Abstract
Histone deacetylases constitute a group of enzymes that participate in several biological processes. Notably, inhibiting HDAC8 has become a therapeutic strategy for various diseases. The current inhibitors for HDAC8 lack selectivity and target multiple HDACs. Consequently, there is a growing recognition of the need for selective HDAC8 inhibitors to enhance the effectiveness of therapeutic interventions. In our current study, we have utilized a multi-faceted approach, including Quantitative Structure–Activity Relationship (QSAR) combined with Quantitative Read-Across Structure–Activity Relationship (q-RASAR) modeling, pharmacophore mapping, molecular docking, and molecular dynamics (MD) simulations. The developed q-RASAR model has a high statistical significance and predictive ability (Q2F1:0.778, Q2F2:0.775). The contributions of important descriptors are discussed in detail to gain insight into the crucial structural features in HDAC8 inhibition. The best pharmacophore hypothesis exhibits a high regression coefficient (0.969) and a low root mean square deviation (0.944), highlighting the importance of correctly orienting hydrogen bond acceptor (HBA), ring aromatic (RA), and zinc-binding group (ZBG) features in designing potent HDAC8 inhibitors. To confirm the results of q-RASAR and pharmacophore mapping, molecular docking analysis of the five potent compounds (44, 54, 82, 102, and 118) was performed to gain further insights into these structural features crucial for interaction with the HDAC8 enzyme. Lastly, MD simulation studies of the most active compound (54, mapped correctly with the pharmacophore hypothesis) and the least active compound (34, mapped poorly with the pharmacophore hypothesis) were carried out to validate the observations of the studies above. This study not only refines our understanding of essential structural features for HDAC8 inhibition but also provides a robust framework for the rational design of novel selective HDAC8 inhibitors which may offer insights to medicinal chemists and researchers engaged in the development of HDAC8-targeted therapeutics.
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
Introduction
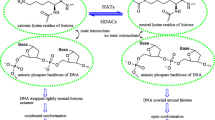
In the modern society, cancer has become an increasingly prevalent disease, characterized by a gradual accumulation of genetic alterations leading to the malfunctioning of tumor suppressor genes and/or the hyperactivity of oncogenes [1,2,3]. The modulation of genes through epigenetic mechanisms is pivotal in reshaping structure of nucleosomes by modifying the interaction between DNA and histones. Among various epigenetic modifications, histone deacetylation, affecting the fundamental packaging units of DNA, has been identified as a contributor to gene expression control. This process results in the formation of more condensed chromatin and subsequent transcriptional suppression [2, 4, 5]. The family of histone deacetylases (HDACs) is primarily responsible for catalyzing deacetylation on lysine residues [6, 7]. Aberrant expression of various HDACs has been implicated in numerous human disorders, including cancer, making them promising targets for therapeutic intervention across a spectrum of human malignancies [8].
HDACs are categorized into four groups based on phylogenetic relationships, with Class III acting as a nicotinamide-adenine dinucleotide-dependent lysine deacetylase, while Classes I, II, and IV are zinc-dependent [7, 9, 10]. Among HDAC enzymes, histone deacetylase 8 (HDAC8) inhibition has emerged as a prominent therapeutic strategy for various diseases. HDAC8, a class I histone deacetylase, is targeted for the treatment of conditions such as parasite infections, cancer, and X-linked intellectual disability [5, 11]. HDAC8 is a class I HDAC found in both the cytoplasm and nucleus of vital organs such as the heart, lung, kidney, and brain. It consists of 377 amino acids, weighing approximately 42 kDa, and lacks a C-terminal protein-binding domain. One of its distinctive features involves an unexpected susceptibility to negative regulation by cAMP-dependent protein kinase (PKA), suggesting the possibility of functional specialization. Notably, HDAC8 targets non-histone proteins, including structural maintenance of chromosome 3 (SMC3) cohesin protein, retinoic acid-induced 1 (RAI1), and the tumor suppressor gene p53 [8,9,10,11,12,13,14]. Moreover, HDAC8 is involved in promoting the proliferation of gastric adenocarcinomas, lung cancers, and cervical malignancies in humans [14, 15]. Additionally, it has been identified as a catalyst responsible for the in vitro deacetylation of several acetylated histone variants [16]. Apart from its implications in cancer, HDAC8 has been recognized for its substantial involvement in conditions like schistosomiasis and influenza-A infections [17].
Ongoing studies suggest that HDAC8 holds promise as a therapeutic target for conditions such as neuroblastoma, T-cell leukemia, and acute myeloid leukemia [18]. At present, the majority of inhibitors targeting HDAC8 exhibit broad-spectrum activity, affecting multiple isoforms from class I, class II, and class IV [19]. However, the primary drawback of these clinically approved pan-HDAC inhibitors lies in their non-selective nature, resulting in various adverse effects. Despite their clinical approval, these medications fall short of fulfilling the criteria for a selective and potent inhibitor, crucial for the effective anticancer treatment of HDAC8-associated diseases [20].
In the recent past, a multitude of QSAR studies [21,22,23,24,25,26,27,28,29,30,31,32] have been employed to decipher the essential structural features that impact HDAC8 inhibition (Table S1). However, despite the widespread use of QSAR techniques, a persistent gap exists due to challenges in statistically significant model generation with a properly curated dataset of HDAC8 inhibitors and also in achieving robust predictions [33]. Furthermore, the diverse chemical structures of HDAC8 inhibitors pose a challenge in identifying a universally applicable set of descriptors that can comprehensively elucidate their inhibitory activity. To address these challenges, we have utilized Quantitative Read-Across Structure–Activity Relationship (q-RASAR) which involves the study of 121 HDAC8 inhibitors with distinct IC50 values sourced from published research [34,35,36,37,38,39,40,41,42,43,44,45].
q-RASAR is a statistical modeling technique that improves the external predictivity of QSAR/QSPR models by including similarity and error-based metrics as descriptors in addition to the standard structural and physicochemical ones [46, 47]. While using similarity-based considerations, this method can produce models that are straightforward, comprehensible, and transferable. The q-RASAR technique holds promise in data gap filling in materials science, food sciences, predictive toxicology, medicinal chemistry, agricultural sciences, nano-sciences, and so on [46]. The present study employs an integrated in silico approach to investigate the critical structural features required for effective HDAC8 inhibition. The research encompasses four primary components (depicted in Fig. 1) aimed at crafting potent HDAC8 inhibitors: (a) Utilizing 2D-QSAR modeling to pinpoint essential structural characteristics of HDAC8 inhibitors, (b) Employing q-RASAR modeling to enhance the external predictability of HDAC8 inhibition, (c) Conducting a pharmacophore mapping study to elucidate potential pharmacophoric features governing HDAC8 inhibitory activity, and (d) Validating these identified structural features through molecular docking and dynamics simulation-based methods. Combining these methods provides a powerful toolkit for understanding, predicting, and validating the structural features crucial for HDAC8 inhibition, offering valuable insights for discovering and developing selective HDAC8 inhibitors.
Materials and methods
2D-QSAR modeling
Data set
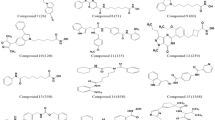
This study focuses on the q-RASAR modeling of 121 HDAC8 inhibitors with specific IC50 values extracted from published research [34,35,36,37,38,39,40,41,42,43,44,45]. The dataset is considered as biologically curated since all the reported inhibitors underwent evaluation for HDAC8 inhibition using the same method. Table S2 displays the reported IC50 values for the 121 compounds. The logarithm of the reciprocal of the half maximal inhibitory concentration (pIC50) of HDAC8 inhibitors was taken as the response variable for the generation of the QSAR model.
