Abstract
Genetic diversity is crucial for successful adaptation and sustained improvement in crops. India is bestowed with diverse agro-climatic conditions which makes it rich in wheat germplasm adapted to various niches. Germplasm repository consists of local landraces, trait specific genetic stocks including introgressions from wild relatives, exotic collections, released varieties, and improved germplasm. Characterization of genetic diversity is done using morpho-physiological characters as well as by analyzing variations at DNA level. However, there are not many reports on array based high throughput SNP markers having characteristics of genome wide coverage employed in Indian spring wheat germplasm. Amongst wheat SNP arrays, 35K Axiom Wheat Breeder’s Array has the highest SNP polymorphism efficiency suitable for genetic mapping and genetic diversity characterization. Therefore, genotyping was done using 35K in 483 wheat genotypes resulting in 14,650 quality filtered SNPs, that were distributed across the B (~ 50%), A (~ 39%), and D (~ 10%) genomes. The total genetic distance coverage was 4477.85 cM with 3.27 SNP/cM and 0.49 cM/SNP as average marker density and average inter-marker distance, respectively. The PIC ranged from 0.09 to 0.38 with an average of 0.29 across genomes. Population structure and Principal Coordinate Analysis resulted in two subpopulations (SP1 and SP2). The analysis of molecular variance revealed the genetic variation of 2% among and 98% within subpopulations indicating high gene flow between SP1 and SP2. The subpopulation SP2 showed high level of genetic diversity based on genetic diversity indices viz. Shannon’s information index (I) = 0.648, expected heterozygosity (He) = 0.456 and unbiased expected heterozygosity (uHe) = 0.456. To the best of our knowledge, this study is the first to include the largest set of Indian wheat genotypes studied exclusively for genetic diversity. These findings may serve as a potential source for the identification of uncharacterized QTL/gene using genome wide association studies and marker assisted selection in wheat breeding programs.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
The future of wheat production program demands a 2.4% increase in yield per year to reach a goal of 70% by 2050 of the current wheat production. However, the current rate of increase in the global average yield is only 0.9% per year [1]. This can be achieved by improvement in crop management practices and genetic improvement of cultivars for better yield [2]. The foundation for such an improvement can be found in genetic diversity [3, 4]. For instance, genetic diversity might be important for the adaptation and acquiring defensive characteristics in wheat varieties against biotic stress. Uniformity in a population will certainly show similar behavior against such a threat and will be unable to withstand an epidemic situation. It has been a long road in wheat evolution from einkorn to bread wheat, where wheat has assimilated several genetic variations. But now, it is reducing due to narrow adaptation, farmer’s selection, repeated cultivation of selected landraces, uniform varietal seed production by industries, etc. [5]. DNA markers such as restriction fragment length polymorphism (RFLP) [6], amplified fragment length polymorphism (AFLP) [7], inter simple sequence repeat (ISSR) [5], simple sequence repeat (SSR) [8], diversity arrays technology (DArT) [9], etc. are well used for genetic variation or diversity analysis in hexaploid wheat. Such study has been reported in Indian hexaploid wheat germplasm using SSRs [10, 11]. The number of markers and genotypes used in these studies seems insufficient to pursue studies like genome-wide association mapping [12]. Moreover, the Indian genotypes used in these reports can simply be classified as cultivars [10], released varieties, elite lines, and genetic stocks [11].
Single Nucleotide Polymorphisms (SNPs) are the best choice for genomic studies which demands a large number of markers with whole genome coverage. A comparative study between SNP and SSR for studying population structure and genetic diversity showed that SNP can be a more valuable tool for genomics approach and crop improvement [13]. The ubiquitous presence, uniform distribution, high heritability, and bi-allelic nature makes SNPs widely accepted molecular marker in terms of being high throughput [14]. It has been widely used for studying population structure and genetic diversity of germplasm collection panels by the means of SNP array or genotyping-by-sequencing (GBS) approach. At present, the application of SNP arrays in many polyploid crops [15] has been extensively reported. Amongst these SNP arrays, 35K has the highest SNP efficiency (94.8%) in terms of polymorphism suitable for genetic mapping and genetic diversity characterization in wheat [15,16,17]. As a brief history, Winfield et al. [16] used the Exom-seq (~ 57 Mb) information to identify 921K SNP variants further narrowing down to 820K. In order to update the 820K SNP array, 35K SNPs were selected with the consideration of high polymorphic rate and more even distribution for constructing genetic maps and characterizing novel genetic diversity [17].
Several genetic diversity studies based on SNPs have been reported in wheat using Genotyping-By-Sequencing (GBS) method [18,19,20]. The major limitations to the GBS approach are large percentage of missing data and uneven genome coverage due to selection of restriction enzyme used for fragment selection [21]. SNP array platform is known to be of high quality, reliable and robust for a multitude of applications in diversity studies and breeding applications [22,23,24]. It has certain advantages over NGS based GBS (Genotyping-By-Sequencing) method. Firstly, SNP array data is relatively easy to analyze and genotypes of SNP markers can be called as per the user guide. Since SNP calling requires read trimming, read alignment, SNP calling and filtering, etc. [25], it requires knowledge and background of bioinformatics. Secondly, genomic region of interest can be probed on SNP array. The number of such probes is flexible in Illumina and Affymetrix platforms. Thirdly, SNP array costs low to moderate per sample, e.g. Affymetrix Axiom array costs around $28–90 (USD) per sample [15]. So far, 35K SNP array has been used to study genetic differentiation between landraces and improved varieties in a panel of 370 durum wheat [26] with 8173 SNPs. In our previous study, this array was used on 404 wheat accessions to perform genome-wide association studies (GWAS) using 14,160 SNPs, with little focus on genetic differentiation studies [27]. This study also used a complete set of filtered markers for the same which may have led to an overestimation of subpopulations due to tightly linked markers [28]. Here we report, genetic diversity and population structure studies on an updated diverse study panel utilizing 14,650 informative SNP markers. The present study was undertaken in the current study panel to characterize the genetic diversity and population structure along with genetic differentiation within and among subpopulations. This study uniquely describes SNP array-based population structure and genetic diversity with the highest number of wheat genotypes in a study panel. This will prove a groundwork for future genomic selection in wheat breeding programs or GWAS.
Materials and methods
Plant material
A study panel of 483 Indian spring wheat (Triticum aestivum) genotypes was used in this study (Supplementary Table S1). These were obtained from Germplasm Resource Unit (GRU), ICAR-IIWBR, Karnal, India. This collection majorly comprised of varieties, improved genotypes, genetic stocks, landraces, exotic lines, etc., procured from different years adapted to different agro-climatic zones of India. A diverse panel is a prerequisite for understanding the population genetics and conducting association mapping studies.
