Abstract
The amount of ultraviolet-B radiation (UV-B: 280–320 nm) reaching Earth’s surface is expected to increase due to stratospheric ozone depletion. This could cause significant biological damage in plants, and serious yield losses in crops. Soybean [Glycine max (L.) Merr.], a major legume crop, is known to be sensitive to UV-B radiation. Thus, developing a UV-B-tolerant soybean is an efficient and economical strategy to avoid putative yield losses through increased UV-B irradiation. The objective of this study is to identify the novel quantitative trait loci (QTLs) for UV-B tolerance in the soybean using high-density genetic linkage mapping. One hundred and fifteen F8-derived F12 recombinant inbred lines developed from a cross between the UV-B susceptible cultivar, Keunol, and a tolerant breeding line, Iksan 10, were used. Three categories of phenotypic traits were scored: degree of leaf color change, degree of leaf shape change and degree of total plant damage. A genome-wide molecular genetic linkage map containing 8691 single nucleotide polymorphism markers was constructed using the recently developed genotyping platform, the 180K Axiom SoyaSNP assay. Using composite interval mapping analysis, one major candidate QTL on chromosome 7 was identified and designated qUVBT1, and is located between two flanking makers, AX-90437826 and AX-90317546, within 1.6 cM, corresponding to a ~24-kb physical region with six annotated gene models. One of them is a homolog of yeast RAD23, which has previously been reported to be a UV excision repair protein. This result could be valuable in breeding new UV-B-tolerant soybean cultivars and elucidating the UV-B response mechanism in soybean plants.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
The depletion of stratospheric ozone has resulted in increased UV-B radiation at ground level that can cause morphological, physiological and biochemical damage in plants (Hidema and Kumagai 2006; Hollósy 2002; Kakani et al. 2003). Soybean [Glycine max (L.) Merr.] plants also show severe reduction in leaf expansion, leaf area, aerial dry biomass and seed yield with elevated UV-B treatment (Reed et al. 1992; Teramura et al. 1990; Yanqun et al. 2003). In particular, exposure to UV-B can induce the formation of covalent bonds between adjacent pyrimidines and creates the cyclobutane pyrimidine dimers and pyrimidine(6–4)pyrimidone photoproducts (Friedberg 2003). During DNA replication and transcription, these lesions disrupt base pairing and cause the mutations which result in phenotypic traits. Therefore, such lesions should be repaired to maintain the DNA integrity.
Recent achievements in soybean genomics research have improved the speed and precision of mapping of genes or quantitative trait loci (QTLs) for agronomically important traits. Since soybean plant genomic information was released (Schmutz et al. 2010), many genomic re-sequencing studies have produced millions of single nucleotide polymorphisms (SNPs). Hyten et al. (2010b) identified 25,047 putative SNPs from the re-sequencing of Glycine soja genotype PI 468916. With the re-sequencing of 17 wild and 14 cultivated soybean accessions, Lam et al. (2010) obtained 6.31 million SNPs. In addition, Chung et al. (2014) sequenced ten Korean cultivated and six wild accessions at ~17× depth and obtained 3.87 million high-quality SNPs. Recently, 302 wild and cultivated soybean accessions have been investigated with re-sequencing, and various novel QTLs have been identified (Zhou et al. 2015). As huge amounts of high-quality SNPs were accumulated, a high-throughput SNP genotyping system was also established. For soybean plants, the GoldenGate assay system was first developed for genotyping 1536 SNPs in 192 DNA samples and then used to construct the Universal Soy Linkage Panels (USLP) (Hyten et al. 2008, 2010a, b). Using an Illumina iSelect Beadchip, the SoySNP50 K array, which enables the high-density genotyping of 24 DNA samples with 52,041 SNPs, was also developed and used to genotype more than 19,000 Glycine accessions in the USDA Soybean Germplasm Collection (Song et al. 2013). Recently, the highest resolution of genotyping array was achieved by a 180K AXIOM SoyaSNP array with Affymetrix platforms (Lee et al. 2015b). This array is composed of more than 170,000 scorable SNPs and provides approximately one SNP per 6.5 kb over the soybean genome. High-density SNP genotyping arrays facilitate the rapid development of high-density genetic maps which permit the more precise localization of QTLs and enable the development of very tightly linked SNP markers for marker-assisted selection in breeding (Ganal et al. 2012).