Molecular descriptor calculation
For the development of QSAR models, we utilized a diverse array of descriptor types [48]. Specifically, we employed 0D−2D descriptors for model generation. This choice was made to reduce the computational load associated with energy minimization and conformational analysis [49]. Several classes of 2D descriptors, including ring descriptors, molecular features, constitutional index, functional group count, and electro-topochemical atom descriptors, were computed using the PaDEL-Descriptor program [50]. The DTC Lab's pre-treatment tool (using Data Pre-Treatment 1.2 from http://teqip.jdvu.ac.in/QSAR_Tools/) eliminated the intercorrelated descriptors (intercorrelation cut off > 0.99) and those with minimal variability in values (variance cut off < 0.0001) [51, 52].
Data set division
Splitting the data set is an important step in QSAR model development. We segregated the dataset into training and test sets. The test set was dedicated to validating the established model externally, whereas the training set primarily facilitated model development. To accomplish this, we utilized the "datasetDivisionGUI1.2_19Feb2019" program [52], which implemented the sorted activity-based division technique in our current work. The data points within a cluster were found to be comparable to each other but distinct from those found outside of it. After the organization of the complete data set by cluster number and related activity levels, we selected approximately 20% of the data points from each cluster to be used as test set compounds (\({N}_{\text{test}}\) = 24, pIC50 range = 4.915 to 7.066), with the remaining 80% treated as the training set compounds (\({N}_{\text{train}}\)= 97, pIC50 range = 4.988 to 7.678) for the QSAR study.
Feature selection and development of the QSAR model
For feature selection, we utilized the genetic algorithm (GA) technique [53], which relies on a fitness function based solely on mean absolute error (MAE) criteria. Using the Genetic Algorithm v4.1 [52], we identified descriptors with the strongest correlation to the response variable. Subsequently, we employed "Partial Least Squares (PLS) regression" to develop the initial QSAR models after pinpointing the significant descriptors.
q-RASAR model development
To increase the QSAR model’s external predictability, a q-RASAR model was developed using RASAR descriptors along with the structural descriptors [54]. The most similar compounds were identified by using the "Readacross v4.2" tool (available from https://sites.google.com/jadavpuruniversity.in/dtc-lab-software/home), which is based on Euclidean distance (ED)-based similarity, Gaussian kernel (GK) function similarity, and Laplacian kernel (LK) function similarity [55]. After splitting the training set in an 80:20 ratio, we got the sub-training and subtest sets, and used the following parameters for the optimization of the method: γ = 1, σ = 1, distance threshold = 1, number of close training compounds = 7, and similarity threshold = 0. Laplacian kernel function similarity read-across is the least error-prone method according to the optimization result (Table S3) and it is used for the RASAR descriptor calculation by using the "RASAR-Desc-Calc-v3.0.2." tool (available from https://sites.google.com/jadavpuruniversity.in/dtc-lab-software/home) [54,55,56] Finally, using the Best Subset Selection 2.1 tool [52], the pooled set of RASAR descriptors and previously selected 2D descriptors has been processed to the best subset selection. Partial Least Squares version 1.0 [52] was used to build the final PLS q-RASAR model.
ML-based qRASAR model development
For the development of a ML regression model, we employed supervised machine learning algorithms such as AdaBoost (ADB), Extreme Gradient Boost (XGB), Linear Support Vector Machine (LSVM) and Support Vector Machine (SVM). These models were developed using the previously mentioned training (\({N}_{train}\)= 97) and test set data (\({N}_{test}\) = 24). There might be opportunities for the model to be improved even more, and in order to do so, hyperparameter tuning was done. In the present study, we used GridSearchCV algorithm for tuning the hyperparameters which were used for the development of ML models. For performing this optimization, we used DTC Lab’s Python-based tool “Optimization and Cross-validation v1.0.” (https://sites.google.com/jadavpuruniversity.in/dtc-lab-software/home/machine-learning-model-development-guis). Using the optimized hyperparameter settings alongside the training and test sets, we developed four machine learning-based q-RASAR models. This was achieved using a Python-based tool called Machine Learning Regressor v2.0, which can be accessed from https://sites.google.com/jadavpuruniversity.in/dtc-lab-software/home/machine-learning-model-development-guis [57].
Pharmacophore mapping
In this investigation, Discovery Studio software [58] was used to create ligand-based 3D QSAR pharmacophore models. Here, the “3D QSAR Pharmacophore Model Generation” module of Discovery Studio 3.0 [59] was applied for model development. The dataset comprising 121 compounds was divided into training set (Ntrain = 24) and a test set (Ntest = 97). The training set consists of molecules with IC50 activity values spanning from 21 nM to 12,170 nM, including highly active (< 350 nM), moderately active (> 350–2500 nM), and inactive (> 2500 nM) compounds. During the hypothesis generation procedure, the module determines the cost function. Equation 1 depicts the formula used for estimating the cost (total cost) of a hypothesis:
where the coefficients for the error (E), weight (W), and configuration (C) components are denoted by the letters e, w, and c, respectively.
Two other important cost values are the fixed cost and the null cost that can be calculated using Eqs. 2 and 3, respectively.
where x basically represents the deviation from the expected values of weight and error.
where \({\chi }_{est}\) is the averaged scaled activity of the training set molecules.
The configuration cost is determined by the complexity of the pharmacophore hypothesis space. An increase in the root mean square (rms) value leads to a corresponding increase in the error cost value. The rms deviations serve as a metric for assessing the correlation quality, primarily between the estimated and actual activity data [60, 61].
Molecular docking study
The molecular docking study was conducted using the zinc metalloenzyme-optimized AutoDock4Zn [62] utilizing the AutoDock Vina program. The crystal structure of HDAC8 (PDB ID: 1T64) sourced from RCSB (https://www.rcsb.org/) was employed to explore the interactions between the protein and ligands. The protein preparation included the removal of water molecules, the application of Gasteiger partial charges, and the addition of polar hydrogens. Five active compounds (44, 54, 82, 102, and 118) and the least active compound (34) were selected from the pharmacophore mapping study for docking analysis. The ligands were imported into the AutoDock Tools suite [63], where polar hydrogens were added. PDBQT files were generated for both the ligands and the HDAC8 enzyme. The protein was placed at the center of a grid box to cover its entire binding region (grid center: 61.3, 73.865, 11.247; grid box volume: 40 × 40 × 40). Molecular docking was performed using the Lamarckian genetic algorithm (LGA). The ligand-binding interactions within the HDAC8 cavity were visualized using Discovery Studio 3.0 [59].
Molecular dynamics study
In this study, we have performed MD simulation study by employing the GROMACS 2021.1 software [64, 65] on the docked complexes of the most active (54) and the least active (34) compounds with HDAC8, implementing CHARM-GUI web server [66] to prepare different inputs. The PDB reader tool was first employed to facilitate protein pre-processing [67]. Additionally, using the OpenBabel program, the best-docked poses of ligands were converted to the “. mol2" format [68]. After that, the ligands were imported into the Modeler and Ligand reader tools to parameterize and generate topology files [69]. Subsequently, each protein–ligand complex was combined into a ‘.pdb’ file, which was then utilized in the "Solution Builder" function to create the GROMACS input system [70]. The entire protein structure was covered by the TIP3P water box system, which is rectangular. To remove steric overlapping, each system was neutralized using a sufficient amount of NaCl ions added using the Monte-Carlo method [71] and then run through 5000 steps of steepest descent energy minimization [72]. The next step involved subjecting the entire system to V-rescale temperature-coupling (constant coupling of 1 ps at 310.15 K temperature) for 125000 steps to achieve NVT equilibration, where the number of particles, volume, and temperature are all constant [73]. The MD simulation was run with a CHARMM36 m forcefield for 500 ns [74]. Using the gmx__rms, gmx_rmsf, and gmx_gyrate programs, respectively, the simulation results were further evaluated for a variety of geometrical properties, including radius of gyration (Rg), root mean square deviation (RMSD), and root mean square fluctuations (RMSF) [75]. With the aid of the gmx_hbond program, the hydrogen bond analysis between ligands of interest and active site amino acid residues was completed [76, 77]. Subsequently, production trajectories were created using PyMOL [78] to analyze the binding poses at various time intervals.