DNA isolation and SNP genotyping
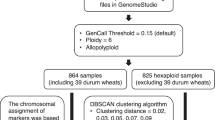
Genomic DNA was isolated from plants at growth stage (GS) 12 [29]. Modified CTAB method was applied for the DNA isolation [30]. Samples were snap frozen and crushed using liquid nitrogen. Then aqueous phase separation using Phenol: Chloroform: Isoamyl alcohol (25:24:1) was done after RNase treatment, at 13,000 g for 10 min at 4 °C. After precipitation with chilled isopropanol washing was done with 70% ethanol and pellet obtained was left to air-dry in a clean environment. The air-dried pellet was dissolved in nuclease free water. DNA isolated was quantified at 260 nm absorbance using NanoQuant Infinite® 200 spectrophotometer (TECAN). DNA samples were genotyped with 35K Axiom® Wheat Breeder’s Array (Affymetrix UK Ltd, UK) [17] as per manufacturer’s guidelines. The SNP array used comprised of 35,143 SNP markers. The SNP calls were visualized and optimized for user handling in Axiom analysis suite v2. Quality filtration was performed on these markers using PLINK v1.07 [31]. Minor allele frequency (MAF) less than 5% (--maf 0.05), individuals with more than 10% missing SNP calls (--mind 0.1) and markers with more than 10% missingness (--geno 0.1) were considered for filtration. Physical map positions of complete set of SNP markers were obtained and studied from Ensembl plants Triticum aestivum database (https://plants.ensembl.org/Triticum_aestivum). Further, on the basis of high-density consensus map provided on CerealDb [17, 32], markers lacking information for genetic distance and consensus chromosome location were removed. Consequently, 14,650 SNP markers and 483 genotypes in the study panel were subjected to further analysis. The collinearity of these filtered SNPs was compared on the basis of their genetic distance and physical position. A pictorial representation using 13,557 SNPs was made using Circos v0.67 software [33] by removing markers (1093) lacking information in terms of their physical position.
Genetic features of markers
Genetic diversity of a population is generally defined by parameters such as gene diversity (GD), Polymorphism Information Content (PIC) and MAF. GD is based on Nei’s gene diversity, which is a probability estimate for two randomly selected markers to be different in a population. GD for a locus is known as expected heterozygosity (He), being the fundamental measure which describes the expected heterozygous genotypes in genetic studies under Hardy–Weinberg Equilibrium [34]. PIC reflects the probability that two arbitrary samples in the analysis are polymorphic in nature. Both parameters were calculated using POWERMARKER v 3.25 [35] with the filtered set of 14,650 SNP markers. The GD [35] and PIC [36] was estimated based on the equations as follows:
where n = number of distinct alleles at any given locus; Pi (i = 1, 2,…, n) = frequency of allele n in the population.
where Pi and Pj are the frequencies of ith and jth alleles for any given marker i, respectively. MAF refers to the occurrence of the second most allele frequency in any given population under study [37] and value below 0.05 is discarded in most genetic studies. MAF for the filtered set of SNPs was calculated in PLINK (–freq). Other parameters such as average pairwise divergence or observed nucleotide diversity, expected nucleotide diversity or estimated mutation rate and Tajima’s D [38] were calculated across genomes using TASSEL v5.2 [39].
Population structure analysis
To determine the population structure, filtered marker set (14,650) was pruned using LD (Linkage Disequilibrium) based pruning method in PLINK (--indep-pairwise 100 5 0.2). The resulted pruned markers (1544) were used for population structure analysis using Bayesian based model method in STRUCTURE 2.3.4 [40]. The command line python program StrAuto [41] was employed for the parallelization of STRUCTURE run under Linux environment. Parallelized run provided an advantage over regular STRUCTURE by saving computational duration and ease of handling. The run parameters were 100,000 iterations of burn-in with 100,000 Monte Carlo Markov Chain (MCMC) iterations. K values were tested from 1 to 10 with five independent runs for each K. Most likely possible numbers of subpopulations (K) was determined by using web-based STRUCTURE HARVESTER [42], a ΔK statistics which depends on the rate of change in log probability [LnP(D)] between consecutive K values. CLUMPP v1.1.2 software [43] was used to generate a consolidated population (Q) matrix from the STRUCTURE runs for the best K value. Genotypes with membership coefficients greater than 0.5 were considered to belong in the same group. MS-Excel 2013 was used to draw a bar graph for the Q matrix. STRUCTURE run outputs were used to determine the fixation index (Fst) of each sub-population. PCoA (Principal Coordinates Analysis) was studied using DARwin v6 [44] on 14,650 SNP markers. The dissimilarity matrix was generated by performing 1000 bootstraps, which was further used in the cluster analysis of all genotypes using weighted neighbor-joining (NJ) method.
Genetic diversity indices and analysis of molecular variance (AMOVA)
In summary statistics, genetic diversity indices such as the number of different alleles (Na), number of effective alleles (Ne), observed heterozygosity (Ho), expected heterozygosity (He), and Shannon’s information index (I) were estimated using GenAlEx v6.5 [45]. For the estimation of genetic differentiation, number of subpopulations determined were used for the analysis of molecular variance (AMOVA) [46] and calculation of Nei’ genetic distance [47]. The analysis was performed on selected markers (8022) having PIC values 0.31 to 0.38. Population pairwise PhiPT value, a Fst analog which calculates population differentiation based on genotypic variance suppressing the within-population variance [48], was calculated along with the estimates of gene flow (Nm, number of migrants per generation = 0.25[(1/PhiPT) − 1]), in GenAlEx 6.5 [45]. A total of 9999 permutations were performed to obtain significance.
Results
SNP marker statistics and distribution
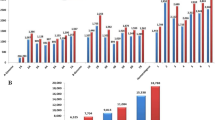
A total of 35,143 SNP markers were used to genotype 483 wheat individuals. Of 35,143 SNPs, monomorphic markers (6041), markers failing minor allele frequency test [MAF < 0.05] (8123) and missingness test [GENO > 0.1] (1412) were removed. Further, 5055 SNPs lacking information in the consensus genetic map for genetic distance and chromosomal location were also removed. No individuals failed for having more than 10 percent missing SNP calls (MIND > 0.1). Therefore, after quality filtration, 483 genotypes with 14,650 markers were used for further analysis. These markers covered a total genetic distance of 4477.85 cM. The B genome was observed to have the maximum numbers of filtered SNP markers (7377, ~ 50%) followed by A genome (5771, ~ 39%) and D genome (1502, ~ 10%). Supporting this, marker density was 1053.85, 824.42 and 214.57 per chromosome for the B, A, and D genome, respectively. Chromosome 2B comprised of a maximum number of genetically mapped SNP markers (1360). The lowest number of SNP markers were genetically mapped to chromosome 4D (61) (Supplementary Table S2, Fig. 1). Physical map positions of the SNP markers were obtained from the Ensembl database. Of 35,143 markers, 32,413 SNPs were found with available information of physical positions, distributed across the genome (Fig. 2a). The distribution of markers was observed in an appropriate window size across chromosomes using the R package CMplot (https://github.com/YinLiLin/R-CMplot). It was found that the markers that have been obtained after filtration are non-randomly distributed except for the central chromosomal region as per their physical position (Fig. 2b). There was a correlation of 0.9 due to the difference in chromosomal location of SNPs based on genetic distance and physical position (Supplementary Table S3). Therefore, we relied on the consensus chromosomal location from the genetic map in this study. The Circos plot (Fig. 2c) indicates that SNPs used in variability and structure analysis of all 483 genotypes of wheat were covered by more than 450 SNP per chromosome. It also depicts the correlation between the genetic and physical map based chromosomal locations of SNP markers graphically. Hence, 14,650 SNP markers were considered good enough to represent genome variability in totality of each genotype for population structure study.
