Abstract
Wheat-Dasypyrum villosum translocations were induced in the progeny of the amphiploid Triticum durum-D. villosum (AABBVV) by pollen irradiation. The rearranged V genome chromosomes were characterized by genomic/fluorescence in situ hybridization (GISH/FISH) and molecular markers. Twenty wheat-D. villosum translocation chromosomes were selected, including four centric, seven large segments, and nine small segments in a Chinese Spring (CS) background. The four centric translocations were subsequently identified by GISH/FISH and by molecular markers specific to chromosome arms of the Triticeae linkage groups. They were T5DL.4VL, T4BL.7VS, and T4BS.7VL as well as the compensating translocation T7AL.7VS. Using a combination of previously developed V chromosome alterations, 52 translocations or deletions that divided V chromosomes into 42 bins were employed for deletion mapping of molecular markers specific to D. villosum in a wheat background. Ninety-five expressed sequence tag (EST)-sequence-tagged site (STS) and seven SSR markers that were previously reported, as well as 72 STS markers screened in the present study, were physically allocated into 37 of 42 chromosome bins of D. villosum. Multiple loci of EST-STS markers were also mapped using CS nullisomic tetrasomic (NT) and ditelosomic (DT) genetic stocks. Most EST-STS homoeoloci were located on homoeologous chromosomes, suggesting a high degree of homology between the genomes of D. villosum and wheat. Four 4VL-specific markers detected homoeoloci on group 7 chromosomes of wheat, indicating that chromosome 4V of D. villosum shows some affinity to both wheat homoeologous groups 4 and 7. This is the first physical map of D. villosum, which will provide insight into the V genome for molecular breeding.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
PCR-based molecular markers have been developed and mapped onto chromosomes in the Triticeae tribe (Harper and Cande 2000). Chromosomal alterations are informative not only for the mapping of genes from specific chromosomal regions but also for structural and functional analyses of the genetic relationships between homoeologous chromosomes of wheat and its relatives (Kojima et al. 2000). Thus far, chromosomal aberrations of wheat and its relatives, barley and rye, have been produced (Endo and Gill 1996; Joshi et al. 2011; Friebe et al. 2000; Tsuchida et al. 2008), and these unique resources have provided excellent tools for estimating the actual physical chromosomal locations of molecular markers and genes, thus facilitating the utility of genetic linkage maps for map-based gene cloning (Harper and Cande 2000; Qi et al. 2003).
The V genome of Dasypyrum villosum (L.) Candargy (Dv), also known as Haynaldia villosa (L.) Schur, an allogamous and annual wild diploid relative of common wheat (Gradzielewska 2006), contains genes conferring various disease resistances (Bizzarri et al. 2009; Xu et al. 2009; Qi et al. 2011; Chen et al. 1995; Yildirim et al. 2000; Li et al. 2002; Zhang et al. 2005a, b; Zhang et al. 2016a, b), increasing seed protein content (Montebove et al. 1987; De Pace et al. 2001; Zhang et al. 2014), increasing grains per spike (Zhang et al. 2015), and enhancing salt tolerance and zinc efficiency (Zhong and Dvorak 1995; Schlegel et al. 1998). D. villosum is therefore recognized as a potentially useful resource for enhancing wheat’s genetic diversity. Several D. villosum accessions have been successfully crossed with tetraploid or hexaploid wheat (reviewed by De Pace et al. 2011) with the goal of transferring the alien genes from the V genome into wheat, but researchers rarely progressed further than the induction of wheat-D. villosum translocation lines. Three sets of wheat-D. villosum disomic addition (DA) lines have been developed by Sears (1953), Chen and Liu (1986), and Lukaszewski (1988) using different D. villosum accessions. Based on the addition lines that Lukaszewski developed, Liu et al. (2011) further developed a set of compensating Triticum aestivum-D. villosum Robertsonian translocation lines in a Chinese Spring genetic background. In addition, the wheat-D. villosum lines developed by Chen and Liu (1986) were used to further produce the V chromosome alterations. Specific potential advantages associated with this V genome have been identified and include powdery mildew resistance genes Pm21 and Pm55, a photoperiod response gene Ppd-V1, a cereal nematode resistance gene CreV, and the wheat yellow mosaic virus resistance gene Wss1, all of which have been physically mapped into special bins associated with DNA markers (Chen et al. 2013; Zhao et al. 2013; Zhang et al. 2015, 2016a, b).
