Abstract
Single nucleotide polymorphisms (SNP) in repair gene DNA such as XPC gene can reduce the DNA repair capacity (DRC). Reduced DRC induce genetic instability and may increase the susceptibility to prostate cancer (PC). We conducted a case-controls study to examine the relationship between XPC Lys939Gln and XPC-PAT polymorphisms and the risk for prostate cancer in Tunisian population. We have also correlated molecular results with clinical parameters (Gleason score and TNM status) and lifestyle factors (tobacco status, alcohol consumption, and exposition to professional risk factors) of prostate cancer patients. We have found that the XPC Lys939Gln polymorphism was not associated with a risk of prostate cancer. However the XPC PAT I/I genotype was found to be associated with 3.83-fold increased risk of prostate cancer compared to controls (p = 0.00006; OR 3.83; 95% CI (1.83–8.05)). The test of linkage disequilibrium showed that XPC-PAT polymorphism is in linkage disequilibrium with XPC Lys939Gln variants. The combined analysis of XPC Lys939Gln and XPC-PAT variants showed that patients who inherited (Lys/Gln + PAT D/D) genotypes were protected against prostate cancer development compared to controls. In the other hand, no significant association has been found between XPC polymorphisms and clinical parameters or between XPC polymorphisms and lifestyle factors.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Prostate cancer (PC) is the most frequently diagnosed malignancy among men in western countries and the sixth leading cause of cancer-specific death worldwide [1]. The International Agency for Research on Cancer reported that PC accounted for 15% of total new cancer cases in 2012. In Tunisia, PC is considered as the third most diagnosed cancer in men with the incidence of 14.1/105 person-years in 2007 [2]. External exposures, including lifestyle factors and genetic predisposition, were the most risk factors for prostate cancer [3,4,5]. Prostate-specific antigen (PSA) testing for PC has been in common practice for more than 20 years. Prognosis of prostate cancer depends on the risk stratification D’Amico classification [6]. This method is based on three factors: prostate specific antigen (PSA), Gleason score and clinical stage [6]. Deficient DNA repair capacity is known to be cancer predisposing factor. Indeed the presence of some Single nucleotide polymorphisms (SNP) in repair gene DNA can change the function of repair enzyme and corrupt their level of transcription or translation. These modifications can reduce the DNA repair capacity (DRC) and induce genetic instability, in turn, activate the susceptibility to carcinogenesis. There are five major important DNA-repair pathways consisting of more than 130 genes: nucleotide excision repair (NER), base excision repair (BER), mismatch repair (MMR), and double-strand break repair (DSBR) and transcription-coupled repair (TCR) [7]. Among these pathways, NER is the most important DNA repair mechanism responsible for various types of DNA damage consisting of oxidative DNA damage, bulky adducts cross-links, alkylating damage and thymidine dimmers [7].
Xeroderma pigmentosum complementary group C (XPC) is an important component of the NER pathway [8]. This gene is localized on chromosome 3p25 and hosts 16 exons (82–882 bp) and 15 introns (0.08–5.4 kb) encodes for a 940 amino acid protein that has a major role in the repair of complex protein formations [8]. The functional DNA-binding domains interact with HR23B to form a complex that recognizes and binds to the sites of DNA damage. Many researchers have confirmed the role of XPC in the regulation of the cell cycle for DNA damage response and oxidative damage on cells [9]. More than one hundred SNPs in the coding regions of XPC have been reported. The most common and studied SNPs are XPC Lys939Gln and XPC poly (AT) [10]. XPC Lys939Gln is a single nucleotide polymorphism (XPC rs2228001 A 33512> C) consisted in the modification of the nucleotide A to C leading to modification from Lysin to Glutamine proteins at 939 positions of the DNA repair XPC gene. Single nucleotide polymorphisms of XPC Lys939Gln can reduce recognition and repair of DNA damage, protein expression, thus leading to more somatic DNA mutations or alteration to occur [11]. XPC-PAT is an insertion/deletion polymorphisms of XPC gene consisting of an insertion of 83 bases of A to T [poly(AT)] and deletion of 5 bases (GTAAC) at positions 1457 to 1461 in intron 9 [12]. This polymorphism can inhibit DRC and increase the risk for tumorigenesis [13]. Although the XPC Lys939 Gln and XPC-PAT polymorphisms have been implicated with altered susceptibility in a number of cancers [14, 15]; studies focusing on their relationship with the risk of prostate cancer have produced the same results. Indeed, Hirata et al. suggested that the 939 Gln mutant allele was significantly higher in PC cases in the Japanese population [16] and Wu et al. have not found a significant relationship between XPC Lys939Gln variant and PC occurrence [17]. With considering the XPC-PAT polymorphism, some studies have reported that the mutant genotype (PAT I/I) is associated with increasing PC risk in different populations such as Indian, Chinese, Japanese, Iranian American and Caucasian [18, 19].