Genome wide SNP marker density on the basis of their physical position for a all available markers of 35 K SNP array and b the filtered set of markers used in this study. Color gradient scale indicates the region rich and poor in SNPs with respect to their numbers. c Schematic representation of relationship between chromosomal location of any one marker based on genetic and physical maps. Gen-1A to Gen-7D represent the 21 wheat chromosomal genetic maps and Phy-1A to Phy-7D represent the 21 wheat chromosomal physical maps
Population structure analysis
To study population structure in the panel of 483 genotypes, delta K (ΔK) values were used to infer the number of subpopulations present. The suitable value of K was obtained from the plot between number of clusters (K) against ΔK where K = 2 showed the maximum value (Fig. 3a). A gradual increase in the assessed log likelihood with an increase in the number of K supports the defined number of sub-populations to be K = 2 (Fig. 3b). This also indicated that the two subpopulations could include all the 483 genotypes with high probability. The two sub-populations were designated as SP1 and SP2, which comprised of 106 and 377 genotypes, respectively. In SP1, the major contribution was observed to be made by the landraces (~ 66%) whereas, in SP2 it was varietal lines (~ 44%) and improved genotypes (~ 30%) collectively (Supplementary Table S1). In addition, contributions made by exotic lines and genetic stocks were 11.32% in SP1 and 17.50% in SP2. Despite the major contributor to SP1, landraces were also observed in SP2 (6.6%). Similarly, varietal lines and improved genotypes majorly representing SP2 were also component of SP1 with 17.9% and 3.8% contributions, respectively. The fixation index defines the overall genetic variation among subpopulations which is essentially determined to test the population substructure. The fixation index values from STRUCTURE runs for SP1 and SP2 were 0.196 and 0.532, respectively. The principal coordinate analysis (PCoA) performed using DARwin v6 tool was found in agreement with the results of STRUCTURE. Two separate clusters were observed in PCoA as shown in Fig. 4a. The first coordinate explained 20.69% and second coordinate explained 5.66% of the variation. The major cluster observed in the PCoA comprised of mainly varietal lines followed by improved genotypes (Fig. 4b). A NJ phylogenetic tree (Supplementary Fig. S1a) was constructed with 14,650 SNPs to represent the genetic distances among the population.
Principal coordinate analysis (PCoA) on 483 genotypes using 14,650 SNPs. a Two separate clusters were observed, marked as red and black colors indicating SP1 and SP2 where each number represents a genotype. b The major cluster (on the right) represents primarily varietal lines (black) and improved genotypes (pink) followed by exotic lines (green), genetic stocks (orange), landraces (red), and mutant lines (cyan). (Color figure online)
Genetic diversity analysis and PIC
Gene diversity (GD) in this study ranged from 0.095 to 0.5 with the highest mean in chromosome 5B (0.39) and lowest in chromosome 5D (0.27). Among the three genomes, the B genome showed the highest mean gene diversity (0.37). Polymorphism information content (PIC) was observed to range from 0.0905 to 0.3750. Highest mean PIC value was observed in chromosome 5B (0.31) and lowest in chromosome 5D (0.23) (Supplementary Table S2, Fig. 1). GD with a value of about 0.5 was observed in maximum number of markers (34.23%) followed by 0.4 (26.88%) and observed least for the value of 0.1 (8.14%). Minor allele frequency from 0.1 to 0.4 was observed in a similar number of markers (approx. 21%). Whereas, MAF of 0.5 was observed in only 10.77% of the total markers. PIC, with a value of 0.3 was observed in most of the markers (38.06%) followed by the value of 0.4 (Fig. 5a). At the genome level, both A and B genomes had more value than the D genome for both GD and PIC (Supplementary Table S2). The following trend was observed for mean MAF in the genomes; D (0.2584) < A (0.2655) < B (0.2795). In a separate analysis, SP1 and SP2 were studied individually for corresponding GD and PIC (Fig. 5b). For each subpopulation, common markers with MAF ≥ 5% were considered for this observation. The average PIC of 0.291 was observed in SP2, higher than that in SP1 (0.245). The same observation was seen with mean GD between SP2 (0.365) and SP1 (0.298). In SP2, chromosome 1D was observed to have the highest value for GD and PIC. All chromosomes except for 5D and 6D in SP1 were observed with lower values for the same when compared to SP2. In this study, landraces representing 20% genotypes in the study panel was observed to have a GD of 0.316. Whereas, the remaining 80% of the study panel showed slightly higher GD of 0.379.
Values of similar magnitude were obtained in diversity summary statistics, even for separate genome (Table 1). Observed nucleotide diversity (π/bp) (Table 1) and gene diversity (GD) (Supplementary Table S2) were found to have similar values and ranged from 0.34 (D genome) to 0.36 (B genome). Expected number of polymorphic sites or expected nucleotide diversity (θ/bp) were also found to have a similar value of 0.14 for all the genomes individually and as a whole. Tajima’s D, a population genetic test which computes a standardized measure of the presence of total number of polymorphic sites or segregating sites in the genotyped samples. This test distinguishes between DNA sites that may have evolved neutrally and those evolved directionally or non-randomly. In our case, Tajima’s D values ranged from 4.1 (D genome) to 4.6 (B genome). This value showed significant deviation from the neutral evolution (D = 0) and the population may have gone through balancing selection. A positive value of D also indicates that rare alleles might be present at low frequency in the population.