Results and discussions
QSAR and q-RASAR models
A dataset of 121 HDAC8 inhibitors was used for 2D-QSAR model development. Initially, a pool of 1444 2D descriptors were generated using the PaDEL-Descriptor tool. This was followed by the pre-treatment of the data, which generated 813 2D descriptors. These descriptors were chosen and put to use in the feature selection and model building. Finally, a PLS regression-based QSAR model (Eq. 4) with four latent variables was generated for the present study.
As per the regulations provided by the OECD, the model's performance was evaluated through stringent internal and external validation. The determination coefficient (R2 = 0.713), leave-one-out cross-validated correlation coefficient (Q2(LOO) = 0.654), and rm2 metrics of the training set [rm2 (train) = 0.540 and Δrm2 (train) = 0.177] are the internal validation metrics that demonstrate the model's robustness and goodness of fit. The mean absolute error of the training set (MAEtrain) for the model is 0.255. Additionally, the external validation metrics was computed, which include external predicted variance (Q2F1 = 0.732 and Q2F2 = 0.727), mean absolute error of test set predictions (MAEtest = 0.249), rm2 (test) = 0.519, Δrm2 (test) = 0.248, and concordance correlation coefficient (CCC = 0.825).
The q-RASAR model was developed to improve the external predictability of the QSAR model. Utilizing PLS regression with four latent variables (4 LVs), we merged the pool of seven structural descriptors of Eq. 4 with the computed read-across-based RASAR descriptors for the optimal subset selection and final model development (Eq. 5). The identified structural descriptors of Eq. 5 and their definition are summarized in Table 1.
Following the OECD guidelines [79], this final model has undergone an exhaustive validation process using several internal and external validation criteria. Table 2 presents a comparison of validation parameters before (Eq. 4) and after (Eq. 5) the read-across strategy. A comparison of internal and external validation metrics before and after the implementation of the read-across strategy is illustrated using bar plots in Fig. 2.
The final q-RASAR model (Eq. 5) exhibits superior statistical significance in terms of both internal and external validation parameters when compared to the QSAR model (Eq. 4). The internal validation parameters like R2 (Train), Q2(LOO) and Scaled Average rm2 are increased, whereas Mean Absolute Errors (MAE-Fitted; Train and MAE-LOO; Train) as well as Scaled Delta rm2 are decreased in the qRASAR model (Eq. 5). The external validation parameters like Q2F1 and Q2F2 as well as concordance correlation coefficient (CCC) are also significantly increased compared to the previous model (Eq. 4). Moreover, Mean Absolute Error (MAE; Test) is decreased suggesting a better external predictivity of the q-RASAR model (Eq. 5).
Thus, external validation metric values (Q2F1 and Q2F2), as well as the model's internal validation metrics, such as R2, Q2(LOO), and rm2 metrics justified better model's performance in the case of q-RASAR model. The coefficient plot, variable importance plot (VIP), score plot, and loading plot of the final q-RASAR model (Eq. 5) were generated by using the SIMCA-P program [80]. The coefficient plot is presented in Fig. 3A, and it demonstrates that AATS7s, maxHaaCH, TIC2, and sm1(LK) are descriptors that contribute positively to the developed model, whereas AATS0i, C3SP2, and sm2(LK) contribute negatively. The variable importance plot (VIP) is shown in Fig. 3B to identify the significant descriptors in the QSAR model for HDAC8 inhibition. As per the VIP, HDAC8 inhibition has the following order of relative relevance for the contributing descriptors: sm1(LK) > sm2(LK) > TIC2 > AATS7s > AATS0i > C3SP2 > maxHaaCH. Figure 3C displays the score plot demonstrating that no molecule is recognized as an outlier. The loading plot in Fig. 3D further indicates that the descriptors sm1(LK), sm2(LK), TIC2, and AATS7s had the largest impact (as mentioned in Fig. 3B) on predicting the HDAC8 inhibition because of their distant placement from the origin.
A Coefficient plot, B variable importance plot (VIP), C score plot, and D loading plot of the final q-RASAR model (Eq. 5)
In addition, we attempted to develop other machine learning-based q-RASAR models, namely AdaBoost, Extreme Gradient Boost, Support Vector Machine, and Linear Support Vector Machine for the prediction of HDAC8 inhibitory activity using seven descriptors, which were identified as significant contributing features in our PLS q-RASAR model. The statistical parameters of the ML models and selected hyperparameters for developing these models are shown in Table 3. It is evident from Table 2 that the statistics of our q-RASAR PLS model outperforms the developed ML-based q-RASAR models. Therefore, we have continued further studies based on the PLS q-RASAR model.
Mechanistic interpretation of HDAC8 inhibition
Mechanistic interpretation is very important for any QSAR model as per OECD Guideline 5. The five structural descriptors (AATS0i, AATS7s, C3SP2, maxHaaCH, and TIC2) and two RASAR descriptors, sm1 (LK) and sm2 (LK) [81], are employed for the development of the final model. The contribution of structural descriptors is important to gain insight into the HDAC8 inhibition. Among the structural descriptors, TIC2 has the highest impact on HDAC8 inhibition (Fig. 3B). The descriptor TIC2, which is a function of molecular structure, basically encapsulates the total information content of the 2-order symmetry. Higher values of this descriptor typically indicate a higher degree of symmetry or balanced relationships, while lower values suggest a more asymmetric or imbalanced structure. The greater values of TIC2 have a positive impact on HDAC8 inhibition. Thus, the compounds 82, 102, and 118 exhibit higher HDAC8 inhibition due to the increased TIC2 values (Fig. 4).
Identified seven 0D-2D descriptors [5 structural descriptors: AATS0i, AATS7s, C3SP2, maxHaaCH, TIC2 and 2 RASAR descriptors: sm1 (LK) and sm2 (LK)] used in the model. Among 7 descriptors, AATS7s, maxHaaCH, TIC2, sm1(LK) are the descriptors responsible for positive contribution. AATS0i, C3SP2 and sm2(LK) are responsible for negative contribution
The descriptor C3SP2 exhibits the greatest negative contribution to the HDAC8 inhibition. The C3SP2 descriptor [82] basically interprets the number of carbons that are doubly bound, and it is further attached to three other carbon atoms. The presence of the above-mentioned type of carbon in the structure was found to be responsible for the decrease in HDAC8 inhibition (compounds 22, 27 and 126). Compound 54 showed significant HDAC8 inhibition due to the low value for the aforementioned descriptor (Fig. 4).
The variable AATS0i [83] also exhibits a significant negative contribution to the model, as observed by its coefficient value in the coefficient plot (Fig. 3A). The descriptor AATS0i is an averaged moreau-broto autocorrelation of lag 0 weighted by ionization potential. This descriptor combines the concepts of ionization potential and the Moreau-Broto autocorrelation to quantify compounds' structural and electronic properties. In simpler terms, it represents how the ionization potential of a compound is related to its internal structural features, specifically focusing on the autocorrelation at lag 0. In the case of compounds 52 and 69, increasing the value of the AATS0i descriptor produces a decrease in HDAC8 inhibition. It is also noticed that compounds 44 exhibit significant HDAC8 activity due to the lower value of the AATS0i descriptor (Fig. 4).
