prenyltransferases as nxC5 transferring enzymes onto diverse acceptors and their natural roles, we begin in this review with a brief overview on different classes of prenyltransferases and then focus on the members of the DMATS superfamily. In another mini-review in this issue, we will discuss the potential usage of DMATS prenyltransferases in biotechnology, which has been demonstrated by a large number of studies in the last years.
Classification of prenyltransferases
IPP and DMAPP, derived from the acetate-mevalonate or methylerythritol phosphate pathway (Boronat and Rodriguez-Concepcion 2015; Chang et al. 2013; Zhao et al. 2013), serve as precursors for prenyl diphosphate (nxC5) biosynthesis (Oldfield and Lin 2012). The prenyl diphosphates can be used for the biosynthesis of different terpenoids by cyclization and oxidation or as prenyl donors for peptide, protein and tRNA prenyltransferases, or aromatic prenyltransferases. The donors, acceptors, products, and functions of different prenyltransferases are summarized in Table 1.
Prenyl diphosphate synthases
Over 63,000 isoprenoid natural products have been identified to date, which bear numerous important biological functions and show an enormous structural diversity based on their different carbon skeletons and substitutions (Liang et al. 2002; Ramamoorthy et al. 2015). In the biosynthesis of isoprenoids, prenyl diphosphate synthases catalyze the formation of the carbon backbone with defined chain length by sequential condensation of DMAPP or other prenyl diphosphates with a given number of IPP. According to the formed double bond configuration in the prenyl units, these chain elongating enzymes can be classified into two major groups, i.e., trans- and cis-prenyltransferases (Fig. 2).
FPP synthases (FPPase) belong to the first group and catalyze the formation of all-trans C15 precursor FPP by utilizing IPP as acceptor for the sequential condensations with DMAPP and GPP (Poulter 2006). FPP could serve as a precursor for a number of essential or important metabolites, e.g., sesquiterpenoids, triterpenoids, sterols, dolichols, and ubiquinones (Anderson et al. 1989).
GGPP synthases also belong to trans-prenyltransferases and catalyze, similar to FPPases, the formation of all-trans configured geranylgeranyl diphosphate. They are involved in the biosynthesis of plant secondary metabolites like diterpenoids or carotenoids (Liu et al. 2014b). Furthermore, octaprenyl diphosphate synthase (OPPS) catalyzes successive condensation reactions of FPP with five IPP to generate C40 products with trans-configured double bonds (Fig. 2). The resulted prenyl diphosphate can be used as donor for quinone prenylation in the biosynthesis of ubiquinone, menaquinone, or plastoquinones (Soballe and Poole 1999). The homodimeric enzymes of trans-prenyltransferases consist of α-helices with two conserved aspartate-rich DDxxD motifs, which allowed the binding of the substrates in complex with Mg2+ (Guo et al. 2004).
In contrast, cis-prenyltransferases produce prenyl diphosphates containing both Z- and E-configured double bonds, although they use the same substrates and their catalyzed reactions are Mg2+ dependent (Fig. 2). Their amino acid sequences and three-dimensional structures completely differ from those of trans-prenyltransferases. Instead of the DDxxD motif, an aspartate in the conserved P-loop of cis-prenyltransferases is involved in the chelating of Mg2+ (Guo et al. 2005). According to the length of their products, they are further classified into short- (C15), medium- (C50–C55), and long-chain (C70–C120) cis-prenyltransferases (Lu et al. 2009; Takahashi and Koyama 2006). Undecaprenyl diphosphate synthase for example catalyzes the condensation of FPP with eight IPPs, resulting in the formation of a C55 product (Fig. 2). Undecaprenyl monophosphate acts as a lipid carrier in the peptidoglycan biosynthesis of bacterial cell wall and therefore serves as a target for antibacterial drugs (Danley et al. 2015).
Prenylated lipids are also found in membranes of archaeal bacteria. The geranylgeranylglyceryl phosphate synthase (GGGP synthase) from Methanobacterium thermoautotrophicum, which shows practically no homology to other prenyltransferases, catalyzes the alkylation of glyceryl phosphate in the presence of GGPP (Soderberg et al. 2001).
Peptide, protein, and tRNA prenyltransferases
The ComX pheromone identified in Bacillus subtilis is a hexapeptide containing a regularly C3-geranylated tryptophan residue. The responsible prenyltransferase ComQ was identified by overproduction in Escherichia coli (E. coli) and incubation with an oligopeptide comprising 58 amino acids in the presence of GPP. The C-terminal geranylated hexapeptide corresponding to the mature ComX pheromone was identified as a minor and a C-terminal geranylated heptapeptide as the main product (Tsuji et al. 2012).
Cyanobactins from Lyngbya aestuarii are cyclic heptapeptides containing regularly C3-prenylated tyrosyl residues. The prenyltransferase LynF uses cyclic heptapeptides as substrates and catalyzes regular and reverse O-prenylations of tyrosyl residues. The reversely prenylated enzyme products then undergo a Claisen rearrangement resulting in C-prenylated derivatives (McIntosh et al. 2011, 2013).
Protein prenyltransferases use polypeptides as substrates. They play an important role in the posttranslational modification of proteins in eukaryotes and represent interesting drug targets (Palsuledesai and Distefano 2015). They transfer either a farnesyl (C15) or a geranylgeranyl (C20) moiety to a conserved cysteine residue in a CaaX motif at the C-terminus of the targeted proteins or peptides (Palsuledesai and Distefano 2015; Perez-Sala 2007). These modifications are responsible for correct cellular localization and activity of several proteins including those from the Ras family. The primary sequences as well as the crystal structures of protein prenyltransferases clearly differ from that of the prenyl diphosphate synthases. For the Ras farnesyltransferase reaction, the formation of Zn2+-activated thiolate by the cysteine residue of the CaaX box is necessary (Long et al. 2002). Due to the role of farnesylated proteins in oncogenic processes or several diseases including progeria, aging, parasitic diseases, and bacterial or viral infections, the underlying prenylation reaction and its inhibition by potential therapeutics have been extensively investigated (Abuhaie et al. 2013; Palsuledesai and Distefano 2015; Sousa et al. 2009; Zhu et al. 2014).
Modifications by prenylations are not only found in proteins, but also in nucleic acids, e.g., tRNAs. Two dimethylallyltransferases (DMATase) from Pseudomonas aeruginosa (P. aeruginosa) and E. coli were proven to prenylate the amino group of adenosine-37 (A37) of all tRNAs with uridine at the beginning (Xie et al. 2007). Recently, SelU from E. coli was reported to catalyze S-geranylation of 5-methylaminomethyl-2-thiouridyl residue in tRNA in the presence of GPP (Dumelin et al. 2012).
Aromatic prenyltransferases
Aromatic prenyltransferases catalyze the transfer reactions of prenyl moieties onto aromatic acceptors such as phenols, phenolic acids, flavonoids, coumarins, naphthalenes, phenazines, or indole derivatives. These enzymes contribute substantially to the large diversity of prenylated secondary metabolites in plants, fungi, and bacteria (Heide 2009b; Li 2009b; Yazaki et al. 2009). They usually catalyze the formation of C–C, C–O, or C–N bonds between the carbon of the prenyl and carbon or functional groups of the aromatic substrates. Membrane-bound and soluble aromatic prenyltransferases were found to exhibit distinct characteristics such as structural fold, substrate binding motifs, or metal ion dependency.
Membrane-bound prenyltransferases for aromatic substrates
Members of this enzyme group are involved in the biosynthesis of both primary metabolites such as ubiquinones and menaquinones (Boronat and Rodriguez-Concepcion 2015; Meganathan and Kwon 2009) and secondary metabolites like microbial and plant natural products (Holm et al. 2014; Wang et al. 2014; Yazaki et al. 2009; Zeyhle et al. 2014a, b).
Ubiquinones and menaquinones function as electron and proton carrier in photosynthesis and cellular respiration and also as antioxidants for prevention of cell damage. Membrane-bound prenyltransferases contain, similar to the aforementioned FPPs, characteristic aspartate-rich motifs, e.g., NDxxDxxxD, and require metal ions such as Mg2+ for their catalytic activity. The underlying prenyl transfer reactions were observed for various aromatic substrates like 4-hydroxybenzoate (4HB), homogentisic acid, coumarines, flavonoids, 1,4-dihydroxy-2-naphthoate, or phenazines (Heide 2009a; Karamat et al. 2014; Yazaki et al. 2009; Zeyhle et al. 2014a, b). Several examples of membrane-bound prenyltransferases and their catalyzed reactions are summarized in Table 2. UbiA from E. coli as a prototype of this family plays an important role in the biosynthesis of ubiquinones and catalyzes the attachment of an all-trans octaprenyl moiety onto 4HB (Melzer and Heide 1994). UbiA shows a broad substrate specificity toward prenyl donors and is able to generate ubiquinones CoQ6 to CoQ10 in different species (Cheng and Li 2014). Recently, the crystal structure of an archeal UbiA was reported (Cheng and Li 2014), which provides new insights into the substrate binding sites and mechanism of the enzyme catalysis. Very recently, a prenyltransferase UbiX from P. aeruginosa involved in the biosynthesis of ubiquinones was demonstrated to use flavin as a prenyl acceptor. In contrast to prenyl diphosphates for other known prenyltransferases, dimethylallyl monophosphate (DMAP) serves as prenyl donor for UbiX reaction (White et al. 2015).