Subpopulation genetic differentiation and Allelic pattern
The two subpopulations observed in the study were considered for the calculation of AMOVA and genetic diversity indices. In AMOVA, genetic differentiation was determined using PhiPT (analog of FST) which represents the total genetic variation among populations. AMOVA estimates the molecular variance at two levels (i) between or among subpopulations, and (ii) within subpopulations. In this study, 2% variation was observed among subpopulations, while the rest of the variation of 98% was observed within subpopulations (Table 2). Therefore, the genetic differentiation observed after 9999 permutations among subpopulations was low and within subpopulations, it was observed to be high. PhiPT value and estimates of gene flow (Nm) were calculated between the two subpopulations SP1 and SP2 in GenAlEx (Table 2). The values observed for PhiPT and Nm were 0.016 and 15.563, respectively. Nei’ genetic distance between SP1 and SP2 was observed to be 0.019. The weighted NJ cluster analysis was performed again with 8022 SNPs with 1000 bootstraps (Supplementary Fig. S1b). The resulted tree was found in agreement with the PCoA and Population structure analysis done on 14,650 SNPs and also suitably represents the genetic differentiation of the study panel in SP1 and SP2 as compared to the NJ tree generated with complete filtered set of markers (Supplementary Fig. S1a). Genetic diversity indices were estimated on the two subpopulations observed in this study are shown in Table 3. The mean value of number of different alleles and number of effective alleles for the two subpopulations were 2.0 and 1.834, respectively. Mean values for I, Ho, He, and uHe were found 0.643, 0.003, 0.451, and 0.452, respectively (Table 3). SP2 showed more diversity as compared to SP1 (with I = 0.648, He = 0.456 and uHe = 0.456). The percentage of polymorphic loci (PPL) per population was observed to be approximately 100%.
Discussion
The genotypes used in this study as genotypic panel are primarily grown in India. Different categories involved in this study panel harbors the characteristics of their own. The objective behind using such material is to infer genetic diversity on the basis of high throughput SNPs in Indian spring wheat germplasm. This may further help in easy identification of diverse source for novel alleles applicable to wheat breeding. It is well known that subsequent domestication and frequent inbreeding poses a major issue for breeders in developing new varieties [49]. Introgressing alien DNA segment from wild relatives has its own limitation referred to as linkage drag of undesirable traits [50]. Novel sources for diversity are expected to be available in less explored genotypes such as exotic lines, wild relatives, and landraces.
We demonstrated the utilization of 35K Axiom Breeder’s Array in this study. This array can be used for genotyping diverse hexaploid wheat derived from various sources [17]. The genetic map for the 35K SNP markers estimated on five bi-parental populations that provided consensus chromosomal location of more than 20K SNPs [17] was used for this study. The information of 35K SNP array markers for their physical position was sorted out of 16,448,754 single nucleotide variations (SNV) available in the Ensembl wheat database. This information was supported by the fully annotated reference wheat genome [51]. In Supplementary Figure S2, the central region of chromosomes in the plot shows less SNV density as compared to the end regions. Out of 35,143 SNPs, information on physical position of 2730 SNPs is not available (Fig. 2a). In the filtered set of SNPs, the SNP density was observed to be very less at centromeric regions as compared to the end regions. The linkage map and physical map are supposed to be collinear and the order of linkage map to be highly correlated with the physical map [52]. Sometimes the linkage group shows concentrated regions of non-collinearity which usually corresponds to the centromeric regions [53]. It is known that crossing over does not occur at centromeres with estimates of suppression ranging from one to many folds depending upon the organism [54]. This non-collinearity might be due to the insufficient number of crossing over events at the centromere to accurately order the distribution of SNP markers. With the availability of fully annotated genome, physical positions of SNP markers were available, yet we followed the genetic distance information. The genetic distance has been reported suitable and as an alternative to physical position for conducting association studies [55].
The average marker density was 3.27 SNP/cM and 0.4924 cM/SNP as the average inter-marker distance. Thus, 35K wheat SNP array provided adequate polymorphic markers to conduct population structure and genetic diversity analysis. This can further be utilized for Linkage Disequilibrium and association analysis. The number of markers was highest in the B genome followed by A genome and least in the D genome. This was in concordance with previous studies where D genome had two [20, 56] to five [23, 57] times less number of markers as compared to A or B genome. An another study reported that a low number of polymorphic markers on the D genome have been put forth as a characteristic of wheat instead of its progenitor Aegilops tauschii [58]. In our findings, polymorphic markers in D genome was 3 times less than A genome and five times less than the B genome. Following the trend reported by Rimbert et al. [59], the homoeologous chromosome group 4 in this study also possessed the least number of filtered markers in all the three genomes (Fig. 1). Chromosome 4D possessed the least number of markers after filtration supported by other studies [20, 59]. As an observation, the average transition/transversion (Ts/Tv) ratio in the filtered set of markers was 2.13 for the three genomes. Ts/Tv ratio was determined as 2.290, 2.219 and 1.883 for genome A, B, and D, respectively (Supplementary Table S4). The average Ts/Tv ratio was found higher than a previous report [20]. It was also found in agreement where genome A had more Ts/Tv ratio followed by genome B and D, suggesting A/G and C/T mutations with high frequencies followed by methylation in the SNP markers used [20]. Transition mutation frequency ranging from 1.59 to 2.80 has been reported in several other studies including hexaploid wheat [60,61,62], Barley [63], and Camelina [64]. Several studies support the fact that transition SNP types are preferred over transversion SNPs, in addition to INDELs or multiple allelic SNPs, for SNP array development [65, 66].
As STRUCTURE assumes loci are at linkage equilibrium within subpopulations, using pruned markers instead of the whole set was found to be more reasonable [67]. This would also reduce the overestimation of sub-populations due to markers in strong LD [28]. Population structure helps in understanding genetic diversity and subsequent association mapping studies in a population, where it might pose as a limiting factor in the interpretation of results [18]. Presence of subgroups in the large populations can be justified by selection and genetic drift [68, 69]. Therefore, testing for population structure should be the first priority while conducting GWAS while identifying association between markers and the trait of interest. It is important because the presence of a structure in a mapping population may cause spurious association results [70]. A population designed for GWAS generally comprises of both population structure and familial relatedness. This is due to local adaptation and trait specific selection breeding [71]. For better inference, PCoA summarizes and represents the relation between a number of objects (in our case, the genotypes), in a simple Euclidean space. In our study, PCoA results concurred with those of STRUCTURE (K = 2), indicating that 483 genotypes could be clustered into two subgroups. As an input PCoA considers a dissimilarity matrix instead of raw data and explains the variability present in the dataset summarized by uncorrelated axes. The extent of variation gets represented by the magnitude of the eigenvalue of an axis. In simple terms, PCoA interprets that the closely ordinated objects will be more similar. The NJ tree, in addition, gave a similar pattern. The results were expected for the reason that the Indian landrace collection that has been less explored for traits like multiple rust resistance, were intentionally introduced in the study panel. Another possible explanation could be the selection of lines for breeding programs for certain specific traits. For instance, genotypes in SP2 had moderate to short plant height, whereas genotypes in SP1 were mostly tall in nature (data not shown). Genotypes that have been involved often in the breeding programs resulted in becoming varieties, improved genotypes, genetic stocks, etc. can be seen mostly in SP2 (Supplementary Table S1, Fig. 3c). The resolution of genetic relatedness is dependent on the number of markers and genotypes used in a study. The lower they are the higher the resolution will be in terms of similarity coefficient, but this will limit the exploitation purposes in finding novel alleles for desirable traits and will be exhausted in short term. The dissimilarity coefficient increases with the increasing number of genotypes and markers, which can give a possible overview of the collection in use. For long term breeding objective it is better to explore the genetic relatedness between the genotypes using larger set of genotypes and subjecting them to high density marker profiling. Our study includes more number of genotypes with high density of polymorphic markers.