Recently, we identified a UV-B-tolerant soybean breeding line, Iksan 10, showing no phenotypic damage in leaf chlorosis, leaf shape change and petiole color change with UV-B radiation (Shim et al. 2015). To identify the QTLs conferring UV-B tolerance, linkage analysis of a 115 recombinant inbred line (RIL) population derived from a cross between Keunol and Iksan 10 with 110 simple sequence repeat (SSR) markers was conducted and four to six QTLs associated with degree of leaf chlorosis, leaf shape change, petiole color change and total plant damage by UV-B radiation were identified. However, because of limited numbers of mapped molecular markers, most QTLs were detected over a 20-cM interval. This low-resolution linkage map might lower the QTL detection power, and therefore there is a need for a fine-mapping study to identify candidate genes or closely linked markers for marker-assisted selection.
The objective of this study is to identify and narrow down the QTLs for UV-B tolerance in soybean plants using the state-of-the-art genotyping platform, 180K Axiom® SoyaSNP assay, and develop SNP markers tightly linked to the genes. Furthermore, it is expected that a limited number of annotated genes may be identified by high-resolution dissection of the UV-B resistance QTLs in the UV-B-tolerant soybean breeding line, Iksan 10.
Materials and methods
Plant materials
The mapping population used in this study has been previously described (Shim et al. 2015). Briefly, 115 F8-derived F12 RILs were advanced by single-seed descent from crosses between a UV-B-susceptible cultivar, Keunol, which was derived from pure line selection of Korean landraces, and a tolerant breeding line, Iksan 10 (KI-RIL), which was derived from the cross between KW552 and Pangsakong (Lee et al. 2015a).
Evaluation of UV-B tolerance
The phenotypic trait for UV-B tolerance was re-evaluated with 115 F8:12 RILs and their parents in a greenhouse according to a previous study with minor modifications (Shim et al. 2015). Briefly, the RILs and their parents were planted in four replicates in 27-cm wide × 53-cm long trays with 50 holes filled with synthetic cultivation soil, and thinned to one plant per slot after germination. The parents, Keunol and Iksan 10, were planted in the center row in each tray, and RILs were planted with a completely randomized design. A UV-B lamp (G40T10E UV-B lamps, Sankyo, Denki, Japan) was wrapped with 0.13 mm of cellulose diacetate film (Cadillac Plastics Co., Baltimore, MD, USA) to filter out the UV-C (<290 nm) light. The cellulose diacetate film was replaced once a week, because of its aging by UV radiation. The distance between the UV lamp and the tops of the plants was maintained to 20–30 cm for constant UV-B dosage. The soybean plants were exposed to supplemental UV-B radiation at the V2 stage (Fehr et al. 1971) for 4 h per day (10:00–14:00) for 4 weeks with 1.5–2.0 W m−2 UV-B intensity. The UV-B and UV-C intensity was monitored using a UV-radiometer (DO 9847, Delta OHM) equipped with LP 471 UVB for UV-B and UV-C, respectively. After 4 weeks of irradiation, three categories of damage degree were scored on a scale of 1–9 [where 1 = no symptom (i.e., same degree of damage to Iksan 10) and 9 = severely damaged]: degree of leaf chlorosis (DLC), degree of leaf shape change (DLS), and degree of total plant damage (DTP) (Supplemental Figure S1). The degree of damage of Keunol was scored at 7. Each plant damage degree was scored by comparing to parents, Keunol and Iksan 10, within each tray.
DNA extraction and 180K Axiom SoyaSNP assay
The non-expanded young trifoliate leaves from three plants per RIL line were harvested for genomic DNA isolation. The genomic DNA was extracted with the modified hexadecyltrimethylammonium bromide (CTAB) method as previously described (Lenis 2011). For genetic linkage map construction, two parental and 115 F8:12 KI-RIL genomic DNA samples were genotyped with 180,961 SNP markers using the 180K Axiom SoyaSNP array (Affymetrix, Santa Clara, CA, USA) and scanned with a GeneTitan® Scanner (Affymetrix). Of 180,961 SNPs, only 169,028 high-quality SNPs (93.4 % of the tiled SNPs), excluding 1185 SNPs in the scaffolds and 10 SNPs in chloroplast, were used for further genetic linkage map construction and QTL identification (Lee et al. 2015b).