In view of that, this work addresses the relationship between XPC Lys939Gln and XPC-PAT polymorphisms and the predisposition to prostate cancer in a case-control study of a previously unstudied Tunisian cohort aiming at clarifying whether genetic variations in XPC represent risk factors for the development of prostate cancer in Tunisia. We have also analyzed the correlation between XPC polymorphisms and tumor characteristics and/or clinical outcome.
Materials and methods
Subjects
This project was approved by a local ethical committee. Informed consent was obtained from all individual participants included in the study. A total of 110 consecutive PC patients were included in this study. All patients had previously undergone prostate biopsy for detection of PC at the Department of Urology at the Charles Nicole Hospital of Tunis; Tunisia. The typical indications for prostate biopsy were: total PSA level > 4.0 ng/ml and abnormal digital rectal examination (DRE) for the prostate nodule and then they were confirmed by the result of the pathology report. The clinical and epidemiological parameters of PC cases were summarized in Table 1. Clinical Characteristics of prostate cancer were obtained from medical records, enrolling PSA level, Gleason pathological score, Tumor stage, and TNM status.
The controls were 266 men matched to PC patients for age and geographic origin. They included 145 individuals with benign prostatic hyperplasia (BPH) and normal PSA levels and 121 healthy individuals with unknown BPH status. BPH samples were identified by their normal pathological reports of transrectal prostate biopsy in the case when PSA was > 4.0 ng/ml. The anatomopathological reviewed the tumor characteristics and screened for the absence of any signs of malignancy in the Formalin Fixed Paraffin Embedded (FFPE) tissues of BPH controls and the follow up of PSA for BPH samples after treatment is < 4 ng/ml. Healthy individuals included in this study are volunteers enrolled in the external consultation and they have normal digital rectal examination (DRE), serum PSA < 0.2 ng/ml and devoid from any forms of cancer.
Molecular analysis
Peripheral blood samples were collected from all subjects into tubes with ethylene diamine tetra-acetic acid (EDTA) at pH 8. Genomic DNA was extracted by a conventional phenol/chloroform protocol [20]. The quantity and quality of extracted DNA were estimated by a Nanodrop.
The XPC–PAT polymorphism was genotyped by Polymerase Chain Reaction (PCR) followed by electrophoresis on 1% agarose gel. The used primers and fragments sizes were summarized in Table 2. The PCR reaction was carried out in a total volume of 25 µl containing 50 ng of genomic DNA, 1 X of DNA polymerase Taq buffer, 2.5 mM of MgCl2, 0.2 mM of dNTP, 0.32 µM of each primer and 1 U of Taq DNA polymerase (Invitrogen™). Thermal cycling conditions were: an initial step of 95 °C for 10 min followed by 35 cycles of denaturing step at 95 °C for 45 s, 63 °C for 45 s and 72 °C for 45 s, followed by a final elongation step at 72 °C for 10 min. The PCR product was resolved by agarose gel electrophoresis (1%) and visualized by UV radiation after ethidium bromide staining. The homozygous wild genotype (PAT D/D) has resulted in a 266-bp amplification. However, the homozygous mutant genotype (PAT I/I) was characterized by the presence of only a 344-bp fragment.