The descriptor maxHaaCH has the least standardized coefficient value and it made the smallest positive contribution to HDAC8 inhibition. The descriptor maxHaaCH indicates the maximum atom-type H and it focuses on the presence and arrangement of hydrogen atoms within the molecular structure [84]. It is used to quantify the maximum occurrence of a specific type of hydrogen atom or a specific configuration of hydrogen atoms in the molecule. It has been found that increasing the value of the maxHaaCH descriptor promotes HDAC8 inhibition as indicated in compounds 44, 112, and 89 (Fig. 4).
The final structural descriptor of the model, AATS7s descriptor indicates the average Broto-Moreau autocorrelation—lag 7/weighted by I-state [85]. Broto-Moreau autocorrelation is a mathematical concept used to analyze the arrangement of atoms in a molecule at different distances, and the lag 7 part indicates that it is specifically looking at the correlation between properties of atoms or substructures separated by a distance of 7 bonds in the molecular structure. The term "weighted by I-state" suggests that the autocorrelation values are adjusted or weighted based on the electronic state of the atoms involved. In the case of compounds 89 and 44, we found that increased values of the AATS7s descriptor promote HDAC8 inhibition, whereas decreased values of the descriptor reduce HDAC8 inhibition (compounds 34 and 52) (Fig. 4).
Notably, two RASAR-based descriptors, sm1(LK) and sm2(LK), emerged as significant in the final model. These descriptors represent similarity coefficients, offering a means to identify compounds or inhibitors with biological activity or HDAC8 inhibition [87]. The mathematical representations of these coefficients are shown below:
where MaxPos and MaxNeg denote the similarity scores of the nearest positive source and negative source compounds, respectively, concerning a specific query compound [81].
where PosAvgSim signifies the average similarity values obtained from the positive close source compounds, whereas NegAvgSim indicates the average similarity values derived from the negative close source compounds [81].
The RASAR descriptor sm1(LK) [86] is positively correlated with HDAC8 inhibition. The higher values of the sm1(LK) coefficient are observed in the case of compounds 54 (pIC50 = 7.678), 78 (pIC50 = 7.456), 104 (pIC50 = 6.987), and 112 (pIC50 = 6.757). For an active compound to be considered ideal, its MaxPos value should be higher than the MaxNeg value. This condition leads to a positive sm1 value. Conversely, a negative sm1 value suggests that the MaxNeg value is higher than the MaxPos value. This indicates that the compound structurally resembles an inactive compound from a close source rather than an active one (e.g., compound 34, pIC50 = 4.915). The RASAR descriptor sm2(LK) shows a negative correlation with HDAC8 inhibition and is determined by the difference between positive and negative average similarity. The higher values of the sm2(LK) coefficient are identified in the case of compounds 60 (pIC50 = 6.076), 69 (pIC50 = 5.883), 124 (pIC50 = 5.699), and 129 (pIC50 = 6.452). This suggests that compounds similar to these in the training set are less effective at inhibiting HDAC8.
Pharmacophore mapping study
A total of 24 training set compounds were employed for the development of pharmacophore models by utilizing the “3D QSAR Pharmacophore Model Generation” module of Discovery Studio 3.0 [59]. Chemical features such as hydrogen bond acceptor (HBA), hydrophobic (HYP), ring aromatic (RA), hydrophobic ring aromatic (HY_RA), and zinc-binding group (ZBG) features were incorporated to generate ten pharmacophore hypotheses, with parameters adjusted for weight variation and uncertainty as detailed in Table 4.
A thorough analysis of Table 4 reveals that HYP1 stands out as the best pharmacophore model, featuring the highest correlation coefficient (r = 0.969), the lowest root mean square (rms) of 0.944, a total cost value of 86.078, and the most significant cost difference value of 146.856. The inhibitory activity (IC50) of the modeling set molecules against HDAC8 using the HYP1 model is illustrated in Table 5.
The statistical significance of the HYP1 model was evaluated using Fischer's randomization test, which involved generating 19 random pharmacophore hypotheses at a 95% confidence level for cross-validation. Comparatively, the cost value of HYP1 was lower (Total cost = 86.078) than that of the 19 randomly generated hypotheses (see Fig. S1, Supplementary file). In essence, this suggests that the superiority of the HYP1 model is not coincidental but rather deliberate and statistically supported.
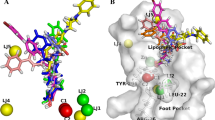
A cost difference (ΔCost) exceeding 60 implies a correlation probability of over 90%. The correlation coefficient (rExternal) of the external set compounds demonstrates promising results, indicating the statistical significance and robustness of the best pharmacophore hypothesis, HYP1. Consequently, HYP1 is chosen as the final pharmacophore model for HDAC8 inhibitors. The identified pharmacophoric features from this model are distinct compared to prior studies [23, 32, 87,88,89,90,91,92,93,94,95,96] (refer to Table S4 of the Supporting Info). Figure 5A illustrates the 3D spatial relationship and geometric parameters of the HYP1 model, along with inter-feature distances. The presence of the ZBG is imperative for HDAC8 inhibitory activity, along with hydrogen bond acceptor features (HBA1, HBA2), and ring aromatic (RA) features. Compound 54, the most potent HDAC8 inhibitor (IC50 = 21 nM), adeptly matches all pharmacophore properties (Fig. 5B), while compound 34, the least potent HDAC8 inhibitor (IC50 = 12,170 nM), fails to align with all features (Fig. 5C).
A Representation of the best pharmacophore (HYP1) model. Inter-feature distances are shown in angstroms. The green and orange contours represent hydrogen bond acceptor (HBA) and ring aromatic (RA) features, respectively. The nevi blue contours represent the zinc-binding group (ZBG) feature. B Mapping of the most active compound (54) onto the selected pharmacophore (HYP1) C Mapping of the least active compound (34) onto the selected pharmacophore (HYP1). The green and orange contours represent hydrogen bond acceptor (HBA) and ring aromatic (RA) features, respectively. The nevi blue contours represent zinc-binding group (ZBG) feature
Molecular docking study
In order to find the significance of different pharmacophoric features in its interaction with the HDAC8 enzyme (PDB: 1T64), a molecular docking study was conducted for the five potent HDAC8 inhibitors (54, 44, 84, 102, and 118). Figure 6 depicts the docking poses and ligand–enzyme interactions of these HDAC8 inhibitors. Notably, five of the most active compounds fit precisely into binding pocket when docked into the active site of HDAC8 using AutoDock Vina program [62]. The 2D interactions and docking score of all these compounds are depicted in Figs. S2–S4 and Table S6, respectively.