The integration of DNA markers into chromosome engineering strategies facilitates the recovery of small alien segments and the physical tagging of genes. Thus far, 103 expressed sequence tag (EST)-sequence-tagged site (STS) and eight SSR markers specific to V chromosomes have been screened, and some of them have been physically mapped onto the chromosome arms 1VS, 2VS, 4VS, 5VS, 6VS, and 6VL (Liu et al. 2004; Zhang et al. 2006; Cao et al. 2006; Wang et al. 2007; Cao et al. 2009a; Qi et al. 2011; Zhao et al. 2013; Chen et al. 2013; Zhang et al. 2014, 2015, 2016a, b). Integration of more physical DNA markers along with complete coverage of the V genome is expected to result in the construction of more precise maps of the D. villosum chromosomes. Therefore, we attempted to develop more V chromosome-specific molecular markers and structural aberrations that mostly focused on 1VL, 2VL, 3VS, 3VL, 4VL, 5VL, 7VS, and 7VL of D. villosum in the present study and to physically map the DNA markers into V chromosome bins using a series of wheat-D. villosum genetic stocks. These wheat-D. villosum translocations, together with V chromosome-specific molecular markers and a V genome physical map, would be useful for comparative genomics and alien gene tagging.
Materials and methods
Plant material and genetic stocks
T. aestivum cv. Chinese Spring (CS), T. durum cv. ZY1286 (AABB), D. villosum GP005 (VV, originally introduced from the Cambridge Botanical Garden, UK), T. durum cv. ZY1286-D. villosum (GP005) amphiploid (Tritpyrum, AABBVV), and other T. aestivum-D. villosum genetic stocks listed in Table 1 were employed to screen the molecular markers specific to V genome chromosomes. All materials were maintained at the Cytogenetic Institution, Nanjing Agricultural University, China (CINAU). Sets of compensating CS nullisomic tetrasomic (NT) lines (Sears 1966) and ditelosomic (DT) lines (Sears and Sears 1978) were used for assigning the homoeoloci of DNA markers to individual chromosomes on the A, B, and D genomes. The NT and DT lines were kindly provided by Prof B. S. Gill of Kansas State University, Kansas, USA.
Wheat-D. villosum translocation development
The flowering spikes of the T. durum-D. villosum amphiploid (AABBVV) were irradiated by 60CO γ-rays at a dose of 1200 Rad to induce the V chromosomal aberrations. Mature, fresh pollen was harvested within 2 days after irradiation from irradiated spikes and was applied to stigmas of emasculated florets of CS plants. Subsequent progenies were backcrossed using CS as recurrent parent. The plants with moderately or fully fertile spikes in the BC2F1 and BC2F2 generations were screened using genomic in situ hybridization (GISH) and V chromosome-specific molecular markers.
Cytogenetic analysis
The protocol used for chromosome in situ hybridization was according to Chen et al. (1995). Genomic in situ hybridization (GISH) was performed using total genomic DNA of D. villosum and labeling with fluorescein-12-dUTP as a probe to detect the chromosomal aberrations. Fluorescence in situ hybridization (FISH) was performed using the oligonucleotide probes pSc119.2, pAs1 and (GAA)10 labeled with digoxigenin-11-dUTP (Roche Diagnostics GmbH, Germany) following the procedures described by Mukai et al. (1993) and Zhang et al. (2004). After hybridization, signals were examined with an Olympus BX60 epifluorescence microscope (Olympus Co., Tokyo, Japan). GISH/FISH images were captured with a SPOT Cooled Color Digital Camera (Diagnostic Instruments, Sterling Heights, MI, USA).
D. villosum-specific EST-STS marker development and analysis
Previously reported molecular markers specific to D. villosum, including 93 STS markers and seven SSR markers, were used to identify the novel translocations and to construct the physical maps (Table S1). To develop more DNA markers specific to chromosomes 3V, 4V, 5V, and 7V of D. villosum, 543 STS primers based on the wheat expressed sequence tags (ESTs) were mapped onto the group 3, 4, 5, and 7 chromosomes of CS, respectively, without paralogous sequences in different homologous groups (http://wheat.pw.usda.gov/cgi-bin/westsql/map_locus.cgi); the primers were designed using the software Primer 3 (http://frodo.wi.mit.edu). In addition, 582 intron-flanking primers based on the wheat Unigene sequences (http://www.ncbi.nlm.nih.gov/unigene) and compared with the Brachypodium genome sequences were designed using Conserved Primers 2.0 software (Frank et al. 2009). The wheat high-molecular-weight glutenin subunit (HMW-GS) gene-specific primer P1/P5 (Pang et al. 2009) and photoperiod response (Ppd) gene-specific primer XHvF11 (Turner et al. 2005) were also mapped onto V chromosomes. All the primers listed in Table S1 were synthesized by Invitrogen Company (Shanghai, China).