Gene diversity (known otherwise as expected heterozygosity, He) and PIC are measures of genetic diversity shedding light on the evolutionary pressure on the alleles, in any given breeding population, and mutation rate at a locus over a time period [36, 72]. The overall Genetic diversity in a population is mainly reflected by the distribution of informative markers [3]. GD provides gene diversity of haploid markers and provides a range of average heterozygosity and genetic distance in a population among individuals [72, 73]. In our study, the overall GD value was found greater than the overall PIC value as expected. In the absence of more number of alleles and increasing evenness of allele frequencies PIC will always be lower than GD [72]. The PIC value of a marker dictates the property of that marker to be informative. Such informative markers with good PIC values can be used for genotyping plant population and genetic diversity studies [74]. There are three categories of PIC values; PIC values > 0.5 are considered highly informative, PIC values ranging 0.25–0.5 are considered moderately informative, and PIC values < 0.25 are considered slightly informative [36]. Since SNP markers are bi-allelic in nature, their PIC values are considered to be moderate or low informative and also restricted to extreme PIC values of 0.5 [18]. In this study, the maximum value of PIC was observed at 0.38 with an overall average of 0.29 between the genomes (Supplementary Table S2). Although it fell under the category of being moderately informative, yet it was observed higher than the average PIC value in previously reported studies in wheat [18, 75,76,77,78]. Studies reported on winter wheat [18], jujube [79] and ryegrass [80], supports that markers used in such studies are acceptable for being moderate to low informative. The results on GD showed that SP2 is more diverse than SP1. Since SP1 comprised mainly of landraces, it was expected to have more diversity as compared to modern germplasm in SP2. There is no denial in the fact that landraces are rich source of novel alleles and genetic diversity [81]. The proportion of landraces to other genotypes in this study was 1:4, even though a competitive GD value of 0.316 was observed in landraces. In Fig. 5b, it is worth noting that despite having less number of markers in the D genome when compared to other genomes, the markers were highly informative. For instance, GD and PIC values in 1D (>), 7D (>), and 4D in SP2; and chromosome 6D (>) and 5D in SP1 showed distinction from the other chromosomes. The genotypes in each subpopulation can be expected for playing a key role in this observation. Another study showed a similar behavior with the GD and PIC values [82]. They found high values for the same in 1D (cultivars) and 6D (landraces). In our study, Tajima’s D values were observed higher than that reported previously on synthetic hexaploid wheat panel [19]. As mentioned earlier, the positive value of Tajima’s D depicts the presence of low frequency of rare alleles in a given population. This might be a possible reason that we could not find any private allele in our study panel. Private alleles are important in identifying unique genetic variability at loci and diverse genotypes that can be used in a breeding program to improve the allele richness in a study panel or population [74]. The method applied in the present study could not capture rare alleles. To enable characterization of private/rare alleles some mechanism has to be devised which should be effective in capturing and harnessing the key adaptive genes.
The result of AMOVA implemented in this study inferred high level of genetic diversity within subpopulations and a low variation among subpopulations (2%). These variations were significant according to the partitioning value (p < 0.001). It also suppresses the intra-individual variation, thus becoming ideal for co-dominant data with a maximum of 10,000 permutations [83]. The possible explanation for high variation within groups is the selection for several agronomically important traits in various breeding programs. A low PhiPT value (0.016) was found between SP1 and SP2, indicating low genetic differentiation between these subpopulations. This coincided with the AMOVA results where only 2% of the total variation was accounted for by among-subpopulation variations. High variation within subpopulations symbolizes more frequent selection for economically important traits. When the value for gene flow is high one could expect a low level of diversity among subpopulation [11]. The value of Nm (15.563) in this study was higher than previous reports [18, 64]. This might be the reason where genetic exchange among subpopulations resulted in a low genetic differentiation among subpopulations. Rigorous efforts by the breeders get manifested in high Nm value indicating frequent gene flow among the subpopulations. The high value of Nm, therefore suggests that newer sources and hitherto less exploited genetic resources such as gene introgressed lines having genomic segments of interest shall be referred. Pre-breeding material utilizing synthetics and wild relatives will find their enhanced role under such circumstances. An indirect estimate of Nm from PhiPT in natural population violates several assumptions including constant population size, random migration, and no selection along with mutation and spatial structure [84]. Caution must be taken while interpreting Nm from indirect estimates, although it can still be useful to know the magnitude of gene flow [85]. The understanding of genetic diversity in Indian spring wheat germplasm will help in future studies as well as in monitoring and maintenance of genetic diversity in a wheat breeding program.
Conclusion
In this study, we performed an array based SNP genotyping to expand the utility of SNP markers for genomic analysis. This study comprised a diverse panel of 483 genotypes. We identified two subpopulations, SP1 and SP2, based on unlinked SNP markers, natural adaptation and selection history for traits of interest. SP2 comprised genotypes that were mostly the result of selection. However, based on GD and PIC analysis, it was identified as genetically more diverse. As compared to SP1, it was identified exhibiting higher values for Shannon’s information index (I), expected heterozygosity (He), and unbiased expected heterozygosity (uHe). This kind of genetic diversity can be utilized for developing biotic and abiotic stress tolerant varieties adaptive to diverse agro-climatic regions. Modern day wheat improvement program involves multi-parent crosses with diverse pedigree, wild germplasm or alien gene introgressed lines, developing a broad genetic based population from which selections are made. This study will be beneficial to the wheat breeders in taking decision about the parental lines to be selected for further genetic improvement. The diversity information made available can be channelized using such approach. These results of genetic diversity and population structure will be crucial for future studies with genomic approaches such as genomic selection, marker-assisted selection, and GWAS.
Data availability
A password protected FTP link will be provided for the genotypic data access from the corresponding author on reasonable request.