From the pharmacophoric mapping study, it has been noted that the zinc-binding group is essential for HDAC8 inhibition. The molecular docking study also explains that all the potent HDAC8 inhibitors show metal (Zn2+) coordination in the HDAC8 binding site. Both the amide oxygen as well as the hydroxyl oxygen of hydroxamate moiety are involved in metal coordination with Zn2+ in compound 44, whereas in the case of compounds 54, 84, 102, and 118, only the amide oxygen is involved. Y306 is involved in various interactions in all the compounds, suggesting that Y306 is a prerequisite for binding with HDAC8 ligands. Y306 participates in H-bonding interactions with the zinc-binding hydroxamate moiety, engaging the amide oxygen and the hydroxyl oxygen in compound 118. In compounds 44 and 54, Y306 forms π–π interactions with F152. In compound 82, Y306 is involved in alkyl interactions with the trimethoxy group. Moreover, in compound 102, hydrogen bonding interactions of Y306 with the NH group of the pyrrolidine ring and π–π stacking with the tetrahydroisoquinoline ring were also seen. Another, important residue F152 showed a significant contribution to ligand-HDAC8 binding. F152 forms π-interactions with the sulfur atom of the dithiolane ring and π–π-T shaped interactions with the phenyl ring in compound 54. As per the pharmacophore mapping study, the sulfur atom of dithiolane ring and the phenyl ring act as hydrogen bond acceptor (HBA1) and ring aromatic (RA) features, respectively, necessary for the HDAC8 inhibition. The importance of ring aromatic feature in the HDAC8 inhibitor structure is also highlighted by the presence of π–π stacking interaction with the residue F152 and the indole ring of compound 82. Residue I34 also forms π–alkyl interactions with the phenyl ring (ring aromatic feature) of compound 54. In compound 44, H143 and K33, are involved in H-bonding interactions, R37 forms π–cation interactions, and P35 is involved in π–alkyl interactions. In compound 54, G304 forms H-bonding with OH of the hydroxamate group. In compound 82, Y100 forms π–alkyl interactions with pyrrolidine ring and L308, P35, and Y306 are involved in alkyl interactions with the trimethoxy group. In compound 102, I34, K33 and F152 are involved in alkyl interactions, M274 forms π–sulfur interactions with the aromatic ring of the tetrahydroisoquinoline ring, and H143 is involved in H-bonding interactions. Residues I34, P35, and F152, along with residue Y306, participate in alkyl interactions with the trimethoxy group within compound 118. The least active compound (34) exhibited poor binding with the HDAC8 enzyme and showed fewer interactions with some amino acid residues. Compound 34 displayed π–π stacking interactions with Y100 and M274. H-bonding interactions with Y100 and van der Waals interaction with H143 were also observed.
Molecular dynamics simulation
A molecular dynamics (MD) simulation study is an approach to assess the fluctuation, and atomic motion of individual atoms or groups, as well as changes in conformations of a molecule with receptor/protein for a specific time [97]. By foreseeing a specific pose in terms of the interactions of the ligand with the macromolecule, the MD simulation aims to verify the stability of the receptor–ligand complex.
In the current study, MD simulation (500 ns) was performed for the most active HDAC8 inhibitor (54) and least active HDAC8 inhibitor (34). Compound 54 has all the pharmacophoric features and it binds nicely in the active site of HDAC8. The least active compound (34) was also docked (Fig. S3) in the active site of HDAC8 and was inspected through MD simulation (500 ns) for comparison. After the MD simulation, we compared the RMSD, Rg, and RMSF values of the protein–ligand complexes stated above to HDAC8 apo form (control system). The average RMSD of the backbone atoms of HDAC8 apo form and HDAC8 complexed with compounds 54 and 34 were 0.20 ± 0.02 nm, 0.15 ± 0.01 nm and 0.17 ± 0.02 nm, respectively during 500 ns. The lower RMSD of HDAC8 complexed with compound 54 implies that this compound induces a more significant conformational change or stabilization in the protein structure than to the apo form. Similarly, the RMSD of HDAC8 complexed with compound 34 indicates that this compound also induces a significant structural change or stabilization in the protein compared to the apo form, though perhaps not as pronounced as compound 54 (Fig. 7A).
To examine the compactness of HDAC8 in the presence and absence of the aforementioned ligands, the radius of gyration (Rg) is plotted in Fig. 7B. The average Rg values for the HDAC8 apo form and complexes with compounds 54 and 34 were uniformly found to be 2.01 ± 0.01 nm, respectively, demonstrating that the compactness of HDAC8 does not significantly change in the presence of compounds 54 and 34 even after 500 ns period (Fig. 7B). Furthermore, the RMSF of individual HDAC8 residues is determined to examine the flexibility or rigidity of various locations within HDAC8 apo form and its complexes with compounds 54 and 34 (Fig. 7C). The average RMSF of backbone atoms in HDAC8 apo form and in the presence of compounds 54 and 34 were found to be 0.09 ± 0.07 nm, 0.08 ± 0.05 nm, and 0.08 ± 0.05 nm, respectively. It is inferred that in the presence of the aforementioned inhibitors, the average residual fluctuation of HDAC8 has reduced with respect to the fluctuation of the apo form.
During MD simulation, snapshots of all complexes were visualized at varied time intervals to investigate HDAC8-inhibitor complex stability within the binding pocket. Interestingly, compound 34 moves out from the catalytic pocket before 100 ns during the simulation period of 500 ns, but compound 54 (pIC50 = 7.678) remains in the binding site until 330 ns of the simulation (Fig. 8). To verify this observation, we have plotted individual distances between the oxygen atom of hydroxamate functionality of each ligand and zinc atom within the binding site as a function of time (Fig. 9). Additionally, to assess the impact of ligand size on HDAC8 inhibition, we conducted a 100 ns molecular dynamics (MD) simulation of a moderately active ligand with intermediate size (compound 37, pIC50 = 6.180) (see Fig. S5, Supplementary file). Compound 37 displays a similar pattern to compound 34 (pIC50 = 4.915), exiting the binding pocket before the 100 ns (Fig. S6, Supplementary file).
Conclusions
This study sequentially employed validated QSAR and q-RASAR models by using the PLS-regression-based method, pharmacophore mapping, molecular docking, and molecular dynamics approaches to identify essential structural features for potential HDAC8 inhibitors. Statistically significant PLS regression-based QSAR model (\({Q}_{F1}^{2}\):0.732, \({Q}_{F2}^{2}\):0.727, MAEtest: 0.249) was developed using four latent variables. q-RASAR strategy was applied to increase the external predictivity of HDAC8 inhibition. The developed q-RASAR model has a high statistical significance and predictive ability (\({Q}_{F1}^{2}\):0.778, \({Q}_{F2}^{2}\):0.775, MAEtest: 0.221). The different descriptors in the final q-RASAR model were discussed to get meaningful insight into the mechanistic aspects of HDAC8 inhibition. Different pharmacophoric features have also been identified through pharmacophore mapping studies, and it revealed that HYP1 was the best pharmacophore model, with the highest correlation coefficient (r = 0.969), and lowest rms of 0.944. The pharmacophore predictions showed that the ring aromatic (RA) feature near the hydrogen bond acceptor feature (HBA2) in HDAC8 inhibitor plays a crucial role in HDAC8 inhibition, while other features, such as the presence of ZBG, are also essential for HDAC8 inhibition. Based on the q-RASAR model and pharmacophore mapping studies, five HDAC8 inhibitors (compounds 44, 54, 82, 102, and 118) were chosen as potent inhibitors to assess the binding interactions by using molecular docking. A molecular docking study validated the results of pharmacophore mapping which demonstrated that the hydroxamate moiety (as ZBG) is involved in metal (Zn2+) coordination in the HDAC8 binding site in all the five potent HDAC8 inhibitors. Moreover, the ring aromatic feature (RA) and hydrogen bond acceptor (HBA1) feature necessary for HDAC8 inhibition of the most active inhibitor (54) were highlighted by the phenyl ring and the sulfur atom of dithiolane ring, respectively, in the molecular docking study (Fig. 10). Lastly, the complex stability of the most (54) and the least active (34) inhibitors were analyzed using MD simulation which indicated that inhibitor 54 exhibited more structural stability of the complex than inhibitor 34. The findings of this study could be useful for future HDAC8 inhibitor design, and the computational strategy used can be broadly applied to different targeted drug designs.
