Abstract
In this study, we aimed to identify the influence of exonuclease 1 (EXO1) single-nucleotide polymorphism rs9350, which is involved in DNA mismatch repair, on prostate cancer risk in Chinese people. In our hospital-based case–control study, 214 prostate cancer patients and 253 cancer-free control subjects were enrolled from three hospitals in China. Genotyping for rs9350 was performed by the SNaPshot® method using peripheral blood samples. Consequently, a significantly higher prostate cancer risk was observed in patients with the CC genotype [odds ratio (OR) = 1.678, 95 % confidence interval (CI) = 1.130–2.494, P = 0.010] than in those with the CT genotype. Further, the CT/TT genotypes were significantly associated with increased prostate cancer risk (adjusted OR = 1.714, 95 % CI = 1.176–2.500, P = 0.005), and the C allele had a statistically significant compared with T allele (P = 0.009) of EXO1 (rs9350). Through stratified analysis, significant associations were revealed for the CT/TT genotype in the subgroup with diagnosis age >72 (adjusted OR = 1.776, 95 % CI = 1.051–3.002, P = 0.032) and in patients with localized disease subgroup (adjusted OR = 1.798, 95 % CI = 1.070–3.022, P = 0.027). In addition, we observed that patients with prostate-specific antigen (PSA) levels of ≤10 ng/mL were more likely to have the CT/TT genotypes than those with PSA levels of >10 ng/mL (P = 0.006). For the first time, we present evidence that the inherited EXO1 polymorphism rs9350 may have a substantial influence on prostate cancer risk in Chinese people. We believe that the rs9350 could be a useful biomarker for assessing predisposition for and early diagnosis of prostate cancer.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Prostate cancer is a common malignancy among men worldwide. With approximately 899,000 new cases and 29,720 deaths reported annually, it is also the most frequently diagnosed cancer and the sixth leading cause of cancer-associated deaths in males worldwide [1]. The exact mechanism of its carcinogenesis has not yet been fully elucidated; nevertheless, genetic, lifestyle-related, and environmental factors are may be involved [2–5]. Genetically, prostate cancer develops as a result of mutational events involving the activation of proto-oncogenes and inactivation of tumor suppressor genes [6]. A key element in the development of cancer reportedly is the accumulation of mutations in DNA [6, 7]. The responses of the cell to genetic injuries and its ability to maintain genomic stability through various DNA repair mechanisms are essential for preventing tumor initiation and progression [7].
DNA mismatch repair (MMR) pathways help maintain genetic stability. Exonuclease 1 (EXO1) and MutS homologs 1, 2, and 3 play important roles in this pathway and can recognize DNA damage induced by carcinogens [7, 8]. The exonuclease 1 (EXO1) gene is a member of both the DNA MMR system and the RAD2 nuclease family [9]. Along with maintaining genomic stability, EXO1 contributes in modulation of DNA recombination and mediation of cell cycle arrest. EXO1 is located at chromosome 1q42–43 and contains one untranslated exon, followed by 13 coding exons encoding an 846 amino acid protein [8, 9].
Single-nucleotide polymorphisms (SNPs) of the DNA MMR system genes are reportedly associated with susceptibility to several cancers [7]. SNP of this system may affect gene function or expression, along with the effectiveness of gene-related systems [6, 8, 9]. These findings suggest that cancer susceptibility may vary in people who carry different genotypes. Several EXO1 SNP have been reported as genetic risk factors of cancer. In 2011, a study on an Iranian population showed that some EXO1 SNPs are associated with colorectal cancer risk [10]. However, the mechanisms of the associations between specific SNP with prostate cancer have not yet been elucidated. In this study, we aimed to investigate the genotype frequencies of an EXO1 SNP, L757P (rs9350), in a Chinese population for investigating its influence on prostate cancer risk.