In human cells, COQ2 is involved in the biosynthesis of ubiquinone for mitochondrial respiration, while UBIAD1 in the biosynthesis of vitamin K for maintaining vascular homeostasis (Hegarty et al. 2013; Nakagawa et al. 2010). UbiA homologs, e.g., AtPPT1 from Arabidopsis thaliana (A. thaliana) and OsPPT1 from Oryza sativa, were found to be involved in primary metabolism of plants (Okada et al. 2004; Ohara et al. 2006). In plants, membrane-bound aromatic prenyltransferases are also involved in the biosynthesis of secondary metabolites. The 4HB geranyltransferases LePGT-1 and LePGT-2 from Lithospermum erythrorhizon are involved in the biosynthesis of the naphthoquinone shikonin (Yazaki et al. 2002). In contrast to the 4HB prenyltransferases for ubiquinone biosynthesis, these enzymes are localized in the endoplasmatic reticulum rather than in mitochondria. They show strict substrate specificity for GPP as prenyl donor. Several members of the membrane-bound prenyltransferases are involved in the biosynthesis of prenylated flavonoids and isoflavonoids in plants. N8DT, G6DT, and SfiLDT from Sophora flavescens catalyze the prenylations of naringenin, genistein, and isoliquiritigenin, respectively (Chen et al. 2013; Sasaki et al. 2011). Furthermore, the formation of C3′-prenylated genistein was detected in vitro with LaPT1 from Lupinus albus (Shen et al. 2012). Pterocarpan 4-dimethylallytransferase (G4DT) catalyzes the transfer of a dimethylallyl moiety onto pterocarpan skeleton and is therefore responsible for the formation of the soybean phytoalexin glyceollin (Akashi et al. 2009). In plants, three kinds of homogentisic acid prenyltransferases use solanesyl (C45), geranylgeranyl (C20), and phytyl (C20, partially saturated) diphosphate for their prenylation reactions in the biosynthesis of plastoquinones, tocotrienols, and tocopherols, respectively (Heide 2009a). The last two compounds are also known as vitamin E. Moreover, 1,4-dihydroxy-2-naphthoate serves as substrate for the octaprenyltransferase MenA in E. coli and for the phytyltransferase ABC4 in phylloquinone biosynthesis in A. thaliana (Shimada et al. 2005; Suvarna et al. 1998). In addition, chlorophyllide and protoheme IX were used by chlorophyll synthase ATG4 (Eckhardt et al. 2004) and protoheme IX farnesyltransferase COX10, both from A. thaliana, respectively (Saiki et al. 1993). Furthermore, two membrane-bound prenyltransferases were found to catalyze three sequential prenylation steps in the biosynthesis of bitter acid in hop (Li et al. 2015). PcPT from parsley was found to be a key enzyme in the biosynthesis of linear and angular furanocoumarins and to catalyze prenylations of umbelliferon at both C-6 and C-8 positions (Karamat et al. 2014).
Membrane-bound aromatic prenyltransferases were also identified for the secondary metabolism in bacteria. One example is the farnesyltransferase AuaA from the myxobacterium Stigmatella aurantiaca in the biosynthesis of aurachins, which catalyzes the farnesylation of 2-methyl-4-hydroxyquinoline (Stec et al. 2011). Very recently, two membrane-bound aromatic prenyltransferases were identified in Streptomyces and found to be responsible for phenazine prenylations (Zeyhle et al. 2014a, b).
Soluble prenyltransferases with PT barrel mostly for aromatic substrates
The large enzyme group for prenylation of aromatic substrates comprises the CloQ/NphB group and the extensively investigated DMATS superfamily, which are soluble proteins from bacteria and fungi. One common structural feature of these enzymes is their aßßa-fold (ABBA), termed PT-barrel and firstly observed for the naphthalene geranyltransferase NphB (Kumano et al. 2008; Kuzuyama et al. 2005).
Enzymes of the CloQ/NphB subgroup
Known prenyltransferases of the CloQ/NphB group use only aromatic compounds as substrates and catalyze prenylations of naphthalenes, phenazines, quinones, and phenolic compounds (Table 3) (Heide 2009a). They were found in both bacteria and fungi and differ strongly from the aforementioned membrane-bound prenyltransferases (Haug-Schifferdecker et al. 2010). They do not contain the aspartate-rich NDxxD motif, and their reactions are with the exception of that for NphB independent of divalent metal ions (Bonitz et al. 2011; Heide 2009a). The notation of the subgroup referred to the first identified enzyme CloQ from Streptomyces roseochromogenes and the 2 years later reported NphB from Streptomyces sp. (Kuzuyama et al. 2005; Pojer et al. 2003). CloQ and its ortholog NovQ from Streptomyces spheroides catalyze the prenylation of 4-hydroxyphenylpyruvic acid in the biosynthesis of clorobiocin (Pojer et al. 2003) and novobiocin (Steffensky et al. 2000) (Fig. 1). NphB is involved in the biosynthesis of the geranylated derivative naphterpin (Kuzuyama et al. 2005). The crystal structures of NphB, CloQ, and EpzP share an ABBA barrel in common (Kuzuyama et al. 2005; Metzger et al. 2010; Wierenga et al. 2010; Zocher et al. 2012). These structures can serve as basis for molecular modeling studies and therefore provide valuable contributions to our knowledge on mechanisms of the prenyl transfer reactions (Bayse and Merz 2014; Yang et al. 2012). Furthermore, NphB shows a broad substrate specificity toward several phenolic compounds, e.g., resveratrol, flavonoids, and 4-HPP (Heide 2009a; Kumano et al. 2008; Kuzuyama et al. 2005). In the last years, several additional prenyltransferases of the CloQ/NphB group have been identified. SCO7190, a homolog of NphB from Streptomyces coelicolor A3(2), catalyzes the attachment of the dimethylallyl from DMAPP, but not geranyl moiety from GPP onto 1,6-dihydroxynaphthalene (Kumano et al. 2008; Kuzuyama et al. 2005). SCO7190 and NovQ were successfully used for production of novel prenylated polyphenols in transgenic plants (Sugiyama et al. 2011). Fnq26 from Streptomyces cinnamonensis DSM 1042 shares a sequence identity of 40 % with NphB and catalyzes reverse and regular C- as well as regular O-prenylations of several phenolic substrates (Haagen et al. 2007). Recently, McI23 from Streptomyces sp. CNH-189 was found to catalyze the formation of a prenylated precursor of merochlorin by utilizing a diphosphate of an unusual branched C15 unit as prenyl donor, which was formed by addition of a dimethylallyl moiety to GPP in a reverse manner catalyzed by Mcl22 (Teufel et al. 2014). In the secondary metabolism of Streptomyces cinnamonensis and Streptomyces anulatus, EpzP and PpzP catalyze the regiospecific C9-prenylation of 5,10-dihydrophenazine-1-carboxylic acid (Saleh et al. 2009; Seeger et al. 2011). As aforementioned, prenylation of phenazine derivatives can also be catalyzed by membrane-bound prenyltransferases (Table 3) (Zeyhle et al. 2014a, b). DzmP from Micromonospora sp. RV115 is the first member of this subgroup which utilize FPP instead of DMAPP and GPP as prenyl donor and catalyzes the unusual N-farnesylation of dibenzodiazepinone (Bonitz et al. 2013).
Enzymes of the DMATS superfamily
The DMATS superfamily is the most investigated subgroup among the prenyltransferases. In the last years, enormous advances have been achieved on the biochemical, molecular, and structural biological investigations of these soluble enzymes. So far, more than 40 such enzymes from fungi and bacteria have been identified mostly by genome mining and characterized biochemically by using the recombinant proteins (Fan et al. 2014; Pockrandt et al. 2014; Winkelblech and Li 2014; Wunsch et al. 2015; Yu et al. 2012; Yu and Li 2012). They mainly catalyze the prenylation of indole derivatives including tryptophan and tryptophan-containing cyclic dipeptides. DMATS prenyltransferases carry no aspartate-rich motifs and their catalysis is independent of the presence of metal ions, although Ca2+, Mg2+, or other metal ions strongly enhance their activities in several cases (Li 2009a; Pockrandt et al. 2012; Yu and Li 2012).