The XPC Lys939Gln was genotyped by PCR-RFLP (restriction fragment length polymorphism). The used PCR primers for XPC Lys939Gln polymorphism were summarized in Table 2. The volume of PCR reaction was carried out in a final volume of 25 µl containing 50 ng of genomic DNA, 0.24 µM of each primer, 1X of Taq DNA polymerase buffer, 2.5 mM of MgCl2, 0.20 mM of dNTP, and 1U of Taq DNA polymerase (Invitrogen™). After initial denaturation at 95 °C for 5 min, PCR reactions were carried out for 35 cycles at 95 °C for 45 s, 64 °C for 45 s and 72 °C for 45 s, followed by a final elongation step at 72 °C for 10 min. The PCR product was digested by 10U of a PvuII enzyme (New England Biolabs) for 4 h at 37 °C and analyzed by agarose gel electrophoresis at 1.5%. The wild genotype XPC Lys/Lys was characterized by the presence of only one fragment of 244 pb. However, the homozygous and heterozygous genotypes (Gln/Gln) and (Lys/Gln) were respectively marked by the presence of two fragments of 189 and 55 pb and three fragments of 244.189 and 55 pb.
Statistical analysis
The Hardy Weinberg equilibrium test as calculated by the software package Arlequin (version 3.01) [21]. The difference of genotypes frequencies between cases and controls were determined by the χ2 test. In addition odds ratios (ORs) and their 95% Confidence Intervals (95% CI) were calculated as a measure of the association of the polymorphic sites with prostate cancer risk. A p-value was considered significant at < 0.05.
Results
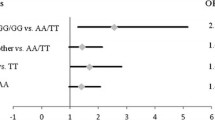
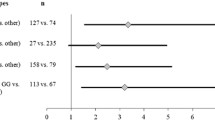
Hardy Weinberg equilibrium for XPC Lys939Gln and XPC-PAT polymorphisms for control subjects and prostate cancer patients are shown in Table 3. All samples were found to be in Hardy Weinberg equilibrium (p > 0.05) except for prostate cancer patients among whom the genotypic distribution for XPC-PAT polymorphism showed a statistical difference from Hardy Weinberg expectations (p = 0.03357). The frequencies of XPC*C allele (XPC 939*Gln) were estimated at 0.395 in both control group and prostate cancer patients. The frequencies of XPC*I allele were estimated at 0.256 and 0.400 in respectively control group and prostate cancer patients. The percentages of XPC PAT I/I genotypes in control group and prostate cancer patients were respectively estimated at 7.52% and 20.91%. The comparison of XPC Lys939Gln genotypes frequencies between control group and prostate cancer patients has not reported a significant difference (Table 4). However the XPC PAT I/I genotype has been found to be associated with 3.83 increased risk of prostate cancer compared to control group (p = 0.00006; OR 3.83; 95% CI 1.83–8.05). The test of linkage disequilibrium showed that XPC-PAT polymorphism is in linkage disequilibrium with XPC Lys939Gln variants. The combined analysis of XPC Lys939Gln and XPC-PAT variants showed that patients who inherited Lys/Gln + PAT D/D genotypes are protected against prostate cancer development compared to controls (Table 5).
The comparison between PC patients according to tobacco status and XPC Lys939Gln or XPC PAT polymorphisms doesn’t report any significant association (Table 6). This result suggests that there isn’t an additive effect between tobacco and the studied polymorphisms. The same result was obtained when we compare the distribution of XPC Lys939Gln or XPC-PAT polymorphism between drinkers and non-drinkers of alcohol (Table 6). In the other hand, the correlations between XPC gene polymorphisms and clinical parameters of prostate cancer patients (Gleason Score and TNM) don’t report any significant association (Tables 7, 8).