Overall, this comprehensive exploration sheds light on the intricate molecular aspects influencing HDAC8 inhibition, offering a foundation for future research endeavors in the pursuit of novel and effective HDAC8 inhibitors.
References
Hanahan D, Weinberg RA (2000) The hallmarks of cancer. Cell 100:57–70. https://doi.org/10.1016/S0092-8674(00)81683-9
Baylin SB, Ohm JE (2006) Epigenetic gene silencing in cancer—a mechanism for early oncogenic pathway addiction? Nat Rev Cancer 6:107–116. https://doi.org/10.1038/nrc1799
Wang LH, Wu CF, Rajasekaran N, Shin YK (2018) Loss of tumor suppressor gene function in human cancer: an overview. Cell Physiol Biochem 51:2647–2693. https://doi.org/10.1159/000495956
Handy DE, Castro R, Loscalzo J (2011) Epigenetic modifications: basic mechanisms and role in cardiovascular disease. Circulation 123:2145–2156. https://doi.org/10.1161/CIRCULATIONAHA.110.956839
Walewska A, Janucik A, Tynecka M, Moniuszko M, Eljaszewicz A (2023) Mesenchymal stem cells under epigenetic control—the role of epigenetic machinery in fate decision and functional properties. Cell Death Dis 14:720. https://doi.org/10.1038/s41419-023-06239-4
Aramsangtienchai P, Spiegelman NA, He B, Miller SP, Dai L, Zhao Y, Lin H (2016) HDAC8 catalyzes the hydrolysis of long chain fatty acyl lysine. ACS Chem Biol 11:2685–2692. https://doi.org/10.1021/acschembio.6b00396
Ruzic D, Djoković N, Srdić-Rajić T, Echeverria C, Nikolic K, Santibanez JF (2022) Targeting histone deacetylases: opportunities for cancer treatment and chemoprevention. Pharmaceutics 14:209. https://doi.org/10.3390/pharmaceics14010209
Gallinari P, Di Marco S, Jones P, Pallaoro M, Steinkühler C (2007) HDACs, histone deacetylation and gene transcription: from molecular biology to cancer therapeutics. Cell Res 17:195–211. https://doi.org/10.1038/sj.cr.7310149
Vannini A, Volpari C, Gallinari P, Jones P, Mattu M, Carfí A, De Francesco R, Steinkühler C, Di Marco S (2007) Substrate binding to histone deacetylases as shown by the crystal structure of the HDAC8—substrate complex. EMBO J Rep 8:879–884. https://doi.org/10.1038/sj.embor.7401047
Seto E, Yoshida M (2014) Erasers of histone acetylation: the histone deacetylase enzymes. Cold Spring Harb Perspect Biol 6:a018713. https://doi.org/10.1101/cshperspect.a018713
Chakrabarti A, Oehme I, Witt O, Oliveira G, Sippl W, Romier C, Pierce RJ, Jung M (2015) HDAC8: a multifaceted target for therapeutic interventions. Trends Pharmacol Sci 36:481–492. https://doi.org/10.1016/j.tips.2015.04.013
Karolczak-Bayatti M, Sweeney M, Cheng J, Edey L, Robson SC, Ulrich SM, Treumann A, Taggart MJ, Europe-Finner GN (2011) Acetylation of heat shock protein 20 (Hsp20) regulates human myometrial activity. J Biol Chem 286:34346–34355. https://doi.org/10.1074/jbc.M111.278549
Spreafico M, Gruszka AM, Valli D, Mazzola M, Deflorian G, Quintè A, Totaro MG, Battaglia C, Alcalay M, Marozzi A (2020) HDAC8: a promising therapeutic target for acute myeloid leukemia. Front Cell Dev Biol 8:844. https://doi.org/10.3389/fcell.2020.00844
Kim JY, Cho H, Yoo J, Kim GW, Jeon YH, Lee SW, Kwon SH (2022) Pathological role of HDAC8: cancer and beyond. Cells 11:3161. https://doi.org/10.3390/cells11193161
Wolfson NA, Pitcairn CA, Fierke CA (2013) HDAC8 substrates: histones and beyond. Biopolymers 99:112–126. https://doi.org/10.1002/bip.22135
Alam N, Zimmerman L, Wolfson NA, Joseph CG, Fierke CA, Schueler-Furman O (2016) Structure-based identification of HDAC8 non-histone substrates. Structure 24:458–468. https://doi.org/10.1016/j.str.2016.02.002
Fontana A, Cursaro I, Carullo G, Gemma S, Butini S, Campiani G (2022) A therapeutic perspective of HDAC8 in different diseases: an overview of selective inhibitors. Int J Mol Sci 23:10014. https://doi.org/10.3390/ijms231710014
Kuo YH, Qi J, Cook GJ (2016) Regain control of p53: targeting leukemia stem cells by isoform-specific HDAC inhibition. Exp Hematol 44:315–321. https://doi.org/10.1016/j.exphem.2016.02.007
Morris MJ, Monteggia LM (2013) Unique functional roles for class I and class II histone deacetylases in central nervous system development and function. Int J Dev Neurosci 31:370–381. https://doi.org/10.1016/j.ijdevneu.2013.02.005
Ghosh S, Chatterjee M, Roy K (2023) Predictive quantitative read-across structure-property relationship modeling of the retention time (Log t R) of pesticide residues present in foods and vegetables. J Agric Food Chem 71:9538–9548. https://doi.org/10.1021/acs.jafc.3c01438
Noor Z, Afzal N, Rashid S (2015) Exploration of novel inhibitors for class I histone deacetylase isoforms by QSAR modeling and molecular dynamics simulation assays. PLoS ONE 10:e0139588. https://doi.org/10.1371/journal.pone.0139588
Cao GP, Thangapandian S, Son M, Kumar R, Choi YJ, Kim Y, Kwon YJ, Kim HH, Suh JK, Lee KW (2016) QSAR modeling to design selective histone deacetylase 8 (HDAC8) inhibitors. Arch Pharm Res 39:1356–1369. https://doi.org/10.1007/s12272-015-0705-5
Manal M, Manish K, Sanal D, Selvaraj A, Devadasan V, Chandrasekar MJN (2017) Novel HDAC8 inhibitors: a multi-computational approach. SAR QSAR Environ Res 28:707–733. https://doi.org/10.1080/1062936X.2017.1375978
Martínez-Pacheco H, Ramírez-Galicia G, Vergara-Arias M, Gertsch J, Fragoso-Vazquez JM, Mendez-Luna D, Abujamra AL, Cristina CL, Cecilia RM, Mendoza-Lujambio I, Correa-Basurto J (2017) Docking and QSAR studies of aryl-valproic acid derivatives to identify antiproliferative agents targeting the HDAC8. Anticancer Agents Med Chem 17:927–940. https://doi.org/10.2174/1871520616666161019143219
Cao GP, Arooj M, Thangapandian S, Park C, Arulalapperumal V, Kim Y, Kwon YJ, Kim HH, Suh JK, Lee KW (2015) A lazy learning-based QSAR classification study for screening potential histone deacetylase 8 (HDAC8) inhibitors. SAR QSAR Environ Res 26:397–420. https://doi.org/10.1080/1062936X.2015.1040453
Amin SA, Adhikari N, Jha T (2018) Diverse classes of HDAC8 inhibitors: in search of molecular fingerprints that regulate activity. Future Med Chem 10:1589–1602. https://doi.org/10.4155/fmc-2018-0005
Banerjee S, Amin SA, Adhikari N, Jha T (2020) Essential elements regulating HDAC8 inhibition: a classification based structural analysis and enzyme-inhibitor interaction study of hydroxamate based HDAC8 inhibitors. J Biomol Struct Dyn 38:5513–5525. https://doi.org/10.1080/07391102.2019.1704881
Amin SA, Adhikari N, Jha T (2021) Development of decision trees to discriminate HDAC8 inhibitors and non-inhibitors using recursive partitioning. J Biomol Struct Dyn 39:1–8. https://doi.org/10.1080/07391102.2019.1661876