Genomic DNA was isolated from young leaves according to instructions accompanying the DNAsecure Plant Kit (Tiangen Biotech Co., Ltd., Beijing, China). The polymerase chain reaction (PCR) amplifications were conducted in a 25-μL reaction mixture containing 1× Taq DNA polymerase buffer, 0.8 mmol/L MgCl2, 0.8 mmol/L dNTPs, 0.5 μL (10 μM) of each primer, 2 U of DNA polymerase, and 30–50 ng of genomic DNA as a template. The samples were denaturated at 94 °C for 5 min and subjected to 33 cycles of 30 s of denaturation at 94 °C, 53–60 °C (depending on the specific primers) for 40 s and 2 min of elongation at 72 °C, with a final extension at 72 °C for 8 min. The PCR products were analyzed on 10% non-denaturing polyacrylamide gels with a 39:1 ratio of acrylamide/bisacrylamide.
Physical mapping of the specific molecular markers into V chromosome bins
For physical mapping of the V chromosome-specific molecular markers, 30 wheat-D. villosum translocations with different breakpoints and two deletions previously reported (Table 1) were used. Markers were first allocated to their respective arms using wheat-D. villosum whole-arm translocations or ditelosomic additions, and the markers were then assigned to arm bins according to whether or not the expected PCR-amplified band was present when the genomic DNA from a particular cytogenetic stock was used as a template. Tests with each marker were repeated twice.
Results
Identification of STS markers specific to V genome chromosomes
Of 1125 STS-containing wheat ESTs, 197 (17.5%) produced PCR amplicons using DNA from D. villosum, indicating a high rate of transferability between wheat and D. villosum. These markers were subsequently used to detect the polymorphisms among the D. villosum (GP005) and T. durum cv. ZY1286 plants, the T. durum cv. ZY1286-D. villosum (GP005) amphiploid, the CS and CS-D. villosum 1V to 7V addition lines, and the wheat-D. villosum ditelosomic addition and whole-arm translocation lines. Seventy-two primers produced stable and clear polymorphic bands that could be allocated to individual V chromosomes or to chromosome arms, and the remaining 125 exhibited no polymorphism between wheat and D. villosum (Table S1). Of the 72 markers, 14 were distributed on 1VL and one was on 2VL. Six markers were assigned on 3VS, and seven were on 3VL. One marker was on 4VS, and seven were assigned to 4VL. Seventeen 5V-specific markers were all located on chromosome arm of 5VL, and three 6V-specific markers were all on 6VS. Nine of the 16 7V-specific markers were on 7VS and the remaining seven were on 7VL.
Development of V chromosome structural aberrations
Twenty V chromosomal aberrations mostly found on 1VL, 2VL, 3VS, 3VL, 4VL, 5VL, 7VS, and 7VL were detected in the progenies following the irradiated pollen treatment. Four aberrations, 4V-4, 7V-1, 7V-2, and 7V-3, were centric; aberrations 2V-7, 3V-2, 4V-5, 4V-6, 6V-10, 7V-5, and 7V-7 were large-segment and the remaining aberrations, 1V-9, 1V-10, 1V-11, 2V-5, 2V-6, 4V-7, 5V-8, 7V-4, and 7V-6, were small-segment translocations (Fig. 1a). Based on the chromosome-specific molecular markers amplified in the structural aberrations and the V chromosome pSc119.2/pAs1 FISH patterns described by Zhang et al. (2013), the translocation breakpoint of 1V-9 on 1VS contained a terminal segment covering approximately 50% of the physical length of arm, and translocation segments of 1V-10 and 1V-11 constituted approximately 50 and 25% of the 1VL terminal segment, respectively. The breakpoint of 2V-5 retained approximately 30% of the terminal region of 2VS, and the 2V-6 translocation segment covered approximately 25% of the terminal region of 2VL. Line 2V-7 consisted of an entire 2VS arm and approximately 50% of the 2VL arm. The line 3V-2 was a T2BS-3VL.3VS(del) translocation confirmed by FISH of (GAA)10 (Fig. 1b) and pSc119.2 (Fig. 1c), and the breakpoints were located approximately 30% of 3VS and 70% of 3VL away from the centromere, respectively. The whole-arm translocation 4V-4 was a T5DL.4VL translocation based on the results of the pAs1 FISH patterns (Fig. 1d) and the 5D-marker analysis (Fig. S2). The breakpoints of 4V-5 and 4V-6 were located approximately 45% of 4VL and 78% of 4VS away from the centromere, respectively, and the 4V-7 translocated segment covered approximately 35% of the terminal region of 4VL. The translocation breakpoint of 5V-8 was located on 5VL covering approximately 30% of the terminal physical length of arm, and 6V-10 was located approximately 70% of 6VS away from the centromere. Of the seven 7V structural aberrations, the centric translocation 7V-1 was T7AL.7VS, confirmed by FISH results using pSc119.2 (Fig. 1e) and the molecular marker CINAU44 (Fig. S2). Another two centric fusion lines, 7V-2 and 7V-3, were T4BL.7VS and T4BS.7VL, respectively, according to the pSc119.2 patterns (Fig. 2f, g). The breakpoint of 7V-4 was located in approximately 20% of the 7VS terminal region, 7V-5 consisted of the entire 7VS arm and approximately 55% of the proximal portion of 7VL, 7V-6 carried approximately 35% of the terminal region of 7VL, and 7V-7 consisted of the entire 7VL arm and approximately 35% of the proximal portion of 7VS.