QTL analysis
A genetic linkage map was constructed using JoinMap 4.1 software (Van Ooijen 2006) according to the manufacturer’s protocol. Recombination fractions were converted to genetic map distance by the Kosambi function (Kosambi 1943). Of 169,028 scorable SNP markers, a total of 26,054 SNP markers showed polymorphism. After removing segregation distortion and redundant markers, only 8691 SNP markers were used for genetic linkage map construction.
Based on the segregation of 8691 SNP markers from KI-RILs and phenotypic traits (i.e., DLS, DLC and DTP), composite interval mapping analysis was performed to identify candidate QTL regions that are responsible for UV-B tolerance, executed by MapQTL 6 software (Van Ooijen 2009). The critical significance threshold was determined by 1000 permutations with a walk speed of 1 cM and a significance level of 0.05. The critical thresholds for logarithm of odds (LOD) were set to 4.0, 3.8 and 3.9 for DLS, DLC and DTP, respectively. The physical locations of two closest markers were identified using the manufacturer’s instructions based on the Wm82.a2 soybean assembly (Schmutz et al. 2010). The annotated genes within the two closest markers were identified through the SoyBase DB Sequence map viewer (Grant et al. 2010).
Results
Phenotypic trait evaluation under supplemental UV-B radiation
The three types of morphological damage caused by supplemental UV-B radiation, DLC, DLS and DTP, were evaluated in the RIL mapping population and their parents, Keunol, and Iksan 10, in greenhouse conditions (Supplemental Figure S1). The UV-B-susceptible parent, Keunol, was assigned a score of 7, while the tolerant parent, Iksan 10, did not show morphological damage and was scored 1 in all traits (Table 1). The average of all RILs’ traits was 4.5 with range distribution from 1 (no symptoms) to 9 (severely damaged). In addition, transgressive segregants, which had higher damage scores than Keunol, were observed. The phenotypic correlation coefficients were estimated between DLC, DLS and DTP (data not shown). All morphological damages showed positive and highly significant associations with each other (r > 0.95). These results suggest that common genes are responsible for three UV-B-induced morphological damage grades, DLC, DLS and DTP. To evaluate the traits’ normality, the Shapiro–Wilk normality test was conducted. None of the three traits showed a normal distribution (p < 0.001), but showed the bimodal shape of phenotype frequency distribution in the 115 KI-RILs. This suggests that a limited number of genes may govern UV-B tolerance in DLC, DLS and DTP (Fig. 1).
Phenotype frequency distribution of the 115 RILs derived from Keunol × Iksan 10. The distribution of degrees of leaf chlorosis (a), leaf shape (b) and total plant damage (c) are presented. Score 1 = no symptoms and 9 = severely damaged. The location of parent Keunol is marked with an open arrow and that of parent Iksan 10 with a closed arrow
Composite interval mapping of QTLs associated with UV-B tolerance
The high-density linkage map for 115 KI-RILs consisted of 20 chromosomes, which were defined by 8691 SNP markers (Table 2). On average, the constructed map revealed a marker density of one SNP marker per 0.5 cM. The increased marker density was 27 times greater than the previously reported map using the same KI-RILs (15.9 cM average marker distance) (Kim et al. 2005).
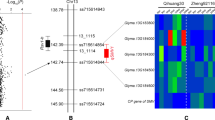
For identification of candidate QTL regions conferring UV-B tolerance, composite interval mapping was performed using the three phenotype traits DLC, DLS and DTP. Based on the composite interval mapping analysis, a single QTL near AX-90312571, which was positioned on a 221,548-bp region on chromosome 7 based on the Wm82.a2 soybean genome assembly, was identified for all the traits (Table 3). The major QTL on chromosome 7 was located between two flanking SNP markers, AX-90437826 and AX-90317546, as shown in Fig. 2. This accounts for approximately 60 % of the phenotypic variation, with a high LOD score of 23. In addition, the Iksan 10 allele was associated with UV-B tolerance in all three phenotypic traits, DLC, DLS and DTP . Currently, there are no reports regarding UV-B tolerance QTLs on chromosome 7 in soybean plants. We designated this newly identified novel QTL as qUVBT1. The physical distance between the two flanking SNP markers, AX-90437826 and AX-90317546, is 24 kb, and there are only six annotated genes based on the Williams 82 reference genome according to the USDA-ARS soybean genetic database (Grant et al. 2010). Figure 2c shows the qUVBT1 region and annotated genes with their physical position information. Table 4 shows the list of potential candidate genes and their Arabidopsis thaliana homologs in the identified qUVBT1 region. Taken together, the previously reported QTL region responsible for UV-B tolerance was narrowed down to six candidate genes within a 24-kb interval and designated qUVBT1.