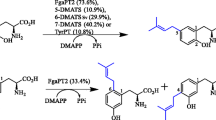
The first member of the DMATS superfamily was the tryptophan C4-prenyltransferase DmaW involved in the biosynthesis of ergot alkaloids in Claviceps fusiformis, reported as Claviceps purpurea (C. purpurea) (Gebler and Poulter 1992; Tsai et al. 1995). Its ortholog FgaPT2 from Aspergillus fumigatus (A. fumigatus) was identified in 2005 by genome mining using the DmaW sequence from C. purpurea (Tudzynski et al. 1999; Unsöld and Li 2005). Until now, 12 enzymes were identified, which catalyze different prenylations of tryptophan at N-1 and C-4 to C-7 (Table 4) (Li 2010; Yu and Li 2012). Cyclic (di)peptide or related prenyltransferases with 14 members build the largest group, which catalyze the prenylations at N-1, C-2, C-3, and C-7 of the indole ring (Table 5). In addition, a large number of indole derivatives were identified as substrates of these enzymes (Table 6). Two members of the DMATS superfamily, SirD and TyrPT, use tyrosine as substrate and catalyze O-prenylation (Table 6) (Fan et al. 2014; Kremer and Li 2010). Since 2010, nitrogen-free or nonaromatic compounds were identified as substrates of additional 16 prenyltransferases of the DMATS superfamily (Table 6). In total, 19 such enzymes have been characterized biochemically so far.
Tryptophan prenyltransferases
As summarized in Table 4, 12 tryptophan prenyltransferases, five from bacteria and seven from fungi, were identified and characterized biochemically. These enzymes catalyze the formation of dimethylallyltryptophan and therefore also termed DMATSs. As aforementioned, DmaW from C. fusiformis was identified as the first enzyme from the DMATS superfamily and catalyzes the first pathway-specific step in the biosynthesis of ergot alkaloids, i.e., the prenylation of tryptophan at C-4 of the indole ring (Gerhards et al. 2014; Tsai et al. 1995). With the identification and characterization of its ortholog, FgaPT2 from A. fumigatus in 2005 began the systematic study on the enzymes of the DMATS superfamily by genome mining and biochemical investigation (Unsöld and Li 2005). Later, two additional fungal 4-DMATS, i.e., DmaW-Cs from Periglandula (Markert et al. 2008; Steiner et al. 2011) and Malbranchea (Ding et al. 2008), were also identified. Identification of 7-DMATS from A. fumigatus as a tryptophan C7-prenylating enzyme obligated to donate the position of the prenyl moiety at the indole ring of tryptophan for prenyltransferase names. 7-DMATS was proven to be involved in the biosynthesis of hexadehydroastechrome (Yin et al. 2013c). Identification of 5-DMATS from A. clavatus filled the last gap of prenylation positions of fungal tryptophan and tryptophan-containing cyclic dipeptide prenyltransferases (Yu et al. 2012).
The first bacterial tryptophan prenyltransferase CymD was identified in Salinispora arenicola and catalyzes the N1-prenylation at the indole ring in the biosynthesis of the anti-inflammatory cyclomarin A and the antibacterial cyclomarazine A (Schultz et al. 2010). Later, four further bacterial DMATSs catalyzing the prenylations at C-5 and C-6 of the indole ring were identified from different Streptomyces strains (Subramanian et al. 2012; Takahashi et al. 2010; Winkelblech and Li 2014). Heterologous expression of the 5-DMATS gene SCO7467 with the flavin-dependent monooxygenase gene SCO7468 from Streptomyces coelicolor in Streptomyces lividans resulted in the formation of 5-dimethylallylindole-3-acetonitrile (Ozaki et al. 2013). IptA from Streptomyces sp. SN-593 was found to prenylate tryptophan at C-6 in the biosynthesis of 6-DMAI-3-carbaldehyde (Takahashi et al. 2010). 6-DMATSSa from Streptomyces ambofaciens and 6-DMATSSv from Streptomyces violaceusniger were identified as homologs of IptA. 6-DMATSSa and 6-DMATSSv can also utilize GPP as a prenyl donor and catalyze prenylations at the same position as for DMAPP, which had not been reported for tryptophan prenyltransferases before (Winkelblech and Li 2014).
Tryptophan-containing cyclic dipeptide and related prenyltransferases
Tryptophan-containing cyclic dipeptide prenyltransferases catalyze regiospecific prenylations at different positions of the indole ring, especially at N-1, C-2, C-3, and C-7 (Grundmann and Li 2005; Wunsch et al. 2015; Yin et al. 2009, 2010, 2013a; Zou et al. 2010). Some peptide-related substances like cyclo-acetoacetyl-l-tryptophan (cAATrp) or ardeemin FQ, a derivative of the cyclic tripeptide of anthranilic acid, alanine, and tryptophan, were also identified as substrates of this subgroup (Haynes et al. 2013; Liu and Walsh 2009). Until now, 14 enzymes from this group have been characterized biochemically (Table 5). With the exception of the two geranyltransferases, LtxC from the cyanobacterium Lyngbya majuscule and TleC from the actinomycetes Streptomyces blastmyceticus, all other 12 enzymes are identified in fungi of Aspergillus and Neosartorya species.
The first identified cyclic dipeptide prenyltransferase was FtmPT1 from A. fumigatus, which catalyzes a regular C2-prenylation of brevianamide F (cyclo-l-Trp-l-Pro) in the biosynthesis of verruculogen/fumitremorgins (Grundmann and Li 2005; Li 2011). Three cyclic dipeptide reverse C2-prenyltransferases, NotF from an Aspergillus sp. and BrePT from Aspergillus versicolor involved in the biosynthesis of notoamides as well as CdpC2PT from Neosartorya fischeri (N. fischeri), were identified 5 years later (Table 5) (Ding et al. 2010; Mundt and Li 2013; Yin et al. 2013a). CdpC2PT was speculated to be involved in the biosynthesis of fellutanine (Mundt and Li 2013). Recently, a regular cyclic dipeptide C7-prenyltransferase CdpC7PT was identified in Aspergillus terreus (A. terreus) (Wunsch et al. 2015), with much higher regioselectivity and substrate flexibility toward cyclic dipeptides than CTrpPT, an N1- and C7-prenyltransferase from Aspergillus oryzae (A. oryzae) (Zou et al. 2010). CdpC7PT also accepted cyclo-l-Tyr-l-Tyr as substrate and catalyzed an O-prenylation at the tyrosyl residue, providing the first example from the DMATS superfamily with an O-prenyltransferase activity toward tyrosine-containing dipeptides (Wunsch et al. 2015). CdpNPT from A. fumigatus was reported to catalyze an N1-prenylation of cyclic dipeptides (Yin et al. 2007). The prenylation position was later revised to C-3 of the indoline ring (Schuller et al. 2012; Yu et al. 2013).
The DMATS enzymes catalyze not only regiospecific, but also stereospecific prenylations. For example, AnaPT from N. fischeri is involved in the biosynthesis of acetylaszonalenin and catalyzes a reverse C3α-prenylation of (R)-benzodiazepinedinone, while CdpNPT from A. fumigatus and CdpC3PT from N. fischeri the reverse C3ß-prenylations of cyclic dipeptides (Schuller et al. 2012; Yin et al. 2009, 2010). ArdB from Aspergillus fischeri catalyzes C3ß-prenylation of ardeemin FQ at the indole ring (Haynes et al. 2013). Investigation with stereoisomers of cyclic dipeptides of tryptophan with proline or alanine revealed that AnaPT and CdpC3PT catalyze reverse anti-cis and syn-cis C3-prenylation, respectively. In contrast, CdpNPT produced both anti-cis and syn-cis C3-prenylated derivatives (Yu et al. 2013).
The product of a NRPS-PKS hybrid cAATrp was found to be the substrate of CpaD in the biosynthesis of the fungal neurotoxin α-cyclopiazonic acid in Aspergillus flavus (A. flavus) and A. oryzae (Liu and Walsh 2009). CpaD catalyzes the regiospecific C4-prenylation of cAATrp (Table 6) (Liu and Walsh 2009).
Several cyclic dipeptide prenyltransferases from bacteria use indolactams as substrates and catalyze prenylations at C-7 of the indole ring. In addition to LtxC mentioned above (Edwards and Gerwick 2004), the function of TleC in the biosynthetic pathway of teleocidin B in Streptomyces blastmyceticus was also characterized biochemically (Awakawa et al. 2014). Gene deletion experiments revealed the involvement of MpnD in the biosynthesis of methylpendolmycin in Marinactinospora thermotolerans (Ma et al. 2012).