Discussion
In this population based case-control study, we investigated the effect of environmental risk factor and gene repair polymorphisms on prostate cancer (PC) development and their association with clinical and epidemiological parameters.
The allele frequency for XPC*C (XPC*Gln) in Tunisian control group is estimated at 0.395. This frequency is lower to which reported for the Caucasian and African populations [18, 22]. The comparison of PC and control group according to XPC Lys939Gln polymorphism has not reported a significant difference. This result confirms many recent meta-analyzes. Indeed, in the meta-analysis of Wu et al., authors have found that the XPC Lys939Gln polymorphism was not associated with PC susceptibility [17]. In this study, authors have also reported that in the subgroup analysis by ethnicity, no significant association was found in three ethnic groups (Asian, Caucasian, and African). Moreover, in the meta-analysis of He et al. authors have found an increased cancer risk associated with this polymorphism in the homozygous genetic model for Asian populations, but not for other ethnic groups [7]. Conversely, Hirata et al. have found that the frequency of 939Gln variant at XPC Lys939Gln was significantly lower in PC cases and was associated with a protective effect (OR 0.39, p = 0.016) [16].
The comparison of PC cases with controls according to XPC-PAT polymorphism, suggests that individuals inheriting XPC I/I genotype were at significantly increased risk of PC malignancy. This result confirms others reported studies and was explained by the fact that the homozygous variant genotype of the PAT polymorphism (I/I) exhibited lower DRC as compared to wild-type carriers (D/D) [16, 23]. To the best of our knowledge, it is the first study in which we report a high association between prostate cancer and XPC I/I genotype (OR 3.83). Indeed, all of the previously reported studies interested in the analysis of the association between XPC-PAT polymorphism and prostate cancer suggest a moderate risk association which doesn’t exceed 2.5-fold increased risk [24]. The observed high-risk association between XPC I/I genotype and prostate cancer in Tunisian population in comparison to other populations could be associated to the difference in ethnicity and may be also explained by the additive effect between the inheritance of XPC I/I genotype, tobacco and professional risk factors. Indeed 74.75% (82/110) of PC patients were smokers and 54.54% of them (60/110) were highly smokers. Moreover it has been found that more than 52% of PC patients are exposed to professional risk factors. Among the professional risk factors we cited aromatic amines (benzidine, 4-aminobiphenyl,2-naphthylamine, 4-chloro-o-toluidine), chlorinated hydrocarbon or polycyclic aromatic hydrocarbons, or pesticides use mainly in industrial areas processing paint, metal, dye, petroleum derivates or agriculture fields. These observations highlight the imperative need to enlarge the predispositions studies in the Tunisian populations in this concern in order to raise the awareness about the use and composition of chemical products. Our result is supported by the study of However, Liu et al. Which suggest that XPC-PAT polymorphisms may contribute to the risk of developing PC and have found an elevated risk of PC associated with a gene-environment interaction [8].
In accordance with previous reports, we detected linkage disequilibrium between XPC Lys939Gln and XPC PAT polymorphisms [25]. We have found that the inheritance of the (Lys/Gln + PAT D/D) haplotype had a protective effect. This result was explained by the presence of the wild allele PAT*D (dominant allele) which modulate the effect of the XPC* 939Lys or *939Gln allele. However, we have not found an additive effect between XPC Gln/Gln and I/I genotypes. This result is in contradiction with previous studies which have found that the mutant diplotype (Gln/I) could be considered as an aggravating marker for prostate cancer [25, 26]. Moreover we have not found a significant association between Lys/Lys + I/I or Lys/Gln + I/I and PC risk, although the inheritance of the risk genotype PAT I/I. These results could be explained by the low number of subjects with Lys/Lys + I/I genotype (only 0.5% of patients and controls) and also by the interaction among the the XPC polymorphisms. Indeed it has been reported that the obtained results for the XPC haplotype could be explained by contradictory results frequently found in studies based on individual-SNP analyzes. Indeed if the interactions between individual-SNP contribute to tumor risk, then the haplotype/diplotypes constructed by these polymorphisms may exert different effects on tumorigenesis in comparison to individual SNPs [27, 28]. Moreover the interactions between polymorphisms, environmental factors and host characteristics of PC patients might contribute to the discrepancies.