Composite interval map and physical region of the major QTL associated with UV-B resistance on soybean chromosome 7 in the 115 RILs derived from Keunol × Iksan 10. a The qUVBT1 on the entire region of chromosome 7. b The qUVBT1 region with adjacent SNP markers. c The annotated genes and their physical position within qUVBT1 based on Glyma2.0 from Soybase DB
Discussion
Soybean is the most important legume crops for humans and livestock, but it is known to be sensitive to UV-B radiation. This may cause a reduction in plant biomass and yield, inhibition of photosynthesis and damage to DNA, resulting in more than 20 % yield reduction in susceptible soybean cultivars (Hollósy 2002; Kakani et al. 2003; Hidema and Kumagai 2006; Reed et al. 1992; Teramura et al. 1990). Thus, developing a UV-B-tolerant soybean cultivar is the most efficient and economical strategy for avoiding significant yield losses of soybean plants.
There have been several previous reports on UV-B responses in the model plant Arabidopsis thaliana. A number of knock-out mutants for photoreceptor (UVR8), photolyase (UVR2) and subunits of nuclear excision repair complex (UVR7 and UVH1) showed hypersensitive phenotypes to UV-B supplemental radiation, even though their overexpression mutants did not showed increased UV-B tolerance (Kliebenstein et al. 2002; Liu et al. 2000; Landry et al. 1997; Hefner et al. 2003). Recent research has demonstrated enhanced UV-B tolerance in rice and Arabidopsis. In rice, overexpression of WRKY89 enhanced UV-B tolerance with increased wax deposition on leaf surfaces (Wang et al. 2007). The HAG3 (histone acetyltransferase) knock-down mutant showed a lower inhibition of leaf and root growth by UV-B due to increased levels of DNA repair genes in the UV-B response (Fina and Casati 2015). In soybean, the QTL conferring UV-B tolerance was recently identified on chromosomes 7, 11, 13 and 19 from the Iksan 10 allele in a RIL population from the cross between Keunol and Iksan 10 using only 110 SSR markers (Shim et al. 2015). However, the limited number of markers mapped caused the wide interval of the QTL region identified and this could be a bottleneck for gene function identification research, because of a number of annotated gene models within the QTL interval identified resulting in difficulties in narrowing down the candidate gene models.
Recent efforts in soybean genomics have identified a huge number of SNPs based on the soybean reference genome assembly (Schmutz et al. 2010; Hyten et al. 2010b; Zhou et al. 2015), and the SNPs identified could be employed to construct a high-density genetic linkage map which is equivalent to a physical sequence map.
In the current research, the most recently developed high-density SNP genotyping platform, 180K Axiom SoyaSNP assay, was employed to narrow down the QTL interval and identify novel QTLs which are responsible for UV-B radiation tolerance. In addition, the QTL interval identified was projected onto the physical sequence map based on the Williams 82 soybean genome assembly to investigate the candidate genes within the QTL interval. The composite interval mapping analysis results showed that the major QTL, qUVBT1, was positioned at around 220 kb on the physical sequence map within a 24-kb interval on chromosome 7 in all three phenotype traits with high LOD score, phenotypic variance explained (%) and additive effect (Table 3). Interestingly, a previous report showed that a major QTL was located on chromosome 19 with minor QTLs on chromosome 7, 11, 13 and 14, explaining 3.8–9.2 % of the phenotypic variance. This significant difference may be due to the limited number of available polymorphic SSR markers used for genotyping and the absence of SSR markers close to QTL. Here, this practical limitation was addressed using the 180K Axiom SoyaSNP assay, which employed more than 80 times more SNP markers than the previous report, to overcome the marker number issue and reveal recombination events within the RILs as precisely as possible. Thus, this study identified the major QTL for UV-B tolerance in soybean plants on chromosome 7, which could not be identified as a major QTL in previous research because of the limited number of molecular markers for genotyping and linkage map construction. Overall, qUVBT1 on chromosome 7 appears to play a key role in tolerance to UV-B radiation in the Iksan 10 soybean breeding line.