Prenyltransferases utilizing other indole or quinolinone derivatives as prenyl acceptor
In addition to tryptophan and tryptophan-containing cyclic dipeptide prenyltransferases listed in Tables 4 and 5, eight members of the DMATS superfamily, which use other indole derivatives as prenylation substrates, are identified in different fungal strains (Table 6). These include FtmPT2 and FtmPT3 in the biosynthesis of verruculogen and fumitremorgin A in A. fumigatus and N. fischeri (Grundmann et al. 2008; Mundt et al. 2012). FgaPT1 from A. fumigatus catalyzes a reverse C2-prenylation, the last step in the biosynthesis of the ergot alkaloid fumigaclavine C (Unsöld and Li 2006). In addition to the cyclic dipeptide prenyltransferase NotF, NotC is also involved in the biosynthesis of stephacidin/notoamides (Ding et al. 2010). TdiB from Aspergillus nidulans (A. nidulans) and AstPT from A. terreus are involved in the biosynthesis of prenylated bisindolyl benzoquinones (Balibar et al. 2007; Schneider et al. 2008; Tarcz et al. 2014). AtmD from A. flavus and PaxD from Penicillium paxilli use indole diterpene derivatives as prenylation substrates (Liu et al. 2013b, 2014a). With the exception of FtmPT3, which catalyzes an O-prenylation of a secondary alcohol, all other enzymes from this group carry out reverse or regular prenylations at the indole ring (Table 6).
Very recently, two members of the DMATS superfamily, PenI and PenG, have been identified in Penicillium thymicola and proven to be involved in the biosynthesis of penigequinolone I. Both enzymes use DMAPP as prenyl donor. PenI catalyzes the prenylation of the quinolinone core at C-7, whereas PenG transfers a C5 unit to the dimethylallyl moiety of the product of PenI, i.e., the elongation of the prenyl moiety by addition of a second dimethylallyl unit (Zou et al. 2015).
Prenyltransferases of tyrosine and other aromatic or nonaromatic substrates
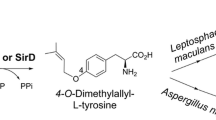
The tyrosine O-prenyltransferase SirD in the biosynthesis of sirodesmin PL in Leptosphaeria maculans was identified in 2010 as the first member of the DMATS superfamily, which use nonindole derivatives as prenylation substrates (Kremer and Li 2010). The second tyrosine O-prenyltransferase TyrPT of an unknown cluster was identified in Aspergillus niger (Fan et al. 2014). Both enzymes share relatively high sequence similarity with 7-DMATS from A. fumigatus and also catalyze the C7-prenylation of tryptophan (Fan et al. 2014; Kremer and Li 2010).
Since 2010, several enzymes of the DMATS superfamily were proven to use even polyketide products, e.g., xanthones by XptB from A. nidulans (Pockrandt et al. 2012), hydroxynaphthalenes by BAE61387 from A. oryzae (Pockrandt et al. 2014), or naphtacenedione by VrtC from Penicillium aethiapicum (Chooi et al. 2010). VrtC utilizes GPP instead of DMAPP as prenyl donor to generate a viridicatumtoxin precursor (Chooi et al. 2010). Interestingly, its homologs from N. fischeri , Microsporum canis, and Trichophyton tonsurans accepted DMAPP as prenyl donor and were proven to be involved in the biosynthesis of neosatoricin (Chooi et al. 2012, 2013; Yin et al. 2013b). More interestingly, PAPT from Phomopsis amygdali catalyzes an O-prenylation of the glucose moiety in the biosynthesis of fusicoccin A (Noike et al. 2012). It should be mentioned that all of the enzymes described here were identified in fungi.
Crystal structures of DMATS prenyltransferases providing basic knowledge for understanding the prenylation reactions
Determination of the crystal structure of the first indole prenyltransferase FgaPT2 was a fundamental step in the advanced understanding of prenyltransferases of the DMATS superfamily and their reaction mechanisms (Metzger et al. 2009). FgaPT2 revealed an unusual PT-barrel fold, formed by ten antiparallel ß-strands surrounded by ten α-helices (Fig. 3). The active site is located in the center of the barrel, and the absence of any metal ion in this area is consistent with the independency of divalent metals for enzyme activity. Interestingly, this architecture has been already observed for the bacterial hydroxynaphthalene prenyltransferase NphB (Kuzuyama et al. 2005), although its primary amino acid sequence and active site differ substantially from that of FgaPT2. The structural information, gained from this work, allowed the first understanding of the enzymatic catalysis and also supported the hypothesis of a common evolutionary origin of the bacterial and the fungal prenyltransferases of the ABBA-family (Luk and Tanner 2009; Metzger et al. 2009). In 2010, the crystal structure of the cyclic dipeptide C2-prenyltransferase FtmPT1 was solved (Jost et al. 2010), followed by the structures of two tryptophan-containing cyclic dipeptide C3-prenyltransferases CdpNPT and AnaPT in 2012 and 2013 (Schuller et al. 2012; Yu et al. 2013). Currently, the crystal structures of FgaPT2, FtmPT1, CdpNPT, and AnaPT are available in their unbound states. In addition, three of them (FgaPT2, FtmPT1, CdpNPT) were crystallized together with their aromatic substrates and an unreactive analog of DMAPP DMSPP (dimethylallyl S-thiolodiphosphate) or SPP (S-thiolodiphosphate). Comparison of the crystal structures revealed that they share a similar secondary core structure with the already described PT-barrel fold (Fig. 3). The amino acids in the DMAPP binding site seem to be strictly conserved in these four structures, whereas the binding sites of the aromatic substrates differ from each other. This feature makes molecular modeling with unknown structures more difficult. Therefore, additional enzyme structures from this superfamily are required. Nevertheless, the detailed structure analysis of the liganded and unliganded structures provides important insight into the catalyzed reaction mechanisms (Jost et al. 2010). Prenylations catalyzed by the enzymes of the DMATS superfamily were proposed as three-step reactions. They begin with the formation of a dimethylallyl cation by removal of the pyrophosphate group, which was proven by a positional isotope exchange with O18-labeled DMAPP (Luk and Tanner 2009). The subsequent nucleophilic attack of this cation by the indole nucleus or other electron-rich atoms to form the σ-complex defined the second step. Then, the resulting arenium intermediate was deprotonated to generate the final product (Fig. 3a). The most controversial issue of this mechanism focused on the second step, whether an electrophilic aromatic substitution mechanism occurs directly at the site of substitution or as results of rearrangements after initial prenylation at C-3 of the indole ring to generate the final product (Luk et al. 2011; Mahmoodi et al. 2013; Mahmoodi and Tanner 2013; Tanner 2014). Furthermore, these results could be applied for generation of modified prenyltransferases by modeling and site-directed mutagenesis experiments. For example, the enzymatic reaction of FtmPT1 was modified by a single point mutation to perform a reverse prenylation at C-3 of the indole nucleus instead of a regular C2-prenylation.
Phylogenic relationships of different prenyltransferases
To get insights into the phylogenetic relationships of prenyltransferases, enzymes listed in Tables 2, 3, 4, 5, and 6 and examples of prenyl diphosphate synthases as well as peptide, protein, and tRNA prenyltransferases were analyzed and illustrated as a phylogenetic tree in Fig. 4. The phylogenetic analysis confirmed the diversity of prenyltransferases, which are located in different clades. The membrane-bound prenyltransferases from different sources are grouped together, while the enzymes of the CloQ/NphB group build a defined clade. The members of the DMATS family are found in two clades, one with fungal and another with bacterial origin. The cis- and trans-prenyltransferases of prenyl diphosphate synthases are clearly separated from each other. The relationships among peptide prenyltransferases appear to be very far. This is also observed for tRNA prenyltransferases.
Phylogenetic relationships of different prenyltransferases (PTs). The phylogenetic tree was created by using the programs ClustalX2 (http://www.clustal.org/) and Treedyn (www.phylogeny.fr). In addition to enzymes listed in Tables 2, 3, 4, 5 and 6, cis- and trans-PTs, peptide and tRNA PTs as well as α- and β-subunits of protein PTs are also included. The accession numbers or pdb codes of the enzymes are given in parenthesis
Conclusion and outlook
In this review, we demonstrated the sequence, biochemical, and structural diversities of prenyltransferases, which catalyze the transfer of nxC5 units to different acceptors such as prenyl moieties, peptides, proteins, tRNAs, and aliphatic or aromatic small molecules (Dumelin et al. 2012; McIntosh et al. 2013; Palsuledesai and Distefano 2015). As given in Fig. 4, enzyme groups mentioned in this review can be found in different clades.
Prenyltransferases play important roles in primary and secondary metabolism as well as in the cellular regulation (Heide 2009a; Palsuledesai and Distefano 2015; Yazaki et al. 2009; Yu and Li 2012). The most intensively studied prenyltransferases belong to the DMATS superfamily, which are mainly identified in fungi, but also in bacteria. They are of great importance for the biosynthesis of secondary metabolites in microorganisms. Until now, no example from this group was known from plants and animals. To date, over 40 members of the DMATS superfamily have been identified and characterized biochemically, and crystal structures of four members are available. Members of this family do not require metal ions for their transfer reactions. Most of these enzymes utilize DMAPP as prenyl donor and an aromatic scaffold as prenyl acceptor, e.g., tryptophan, tyrosine, tryptophan-containing cyclic dipeptides, or other indole derivatives. With the first solved crystal structure of the 4-DMATS and its unexpected PT-barrel, new insights into reaction mechanism, substrate binding, and evolutionary origin were gained.