Materials and methods
Study subjects
Subjects were from three hospitals, one each from Beijing, Hangzhou, and Guangzhou, China, between January 2009 and April 2013. Histologically verified prostate cancer patients (253 cases) were included. For the stratified analysis, we classified localized and advanced prostate cancer according to the following criteria: localized prostate cancer (T1-2 N0M0, Gleason score, 2–7; prostate-specific antigen (PSA) levels ≤50 ng/mL) and advanced prostate cancer [T3-4 or N+ or M+; Gleason score, 8–10; PSA levels >50 ng/mL]. Gleason score was estimated by experienced pathologists. TNM staging was determined by pathological findings after a surgery or based on imaging examinations and biopsy results. The 214 control subjects consisted of cancer-free patients or healthy men, having no history of prostate cancer according to their PSA levels and digital rectal examination. A prostate biopsy was performed to exclude prostate cancer if the total PSA was persistently >4 ng/mL. All controls in this study were randomly selected from hospitals during the recruitment stage and were frequency matched with the patients in terms of age and geographic origin. Informed consent was obtained from each subject after detailed explanation of the study, which was approved by the ethics committees of the involved hospitals.
Blood samples and DNA extraction
A 2-mL peripheral blood sample was obtained from each participant. Blood samples were stored in EDTA vacuum blood tubes at −20 °C (or −80 °C when stored for >1 month prior to analysis). Genomic DNA was extracted using the phenol–chloroform method or using the TIANamp Blood DNA Kit (Tiangen Biotech, Beijing, China). Both methods could extract sufficient DNA for genotyping and fulfill quality control requirements for genotyping.
Genotyping
Genotyping analyses for rs9350 were performed by Life Technologies Corporation (Shanghai, China) using the ABI SNaPshot® Multiplex System.
Multiplex PCR
The SNaPshot® processing genomic DNA methods differed slightly among the three hospitals because the recently built Zhujiang Hospital (Guangzhou) had improved genotyping procedures, which were introduced recently.
Multiplex PCR amplification was executed on a MyCycler™ Thermal Cycler using 1 μL of genomic DNA (10–50 ng) as the template, with 2.5 μL of 10× PCR buffer (Mg2+ free), 0.8 μL of MgCI2 (50 mM), 0.5 μL of deoxynucleotide triphosphates (dNTPs, 2.5 mM each), 0.2 μL of Platinum® Taq (5 U/μL), 1 μL of first primer mix, and ddH2O to a final reaction volume of 25 μL. The program included an initial denaturation at 95 °C for 5 min, followed by 33 cycles at 95 °C for 30 s, 56 °C for 30 s, and 72 °C for 30 s. The final extension was executed at 72 °C for 5 min. The first PCR primers for rs9350 were as follows: upper primer, 5′-CAGAATGGTCTTAAAATGGGTGT-3′, and lower primer, 5′-TTCAGAATAAGAAACAAGGCAAC-3′. The PCR products were run on 2 % agarose gel. To avoid their interference in subsequent primer extension reactions, the primers and unincorporated dNTPs of the preliminary PCR reaction were removed before genotyping; 2 μL of PCR product was incubated with 0.3 μL shrimp alkaline phosphatase (SAP, 1 U/μL), 0.2 μL of EXO1 (20 U/μL), and 7.5 μL of ddH2O for 90 min at 37 °C, followed by 15 min at 75 °C for enzyme inactivation.
SNaPshot® reaction
The typing PCR (second PCR) primer for rs9350 was 5′-TTTTTTTTTTTTTTTTTAGACTCTCTTTCTACAACCAAGATCAAAC-3′. SNaPshot® reactions were executed using 1 μL of the purified products from the first PCR as a template, 1.5 μL of reaction mixture, and 0.5 μL of Probe Mix. The reactions were executed in the MyCycler™ Thermal Cycler at 96 °C for 10 s, 51 °C for 5 s, and 60 °C for 30 s, which were repeated for 25 cycles. The samples were then treated with 0.3 μL of SAP (1 U/μL) at 37 °C for 1 h and heat inactivated at 75 °C for 15 min. In an ABI optical plate, 8.8 μL of Hi-Di™ formamide, 0.2 μL of Genescan™ 120 LIZ™ Size Standard, and 1 μL of the reaction mixture were combined and then denatured at 95 °C for 5 min. Samples were loaded on the ABI PRISM 3730 DNA Analyzer and analyzed using GeneMapper (version 4.1) software.