The combined analysis of XPC gene polymorphisms and Gleason score of PC patients don’t report any significant result which confirms others previously reported studies [11, 29]. Conversely, Mandal et al. reported that XPC-PAT polymorphism is associated with high Gleason score in Indian population and explained this association by the fact that the interactions among genetic polymorphisms in NER pathway genes may affect the DNA damage repair capacity and contribute to increased PC risk [25]. We could explain the absence of the association between XPC polymorphisms and Gleason score of PC in Tunisian population in comparison to the Indian population by the fact that these two populations were not exposed to the same environmental risk factors. The correlation between XPC polymorphisms, lifestyle factors (tobacco status, alcohol consumption, exposition to the professional risk factor) and XPC gene polymorphisms does not report any significant difference. These results suggest that the severity and the progression of prostate cancer in our population don’t depend on environmental and genetic risk factors. The role of these risk factors was only limited to the tumor initiation, however, tumor progression is directly associated with somatic alteration.
Conclusion
In this study, we find that the XPC-PAT I/I genotype may be involved in the susceptibility to PC in the Tunisian population. However, this genotype was not associated with tumors severity.
Data availability
Data sharing not applicable to this article as no datasets were generated or analyzed during the current study.
Abbreviations
- SNP:
-
Single nucleotide polymorphisms
- DRC:
-
DNA repair capacity
- PC:
-
Prostate cancer
- PSA:
-
Prostate-specific antigen
- NER:
-
Nucleotide excision repair
- BER:
-
Base excision repair
- MMR:
-
Mismatch repair
- DSBR:
-
Double-strand break repair
- TCR:
-
Transcription-coupled repair
- XPC:
-
Xeroderma pigmentosum complementary group C
- DRE:
-
Digital rectal examination
- EDTA:
-
Ethylene diamine tetra-acetic acid
- PCR:
-
Polymerase chain reaction
- PCR-RFLP:
-
Restriction Fragment Length polymorphism
- ORs:
-
Odds ratios
- CI:
-
Confidence intervals
References
Bello AP, Masip TC (2014) Prostate cancer epidemiology. Arch Esp de Urol 67(5):373–382
Curado MP, Shin HR, Storm H, Heanue M, Boyle P (2007) Cancer incidence in five continents IARC scientific publications
Forrest MS, Edwards SM, Houlston R, Kote-Jarai Z, Key T, Allen N, Knowles MA, Turner F, Ardern-Jones A, Murkin A, Williams S, Oram R, collaborators C-UBUpcs, Bishop DT, Eeles RA (2005) Association between hormonal genetic polymorphisms and early-onset prostate cancer. Prostate Cancer Prostat Dis 8:95. https://doi.org/10.1038/sj.pcan.4500785
Gong Z, Agalliu I, Lin DW, Stanford JL, Kristal AR (2008) Cigarette smoking and prostate cancer-specific mortality following diagnosis in middle-aged men. Cancer Causes Control: CCC 19(1):25–31. https://doi.org/10.1007/s10552-007-9066-9
Gong Z, Kristal AR, Schenk JM, Tangen CM, Goodman PJ, Thompson IM (2009) Alcohol consumption, finasteride, and prostate cancer risk: results from the prostate cancer prevention trial. Cancer 115(16):3661–3669. https://doi.org/10.1002/cncr.24423