As demonstrated in the last years, mining of available genome sequences has proven to be a convenient approach for identification of putative prenyltransferase genes. It can be expected that additional prenyltransferases with new features will be identified in the next years. Prenyltransferases, especially the members of the DMATS superfamily, show a high potential for production of biologically active low molecular weight compounds by in vitro catalysis or by synthetic biology. This has already been demonstrated by a large number of studies on their substrate and catalytic promiscuity in the last years. This aspect will be discussed in another mini-review in the same issue in detail. Moreover, elucidation of further prenyltransferase structures would provide new chance and challenge for a better understanding of their functionality. This in turn will enable us to create modified enzymes with special features, e.g., changes in substrate specificity, regioselectivity, and stereoselectivity.
References
Abuhaie CM, Ghinet A, Farce A, Dubois J, Rigo B, Bîcu E (2013) Synthesis and biological evaluation of a new series of N-ylides as protein farnesyltransferase inhibitors. Bioorg Med Chem Lett 23:5887–5892
Anderson MS, Yarger JG, Burck CL, Poulter CD (1989) Farnesyl diphosphate synthetase. Molecular cloning, sequence, and expression of an essential gene from Saccharomyces cerevisiae. J Biol Chem 264:19176–19184
Akashi T, Sasaki K, Aoki T, Ayabe S, Yazaki K (2009) Molecular cloning and characterization of a cDNA for pterocarpan 4-dimethylallyltransferase catalyzing the key prenylation step in the biosynthesis of glyceollin, a soybean phytoalexin. Plant Physiol 149:683–693
Awakawa T, Zhang L, Wakimoto T, Hoshino S, Mori T, Ito T, Ishikawa J, Tanner ME, Abe I (2014) A methyltransferase initiates terpene cyclization in teleocidin B biosynthesis. J Am Chem Soc 136:9910–9913
Balibar CJ, Howard-Jones AR, Walsh CT (2007) Terrequinone A biosynthesis through L-tryptophan oxidation, dimerization and bisprenylation. Nat Chem Biol 3:584–592
Bayse CA, Merz KM (2014) Mechanistic insights into Mg2+-independent prenylation by CloQ from classical molecular mechanics and hybrid quantum mechanics/molecular mechanics molecular dynamics simulations. Biochemistry 53:5034–5041
Bonitz T, Alva V, Saleh O, Lupas AN, Heide L (2011) Evolutionary relationships of microbial aromatic prenyltransferases. PLoS One 6, e27336
Bonitz T, Zubeil F, Grond S, Heide L (2013) Unusual N-prenylation in diazepinomicin biosynthesis: The farnesylation of a benzodiazepine substrate is catalyzed by a new member of the ABBA prenyltransferase superfamily. PLoS One 8, e85707
Boronat A, Rodriguez-Concepcion M (2015) Terpenoid biosynthesis in prokaryotes. Adv Biochem Eng Biotechnol 148:3–18
Chang WC, Song H, Liu HW, Liu P (2013) Current development in isoprenoid precursor biosynthesis and regulation. Curr Opin Chem Biol 17:571–579
Chen R, Liu X, Zou J, Yin Y, Ou C, Li J, Wang R, Xie D, Zhang P, Dai J (2013) Regio- and stereospecific prenylation of flavonoids by Sophora flavescens prenyltransferases. Adv Synth Catal 355:1817–1828
Cheng W, Li W (2014) Structural insights into ubiquinone biosynthesis in membranes. Science 343:878–881
Chooi YH, Cacho R, Tang Y (2010) Identification of the viridicatumtoxin and griseofulvin gene clusters from Penicillium aethiopicum. Chem Biol 17:483–494
Chooi YH, Wang P, Fang J, Li Y, Wu K, Wang P, Tang Y (2012) Discovery and characterization of a group of fungal polycyclic polyketide prenyltransferases. J Am Chem Soc 134:9428–9437
Chooi YH, Fang J, Liu H, Filler SG, Wang P, Tang Y (2013) Genome mining of a prenylated and immunosuppressive polyketide from pathogenic fungi. Org Lett 15:780–783
Danley DE, Baima ET, Mansour M, Fennell KF, Chrunyk BA, Mueller JP, Liu S, Qiu X (2015) Discovery and structural characterization of an allosteric inhibitor of bacterial cis-prenyltransferase. Protein Sci 24:20–26
Ding Y, Williams RM, Sherman DH (2008) Molecular analysis of a 4-dimethylallyltryptophan synthase from Malbranchea aurantiaca. J Biol Chem 283:16068–16076
Ding Y, Wet JR, Cavalcoli J, Li S, Greshock TJ, Miller KA, Finefield JM, Sunderhaus JD, McAfoos TJ, Tsukamoto S, Williams RM, Sherman DH (2010) Genome-based characterization of two prenylation steps in the assembly of the stephacidin and notoamide anticancer agents in a marine-derived Aspergillus sp. J Am Chem Soc 132:12733–12740
Dumelin CE, Chen Y, Leconte AM, Chen YG, Liu DR (2012) Discovery and biological characterization of geranylated RNA in bacteria. Nat Chem Biol 8:913–919
Eckhardt U, Grimm B, Hortensteiner S (2004) Recent advances in chlorophyll biosynthesis and breakdown in higher plants. Plant Mol Biol 56:1–14
Edwards DJ, Gerwick WH (2004) Lyngbyatoxin biosynthesis: sequence of biosynthetic gene cluster and identification of a novel aromatic prenyltransferase. J Am Chem Soc 126:11432–11433
Fan A, Chen H, Wu R, Xu H, Li S-M (2014) A new member of the DMATS superfamily from Aspergillus niger catalyzes prenylations of both tyrosine and tryptophan derivatives. Appl Microbiol Biotechnol 98:10119–10129
Gebler JC, Poulter CD (1992) Purification and characterization of dimethylallyl tryptophan synthase from Claviceps purpurea. Arch Biochem Biophys 296:308–313
Gerhards N, Neubauer L, Tudzynski P, Li S-M (2014) Biosynthetic pathways of ergot alkaloids. Toxins (Basel) 6:3281–3295
Grundmann A, Li S-M (2005) Overproduction, purification and characterization of FtmPT1, a brevianamide F prenyltransferase from Aspergillus fumigatus. Microbiology 151:2199–2207
Grundmann A, Kuznetsova T, Afiyatullov SS, Li S-M (2008) FtmPT2, an N-prenyltransferase from Aspergillus fumigatus, catalyses the last step in the biosynthesis of fumitremorgin B. Chembiochem 9:2059–2063
Guo RT, Kuo CJ, Chou CC, Ko TP, Shr HL, Liang PH, Wang AH (2004) Crystal structure of octaprenyl pyrophosphate synthase from hyperthermophilic Thermotoga maritima and mechanism of product chain length determination. J Biol Chem 279:4903–4912
Guo RT, Ko TP, Chen AP, Kuo CJ, Wang AH, Liang PH (2005) Crystal structures of undecaprenyl pyrophosphate synthase in complex with magnesium, isopentenyl pyrophosphate, and farnesyl thiopyrophosphate: roles of the metal ion and conserved residues in catalysis. J Biol Chem 280:20762–20774
Haagen Y, Unsöld I, Westrich L, Gust B, Richard SB, Noel JP, Heide L (2007) A soluble, magnesium-independent prenyltransferase catalyzes reverse and regular C-prenylations and O-prenylations of aromatic substrates. FEBS Lett 581:2889–2893
Haug-Schifferdecker E, Arican D, Brueckner R, Heide L (2010) A new group of aromatic prenyltransferases in fungi, catalyzing a 2,7-dihydroxynaphthalene dimethylallyltransferase reaction. J Biol Chem 285:16487–16494
Haynes SW, Gao X, Tang Y, Walsh CT (2013) Complexity generation in fungal peptidyl alkaloid biosynthesis: A two-enzyme pathway to the hexacyclic MDR export pump inhibitor ardeemin. ACS Chem Biol 8:741–748
Hegarty JM, Yang H, Chi NC (2013) UBIAD1-mediated vitamin K2 synthesis is required for vascular endothelial cell survival and development. Development 140:1713–1719
Heide L (2009a) Prenyl transfer to aromatic substrates: genetics and enzymology. Curr Opin Chem Biol 13:171–179
Heide L (2009b) The aminocoumarins: biosynthesis and biology. Nat Prod Rep 26:1241–1250