The GISH/FISH patterns of the wheat-D. villosum translocations. GISH used Dasypyrum villosum genomic DNA labeled with digoxigenin-11-dUTP as a probe and D. villosum chromatin fluoresced with a yellowish-green color. Dual-color FISH used pSc119.2, pAs1 or (GAA)10 labeled with digoxigenin-11-dUTP (red) and total genomic DNA of D. villosum labeled with fluorescein-12-dUTP (green) as probes, and chromosomes were counterstained with DAPI (blue). a GISH patterns of the 20 wheat-D. villosum translocated chromosomes developed in the present study. b Dual-color GISH/GAA-FISH patterns of 3V-2. Signal patterns present on the short arm of the translocated chromosome confirmed the assignment to 3VS. c Dual-color GISH/pSc119.2-FISH patterns of 3V-2. B-genome chromosomes colored by the pSc119.2 probe (red) are marked, and the patterns show the translocated chromosome in 3V-2 is T2BS-3VL.3VS(del). d Dual-color GISH/pAs1-FISH patterns of 4V-4. GISH used D. villosum DNA as a probe (green), and D-genome chromosomes colored by the pAs1 probe (red) are marked, indicating that the translocated chromosome in 4V-4 is T5DL.4VL. e, f, g Dual-color pSc119.2-FISH patterns of 7V-1, 7V-2, and 7V-3. The patterns suggest that the translocations in 7V-1, 7V-2, and 7V-3 are TW.7VS, T4BL.7VS, and T4BS.7VL, respectively (color figure online)
The rough physical map of Dasypyrum villosum. 174 V genome-specific molecular markers are integrated into 37 bins covering 14 V chromosome arms. The alien genes (red) are also mapped into the specific bins that refer to the previous reports of Zhang et al. (2014, 2015, 2016a, 2016b), Zhao et al. (2013), and Chen et al. (2013)
Distribution of molecular marker loci in V chromosome bins
The DNA markers, including the 72 EST-STS primer pairs developed in this study and the 102 previously reported markers, were allocated to V chromosome bins (Fig. 2) using 50 translocations and two deletions (Table 1). Physical maps of 1VS, 2VS, 5VS, and 6VL are the same as those of earlier reports (Zhang et al. 2014, 2015, 2016a, b). However, when comparing the map with the 4VS physical map reported by Zhao et al. (2013), a novel STS marker, CINAU202, was physically positioned onto 4VS FL0.78-1.00, which is associated with the wheat yellow mosaic virus resistance gene, Wss1. In addition, three new STS markers, 6EST802, 6EST928, and 6EST970, were mapped onto 6VS FL0.45-0.58, which is associated with Pm21 when compared with the 6VS physical map constructed by Chen et al. (2013). The remaining physical maps of 1VL, 2VL, 3VS, 3VL, 4VL, 5VL, 7VS, and 7VL were constructed in the present study.
Of the 174 V chromosome-specific markers, 114 have been allocated to special bins of wheat chromosomes (Qi et al. 2004), and the remaining 60 STS markers were mapped onto the CS chromosomes using NT and DT genetic stocks in the present study. Among the 60 STS markers, 56 markers provided homoeoloci that can be assigned onto the chromosome arms of CS. Two markers, CINAU17 and CINAU18 (mapping to 6 VS), only had single locus in D. villosum and none in CS. Another two markers, 6EST245 and 6EST267 (also mapping to 6VS), produced a single band in CS, but their homoeoloci lacked polymorphism. Most EST-STS homoeoloci were allocated to homoeologous chromosomes of CS and D. villosum, with the exception of the 4VL-specific markers CINAU206, CINAU207, CINAU208, and CINAU209, which mapped onto group 7 chromosomes of CS.
Discussion
Several chromosome engineering approaches have been used to produce alien introgressions from D. villosum into common wheat. These include pairing and recombination of alien chromosomes with their wheat homoeologues (Qi et al. 2011; Liu et al. 2011; Li et al. 2011; Zhao et al. 2013), random chromosome breakage by irradiation or gametocidal chromosomes (Chen et al. 2002; Chen et al. 2008; Bie et al. 2007; Cao et al. 2009b), and even chromosome aberrations occurring in tissue culture (Li et al. 2000; Li et al. 2005). Introgressions via chromosome fragmentation irradiation are effective; however, the chromosome breakpoints are random; therefore, these types of introgressions usually are unsuitable for agriculture but still are useful genetic stocks to physically map if they are stable in the wheat background. The 20 wheat-D. villosum translocations developed in the present study may be stable in the wheat background because these plants have homozygous translocated chromosomes with moderate or full spike fertility after two generations of backcrossing using CS as a recurrent parent. Of the four novel whole-arm translocations, T5DL.4VL, T4BL.7VS, and T4BS.7VL were not compensating, while translocation T7AL.7VS was compensating, which may be useful for wheat breeding. Yildirim et al. (1998) mapped the eyespot (Pseudocercosporella herpotrichoides) resistance gene onto chromosome 4VL, and Liu et al. (1989) located the water-soluble endosperm protein gene Wsp-1 onto chromosome 7V. Therefore, these new whole-arm translocation lines will be promising donors for the production of useful small-fragment translocations to aid in using alien genes for wheat breeding.