Rozet F, Hennequin C, Beauval JB, Beuzeboc P, Cormier L, Fromont G, Mongiat-Artus P, Ouzzane A, Ploussard G, Azria D, Brenot-Rossi I, Cancel-Tassin G, Cussenot O, Lebret T, Rebillard X, Soulié M, Renard-Penna R, Méjean A (2016) Recommandations en onco-urologie 2016–2018 du CCAFU: cancer de la prostate. Progrès en Urol 27:S95–S143. https://doi.org/10.1016/s1166-7087(16)30705-9
He J, Shi TY, Zhu ML, Wang MY, Li QX, Wei QY (2013) Associations of Lys939Gln and Ala499Val polymorphisms of the XPC gene with cancer susceptibility: a meta-analysis. Int J Cancer 133(8):1765–1775. https://doi.org/10.1002/ijc.28089
Liu Y, Wang H, Lin T, Wei Q, Zhi Y, Yuan F, Song B, Yang J, Chen Z (2012) Interactions between cigarette smoking and XPC-PAT genetic polymorphism enhance bladder cancer risk. Oncol Rep 28(1):337–345. https://doi.org/10.3892/or.2012.1759
Brown KL, Roginskaya M, Zou Y, Altamirano A, Basu AK, Stone MP (2010) Binding of the human nucleotide excision repair proteins XPA and XPC/HR23B to the 5R-thymine glycol lesion and structure of the cis-(5R,6S) thymine glycol epimer in the 5′-GTgG-3′ sequence: destabilization of two base pairs at the lesion site. Nucleic Acids Res 38(2):428–440. https://doi.org/10.1093/nar/gkp844
Jiang X, Zhou L-t, Zhang S-c, Chen K (2012) XPC polymorphism increases risk of digestive system cancers: current evidence from A meta-analysis. Chin J Cancer Res 24(3):181–189. https://doi.org/10.1007/s11670-012-0181-0
Zhu Y, Yang H, Chen Q, Lin J, Grossman HB, Dinney CP, Wu X, Gu J (2008) Modulation of DNA damage/DNA repair capacity by XPC polymorphisms. DNA Repair 7(2):141–148. https://doi.org/10.1016/j.dnarep.2007.08.006
Marin MS, Lopez-Cima MF, Garcia-Castro L, Pascual T, Marron MG, Tardon A (2004) Poly (AT) polymorphism in intron 11 of the XPC DNA repair gene enhances the risk of lung cancer. Cancer Epidemiol, Biomark Prev 13 (11 Pt 1):1788–1793
Qiao Y, Spitz MR, Shen H, Guo Z, Shete S, Hedayati M, Grossman L, Mohrenweiser H, Wei Q (2002) Modulation of repair of ultraviolet damage in the host-cell reactivation assay by polymorphic XPC and XPD/ERCC2 genotypes. Carcinogenesis 23(2):295–299
Sak SC, Barrett JH, Paul AB, Bishop DT, Kiltie AE (2005) The polyAT, intronic IVS11-6 and Lys939Gln XPC polymorphisms are not associated with transitional cell carcinoma of the bladder. Br J Cancer 92(12):2262–2265. https://doi.org/10.1038/sj.bjc.6602616
Zhang L, Zhang Z, Yan W (2005) Single nucleotide polymorphisms for DNA repair genes in breast cancer patients. Clin Chim Acta 359(1–2):150–155. https://doi.org/10.1016/j.cccn.2005.03.047
Hirata H, Hinoda Y, Tanaka Y, Okayama N, Suehiro Y, Kawamoto K, Kikuno N, Majid S, Vejdani K, Dahiya R (2007) Polymorphisms of DNA repair genes are risk factors for prostate cancer. Eur J Cancer (Oxford, England 1990) 43(2):231–237. https://doi.org/10.1016/j.ejca.2006.11.005
Wu H, Lv Z, Wang X, Zhang L, Mo N (2015) Lack of association between XPC Lys939Gln polymorphism and prostate cancer risk: an updated meta-analysis based on 3039 cases and 3253 controls. Int J Clin Exp Med 8(10):17959–17967
Mirecka A, Paszkowska-Szczur K, Scott RJ, Gorski B, van de Wetering T, Wokolorczyk D, Gromowski T, Serrano-Fernandez P, Cybulski C, Kashyap A, Gupta S, Golab A, Slojewski M, Sikorski A, Lubinski J, Debniak T (2014) Common variants of xeroderma pigmentosum genes and prostate cancer risk. Gene 546(2):156–161. https://doi.org/10.1016/j.gene.2014.06.026