Holm DK, Petersen LM, Klitgaard A, Knudsen PB, Jarczynska ZD, Nielsen KF, Gotfredsen CH, Larsen TO, Mortensen UH (2014) Molecular and chemical characterization of the biosynthesis of the 6-MSA-derived meroterpenoid yanuthone D in Aspergillus niger. Chem Biol 21:519–529
Jost M, Zocher G, Tarcz S, Matuschek M, Xie X, Li S-M, Stehle T (2010) Structure-function analysis of an enzymatic prenyl transfer reaction identifies a reaction chamber with modifiable specificity. J Am Chem Soc 132:17849–17858
Karamat F, Olry A, Munakata R, Koeduka T, Sugiyama A, Paris C, Hehn A, Bourgaud F, Yazaki K (2014) A coumarin-specific prenyltransferase catalyzes the crucial biosynthetic reaction for furanocoumarin formation in parsley. Plant J 77:627–638
Kremer A, Li S-M (2010) A tyrosine O-prenyltransferase catalyses the first pathway-specific step in the biosynthesis of sirodesmin PL. Microbiology 156:278–286
Kremer A, Westrich L, Li S-M (2007) A 7-dimethylallyltryptophan synthase from Aspergillus fumigatus: overproduction, purification and biochemical characterization. Microbiology 153:3409–3416
Kumano T, Richard SB, Noel JP, Nishiyama M, Kuzuyama T (2008) Chemoenzymatic syntheses of prenylated aromatic small molecules using Streptomyces prenyltransferases with relaxed substrate specificities. Bioorg Med Chem 16:8117–8126
Kuzuyama T, Noel JP, Richard SB (2005) Structural basis for the promiscuous biosynthetic prenylation of aromatic natural products. Nature 435:983–987
Li S-M (2009a) Applications of dimethylallyltryptophan synthases and other indole prenyltransferases for structural modification of natural products. Appl Microbiol Biotechnol 84:631–639
Li S-M (2009b) Evolution of aromatic prenyltransferases in the biosynthesis of indole derivatives. Phytochemistry 70:1746–1757
Li S-M (2010) Prenylated indole derivatives from fungi: structure diversity, biological activities, biosynthesis and chemoenzymatic synthesis. Nat Prod Rep 27:57–78
Li S-M (2011) Genome mining and biosynthesis of fumitremorgin-type alkaloids in ascomycetes. J Antibiot 64:45–49
Li H, Ban Z, Qin H, Ma L, King AJ, Wang G (2015) A heteromeric membrane-bound prenyltransferase complex from hop catalyzes three sequential aromatic prenylations in the bitter acid pathway. Plant Physiol 167:650–659
Liang PH, Ko TP, Wang AH (2002) Structure, mechanism and function of prenyltransferases. Eur J Biochem 269:3339–3354
Liu X, Walsh CT (2009) Characterization of cyclo-acetoacetyl-L-tryptophan dimethylallyltransferase in cyclopiazonic acid biosynthesis: substrate promiscuity and site directed mutagenesis studies. Biochemistry 48:11032–11044
Liu AH, Liu DQ, Liang TJ, Yu XQ, Feng MT, Yao LG, Fang Y, Wang B, Feng LH, Zhang MX, Mao SC (2013a) Caulerprenylols A and B, two rare antifungal prenylated para-xylenes from the green alga Caulerpa racemosa. Bioorg Med Chem Lett 23:2491–2494
Liu C, Minami A, Noike M, Toshima H, Oikawa H, Dairi T (2013b) Regiospecificities and prenylation mode specificities of the fungal indole diterpene prenyltransferases AtmD and PaxD. Appl Environ Microbiol 79:7298–7304
Liu C, Noike M, Minami A, Oikawa H, Dairi T (2014a) Functional analysis of a prenyltransferase gene (paxD) in the paxilline biosynthetic gene cluster. Appl Microbiol Biotechnol 98:199–206
Liu C, Sun Z, Shen S, Lin L, Li T, Tian B, Hua Y (2014b) Identification and characterization of the geranylgeranyl diphosphate synthase in Deinococcus radiodurans. Lett Appl Microbiol 58:219–224
Long SB, Casey PJ, Beese LS (2002) Reaction path of protein farnesyltransferase at atomic resolution. Nature 419:645–650
Lu YP, Liu HG, Liang PH (2009) Different reaction mechanisms for cis- and trans-prenyltransferases. Biochem Biophys Res Commun 379:351–355
Luk LYP, Tanner ME (2009) Mechanism of dimethylallyltryptophan synthase: evidence for a dimethylallyl cation intermediate in an aromatic prenyltransferase reaction. J Am Chem Soc 131:13932–13933
Luk LY, Qian Q, Tanner ME (2011) A cope rearrangement in the reaction catalyzed by dimethylallyltryptophan synthase? J Am Chem Soc 133:12342–12345
Ma J, Zuo D, Song Y, Wang B, Huang H, Yao Y, Li W, Zhang S, Zhang C, Ju J (2012) Characterization of a single gene cluster responsible for methylpendolmycin and pendolmycin biosynthesis in the deep sea bacterium Marinactinospora thermotolerans. Chembiochem 13:547–552
Mahmoodi N, Tanner ME (2013) Potential rearrangements in the reaction catalyzed by the indole prenyltransferase FtmPT1. Chembiochem 14:2029–2037
Mahmoodi N, Qian Q, Luk LYP, Tanner ME (2013) Rearrangements in the mechanisms of the indole alkaloid prenyltransferases. Pure Appl Chem 85:1935–1948
Markert A, Steffan N, Ploss K, Hellwig S, Steiner U, Drewke C, Li S-M, Boland W, Leistner E (2008) Biosynthesis and accumulation of ergoline alkaloids in a mutualistic association between Ipomoea asarifolia (Convolvulaceae) and a clavicipitalean fungus. Plant Physiol 147:296–305
McIntosh JA, Donia MS, Nair SK, Schmidt EW (2011) Enzymatic basis of ribosomal peptide prenylation in cyanobacteria. J Am Chem Soc 133:13698–13705
McIntosh JA, Lin Z, Tianero MD, Schmidt EW (2013) Aestuaramides, a natural library of cyanobactin cyclic peptides resulting from isoprene-derived Claisen rearrangements. ACS Chem Biol 8:877–883
Meganathan R, Kwon O (2009) Biosynthesis of menaquinone (vitamin K) and ubiquinone (coenzyme Q). Ecosal Plus 3:DOI: 10.1128/ecosalplus.3.6.3.3
Melzer M, Heide L (1994) Characterization of polyprenyldiphosphate: 4-hydroxybenzoate polyprenyltransferase from Escherichia coli. Biochim Biophys Acta 1212:93–102
Metzger U, Schall C, Zocher G, Unsöld I, Stec E, Li S-M, Heide L, Stehle T (2009) The structure of dimethylallyl tryptophan synthase reveals a common architecture of aromatic prenyltransferases in fungi and bacteria. Proc Natl Acad Sci U S A 106:14309–14314
Metzger U, Keller S, Stevenson CE, Heide L, Lawson DM (2010) Structure and mechanism of the magnesium-independent aromatic prenyltransferase CloQ from the clorobiocin biosynthetic pathway. J Mol Biol 404:611–626
Miyamoto K, Ishikawa F, Nakamura S, Hayashi Y, Nakanishi I, Kakeya H (2014) A 7-dimethylallyl tryptophan synthase from a fungal Neosartorya sp.: Biochemical characterization and structural insight into the regioselective prenylation. Bioorg Med Chem 22:2517–2528
Mundt K, Li S-M (2013) CdpC2PT, a reverse prenyltransferase from Neosartorya fischeri with distinct substrate preference from known C2-prenyltransferases. Microbiology 159:2169–2179
Mundt K, Wollinsky B, Ruan HL, Zhu T, Li S-M (2012) Identification of the verruculogen prenyltransferase FtmPT3 by a combination of chemical, bioinformatic and biochemical approaches. Chembiochem 13:2583–2592
Nakagawa K, Hirota Y, Sawada N, Yuge N, Watanabe M, Uchino Y, Okuda N, Shimomura Y, Suhara Y, Okano T (2010) Identification of UBIAD1 as a novel human menaquinone-4 biosynthetic enzyme. Nature 468:117–121
Noike M, Liu C, Ono Y, Hamano Y, Toyomasu T, Sassa T, Kato N, Dairi T (2012) An enzyme catalyzing O-prenylation of the glucose moiety of fusicoccin A, a diterpene glucoside produced by the fungus Phomopsis amygdali. Chembiochem 13:566–573
Ohara K, Yamamoto K, Hamamoto M, Sasaki K, Yazaki K (2006) Functional characterization of OsPPT1, which encodes p-hydroxybenzoate polyprenyltransferase involved in ubiquinone biosynthesis in Oryza sativa. Plant Cell Physiol 47:581–590