An abundance of EST sequence polymorphisms exist between wheat and the homoeologous chromosomes belonging to closely related species (Zhang et al. 2005b). Deletion-based physical mapping using chromosome structural changes enabling the physical allocation of EST-PCR markers has been used in wheat (Qi et al. 2004), rye (Lukaszewski et al. 2004) and barley (Harper and Cande 2000). Using 52V chromosome alterations, we integrated 174 markers, including 167 EST-STS and seven SSR markers, into 37 bins of the V chromosomes and provided the first rough physical map of D. villosum in the present study (Fig. 2). The homoeoloci of most markers were distributed on the chromosome arms of the V genome and the A, B, and D genomes, revealing a high degree of homoeology between the genomes of D. villosum and wheat (Table S1). However, homoeoloci of wheat group 7 markers CINAU206, CINAU207, CINAU208, and CINAU209 were specific to 4VL of D. villosum, suggesting that D. villosum chromosome 4V showed some affinity to both wheat homoeologous groups 4 and 7. The physical mapping of EST-STS markers in V chromosome bins will help in further elucidating the relationships between the wheat and D. villosum genomes, making them a valuable source for comparative genomic research.
Physical mapping of DNA markers in V chromosome bins also facilitates the linkage between those EST sequences and alien genes. On the terminal end of chromosome arm 1VS, there are genes at complex loci coding for high-molecular-weight glutenins (Glu-V1) and prolamins (Gli-V1) and low-molecular-weight (LMW) polymeric prolamin proteins (Glu-V3) detected by SDS-PAGE (Zhang et al. 2014). However, the homoeoloci of the HMW-GS gene-specific marker P1/P5 were present on 1VL, 1BL, and 1DL (Fig. S1), indicating that an HMW-GS pseudogene orthologous to the Glu-A1, Glu-B1, and Glu-D1 loci of hexaploid wheat may be present on 1VL. Three wheat group 6 EST-STS markers, 6EST802, 6EST928, and 6EST970, were added to the 6VS FL0.45-0.58, where a powdery mildew resistance gene Pm21 has been physically mapped (Chen et al. 2013). These markers provide the reference sequences for the assignment of alien candidate genes through the microcollinearity between these regions of D. villosum and the homoeologous chromosome sequences of wheat.
References
Bie TD, Cao YP, Chen PD (2007) Mass production of intergeneric chromosomal translocations through pollen irradiation of Triticum durum–Haynaldia villosa amphiploid. J Integr Plant Biol 49:1619–1626
Bie TD, Zhao RH, Jiang ZN, Gao DR, Zhang BQ, He HG (2015) Efficient marker-assisted screening of structural changes involving Haynaldia villosa chromosome 6V using a double-distal-marker strategy. Mol Breed 35:34–43
Bizzarri M, Pasquini M, Matere A, Sereni L, Vida G, Sepsi A, Molnar-Lang M, De Pace C (2009) Dasypyrum villosum 6V chromosome as source of adult plant resistance to Puccinia triticina in wheat. In: Proceedings of the 53rd Italian society of agricultural genetics annual congress, Torino, Italy, 16–19 Sep 2009, Abstract
Cao AZ, Wang XE, Chen YP, Zou XW, Chen PD (2006) A sequence-specific PCR marker linked with Pm21 distinguishes chromosomes 6AS, 6BS, 6DS of Triticum aestivum and 6VS of Haynaldia villosa. Plant Breed 125:201–205
Cao YP, Cao AZ, Wang XE, Chen PD (2009a) Screening and application of EST-based PCR markers specific to individual chromosomes of Haynaldia villosa. Acta Agron Sin 35:1–10
Cao YP, Bie TD, Wang XE, Chen PD (2009b) Induction and transmission of wheat–Haynaldia villosa chromosomal translocations. J Genet Genomics 36:313–320
Chen PD, Liu DJ (1986) Identification of Haynaldia villosa chromosomes in wheat alien addition lines. In: Zhensheng L, Swaminathan MS (eds) Proceedings of the 1st International Symposium on Chromosome Engineering in Plants, Xian. pp. 31–32
Chen PD, Qi LL, Zhou B, Zhang SZ, Liu DJ (1995) Development and molecular cytogenetic analysis of wheat-Haynaldia villosa 6VS/6AL translocation lines specifying resistance to powdery mildew. Theor Appl Genet 91:1125–1128