There are six annotated genes models within the qUVBT1 interval, based on the Wm82.a2.v1 soybean reference genome assembly and annotation. Interestingly, one of the candidate genes encodes a UV excision repair protein homolog of RADIATION SENSITIVE23B (RAD23b) in Arabidopsis thaliana. RAD23b in Arabidopsis is the homolog of RAD23, which is associated with the nucleotide excision repair factor 2 (NEF2) complex, originally discovered in yeast (Saccharomyces cerevisiae) (Guzder et al. 1998). A mutation in RAD23 family members shows hypersensitivity to UV light in yeast (Lambertson et al. 1999). In Arabidopsis, there are four homologs of the RAD23 yeast gene: RAD23a, RAD23b, RAD23c and RAD23d. The Arabidopsis homolog of the candidate gene is RAD23b, with 76 % protein sequence similarity. However, RAD23b-1 seedlings, which were Arabidopsis knock-out mutants for RAD23b by T-DNA insertion, failed to display a hyper- or hyposensitive phenotype to several DNA-damaging agents, including UV light (Farmer et al. 2010). However, Sturm and Lienhard (1998) reported that two RAD23 isoforms from carrot (Daucus carota) can rescue the UV-sensitive phenotype of RAD23Δ yeast. These previous reports suggest that RAD23 in soybean plants might participate in the DNA damage repair process.
A soybean plant has 13 RAD23 homologs, and one of them was identified within the qUVBT1 interval. Thus, the investigation of RAD23b homologs in soybean plants (Glyma.07g002800) for a UV-B tolerance phenotype in Iksan 10 may be interesting and valuable work for developing UV-B-tolerant varieties of soybean plants, though the functional analyses of these candidate genes are ongoing and will be the subject of future studies.
In summary, a novel QTL, qUVBT1, for tolerance to supplemental UV-B radiation in soybean has been identified with 8691 polymorphic SNP markers using the 180K Axiom SoyaSNP assay and composite interval mapping analysis. qUVBT1 is on chromosome 7 between the two flanking markers AX-90437826 and AX-90317546, located at 198,735 and 223,377 bps, respectively. Within this approximately 24-kb region, there are six annotated candidate genes. Since soybean plants are a UV-B-susceptible crop, UV-B may cause serious yield losses, and therefore breeding for new UV-B-tolerant soybean cultivars is necessary. Thus, identification of qUVBT1 and its two flanking SNP markers will contribute to marker-assisted selection for UV-B-tolerant soybean cultivars. Certainly, continued gene-functional analysis for candidate genes in mutant soybean plants and construction of near-isogenic lines to elucidate the UV-B tolerance mechanism in soybean plants is required and will be the subject of subsequent research.
Abbreviations
- QTL:
-
Quantitative trait locus
- SNP:
-
Single nucleotide polymorphism
- RIL:
-
Recombinant inbred line
- DTP:
-
Degree of total plant damage
- DLC:
-
Degree of leaf color chlorosis
- DLS:
-
Degree of leaf shape change
References
Chung WH, Jeong N, Kim J, Lee WK, Lee YG, Lee SH, Yoon W, Kim JH, Choi IY, Choi HK, Moon JK, Kim N, Jeong SC (2014) Population structure and domestication revealed by high-depth resequencing of Korean cultivated and wild soybean genomes. DNA Res 21(2):153–167. doi:10.1093/dnares/dst047
Farmer LM, Book AJ, Lee KH, Lin YL, Fu H, Vierstra RD (2010) The RAD23 family provides an essential connection between the 26S proteasome and ubiquitylated proteins in Arabidopsis. Plant Cell 22(1):124–142. doi:10.1105/tpc.109.072660
Fehr WR, Caviness CE, Burmood DT, Pennington JS (1971) Stage of development descriptions for soybeans, Glycine max (L.) Merrill. Crop Sci. doi:10.2135/cropsci1971.0011183X001100060051x
Fina JP, Casati P (2015) HAG3, a histone acetyltransferase, affects UV-B responses by negatively regulating the expression of DNA repair enzymes and sunscreen content in Arabidopsis thaliana. Plant Cell Physiol 56(7):1388–1400. doi:10.1093/pcp/pcv054