Okada K, Ohara K, Yazaki K, Nozaki K, Uchida N, Kawamukai M, Nojiri H, Yamane H (2004) The AtPPT1 gene encoding 4-hydroxybenzoate polyprenyl diphosphate transferase in ubiquinone biosynthesis is required for embryo development in Arabidopsis thaliana. Plant Mol Biol 55:567-577
Oldfield E, Lin FY (2012) Terpene biosynthesis: modularity rules. Angew Chem Int Ed Engl 51:1124–1137
Oya A, Tanaka N, Kusama T, Kim SY, Hayashi S, Kojoma M, Hishida A, Kawahara N, Sakai K, Gonoi T, Kobayashi J (2015) Prenylated benzophenones from Triadenum japonicum. J Nat Prod 78:258–264
Ozaki T, Mishima S, Nishiyama M, Kuzuyama T (2009) NovQ is a prenyltransferase capable of catalyzing the addition of a dimethylallyl group to both phenylpropanoids and flavonoids. J Antibiot 62:385–392
Ozaki T, Nishiyama M, Kuzuyama T (2013) Novel tryptophan metabolism by a potential gene cluster that is widely distributed among actinomycetes. J Biol Chem 288:9946–9956
Palsuledesai CC, Distefano MD (2015) Protein prenylation: enzymes, therapeutics, and biotechnology applications. ACS Chem Biol 10:51–62
Perez-Sala D (2007) Protein isoprenylation in biology and disease: general overview and perspectives from studies with genetically engineered animals. Front Biosci 12:4456–4472
Pockrandt D, Ludwig L, Fan A, König GM, Li S-M (2012) New insights into the biosynthesis of prenylated xanthones: XptB from Aspergillus nidulans catalyses an O-prenylation of xanthones. Chembiochem 13:2764–2771
Pockrandt D, Sack C, Kosiol T, Li S-M (2014) A promiscuous prenyltransferase from Aspergillus oryzae catalyses C-prenylations of hydroxynaphthalenes in the presence of different prenyl donors. Appl Microbiol Biotechnol 98:4987–4994
Pojer F, Wemakor E, Kammerer B, Chen H, Walsh CT, Li S-M, Heide L (2003) CloQ, a prenyltransferase involved in clorobiocin biosynthesis. Proc Natl Acad Sci U S A 100:2316–2321
Poulter CD (2006) Farnesyl diphosphate synthase. A paradigm for understanding structure and function relationships in E-polyprenyl diphosphate synthases. Phytochem Rev 5:17–26
Raju R, Piggott AM, Huang XC, Capon RJ (2011) Nocardioazines: a novel bridged diketopiperazine scaffold from a marine-derived bacterium inhibits p-glycoprotein. Org Lett 13:2770–2773
Ramamoorthy G, Pugh ML, Tian BX, Phan RM, Perez LB, Jacobson MP, Poulter CD (2015) Synthesis and enzymatic studies of bisubstrate analogs for farnesyl diphosphate synthase. J Org Chem 80:3902–3913
Saiki K, Mogi T, Ogura K, Anraku Y (1993) In vitro heme O synthesis by the cyoE gene product from Escherichia coli. J Biol Chem 268:26041–26044
Saleh O, Gust B, Boll B, Fiedler H-P, Heide L (2009) Aromatic prenylation in phenazine biosynthesis: dihydrophenazine-1-carboxylate dimethylallyltransferase from Streptomyces anulatus. J Biol Chem 284:14439–14447
Sasaki K, Tsurumaru Y, Yamamoto H, Yazaki K (2011) Molecular characterization of a membrane-bound prenyltransferase specific for isoflavone from Sophora flavescens. J Biol Chem 286:24125–24134
Schneider P, Weber M, Hoffmeister D (2008) The Aspergillus nidulans enzyme TdiB catalyzes prenyltransfer to the precursor of bioactive asterriquinones. Fungal Genet Biol 45:302–309
Schuller JM, Zocher G, Liebhold M, Xie X, Stahl M, Li S-M, Stehle T (2012) Structure and catalytic mechanism of a cyclic dipeptide prenyltransferase with broad substrate promiscuity. J Mol Biol 422:87–99
Schultz AW, Lewis CA, Luzung MR, Baran PS, Moore BS (2010) Functional characterization of the cyclomarin/cyclomarazine prenyltransferase CymD directs the biosynthesis of unnatural cyclic peptides. J Nat Prod 73:373–377
Seeger K, Flinspach K, Haug-Schifferdecker E, Kulik A, Gust B, Fiedler HP, Heide L (2011) The biosynthetic genes for prenylated phenazines are located at two different chromosomal loci of Streptomyces cinnamonensis DSM 1042. Microb Biotechnol 4:252–262
Shen G, Huhman D, Lei Z, Snyder J, Sumner LW, Dixon RA (2012) Characterization of an isoflavonoid-specific prenyltransferase from Lupinus albus. Plant Physiol 159:70–80
Shimada H, Ohno R, Shibata M, Ikegami I, Onai K, Ohto MA, Takamiya K (2005) Inactivation and deficiency of core proteins of photosystems I and II caused by genetical phylloquinone and plastoquinone deficiency but retained lamellar structure in a T-DNA mutant of Arabidopsis. Plant J 41:627–637
Soballe B, Poole RK (1999) Microbial ubiquinones: multiple roles in respiration, gene regulation and oxidative stress management. Microbiology 145(Pt 8):1817–1830
Soderberg T, Chen A, Poulter CD (2001) Geranylgeranylglyceryl phosphate synthase. Characterization of the recombinant enzyme from Methanobacterium thermoautotrophicum. Biochemistry 40:14847–14854
Sousa SF, Fernandes PA, Ramos MJ (2009) Molecular dynamics simulations on the critical states of the farnesyltransferase enzyme. Bioorg Med Chem 17:3369–3378
Stec E, Pistorius D, Müller R, Li S-M (2011) AuaA, a membrane-bound farnesyltransferase from Stigmatella aurantiaca, catalyzes the prenylation of 2-methyl-4-hydroxyquinoline in the biosynthesis of aurachins. Chembiochem 12:1724–1730
Steffan N, Grundmann A, Yin W-B, Kremer A, Li S-M (2009) Indole prenyltransferases from fungi: a new enzyme group with high potential for the production of prenylated indole derivatives. Curr Med Chem 16:218–231
Steffensky M, Mühlenweg A, Wang Z-X, Li S-M, Heide L (2000) Identification of the novobiocin biosynthetic gene cluster of Streptomyces spheroides NCIB 11891. Antimicrob Agents Chemother 44:1214–1222
Steiner U, Leibner S, Schardl CL, Leuchtmann A, Leistner E (2011) Periglandula, a new fungal genus within the Clavicipitaceae and its association with Convolvulaceae. Mycologia 103:1133–1145
Subramanian S, Shen X, Yuan Q, Yan Y (2012) Identification and biochemical characterization of a 5-dimethylallyltryptophan synthase in Streptomyces coelicolor A3(2). Process Biochem 47:1419–1422
Sugiyama A, Linley PJ, Sasaki K, Kumano T, Yamamoto H, Shitan N, Ohara K, Takanashi K, Harada E, Hasegawa H, Terakawa T, Kuzuyama T, Yazaki K (2011) Metabolic engineering for the production of prenylated polyphenols in transgenic legume plants using bacterial and plant prenyltransferases. Metab Eng 13:629–637
Sunassee SN, Davies-Coleman MT (2012) Cytotoxic and antioxidant marine prenylated quinones and hydroquinones. Nat Prod Rep 29:513–535
Suvarna K, Stevenson D, Meganathan R, Hudspeth ME (1998) Menaquinone (vitamin K2) biosynthesis: localization and characterization of the menA gene from Escherichia coli. J Bacteriol 180:2782–2787
Tagami K, Liu C, Minami A, Noike M, Isaka T, Fueki S, Shichijo Y, Toshima H, Gomi K, Dairi T, Oikawa H (2013) Reconstitution of biosynthetic machinery for indole-diterpene paxilline in Aspergillus oryzae. J Am Chem Soc 135:1260–1263
Takahashi S, Koyama T (2006) Structure and function of cis-prenyl chain elongating enzymes. Chem Rec 6:194–205
Takahashi S, Takagi H, Toyoda A, Uramoto M, Nogawa T, Ueki M, Sakaki Y, Osada H (2010) Biochemical characterization of a novel indole prenyltransferase from Streptomyces sp. SN-593. J Bacteriol 192:2839–2851
Tanner ME (2014) Mechanistic studies on the indole prenyltransferases. Nat Prod Rep 32:88–101
Tarcz S, Ludwig L, Li S-M (2014) AstPT catalyses both reverse N1- and regular C2-prenylation of a methylated bisindolyl benzoquinone. Chembiochem 15:108–116