Chen QZ, Qi ZJ, Feng YG, Wang SL, Chen PD (2002) Structural changes of 4V chromosome of Haynaldia villosa induced by gametocidal chromosome 3C of Aegilops triuncialis. J Genet Genom 29:355–358
Chen SW, Chen PD, Wang XE (2008) Inducement of chromosome translocation with small alien segments by irradiating mature female gametes of the whole arm translocation line. Sci China Ser C Life Sci 51:346–352
Chen PD, You CF, Hu Y, Chen SW, Zhou B, Cao AZ, Wang XE (2013) Radiation-induced translocations with reduced Haynaldia villosa chromatin at the Pm21 locus for powdery mildew resistance in wheat. Mol Breed 31:477–484
De Pace C, Snidaro D, Ciaffi M, Vittori D, Ciofo A, Cenci A, Tanzarella OA, Qualset CO, Scarascia Mugnozza GT (2001) Introgression of Dasypyrum villosum chromatin into common wheat improves grain protein quality. Euphytica 117:67–75
De Pace C, Vaccino P, Cionini G, Pasquini M, Bizzarri M, Qualset CO (2011) Dasypyrum. In: Kole C (ed) Wild crop relatives: genomic and breeding resources, cereals, vol 1, chapter 4. Springer, Heidelberg, pp 185–292
Endo TR, Gill BS (1996) The deletion stocks of common wheat. J Hered 87:295–307
Frank MY, Naxin H, Yong QG, Gerard RL, Dvorak J, Anderson OD (2009) ConservedPrimers 2.0: a high-throughput pipeline for comparative genome referenced intron-flanking PCR primer design and its application in wheat SNP discovery. BMC Bioinformatics 10:331–341
Friebe B, Kynast RG, Gill BS (2000) Gametocidal factor-induced structural rearrangements in rye chromosomes added to common wheat. Chromosom Res 8:501–511
Gradzielewska A (2006) The genus Dasypyrum—part 2. Dasypyrum villosum—a wild species used in wheat improvement. Euphytica 152:441–454
Harper LC, Cande WZ (2000) Mapping a new frontier, development of integrated cytogenetic maps in plants. Funct Integr Genomics 1:89–98
Joshi GP, Nasuda S, Endo TR (2011) Dissection and cytological mapping of barley chromosome 2H in the genetic background of common wheat. Genes Genet Syst 86:231–248
Kojima T, Habu Y, Iida S, Ogihara Y (2000) Direct isolation of differentially expressed genes from a specific chromosome region of common wheat: application of the amplified fragment length polymorphism-based mRNA fingerprinting (AMF) method in combination with a deletion line of wheat. Mol Gen Genet 263:635–641
Li HJ, Guo BH, Li YW, Du LQ, Jia X, Chu CC (2000) Molecular cytogenetic analysis of intergeneric chromosomal translocations between wheat (Triticum aestivum L.) and Dasypyrum villosum arising from tissue culture. Genome 43:756–762
Li HJ, Conner RL, Chen Q, Jia X, Li H, Graf RJ, Laroche A, Kuzyk AD (2002) Different reactions to the wheat curl mite and wheat streak mosaic virus in various wheat–Haynaldia villosa 6V and 6VS lines. Plant Dis 86:423–428
Li H, Chen X, Xin ZY, Ma YZ, Xu HJ, Chen XY, Jia X (2005) Development and identification of wheat–Haynaldia villosa T6DL·6VS chromosome translocation lines conferring resistance to powdery mildew. Plant Breed 124:203–205
Li HF, Gill BS, Wang XE, Chen PD (2011) A Tal-PhI wheat genetic stock facilitates efficient alien introgression. Genet Resour Crop Evol 58:667–678
Liu CJ, Chao S, Gale MD (1989) Wsp-1, a set of genes controlling water-soluble proteins in wheat and related species. Genet Res 54:173–181
Liu SB, Tang ZH, You MS, Li BY, Song JM, Liu GT (2004) Characterization of chromosome 1V specific SSR molecular markers for Haynaldia villosa. Acta Agron Sin 30:138–142
Liu C, Qi LL, Liu WX, Zhao WC, Wilson J, Friebe B, Gill BS (2011) Development of a set of compensating Triticum aestivum–Dasypyrum villosum Robertsonian translocation lines. Genome 54:836–844
Lukaszewski AJ (1988) A comparison of several approaches in the development if disomic alien addition lines of wheat. In: Miller TE, Koebner RMD (eds) Proc 7th Int Wheat Genet Symp, Cambridge, pp 363–367
Lukaszewski AJ, Rybka K, Korzun V, Malyshev SV, Lapinski B, Whitkus R (2004) Genetic and physical mapping of homoeologous recombination points involving wheat chromosome 2B and rye chromosome 2R. Genome 47:36–45