Zou Y-F, Tao J-H, Ye Q-L, Pan H-F, Pan F-M, Su H, Ye D-Q (2013) Association of XPC gene polymorphisms with susceptibility to prostate cancer: evidence from 3936 subjects. Genet Test Mol Biomark 17(12):926–931. https://doi.org/10.1089/gtmb.2013.0267
Evans GA (1990) Molecular cloning: a laboratory manual. 2nd edn. Volumes 1, 2, and 3. Current protocols in molecular biology. Cell 61 (1):17–18. https://doi.org/10.1016/0092-8674(90)90210-6
Excoffier L, Laval G, Schneider S (2005) Arlequin (version 3.0): an integrated software package for population genetics data analysis. Evol Bioinf Online 1:47–50
Agalliu I, Kwon EM, Salinas CA, Koopmeiners JS, Ostrander EA, Stanford JL (2010) Genetic variation in DNA repair genes and prostate cancer risk: results from a population-based study. Cancer Causes Control 21(2):289–300. https://doi.org/10.1007/s10552-009-9461-5
Yoshino Y, Takeuchi S, Katoh T, Kuroda Y (2016) XPC intron11 C/A polymorphism as a risk factor for prostate cancer. Environ Health Prev Med 21(2):100–104. https://doi.org/10.1007/s12199-015-0505-z
Dai QS, Hua RX, Zhang R, Huang YS, Hua ZM, Yun CT, Zeng RF, Long JT (2013) Poly (AT) deletion/insertion polymorphism of the XPC gene contributes to urinary system cancer susceptibility: a meta-analysis. Gene 528(2):335–342. https://doi.org/10.1016/j.gene.2013.06.092
Mandal RK, Gangwar R, Kapoor R, Mittal RD (2012) Polymorphisms in base-excision and nucleotide-excision repair genes and prostate cancer risk in north Indian population. Indian J Med Res 135:64–71
Mittal RD, Mandal RK (2012) Genetic variation in nucleotide excision repair pathway genes influence prostate and bladder cancer susceptibility in North Indian population. Indian J Hum Genet 18(1):47–55. https://doi.org/10.4103/0971-6866.96648
Wu X, Gu J, Grossman HB, Amos CI, Etzel C, Huang M, Zhang Q, Millikan RE, Lerner S, Dinney CP et al (2006) Bladder cancer predisposition: a multigenic approach to DNA-repair and cell-cycle-control genes. Am J Hum Genet 78(3):464–479
Zhu Y, Lai M, Yang H, Lin J, Huang M, Grossman HB, Dinney CP, Wu X (2007) Genotypes, haplotypes and diplotypes of XPC and risk of bladder cancer. Carcinogenesis 28(3):698–703
Kahnamouei SA, Narouie B, Sotoudeh M, Mollakouchekian MJ, Simforoosh N, Ziaee SA, Samzadeh M, Afshari M, Jamaldini SH, Imeni M, Hasanzad M (2016) Association of XPC Gene Polymorphisms with Prostate Cancer Risk. Clin Lab 62(6):1009–1015
Acknowledgements
The team work would like to express their thanks and gratitude to the medical team of Urology department, Charles Nicolle Hospital, Tunis—Tunisia.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Ethics approval
This project was approved by a Charles Nicolle ethical committee, Tunis; Tunisia.
Informed consent
Informed consent was obtained from all individual participants included in the study.
Additional information
Publisher's Note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Said, R., Bougatef, K., Setti Boubaker, N. et al. Polymorphisms in XPC gene and risk for prostate cancer. Mol Biol Rep 46, 1117–1125 (2019). https://doi.org/10.1007/s11033-018-4572-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11033-018-4572-2