Statistical analysis
Deviations in the genotype distribution from Hardy–Weinberg equilibrium in the controls were calculated using the χ 2 test. The differences of allele and genotype data were analyzed with the homozygotes of the common allele as the reference group. Odds ratio (OR), 95 % confidence interval (CI), and P values for the association between prostate cancer risk and genotypes (or alleles) were computed using logistic regression analysis between patients and controls with or without adjustment for potential confounders. In the stratified analyses, we further calculated stratification factors using age at diagnosis (≤72 and >72 years) and aggressiveness (localized and advanced). For all analyses, genetic effects were adjusted for age (at diagnosis) and smoking and drinking statuses. Continuous variables were analyzed using t tests, and categorical variables were analyzed using the χ 2 test to assess any differences in the frequency distributions of clinical variables.
All statistical analyses were performed using SPSS 20.0 (IBM SPSS Statistics; IBM Corp., Armonk, NY, USA). P values of <0.05 were considered statistically significant (two-tailed).
Results
Clinical characteristics
The clinical characteristics of 467 Chinese men (253 prostate cancer patients and 214 control subjects) are shown in Table 1. The mean ages of the patients (at diagnosis) and control subjects (at inclusion into study) were 71.4 and 70.2 years, respectively. We analyzed data from all subjects in this study. No significant difference was found in the genotype frequency of rs9350 in populations living in different areas in China (Table 2). The Hardy–Weinberg principle was true for the genotype frequency of rs9350 in the control group (P ≥ 0.05).
SNP analysis of rs9350 in prostate cancer
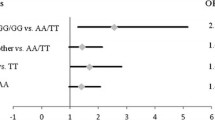
The allele and genotype frequencies of rs9350 among cases and control subjects are listed in Table 3. The same proportion of rs9350 CC allele (32.8 vs. 45.3 %) was observed in prostate cancer patients and control subjects. The proportions of rs9350 CT and TT genotypes in prostate cancer patients versus control subjects were 53.0 versus 44.4 % and 14.2 versus 10.3 %, respectively. Compared with the CC genotype, the CT genotype may increase prostate cancer risk (OR = 1.648, 95 % CI = 1.112–2.444, P = 0.013 and adjusted OR = 1.678, 95 % CI = 1.130–2.494, P = 0.010). Further, the CT/TT genotypes were significantly associated with increased prostate cancer risk (OR = 1.698, 95 % CI = 1.166–2.473, P = 0.006 and adjusted OR = 1.714, 95 % CI = 1.176–2.500, P = 0.005), and the C allele had a significant statistically compared with T allele (P = 0.009) of EXO1 (rs9350).
SNP analysis of rs9350 for different ages at diagnosis
In stratified analysis, we selected an age at diagnosis of 72 years as the cutoff, with similar numbers of cases above and below the cutoff age (Table 4). In the >72 years group, the CT/TT genotypes of rs9350 were statistically significant (P = 0.032) as well as the C allele compared with the T allele (adjusted OR = 1.464, 95 % CI = 0.998–12.147, P = 0.041) for prostate cancer risk.
SNP analysis of rs9350 in different prostate cancer classes
In another stratified analysis, we classified the prostate cancer patients into localized and advanced disease in terms of the aggressiveness (Table 5). In the localized disease subgroup (controls: all cancer-free patients or healthy men), the CT/TT genotypes of rs9350 were associated with significantly increased prostate cancer risk (adjusted OR = 1.798, 95 % CI = 1.070–3.022, P = 0.027).
SNP analysis of rs9350 for different patient factors
Except for PSA levels at diagnosis, no significant association was observed between rs9350 and clinical factors, including aggressiveness, Gleason scores, age at diagnosis, smoking status, or drinking status (Table 6). Subjects with PSA levels of ≤10 ng/mL were more likely to have the CT/TT genotype than those with PSA levels of >10 ng/mL (P = 0.006).