Montebove L, De Pace C, Jan CC, Qualset CO, Scarascia Mugnozza GT (1987) Chromosomal location of isozyme and seed storage protein genes in Dasypyrum villosum (L.) Candargy. Theor Appl Genet 73:836–845
Mukai Y, Nakaharya Y, Yamamotmo M (1993) Simultaneous discrimination of the three genomes in hexaploid wheat by multicolor fluorescence in situ hybridization using total genomic and highly repeated DNA probes. Genome 36:489–494
Pang YH, Chen XH, Zhao JX, Wu J, Chen XN, Liu SH, Yang QH, Du WL, Chen IG (2009) Cloning and sequence analysis of the HMW-GS gene and its promoter from Dasypyrum villosum. Acta Bot Boreal Sin 29:0859–0866
Qi LL, Wang SL, Chen PD, Liu DJ, Gill BS (1998) Identification and physical mapping of three Haynaldia villosa chromosome-6V deletion lines. Theor Appl Genet 97:1042–1046
Qi LL, Echalier B, Friebe B et al (2003) Molecular characterization of a set of wheat deletion stocks for use in chromosome bin mapping of ESTs. Funct Integr Genomics 3:39–55
Qi LL, Echalier B, Chao S et al (2004) A chromosome bin map of 16,000 expressed sequence tag loci and distribution of genes among the three genomes of polyploid wheat. Genetics 168:701–712
Qi LL, Pumphrey MO, Friebe B, Zhang P, Qian C, Bowden RL, Rouse MN, Jin Y, Gill BS (2011) A novel Robertsonian translocation event leads to transfer of a stem rust resistance gene (Sr52) effective against race Ug99 from Dasypyrum villosum into bread wheat. Theor Appl Genet 123:159–167
Schlegel R, Cakmak I, Torun B, Eker S, Tolay I, Ekiz H, Kalayci M, Braun HJ (1998) Screening for zinc efficiency among wheat relatives and their utilization for alien gene transfer. Euphytica 100:281–286
Sears ER (1953) Addition of the genome of Haynaldia villosa to Triticum aestivum. American J Botany 40:168–174
Sears ER (1966) Nullisomic–tetrasomic combinations in hexaploid wheat. In: Riley R, Lewis KR (eds) Chromosome manipulations and plant genetics. Oliver and Boyd, Edinburgh, pp 29–45
Sears ER, Sears LMS (1978) The telocentric chromosomes of common wheat. Ramanujam S (ed) Proceedings of the 5th international wheat genetics symposium, New Delhi, pp 389–407
Tsuchida M, Fukushima T, Nasuda S, Masoudi-Nejad A, Ishikawa G, Nakamura T, Endo TR (2008) Dissection of rye chromosome 1R in common wheat. Genes Genet Syst 83:43–53
Turner A, Beales J, Faure S, Dunford RP, Laurie DA (2005) The pseudo-response regulator Ppd-H1 provides adaptation to photoperiod in barley. Science 310:1031–1034
Wang CM, Feng YG, Zhuang LF, Cao YP, Qi ZJ, Bie TD, Cao AZ, Chen PD (2007) Screening of chromosome-specific markers for chromosome 1R of Secale cereale, 1V of Haynaldia villosa and 1Rk# 1 of Roegneria kamoji. Acta Agron Sin 33:1741–1747
Xu SS, Jin Y, Klindworth DL, Wang RR-C, Cai X (2009) Evaluation and characterization of seedling resistances to stem rust Ug99 races in wheat–alien species derivatives. Crop Sci 49:2167–2175
Yildirim A, Jones SS, Murray TD (1998) Mapping a gene conferring resistance to Pseudocercosporella herpotrichoides on chromosome 4V of Dasypyrum villosum in a wheat background. Genome 41:1–6
Yildirim A, Jones SS, Murray TD, Line RF (2000) Evaluation of Dasypyrum villosum populations for resistance to cereal eyespot and stripe rust pathogens. Plant Dis 84:40–44
Zhang P, Li WL, Friebe B, Gill BS (2004) Simultaneous painting of three genomes in hexaploid wheat by BAC FISH. Genome 7:979–987
Zhang QP, Li Q, Wang XE, Wang HY, Lang SP, Wang Y, Wang SL, Chen PD, Liu DJ (2005a) Development and characterization of a Triticum aestivum–Haynaldia villosa translocation line T4VS/4DL conferring resistance to wheat spindle streak mosaic virus. Euphytica 145:317–320
Zhang LY, Bernard M, Leroy P, Feuillet C, Sourdille P (2005b) High transferability of bread wheat EST-derived SSRs to other cereals. Theor Appl Genet 111:677–687
Zhang W, Gao AL, Zhou B, Chen PD (2006) Screening and applying wheat microsatellite markers to trace individual Haynaldia villosa chromosome. Acta Genet Sin 33:236–241