References
Ray DK, Mueller ND, West PC, Foley JA (2013) Yield trends are insufficient to double global crop production by 2050. PLoS ONE. https://doi.org/10.1371/journal.pone.0066428
Sener O, Arslan M, Soysal Y, Erayman M (2009) Estimates of relative yield potential and genetic improvement of wheat cultivars in the Mediterranean region. J Agric Sci 147:323–332. https://doi.org/10.1017/S0021859609008454
Nielsen NH, Backes G, Stougaard J et al (2014) Genetic diversity and population structure analysis of European hexaploid bread wheat (Triticum aestivum L.) varieties. PLoS ONE. https://doi.org/10.1371/journal.pone.0094000
Govindaraj M, Vetriventhan M, Srinivasan M (2015) Importance of genetic diversity assessment in crop plants and its recent advances: an overview of its analytical perspectives. Genet Res Int. https://doi.org/10.1155/2015/431487
Khan MK, Pandey A, Thomas G et al (2015) Genetic diversity and population structure of wheat in India and Turkey. AoB Plants 7:plv083. https://doi.org/10.1093/aobpla/plv083
Kim HS, Ward RW (2000) Patterns of RFLP-based genetic diversity in germplasm pools of common wheat with different geographical or breeding program origins. Euphytica 115:197–208
Talebi R, Fayyaz F (2012) Quantitative evaluation of genetic diversity in Iranian modern cultivars of wheat (Triticum aestivum L.) using morphological and amplified fragment length polymorphism (AFLP) markers. Biharean Biol 6:14–18
Chao S, Zhang W, Dubcovsky J, Sorrells M (2007) Evaluation of genetic diversity and genome-wide linkage disequilibrium among US wheat (Triticum aestivum L.) germplasm representing different market classes. Crop Sci 47:1018–1030
Sohail Q, Manickavelu A, Ban T (2015) Genetic diversity analysis of Afghan wheat landraces (Triticum aestivum) using DArT markers. Genet Resour Crop Evol 62:1147–1157. https://doi.org/10.1007/s10722-015-0219-5
Mir RR, Kumar J, Balyan HS, Gupta PK (2011) A study of genetic diversity among Indian bread wheat (Triticum aestivum L.) cultivars released during last 100 years. Genet Resour Crop Evol 59:717–726. https://doi.org/10.1007/s10722-011-9713-6
Arora A, Kundu S, Dilbaghi N et al (2014) Population structure and genetic diversity among Indian wheat varieties using microsatellite (SSR) markers. Aust J Crop Sci 8:1281–1289
Nordborg M, Weigel D (2008) Next-generation genetics in plants. Nature 456:720
Wurschum T, Langer SM, Longin CFH et al (2013) Population structure, genetic diversity and linkage disequilibrium in elite winter wheat assessed with SNP and SSR markers. Theor Appl Genet 126:1477–1486. https://doi.org/10.1007/s00122-013-2065-1
Verma S, Gupta S, Bandhiwal N et al (2015) High-density linkage map construction and mapping of seed trait QTLs in chickpea (Cicer arietinum L.) using genotyping-by-sequencing (GBS). Sci Rep. https://doi.org/10.1038/srep17512
You Q, Yang X, Peng Z et al (2018) Development and applications of a high throughput genotyping tool for polyploid crops: single nucleotide polymorphism (SNP) array. Front Plant Sci. https://doi.org/10.3389/fpls.2018.00104
Winfield MO, Allen AM, Burridge AJ et al (2016) High-density SNP genotyping array for hexaploid wheat and its secondary and tertiary gene pool. Plant Biotechnol J 14:1195–1206. https://doi.org/10.1111/pbi.12485
Allen AM, Winfield MO, Burridge AJ et al (2017) Characterization of a wheat breeders’ array suitable for high-throughput SNP genotyping of global accessions of hexaploid bread wheat (Triticum aestivum). Plant Biotechnol J 15:390–401. https://doi.org/10.1111/pbi.12635
Eltaher S, Sallam A, Belamkar V et al (2018) Genetic diversity and population structure of F3:6 nebraska winter wheat genotypes using genotyping-by-sequencing. Front Genet 9:1–9. https://doi.org/10.3389/fgene.2018.00076
Bhatta M, Morgounov A, Belamkar V et al (2018) Unlocking the novel genetic diversity and population structure of synthetic Hexaploid wheat. BMC Genom. https://doi.org/10.1186/s12864-018-4969-2
Alipour H, Bihamta MR, Mohammadi V et al (2017) Genotyping-by-sequencing (GBS) revealed molecular genetic diversity of Iranian wheat landraces and cultivars. Front Plant Sci 8:1293. https://doi.org/10.3389/fpls.2017.01293
Peterson G, Dong Y, Horbach C, Fu Y-B (2014) Genotyping-by-sequencing for plant genetic diversity analysis: a lab guide for SNP genotyping. Diversity 6:665–680
Elbasyoni IS, Lorenz AJ, Guttieri M et al (2018) A comparison between genotyping-by-sequencing and array-based scoring of SNPs for genomic prediction accuracy in winter wheat. Plant Sci 270:123–130. https://doi.org/10.1016/j.plantsci.2018.02.019
Cavanagh CR, Chao S, Wang S et al (2013) Genome-wide comparative diversity uncovers multiple targets of selection for improvement in hexaploid wheat landraces and cultivars. Proc Natl Acad Sci 110:8057–8062. https://doi.org/10.1073/pnas.1217133110
Chen H, Xie W, He H et al (2014) A high-density snp genotyping array for rice biology and molecular breeding. Mol Plant 7:541–553. https://doi.org/10.1093/mp/sst135
Clevenger J, Chu Y, Chavarro C et al (2017) Genome-wide SNP genotyping resolves signatures of selection and tetrasomic recombination in peanut. Mol Plant. https://doi.org/10.1016/j.molp.2016.11.015
Kabbaj H, Sall AT, Al-Abdallat A et al (2017) Genetic diversity within a global panel of durum wheat (Triticum durum) landraces and modern germplasm reveals the history of alleles exchange. Front Plant Sci 8:1277
Sheoran S, Jaiswal S, Kumar D et al (2019) Uncovering genomic regions associated with 36 agro-morphological traits in Indian spring wheat using GWAS. Front Plant Sci. https://doi.org/10.3389/fpls.2019.00527
Falush D, Stephens M, Pritchard JK (2003) Inference of population structure using multilocus genotype data: linked loci and correlated allele frequencies. Genetics 164:1567–1587
Zadoks J, Chang T, Konzak C (1974) A decimal growth code for the growth stages of cereals. Weed Res 14:415–421
Saghai-Maroof MA, Soliman KM, Jorgensen RA, Allard RW (1984) Ribosomal DNA spacer-length polymorphisms in barley: Mendelian inheritance, chromosomal location, and population dynamics. Proc Natl Acad Sci 81:8014–8018. https://doi.org/10.1073/pnas.81.24.8014