Discussion
In order to assess whether mutations accumulate in the DNA in prostate cancer, we investigated the association between rs9350 and the susceptibility of a Chinese population to prostate cancer. Our data showed that certain genotypes of EXO1 were associated with a significantly higher susceptibility for prostate cancer. Our blood samples were obtained from prostate cancer patients and healthy subjects from different geographical locations—Beijing, Hangzhou, and Guangzhou—located in northern, central, and southern China, respectively; this variation in location could have helped obtain convincing results, although the sample size was small. Other studies have demonstrated a significant association between rs9350 and development of pancreatic cancer in American [11] and Taiwanese [12] populations.
To our knowledge, our study provides the first evidence for the association between the EXO1 SNP rs9350 and prostate cancer risk in Chinese people.
EXO1, a structure-specific 5′ nuclease superfamily protein, recognizes single-stranded DNA (ssDNA)–double-stranded DNA (dsDNA) junctions and cleaves one nucleotide into the dsDNA. EXO1 is a processive 5′–3′ exonuclease that plays a role in MMR, double-strand break repair, and telomere maintenance. The challenge for EXO1 is that they should recognize structure-specific damage to distinguish their substrate from among ssDNA, RNA, and dsDNA [13]. For EXO1, substrate- and product-bound structures have been determined in previous studies; EXO1 is associated with multiple types of human cancers, including gastric [14], oral [15], and lung cancers [16] and head and neck squamous cell carcinoma [17].
Exposure to environmental factors is considered the most important etiological factor in the development of carcinomas. For example, carcinogens in tobacco, such as benzo(a)pyrene, can bind to DNA in the epithelial cells of the upper aerodigestive tract, forming covalent DNA adducts and inducing replication errors, associated with cancer origin [17]. Recently, a Brazilian study showed that the EXO1 abnormalities in the DNA MMR system are important determinants of head and neck squamous cell carcinoma, particularly among smokers, and they can be used to predict patient outcomes [17].
In addition to the association between rs9350 and prostate cancer, we included environmental exposure factors and the family history of the subjects in our analyses. The results showed that age at diagnosis (72 years as the cutoff age), cancer type (localized or advanced disease), and PSA levels at diagnosis (10 ng/mL as the cutoff value) were all significantly associated with an increased prostate cancer risk. In contrast, no significant association was found between prostate cancer risk and other factors such as Gleason scores, smoking status, or drinking status.
Our study had some limitations. First, the control and patient sample size might limit the statistical power of this study, especially for subgroups with different ages of diagnosis and different factors. Next, our prostate cancer patients were from hospitals and the controls were randomly selected from among cancer-free patients in these same hospitals; therefore, whether they formed a representative control population needs to be assessed. The results of our study represent three different areas from China, which is not an accurate expression of entire China.
In conclusion, we provided evidence of independent replication for the EXO1 SNP rs9350 associated with prostate cancer risk in China. Although additional studies with larger and more diverse populations and functional analysis of the SNP are necessary to confirm and extend our findings, we believe that the EXO1 SNP rs9350 could be a useful biomarker for assessing predisposition to prostate cancer and early diagnosis of the disease.
References
Siegel RL, Miller KD, Jemal A. Cancer statistics, 2015. CA Cancer J Clin. 2015;65(1):5–29.
Ma C, Liu C, Huang P, Kaku H, Chen J, Guo K, et al. Significant association between the Axin2 rs2240308 single nucleotide polymorphism and the incidence of prostate cancer. Oncol Lett. 2014;8(2):789–94.
Patel CJ, Rehkopf DH, Leppert JT, Bortz WM, Cullen MR, Chertow GM, et al. Systematic evaluation of environmental and behavioral factors associated with all-cause mortality in the United States national health and nutrition examination survey. Int J Epidemiol. 2013;42(6):1795–810.
Siegel RL, Sahar L, Portier KM, Ward EM, Jemal A. Cancer death rates in US congressional districts. CA Cancer J Clin. 2015. doi:10.3322/caac.21292.