Friedberg EC (2003) DNA damage and repair. Nature 421(6921):436–440. doi:10.1038/nature01408
Ganal MW, Polley A, Graner EM, Plieske J, Wieseke R, Luerssen H, Durstewitz G (2012) Large SNP arrays for genotyping in crop plants. J Biosci 37(5):821–828
Grant D, Nelson RT, Cannon SB, Shoemaker RC (2010) SoyBase, the USDA-ARS soybean genetics and genomics database. Nucleic Acids Res 38(Database issue):D843–D846. doi:10.1093/nar/gkp798
Guzder SN, Sung P, Prakash L, Prakash S (1998) Affinity of yeast nucleotide excision repair factor 2, consisting of the Rad4 and Rad23 proteins, for ultraviolet damaged DNA. J Biol Chem 273(47):31541–31546
Hefner E, Preuss SB, Britt AB (2003) Arabidopsis mutants sensitive to gamma radiation include the homologue of the human repair gene ERCC1. J Exp Bot 54(383):669–680
Hidema J, Kumagai T (2006) Sensitivity of rice to ultraviolet-B radiation. Ann Bot 97(6):933–942. doi:10.1093/aob/mcl044
Hollósy F (2002) Effects of ultraviolet radiation on plant cells. Micron 33(2):179–197. doi:10.1016/S0968-4328(01)00011-7
Hyten D, Song Q, Choi I-Y, Yoon M-S, Specht J, Matukumalli L, Nelson R, Shoemaker R, Young N, Cregan P (2008) High-throughput genotyping with the GoldenGate assay in the complex genome of soybean. Theor Appl Genet 116(7):945–952. doi:10.1007/s00122-008-0726-2
Hyten D, Cannon S, Song Q, Weeks N, Fickus E, Shoemaker R, Specht J, Farmer A, May G, Cregan P (2010a) High-throughput SNP discovery through deep resequencing of a reduced representation library to anchor and orient scaffolds in the soybean whole genome sequence. BMC Genom 11(1):38
Hyten DL, Choi I-Y, Song Q, Specht JE, Carter TE, Shoemaker RC, Hwang E-Y, Matukumalli LK, Cregan PB (2010b) A high density integrated genetic linkage map of soybean and the development of a 1536 universal soy linkage panel for quantitative trait locus mapping. Crop Sci 50(3):960–968. doi:10.2135/cropsci2009.06.0360
Kakani VG, Reddy KR, Zhao D, Sailaja K (2003) Field crop responses to ultraviolet-B radiation: a review. Agric For Meteorol 120(1–4):191–218. doi:10.1016/j.agrformet.2003.08.015
Kim HK, Kang ST, Cho JH, Choung MG, Suh DY (2005) Quantitative trait loci associated with oligosaccharide and sucrose contents in soybean (Glycine max L.). J Plant Biol 48(1):106–112
Kliebenstein DJ, Lim JE, Landry LG, Last RL (2002) Arabidopsis UVR8 regulates ultraviolet-B signal transduction and tolerance and contains sequence similarity to human regulator of chromatin condensation 1. Plant Physiol 130(1):234–243. doi:10.1104/pp.005041
Kosambi DD (1943) The estimation of map distances from recombination values. Ann Eugen 12(1):172–175. doi:10.1111/j.1469-1809.1943.tb02321.x
Lam HM, Xu X, Liu X, Chen W, Yang G, Wong FL, Li MW, He W, Qin N, Wang B, Li J, Jian M, Wang J, Shao G, Wang J, Sun SS, Zhang G (2010) Resequencing of 31 wild and cultivated soybean genomes identifies patterns of genetic diversity and selection. Nat Genet 42(12):1053–1059. doi:10.1038/ng.715
Lambertson D, Chen L, Madura K (1999) Pleiotropic defects caused by loss of the proteasome-interacting factors Rad23 and Rpn10 of Saccharomyces cerevisiae. Genetics 153(1):69–79
Landry LG, Stapleton AE, Lim J, Hoffman P, Hays JB, Walbot V, Last RL (1997) An Arabidopsis photolyase mutant is hypersensitive to ultraviolet-B radiation. Proc Natl Acad Sci USA 94(1):328–332
Lee C, Choi M-S, Kim H-T, Yun H-T, Lee B, Chung Y-S, Kim RW, Choi H-K (2015a) Soybean [Glycine max (L.) Merrill]: importance as a crop and pedigree reconstruction of Korean varieties. Plant Breed Biotechnol 3(3):179–196
Lee Y-G, Jeong N, Kim JH, Lee K, Kim KH, Pirani A, Ha B-K, Kang S-T, Park B-S, Moon J-K, Kim N, Jeong S-C (2015b) Development, validation and genetic analysis of a large soybean SNP genotyping array. Plant J 81(4):625–636. doi:10.1111/tpj.12755