Amin SA, Adhikari N, Jha T (2020) Exploration of histone deacetylase 8 inhibitors through classification QSAR study: Part II. J Mol Struct 1204:127529. https://doi.org/10.1016/j.molstruc.2019.127529
Amin SA, Banerjee S, Adhikari N, Jha T (2020) Discriminations of active from inactive HDAC8 inhibitors part II: bayesian classification study to find molecular fingerprints. SAR QSAR Environ Res 31:245–260. https://doi.org/10.1080/1062936X.2020.1723136
Banerjee S, Adhikari N, Amin SA, Jha T (2020) Structural exploration of tetrahydroisoquinoline derivatives as HDAC8 inhibitors through multi-QSAR modeling study. J Biomol Struct Dyn 38:1551–1564. https://doi.org/10.1080/07391102.2019.1617782
Banerjee S, Baidya SK, Adhikari N, Jha T (2022) A comparative quantitative structural assessment of benzothiazine-derived HDAC8 inhibitors by predictive ligand-based drug designing approaches. SAR QSAR Environ Res 33:987–1011. https://doi.org/10.1080/1062936X.2022.2155241
Veerasamy R (2022) QSAR—an important in-silico tool in drug design and discovery. Advances in computational modeling and simulation. Springer, Berlin
Jiang Y, Li X, Wang X, Wang Z, Zhang J, Wu J, Xu W (2016) Design, synthesis, and biological characterization of tamibarotene analogs as anticancer agents. Chem Biol Drug Des 88:542–555. https://doi.org/10.1111/cbdd.12778
Wang X, Li X, Li J, Hou J, Qu Y, Yu C, He F, Xu W, Wu J (2017) Design, synthesis, and preliminary bioactivity evaluation of N1-hydroxyterephthalamide derivatives with indole cap as novel histone deacetylase inhibitors. Chem Biol Drug Des 89:38–46. https://doi.org/10.1111/cbdd.12819
Li J, Li X, Wang X, Hou J, Zang J, Gao S, Xu W, Zhang Y (2016) PXD 101 analogs with L-phenylglycine-containing branched cap as histone deacetylase inhibitors. Chem Biol Drug Des 88:574–584. https://doi.org/10.1111/cbdd.12787
Li X, Inks ES, Li X, Hou J, Chou CJ, Zhang J, Jiang Y, Zhang Y, Xu W (2014) Discovery of the first N-hydroxycinnamamide-based histone deacetylase 1/3 dual inhibitors with potent oral antitumor activity. J Med Chem 57:3324–3341. https://doi.org/10.1021/jm401877m
Zhang Y, Fang H, Feng J, Jia Y, Wang X, Xu W (2011) Discovery of a tetrahydroisoquinoline-based hydroxamic acid derivative (ZYJ-34c) as histone deacetylase inhibitor with potent oral antitumor activities. J Med Chem 54:5532–5539. https://doi.org/10.1021/jm200577a
Zhang Y, Feng J, Jia Y, Wang X, Zhang L, Liu C, Fang H, Xu W (2011) Development of tetrahydroisoquinoline-based hydroxamic acid derivatives: potent histone deacetylase inhibitors with marked in vitro and in vivo antitumor activities. J Med Chem 54:2823–2838. https://doi.org/10.1021/jm101605z
Duan W, Li J, Inks ES, Chou CJ, Jia Y, Chu X, Li X, Xu W, Zhang Y (2015) Design, synthesis, and antitumor evaluation of novel histone deacetylase inhibitors equipped with a phenylsulfonylfuroxan module as a nitric oxide donor. J Med Chem 58:4325–4338. https://doi.org/10.1021/acs.jmedchem.5b00317
Li X, Hou J, Li X, Jiang Y, Liu X, Mu W, Jin Y, Zhang Y, Xu W (2015) Development of 3-hydroxycinnamamide-based HDAC inhibitors with potent in vitro and in vivo anti-tumor activity. Eur J Med Chem 89:628–637. https://doi.org/10.1016/j.ejmech.2014.10.077
Zhang Y, Feng J, Jia Y, Xu Y, Liu C, Fang H, Xu W (2011) Design, synthesis and primary activity assay of tripeptidomimetics as histone deacetylase inhibitors with linear linker and branched cap group. Eur J Med Chem 46:5387–5397. https://doi.org/10.1016/j.ejmech.2011.08.045
Liu R, Wang J, Tang W, Fang H (2016) Design and synthesis of a new generation of substituted purine hydroxamate analogs as histone deacetylase inhibitors. Bioorg Med Chem 24:1446–1454. https://doi.org/10.1016/j.bmc.2016.02.005
Zhang Y, Feng J, Liu C, Fang H, Xu W (2011) Design, synthesis and biological evaluation of tyrosine-based hydroxamic acid analogs as novel histone deacetylases (HDACs) inhibitors. Bioorg Med Chem 19:4437–4444. https://doi.org/10.1016/j.bmc.2011.06.046
Zhang Y, Feng J, Liu C, Zhang L, Jiao J, Fang H, Su L, Zhang X, Zhang J, Li M (2010) Design, synthesis and preliminary activity assay of 1, 2, 3, 4-tetrahydroisoquinoline-3-carboxylic acid derivatives as novel Histone deacetylases (HDACs) inhibitors. Bioorg Med Chem 18:1761–1772. https://doi.org/10.1016/j.bmc.2010.01.060
Roy K, Banerjee A (2024) Tools, applications, and case studies (q-RA and q-RASAR). q-RASAR SpringerBriefs in molecular science. Springer, Cham
Kumar V, Banerjee A, Roy K (2023) Machine learning-based q-RASAR approach for the in silico identification of novel multi-target inhibitors against Alzheimer’s disease. Chemom Intell Lab Syst 245:105049. https://doi.org/10.1016/j.chemolab.2023.105049
Todeschini R, Consonni V (2008) Handbook of molecular descriptors. JWS, Cincinnati
Roy K, Kar S, Das RN (2015) Understanding the basics of QSAR for applications in pharmaceutical sciences and risk assessment. Elsevier, Amsterdam
Yap CW (2011) PaDEL-descriptor: an open source software to calculate molecular descriptors and fingerprints. J Comput Chem 32:1466–1474
Ambure P, Aher RB, Gajewicz A, Puzyn T, Roy K (2015) “NanoBRIDGES” software: open access tools to perform QSAR and nano-QSAR modeling. Chemometr Intell Lab Syst 147:1–13. https://doi.org/10.1016/j.chemolab.2015.07.007
QSAR Tools, DTC Laboratory, India (2023) Software available at http://teqip.jdvu.ac.in/QSAR_Tools/. Accessed 19 Dec 2023
Rogers D, Hopfinger AJ (1994) Application of genetic function approximation to quantitative structure-activity relationships and quantitative structure-property relationships. J Chem Inf Comput Sci 34:854–866. https://doi.org/10.1021/ci00020a020
Banerjee A, Roy K (2022) First report of q-RASAR modeling toward an approach of easy interpretability and efficient transferability. Mol Divers 26:2847–2862. https://doi.org/10.1007/s11030-022-10478-6
Chatterjee M, Banerjee A, De P, Gajewicz-Skretna A, Roy K (2022) A novel quantitative read-across tool designed purposefully to fill the existing gaps in nanosafety data. Environ Sci Nano 9:189–203. https://doi.org/10.1039/D1EN00725D
Banerjee A, Chatterjee M, De P, Roy K (2022) Quantitative predictions from chemical read-across and their confidence measures. Chemometr Intell Lab Syst 227:104613. https://doi.org/10.1016/j.chemolab.2022.104
Banerjee A, Kar S, Pore S, Roy K (2023) Efficient predictions of cytotoxicity of TiO2-based multi component nanoparticles using a machine learning-based q-RASAR approach. Nanotoxicology 17:78–93. https://doi.org/10.1080/17435390.2023.2186280
Clement O, Mehl AT (2003) Accelrys, San Diego, California, USA 1 background, Protein Struct: Determin Anal Appl Drug Discov. 378
Discovery Studio 3.0 (DS 3.0) (2015) Accelrys Inc., San Diego, USA.