Purcell S, Neale B, Todd-Brown K et al (2007) PLINK: a tool set for whole-genome association and population-based linkage analyses. Am J Hum Genet 81:559–575. https://doi.org/10.1086/519795
Wilkinson PA, Winfield MO, Barker GLA et al (2012) CerealsDB 2.0: an integrated resource for plant breeders and scientists. BMC Bioinform 13:219. https://doi.org/10.1186/1471-2105-13-219
Krzywinski M, Schein J, Birol I et al (2009) Circos: an information aesthetic for comparative genomics. Genome Res. https://doi.org/10.1101/gr.092759.109
Nei M (1973) Analysis of gene diversity in subdivided populations. Proc Natl Acad Sci USA 70:3321–3323. https://doi.org/10.1073/pnas.70.12.3321
Liu K, Muse SV (2005) PowerMarker: an integrated analysis environment for genetic marker analysis. Bioinformatics 21:2128–2129
Botstein D, White RL, Skolnick M, Davis RW (1980) Construction of a genetic linkage map in man using restriction fragment length polymorphisms. Am J Hum Genet 32:314–331
Tabangin ME, Woo JG, Martin LJ (2009) The effect of minor allele frequency on the likelihood of obtaining false positives. BMC Proc. https://doi.org/10.1186/1753-6561-3-s7-s41
Tajima F (1989) Statistical method for testing the neutral mutation hypothesis by DNA polymorphism. Genetics 123:585–595
Bradbury PJ, Zhang Z, Kroon DE et al (2007) TASSEL: software for association mapping of complex traits in diverse samples. Bioinformatics 23:2633–2635. https://doi.org/10.1093/bioinformatics/btm308
Pritchard JK, Stephens M, Donnelly P (2000) Inference of population structure using multilocus genotype data. Genetics 155:945–959. https://doi.org/10.1111/j.1471-8286.2007.01758.x
Chhatre VE, Emerson KJ (2017) StrAuto: automation and parallelization of STRUCTURE analysis. BMC Bioinform 18:192. https://doi.org/10.1186/s12859-017-1593-0
Earl D, VonHoldt B (2011) STRUCTURE HARVESTER: a website and a program for visualizing STRUCTURE output and implementing the Evanno method. Conserv Genet Resour 1–3:359–361
Jakobsson M, Rosenberg NA (2007) CLUMPP: a cluster matching and permutation program for dealing with label switching and multimodality in analysis of population structure. Bioinformatics 23:1801–1806. https://doi.org/10.1093/bioinformatics/btm233
Perrier X, Jacquemoud-Collet J (2006) DARwin software. http://darwin.cirad.fr/darwin
Peakall R, Smouse PE (2012) GenALEx 6.5: genetic analysis in Excel. Population genetic software for teaching and research-an update. Bioinformatics 28:2537–2539. https://doi.org/10.1093/bioinformatics/bts460
Excoffier L, Smouse PE, Quattro JM (1992) Analysis of molecular variance inferred from metric distances among DNA haplotypes: application to human mitochondrial DNA restriction data. Genetics 131:479–491
Nei M (1972) Genetic distance between populations. Am Nat 106:283–292
Yamasaki M, Ideta O (2013) Population structure in Japanese rice population. Breed Sci 63:49–57. https://doi.org/10.1270/jsbbs.63.49
White J, Law JR, MacKay I et al (2008) The genetic diversity of UK, US and Australian cultivars of Triticum aestivum measured by DArT markers and considered by genome. Theor Appl Genet 116:439–453. https://doi.org/10.1007/s00122-007-0681-3
Klindworth DL, Hareland GA, Elias EM, Xu SS (2013) Attempted compensation for linkage drag affecting agronomic characteristics of durum wheat 1AS/1DL translocation lines. Crop Sci 53:422–429. https://doi.org/10.2135/cropsci2012.05.0310
Appels R, Eversole K, Stein N et al (2018) Shifting the limits in wheat research and breeding using a fully annotated reference genome. Science 361:eaar7191. https://doi.org/10.1126/science.aar7191
Maughan PJ, Yourstone SM, Byers RL et al (2010) Single-nucleotide polymorphism genotyping in mapping populations via genomic reduction and next-generation sequencing: proof of concept. Plant Genome J 3:166. https://doi.org/10.3835/plantgenome2010.07.0016
Hosouchi T, Kumekawa N, Tsuruoka H, Kotani H (2002) Physical map-based sizes of the centromeric regions of Arabidopsis thaliana chromosomes 1, 2, and 3. DNA Res 9:117–121. https://doi.org/10.1093/dnares/9.4.117
Talbert PB, Henikoff S (2010) Centromeres convert but don’t cross. PLoS Biol 8:e1000326. https://doi.org/10.1371/journal.pbio.1000326
Rodriguez-Fontenla C, Calaza M, Gonzalez A (2014) Genetic distance as an alternative to physical distance for definition of gene units in association studies. BMC Genom 15:1–9. https://doi.org/10.1186/1471-2164-15-408
Iehisa JCM, Shimizu A, Sato K et al (2014) Genome-wide marker development for the wheat D genome based on single nucleotide polymorphisms identified from transcripts in the wild wheat progenitor Aegilops tauschii. Theor Appl Genet 127:261–271. https://doi.org/10.1007/s00122-013-2215-5
Allen AM, Barker GLA, Wilkinson P et al (2013) Discovery and development of exome-based, co-dominant single nucleotide polymorphism markers in hexaploid wheat (Triticum aestivum L.). Plant Biotechnol J 11:279–295
Akhunov ED, Akhunova AR, Anderson OD et al (2010) Nucleotide diversity maps reveal variation in diversity among wheat genomes and chromosomes. BMC Genom. https://doi.org/10.1186/1471-2164-11-702
Rimbert H, Darrier B, Navarro J et al (2018) High throughput SNP discovery and genotyping in hexaploid wheat. PLoS ONE 13:1–19. https://doi.org/10.1371/journal.pone.0186329
Lorenc MT, Hayashi S, Stiller J et al (2012) Discovery of single nucleotide polymorphisms in complex genomes using SGSautoSNP. Biology 1:370–382
Winfield MO, Wilkinson PA, Allen AM et al (2012) Targeted re-sequencing of the allohexaploid wheat exome. Plant Biotechnol J 10:733–742
Manickavelu A, Jighly A, Ban T (2014) Molecular evaluation of orphan Afghan common wheat (Triticum aestivum L.) landraces collected by Dr. Kihara using single nucleotide polymorphic markers. BMC Plant Biol 14:320
Turuspekov Y, Ormanbekova D, Rsaliev A, Abugalieva S (2016) Genome-wide association study on stem rust resistance in Kazakh spring barley lines. BMC Plant Biol 16:6
Luo Z, Brock J, Dyer JM et al (2019) Genetic diversity and population structure of a camelina sativa spring panel. Front Plant Sci. https://doi.org/10.3389/fpls.2019.00184
Bianco L, Cestaro A, Linsmith G et al (2016) Development and validation of the Axiom®Apple480 K SNP genotyping array. Plant J 86:62–74. https://doi.org/10.1111/tpj.13145