Lenis JM (2011) Genetics of soybean seed lipoxygenases and linolenic acid content in seeds of the soybean wild ancestor. University of Missouri-Columbia
Liu Z, Hossain GS, Islas-Osuna MA, Mitchell DL, Mount DW (2000) Repair of UV damage in plants by nucleotide excision repair: arabidopsis UVH1 DNA repair gene is a homolog of Saccharomyces cerevisiae Rad1. Plant J Cell Mol Biol 21(6):519–528
Reed HE, Teramura A, Kenworthy WJ (1992) Ancestral U.S. soybean cultivars characterized for tolerance to ultravioletB radiation. Crop Sci 32(5):1214–1219
Schmutz J, Cannon SB, Schlueter J, Ma J, Mitros T, Nelson W, Hyten DL, Song Q, Thelen JJ, Cheng J, Xu D, Hellsten U, May GD, Yu Y, Sakurai T, Umezawa T, Bhattacharyya MK, Sandhu D, Valliyodan B, Lindquist E, Peto M, Grant D, Shu S, Goodstein D, Barry K, Futrell-Griggs M, Abernathy B, Du J, Tian Z, Zhu L, Gill N, Joshi T, Libault M, Sethuraman A, Zhang XC, Shinozaki K, Nguyen HT, Wing RA, Cregan P, Specht J, Grimwood J, Rokhsar D, Stacey G, Shoemaker RC, Jackson SA (2010) Genome sequence of the palaeopolyploid soybean. Nature 463(7278):178–183. doi:10.1038/nature08670
Shim H-C, Ha B-K, Yoo M, Kang S-T (2015) Detection of quantitative trait loci controlling UV-B resistance in soybean. Euphytica 202(1):109–118. doi:10.1007/s10681-014-1233-y
Song Q, Hyten DL, Jia G, Quigley CV, Fickus EW, Nelson RL, Cregan PB (2013) Development and evaluation of SoySNP50K, a high-density genotyping array for soybean. PLoS One 8(1):e54985. doi:10.1371/journal.pone.0054985
Sturm A, Lienhard S (1998) Two isoforms of plant RAD23 complement a UV-sensitive rad23 mutant in yeast. Plant J Cell Mol Biol 13(6):815–821
Teramura AH, Sullivan J, Lydon J (1990) Effects of UV-B radiation on soybean yield and seed quality: a 6 year field study. Physiol Plant 80(1):5–1; (1):5–11
Van Ooijen JW (2006) JoinMap 4.0: software for the calculation of genetic linkage maps in experimental populations. Kyazma B.V., Wageningen, Netherlands
Wang H, Hao J, Chen X, Hao Z, Wang X, Lou Y, Peng Y, Guo Z (2007) Overexpression of rice WRKY89 enhances ultraviolet B tolerance and disease resistance in rice plants. Plant Mol Biol 65(6):799–815. doi:10.1007/s11103-007-9244-x
Yanqun Z, Yuan L, Haiyan C, Jianjun C (2003) Intraspecific differences in physiological response of 20 soybean cultivars to enhanced ultraviolet-B radiation under field conditions. Environ Exp Bot 50(1):87–97. doi:10.1016/s0098-8472(03)00004-2
Zhou Z, Jiang Y, Wang Z, Gou Z, Lyu J, Li W, Yu Y, Shu L, Zhao Y, Ma Y, Fang C, Shen Y, Liu T, Li C, Li Q, Wu M, Wang M, Wu Y, Dong Y, Wan W, Wang X, Ding Z, Gao Y, Xiang H, Zhu B, Lee SH, Wang W, Tian Z (2015) Resequencing 302 wild and cultivated accessions identifies genes related to domestication and improvement in soybean. Nat Biotechnol 33(4):408–414. doi:10.1038/nbt.3096
Acknowledgments
This work was carried out with the support of National Research Foundation of Korea (Project No. NRF-2013R1A1A2009812).
Author contribution
JSL designed and conducted field tests and drafted the manuscript. SK conducted field tests. BKH helped to construct genetic linkage map and QTL analysis. STK designed the experiment and organized the manuscript. All authors read and approved the final manuscript. Authors state that the experiments comply with the current laws of the country in which they were performed.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Lee, J.S., Kim, S., Ha, BK. et al. Positional mapping and identification of novel quantitative trait locus responsible for UV-B radiation tolerance in soybean [Glycine max (L.) Merr.]. Mol Breeding 36, 50 (2016). https://doi.org/10.1007/s11032-016-0471-1
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s11032-016-0471-1