Teufel R, Kaysser L, Villaume MT, Diethelm S, Carbullido MK, Baran PS, Moore BS (2014) One-pot enzymatic synthesis of merochlorin A and B. Angew Chem Int Ed 53:11019–11022
Tsai HF, Wang H, Gebler JC, Poulter CD, Schardl CL (1995) The Claviceps purpurea gene encoding dimethylallyltryptophan synthase, the committed step for ergot alkaloid biosynthesis. Biochem Biophys Res Commun 216:119–125
Tsuji F, Ishihara A, Kurata K, Nakagawa A, Okada M, Kitamura S, Kanamaru K, Masuda Y, Murakami K, Irie K, Sakagami Y (2012) Geranyl modification on the tryptophan residue of ComXRO-E-2 pheromone by a cell-free system. FEBS Lett 586:174–179
Tsurumaru Y, Sasaki K, Miyawaki T, Uto Y, Momma T, Umemoto N, Momose M, Yazaki K (2012) HlPT-1, a membrane-bound prenyltransferase responsible for the biosynthesis of bitter acids in hops. Biochem Biophys Res Commun 417:393–398
Tudzynski P, Holter K, Correia T, Arntz C, Grammel N, Keller U (1999) Evidence for an ergot alkaloid gene cluster in Claviceps purpurea. Mol Gen Genet 261:133–141
Unsöld IA, Li S-M (2005) Overproduction, purification and characterization of FgaPT2, a dimethylallyltryptophan synthase from Aspergillus fumigatus. Microbiology 151:1499–1505
Unsöld IA, Li S-M (2006) Reverse prenyltransferase in the biosynthesis of fumigaclavine C in Aspergillus fumigatus: gene expression, purification and characterization of fumigaclavine C synthase FgaPT1. Chembiochem 7:158–164
Walsh CT (2014) Biological matching of chemical reactivity: pairing indole nucleophilicity with electrophilic isoprenoids. ACS Chem Biol 9:2718–2728
Wang R, Chen R, Li J, Liu X, Xie K, Chen D, Yin Y, Tao X, Xie D, Zou J, Yang L, Dai J (2014) Molecular characterization and phylogenetic analysis of two novel regio-specific flavonoid prenyltransferases from Morus alba and Cudrania tricuspidata. J Biol Chem 289:35815–35825
White MD, Payne KA, Fisher K, Marshall SA, Parker D, Rattray NJ, Trivedi DK, Goodacre R, Rigby SE, Scrutton NS, Hay S, Leys D (2015) UbiX is a flavin prenyltransferase required for bacterial ubiquinone biosynthesis. Nature 522:502–506. doi:10.1038/nature14559
Wierenga RK, Kapetaniou EG, Venkatesan R (2010) Triosephosphate isomerase: a highly evolved biocatalyst. Cell Mol Life Sci 67:3961–3982
Winkelblech J, Li S-M (2014) Biochemical investigations of two 6-DMATS enzymes from Streptomyces revealing novel features of L-tryptophan prenyltransferases. Chembiochem 15:1030–1039
Wunsch C, Zou HX, Linne U, Li S-M (2015) C7-prenylation of tryptophanyl and O-prenylation of tyrosyl residues in dipeptides by an Aspergillus terreus prenyltransferase. Appl Microbiol Biotechnol 99:1719–1730
Xie W, Zhou C, Huang RH (2007) Structure of tRNA dimethylallyltransferase: RNA modification through a Channel. J Mol Biol 367:872–881
Yang Y, Miao Y, Wang B, Cui G, Merz KM Jr (2012) Catalytic mechanism of aromatic prenylation by NphB. Biochemistry 51:2606–2618
Yazaki K, Kunihisa M, Fujisaki T, Sato F (2002) Geranyl diphosphate:4-hydroxybenzoate geranyltransferase from Lithospermum erythrorhizon. Cloning and characterization of a key enzyme in shikonin biosynthesis. J Biol Chem 277:6240–6246
Yazaki K, Sasaki K, Tsurumaru Y (2009) Prenylation of aromatic compounds, a key diversification of plant secondary metabolites. Phytochemistry 70:1739–1745
Yin W-B, Ruan H-L, Westrich L, Grundmann A, Li S-M (2007) CdpNPT, an N-prenyltransferase from Aspergillus fumigatus: overproduction, purification and biochemical characterisation. Chembiochem 8:1154–1161
Yin W-B, Grundmann A, Cheng J, Li S-M (2009) Acetylaszonalenin biosynthesis in Neosartorya fischeri: Identification of the biosynthetic gene cluster by genomic mining and functional proof of the genes by biochemical investigation. J Biol Chem 284:100–109
Yin W-B, Yu X, Xie X-L, Li S-M (2010) Preparation of pyrrolo[2,3-b]indoles carrying a ß-configured reverse C3-dimethylallyl moiety by using a recombinant prenyltransferase CdpC3PT. Org Biomol Chem 8:2430–2438
Yin S, Yu X, Wang Q, Liu XQ, Li S-M (2013a) Identification of a brevianamide F reverse prenyltransferase BrePT from Aspergillus versicolor with a broad substrate specificity towards tryptophan-containing cyclic dipeptides. Appl Microbiol Biotechnol 97:1649–1660
Yin WB, Chooi YH, Smith AR, Cacho RA, Hu Y, White TC, Tang Y (2013b) Discovery of cryptic polyketide metabolites from dermatophytes using heterologous expression in Aspergillus nidulans. ACS Synth Biol 2:629–634
Yin W-B, Baccile JA, Bok JW, Chen Y, Keller NP, Schroeder FC (2013c) A nonribosomal peptide synthetase-derived iron(III) complex from the pathogenic fungus Aspergillus fumigatus. J Am Chem Soc 135:2064–2067
Yu X, Li S-M (2012) Prenyltransferases of the dimethylallyltryptophan synthase superfamily. Methods Enzymol 516:259–278
Yu X, Liu Y, Xie X, Zheng X-D, Li S-M (2012) Biochemical characterization of indole prenyltransferases: Filling the last gap of prenylation positions by a 5-dimethylallyltryptophan synthase from Aspergillus clavatus. J Biol Chem 287:1371–1380
Yu X, Zocher G, Xie X, Liebhold M, Schütz S, Stehle T, Li S-M (2013) Catalytic mechanism of stereospecific formation of cis-configured prenylated pyrroloindoline diketopiperazines by indole prenyltransferases. Chem Biol 20:1492–1501
Zeyhle P, Bauer JS, Kalinowski J, Shin-ya K, Gross H, Heide L (2014a) Genome-based discovery of a novel membrane-bound 1,6-dihydroxyphenazine prenyltransferase from a marine actinomycete. PLoS One 9, e99122
Zeyhle P, Bauer JS, Steimle M, Leipoldt F, Rösch M, Kalinowski J, Gross H, Heide L (2014b) A membrane-bound prenyltransferase catalyzes the O-prenylation of 1,6-dihydroxyphenazine in the marine bacterium Streptomyces sp. CNQ-509. Chembiochem 15:2385–2392
Zhao L, Chang WC, Xiao Y, Liu HW, Liu P (2013) Methylerythritol phosphate pathway of isoprenoid biosynthesis. Annu Rev Biochem 82:497–530
Zhu HY, Desai J, Cooper AB, Wang J, Rane DF, Kirschmeier P, Strickland C, Liu M, Nomeir AA, Girijavallabhan VM (2014) New class of azaheptapyridine FPT inhibitors as potential cancer therapy agents. Bioorg Med Chem Lett 24:1228–1231
Zocher G, Saleh O, Heim JB, Herbst DA, Heide L, Stehle T (2012) Structure-based engineering increased the catalytic turnover rate of a novel phenazine prenyltransferase. PLoS One 7, e48427
Zou H-X, Xie X-L, Linne U, Zheng X-D, Li S-M (2010) Simultaneous C7- and N1-prenylation of cyclo-L-Trp-L-Trp catalyzed by a prenyltransferase from Aspergillus oryzae. Org Biomol Chem 8:3037–3044
Zou Y, Zhan Z, Li D, Tang M, Cacho RA, Watanabe K, Tang Y (2015) Tandem prenyltransferases catalyze isoprenoid elongation and complexity generation in biosynthesis of quinolone alkaloids. J Am Chem Soc 137:4980–4983
Acknowledgments
The works in the authors’ laboratory were supported in part by a grant from Deutsche Forschungsgemeinschaft (Li844/4-1 to S.-M. Li). Julia Winkelblech is partially financed by the LOEWE program of the State of Hessen (SynMikro to S.-M. Li). Aili Fan is a recipient of a scholarship from China scholarship council.
Conflict of interest
The authors declare that there are no conflicts of interest.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Winkelblech, J., Fan, A. & Li, SM. Prenyltransferases as key enzymes in primary and secondary metabolism. Appl Microbiol Biotechnol 99, 7379–7397 (2015). https://doi.org/10.1007/s00253-015-6811-y
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00253-015-6811-y