Zhang RQ, Wang XE, Chen PD (2012) Molecular and cytogenetic characterization of a small alien-segment translocation line carrying the softness genes of Haynaldia villosa. Genome 55:639–646
Zhang W, Zhang RQ, Feng YG, Bie TD, Chen PD (2013) Distribution of highly repeated DNA sequences in Haynaldia villosa and its application in the identification of alien chromatin. Chin Sci Bull 58:890–897
Zhang RQ, Zhang MY, Wang XE, Chen PD (2014) Introduction of chromosome segment carrying the seed storage protein genes from chromosome 1V of Dasypyrum villosum showed positive effect on bread-making quality of common wheat. Theor Appl Genet 127:523–533
Zhang RQ, Hou F, Feng YG, Zhang W, Zhang MY, Chen PD (2015) Characterization of a Triticum aestivum–Dasypyrum villosum T2VS·2DL translocation line expressing a longer spike and more kernels traits. Theor Appl Genet 128:2415–2425
Zhang RQ, Sun BX, Chen J, Cao AZ, Xing LP, Feng YG, Lan CX, Chen PD (2016a) Pm55, a developmental-stage and tissue-specific powdery mildew resistance gene introgressed from Dasypyrum villosum into common wheat. Theor Appl Genet 129:1975–1984
Zhang RQ, Feng YG, Li HF, Yuan HX, Dai JL, Cao AZ, Xing LP, Li HL (2016b) Cereal cyst nematode resistance gene CreV effective against Heterodera filipjevi transferred from chromosome 6VL of Dasypyrum villosum to bread wheat. Mol Breed 36:122. doi:10.1007/s11032-016-0549-9
Zhao RH, Wang HY, Xiao J, Bie TG, Cheng SH, Jia Q, Yuan CX, Zhang RQ, Cao AZ, Chen PD, Wang XE (2013) Induction of 4VS chromosome recombinants using the CS ph1b mutant and mapping of the wheat yellow mosaic virus resistance gene from Haynaldia villosa. Theor Appl Genet 126:2921–2930
Zhong GY, Dvorak J (1995) Evidence for common genetic mechanism controlling the tolerance of sudden salt stress in the tribe Triticeae. Plant Breed 114:297–302
Acknowledgments
This project was supported by the National Natural Science Foundation of China (31101142), the State Transgenic Project (2014ZX08009-40B), and the Fundamental Research Funds for the Central Universities (KYZ201303). The authors are grateful to Prof. Peidu Chen, College of Agronomy, Nanjing Agricultural University, Nanjing, China, for critically reviewing the manuscript.
Author information
Authors and Affiliations
Contributions
Ruiqi Zhang conceived and designed the project; Ruonan Yao, Dafei Sun, and Binxiao Sun screened the molecular markers; Yigao Feng, Wei Zhang, and Mingyi Zhang contributed to the creation and identification of the alien introgressions; and Ruiqi Zhang and Ruonan Yao wrote the manuscript.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflicts of interest.
Electronic supplementary material
Supplemental Table S1
The molecular markers physically mapped onto the V genome of D. villosum in this study. (PDF 100 kb)
Supplemental Figure S1
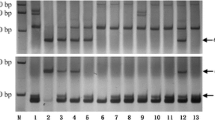
Electrophoresis patterns of the PCR products amplified by the HMW-GS gene-specific marker P1/P5. The straight line on the right indicates the fragment that can be allocated to a certain HMW-GS gene. The specific band of D. villosum is present in T1DL.1VL and 1 V-12 (TW.W-1VL) but is absent in T1BL.1VS. (JPEG 253 kb)
Supplemental Figure S2
Electrophoresis patterns of the PCR products amplified by the SSR markers Xcfd81 a and Xgwm182 b. The 5DS-specific bands are absent in 4 V-4 (T5DL.4VL), but 5DL-specific bands are present. (JPEG 77 kb)
Supplemental Figure S3
Electrophoresis patterns of the PCR products amplified by the STS marker CINAU44. The 7AS-specific band is absent in 7 V-1, indicating that the translocated chromosome of 7 V-1 may be T7AL.7VS. (JPEG 171 kb)
Rights and permissions
About this article
Cite this article
Zhang, R., Yao, R., Sun, D. et al. Development of V chromosome alterations and physical mapping of molecular markers specific to Dasypyrum villosum . Mol Breeding 37, 67 (2017). https://doi.org/10.1007/s11032-017-0671-3
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s11032-017-0671-3