Clarke WE, Higgins EE, Plieske J et al (2016) A high-density SNP genotyping array for Brassica napus and its ancestral diploid species based on optimised selection of single-locus markers in the allotetraploid genome. Theor Appl Genet 129:1887–1899. https://doi.org/10.1007/s00122-016-2746-7
Gao L, Kathryn Turner M, Chao S et al (2016) Genome wide association study of seedling and adult plant leaf rust resistance in elite spring wheat breeding lines. PLoS ONE 11:e0148671. https://doi.org/10.1371/journal.pone.0148671
Buckler ES IV, Thornsberry JM (2002) Plant molecular diversity and applications to genomics. Curr Opin Plant Biol 5:107–111
Breseghello F, Sorrells ME (2006) Association mapping of kernel size and milling quality in wheat (Triticum aestivum L.) cultivars. Genetics 172:1165–1177. https://doi.org/10.1534/genetics.105.044586
Oraguzie NC, Gardiner SE, Rikkerink EHA, Silva HN (2007) Association mapping in plants. Springer, Berlin
Zhu C, Gore M, Buckler ES, Yu J (2008) Status and prospects of association mapping in plants. Plant Genome J 1:5. https://doi.org/10.3835/plantgenome2008.02.0089
Shete S, Tiwari H, Elston RC (2000) On estimating the heterozygosity and polymorphism information content value. Theor Popul Biol 57:265–271
Nei M (1978) Estimation of average heterozygosity and genetic distance from a small number of individuals. Genetics 89:583–590
Salem KFM, Sallam A (2016) Analysis of population structure and genetic diversity of Egyptian and exotic rice (Oryza sativa L.) genotypes. C R Biol 339:1–9. https://doi.org/10.1016/j.crvi.2015.11.003
Allen AM, Barker GLAA, Berry ST et al (2011) Transcript-specific, single-nucleotide polymorphism discovery and linkage analysis in hexaploid bread wheat (Triticum aestivum L.). Plant Biotechnol J 9:1086–1099. https://doi.org/10.1111/j.1467-7652.2011.00628.x
Edwards KJ, Reid AL, Coghill JA et al (2009) Multiplex single nucleotide polymorphism (SNP)-based genotyping in allohexaploid wheat using padlock probes. Plant Biotechnol J 7:375–390. https://doi.org/10.1111/j.1467-7652.2009.00413.x
Chao S, Zhang W, Akhunov E et al (2009) Analysis of gene-derived SNP marker polymorphism in US wheat (Triticum aestivum L.) cultivars. Mol Breed 23:23–33. https://doi.org/10.1007/s11032-008-9210-6
Somers DJ, Kirkpatrick R, Moniwa M, Walsh A (2003) Mining single-nucleotide polymorphisms from hexaploid wheat ESTs. Genome 46:431–437. https://doi.org/10.1139/g03-027
Chen W, Hou L, Zhang Z et al (2017) Genetic diversity, population structure, and linkage disequilibrium of a core collection of Ziziphus jujuba assessed with genome-wide SNPs developed by genotyping-by-sequencing and SSR markers. Front Plant Sci. https://doi.org/10.3389/fpls.2017.00575
Roldán-Ruiz I, Dendauw J, Van Bockstaele E et al (2000) AFLP markers reveal high polymorphic rates in ryegrasses (Lolium spp.). Mol Breed 6:125–134. https://doi.org/10.1023/A:1009680614564
Azeez MA, Adubi AO, Durodola FA (2018) Landraces and crop genetic improvement, Ch. 1. In: Adubi AO (ed) Rediscovery of landraces as a resource for the future. IntechOpen, Rijeka
Muleta KT, Bulli P, Rynearson S et al (2017) Loci associated with resistance to stripe rust (Puccinia striiformis f. sp. tritici) in a core collection of spring wheat (Triticum aestivum). PLoS ONE 12:087. https://doi.org/10.1371/journal.pone.0179087
Assoumane A, Zoubeirou AM, Rodier-Goud M et al (2013) Highlighting the occurrence of tetraploidy in Acacia senegal (L.) Willd. and genetic variation patterns in its natural range revealed by DNA microsatellite markers. Tree Genet Genom 9:93–106. https://doi.org/10.1007/s11295-012-0537-0
Whitlock MC, McCauley DE (1999) Indirect estimation of gene flow and migration: FST ≠ 1(4Nm + 1). Heredity 82:117–125
Neigel JE (2002) Is F st obsolete? Conserv Genet 3:167–173
Acknowledgements
Authors would like to acknowledge the project funding provided by Indian Council of Agricultural Research; Ministry of Agriculture and Farmer’s welfare, Government of India in the form of CABin Grant (F.No. Agril. Edn.4-1/2013-A&P) to ICAR-IASRI and ICAR-IIWBR. Support provided by Department of Biotechnology, Ministry of Science and Technology, Government of India in the form of DBT-JRF fellowship program to the first author is also acknowledged. Additionally, kind support from Dr. Arun Gupta from the GRU is acknowledged. Suggestions provided by Ms. Swati Verma and Mr. Pawan Kumar, PhD scholars at CCSHAU, Hisar, India and Dr. Rakesh Kumar, Senior Research Fellow at ICAR-IIWBR, Karnal, India, are also acknowledged.
Author information
Authors and Affiliations
Contributions
DeK analyzed the data and wrote the manuscript. VC and RT provided guidance and edited the manuscript. SS provided the facility of workstation for computational purposes. Mutant lines used in this study was developed and provided by RS. AJ and JJ contributed in illustrations for the manuscript. RT, DiK, SS, RS, PS, SJ, MAI, UBA, AR and GPS contributed in conceiving and progression of the project and revising the manuscript. All authors read and approved the final version of the manuscript.
Corresponding author
Ethics declarations
Conflict of interest
All authors declare that that they have no conflict of interest.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
11033_2019_5132_MOESM1_ESM.tif
Supplementary material 1 (TIFF 1275 kb) Supplementary Fig. S1 Neighbor-joining cluster analysis has been shown with two subpopulations indicated with red (SP1) and black (SP2) colors. As compared to (a) the NJ tree generated with 14,650 SNP markers, the two clusters can be seen clearly in (b) the NJ tree obtained with 8022 SNP markers used in AMOVA
11033_2019_5132_MOESM2_ESM.jpg
Supplementary material 2 (JPEG 1256 kb) Supplementary Fig. S2 SNP marker density of more than 16 million SNVs in the wheat genome available on Ensembl Wheat database. Un: Unknown
Rights and permissions
About this article
Cite this article
Kumar, D., Chhokar, V., Sheoran, S. et al. Characterization of genetic diversity and population structure in wheat using array based SNP markers. Mol Biol Rep 47, 293–306 (2020). https://doi.org/10.1007/s11033-019-05132-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11033-019-05132-8