Lombardi S, Fuoco I, di Fluri G, Costa F, Ricchiuti A, Biondi G, et al. Genomic instability and cellular stress in organ biopsies and peripheral blood lymphocytes from patients with colorectal cancer and predisposing pathologies. Oncotarget. 2015;6(17):14852–64.
Cussenot O, Cancel-Tassin G. Update on genetic predisposition to prostate cancer. Bull Cancer. 2015;102(1):53–6.
Sperka T, Wang J, Rudolph KL. DNA damage checkpoints in stem cells, aging and cancer. Nat Rev Mol Cell Biol. 2012;13(9):579–90.
Song F, Qureshi AA, Zhang J, Amos CI, Lee JE, Wei Q, et al. Exonuclease 1 (EXO1) gene variation and melanoma risk. DNA Repair (Amst). 2012;11(3):304–9.
Bregenhorn S, Jiricny J. Biochemical characterization of a cancer-associated E109K missense variant of human exonuclease 1. Nucleic Acids Res. 2014;42(11):7096–103.
Haghighi MM, Taleghani MY, Mohebbi SR, Vahedi M, Fatemi SR, Zali N, et al. Impact of EXO1 polymorphism in susceptibility to colorectal cancer. Genet Test Mol Biomarkers. 2010;14(5):649–52.
Tang H, Wei P, Duell EJ, Risch HA, Olson SH, Bueno-de-Mesquita HB, et al. Axonal guidance signaling pathway interacting with smoking in modifying the risk of pancreatic cancer: a gene- and pathway-based interaction analysis of GWAS data. Carcinogenesis. 2014;35(5):1039–45.
Dong X, Li Y, Hess KR, Abbruzzese JL, Li D. DNA mismatch repair gene polymorphisms affect survival in pancreatic cancer. Oncologist. 2011;16(1):61–70.
Tsutakawa SE, Tainer JA. Double strand binding-single strand incision mechanism for human flap endonuclease: implications for the superfamily. Mech Ageing Dev. 2012;133(4):195–202.
Kim YR, Yoo NJ, Lee SH. Somatic mutation of EXO1 gene in gastric and colorectal cancers with microsatellite instability. Acta Oncol. 2010;49(6):859–60.
Tsai MH, Tseng HC, Liu CS, Chang CL, Tsai CW, Tsou YA, et al. Interaction of EXO1 genotypes and smoking habit in oral cancer in Taiwan. Oral Oncol. 2009;45(9):e90–44.
Bayram S. The exonuclease 1 Glu589Lys gene polymorphism and cancer susceptibility: evidence based on a meta-analysis. Asian Pac J Cancer Prev. 2014;15(6):2571–6.
Nogueria GA, Lourenço GJ, Oliveira CB, Marson FA, Lopes-Aguiar L, Costa EF, et al. Association between genetic polymorphisms in DNA mismatch repair-related genes with risk and prognosis of head and neck squamous cell carcinoma. Int J Cancer. 2015;137(4):810–8.
Acknowledgments
This study was supported by scientific research grants from the Pearl River Nova Program of Guangzhou (No. 2013J2200044); the National Natural Scientific Foundation of China (No. 81101559); and the Ministry of Education, Culture, Sports, Science and Technology of Japan (Nos. KAKENHI 25861425 and 15K20093).
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Ethical approval
All procedures performed in studies involving human participants were in accordance with the ethical standards of the institutional and/or national research committee and with the 1964 Helsinki declaration and its later amendments or comparable ethical standards.
Conflicts of interest
None
Additional information
Yiming Zhang, Pengju Li and Abai Xu contributed equally to this work.
Rights and permissions
About this article
Cite this article
Zhang, Y., Li, P., Xu, A. et al. Influence of a single-nucleotide polymorphism of the DNA mismatch repair-related gene exonuclease-1 (rs9350) with prostate cancer risk among Chinese people. Tumor Biol. 37, 6653–6659 (2016). https://doi.org/10.1007/s13277-015-4298-x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s13277-015-4298-x