Debnath AK (2003) Generation of predictive pharmacophore models for CCR5 antagonists: study with piperidine-and piperazine-based compounds as a new class of HIV-1 entry inhibitors. J Med Chem 46:4501–4515. https://doi.org/10.1021/jm030265z
Debnath AK (2002) Pharmacophore mapping of a series of 2, 4-diamino-5-deazapteridine inhibitors of Mycobacterium avium complex dihydrofolate reductase. J Med Chem 45:41–53. https://doi.org/10.1021/jm010360c
Santos-Martins D, Forli S, Ramos MJ, Olson AJ (2014) AutoDock4Zn: an improved AutoDock force field for small-molecule docking to zinc metalloproteins. J Chem Inf Model 54:2371–2379. https://doi.org/10.1021/ci500209e
Morris GM, Huey R, Lindstrom W, Sanner MF, Belew RK, Goodsell DS, Olson AJ (2009) AutoDock4 and AutoDockTools4: automated docking with selective receptor flexibility. J Comput Chem 30:2785–2791. https://doi.org/10.1002/jcc.21256
Abraham MJ, Murtola T, Schulz R, Pall S, Smith JC, Hess B, Lindahl E (2005) GROMACS: high performance molecular simulations through multi-level parallelism from laptops to supercomputers. SoftwareX 1:19–25. https://doi.org/10.1016/j.softx.2015.06.001
Van Der Spoel D, Lindahl E, Hess B, Groenhof G, Mark AE, Berendsen HJ (2005) GROMACS: fast, flexible, and free. J Comput Chem 26:1701–1718. https://doi.org/10.1002/jcc.20291
Jo S, Kim T, Iyer VG, Im W (2008) CHARMM-GUI: a web-based graphical user interface for CHARMM. J Comput Chem 29:1859–1865. https://doi.org/10.1002/jcc.20945
Rodrigues JP, Teixeira JM, Trellet M, Bonvin AM (2018) Pdb-tools: a swiss army knife for molecular structures. F1000Research 7:1961
O’Boyle NM, Banck M, James CA, Morley C, Vandermeersch T, Hutchison GR (2011) Open babel: an open chemical toolbox. J Cheminformatics 3:1–14. https://doi.org/10.1186/1758-2946-3-33
Kim S, Lee J, Jo S, Brooks CL, Lee HS, Im W (2017) CHARMM‐GUI ligand reader and modeler for CHARMM force field generation of small molecules. https://doi.org/10.1002/jcc.24829
Qi Y, Cheng X, Han W, Jo S, Schulten K, Im W (2014) CHARMM-GUI PACE CG builder for solution, micelle, and bilayer coarse-grained simulations. J Chem Inf Model 54:1003–1009. https://doi.org/10.1021/ci500007n
Nagaraj A, Srinivasa Raghavan S, Niraikulam A, Gautham N, Gunasekaran K, Sanggenol B (2023) A plant bioactive, as a safer alternative to tackle cancer by antagonising human FGFR. J Biomol Struct Dyn. https://doi.org/10.1080/07391102.2023.2245047
Sa-Ngiamsuntorn K, Suksatu A, Pewkliang Y, Thongsri P, Kanjanasirirat P, Manopwisedjaroen S, Charoensutthivarakul S, Wongtrakoongate P, Pitiporn S, Chaopreecha J (2021) Anti-SARS-CoV-2 activity of Andrographis paniculata extract and its major component andrographolide in human lung epithelial cells and cytotoxicity evaluation in major organ cell representatives. J Nat Prod 84:1261–1270. https://doi.org/10.1021/acs.jnatprod.0c01324
Nikolsky KS, Kulikova LI, Petrovskiy DV, Rudnev VR, Butkova TV, Malsagova KA, Kopylov AT, Kaysheva AL (2023) Three-helix bundle and SH3-type barrels: autonomously stable structural motifs in small and large proteins. J Biomol Struct Dyn. https://doi.org/10.1080/07391102.2023.2250450
Ashok AK, Gnanasekaran TS, Santosh Kumar HS, Srikanth K, Prakash N, Gollapalli P (2023) High-throughput screening and molecular dynamics simulations of natural products targeting LuxS/AI-2 system as a novel antibacterial strategy for antibiotic resistance in helicobacter pylori. J Biomol Struct Dyn. https://doi.org/10.1080/07391102.2023.2210674
Ilyas A, Poddar NK, Borkotoky S (2023) Insights into the dynamic interactions of RNase a and osmolytes through computational approaches. J Biomol Struct Dyn. https://doi.org/10.1080/07391102.2023.2229445
Moharana M, Pattanayak SK, Khan F (2023) Identification of phytochemicals from Eclipta alba and assess their potentiality against hepatitis C virus envelope glycoprotein: virtual screening, docking, and molecular dynamics simulation study. J Biomol Struct Dyn 41:5328–5344. https://doi.org/10.1080/07391102.2022.2085804
Singh G, Al-Fahad D, Al-Zrkani MK, Chaudhuri TK, Soni H, Tandon S, Narasimhaji CV, Azam F (2023) Patil R (2023) Identification of potential inhibitors of HER2 targeting breast cancer—a structure-based drug design approach. J Biomol Struct Dyn. https://doi.org/10.1080/07391102.2023.2246576
DeLano WL (2002) Pymol: an open-source molecular graphics tool, CCP4 newsl. Protein Crystallogr 40:82–92
De P, Kar S, Ambure P, Roy K (2022) Prediction reliability of QSAR models: an overview of various validation tools. Arch Toxicol 96:1279–1295. https://doi.org/10.1007/s00204-022-03252-y
Wu Z, Li D, Meng J, Wang H (2010) Introduction to SIMCA-P and its application, Handbook of partial least squares: concepts, methods and applications. 757–774.
Banerjee A, Roy K (2023) Prediction-inspired intelligent training for the development of classification read-across structure-activity relationship (c-RASAR) models for organic skin sensitizers: assessment of classification error rate from novel similarity coefficients. Chem Res Toxicol 36:1518–1531. https://doi.org/10.1021/acs.chemrestox.3c00155
Bertato L, Chirico N, Papa E (2023) QSAR models for the prediction of dietary biomagnification factor in fish. Toxics 11:209. https://doi.org/10.3390/toxics11030209
Choi E, Yoo WJ, Jang HY, Kim TY, Lee SK, Oh HB (2023) Machine learning liquid chromatography retention time prediction model augments the dansylation strategy for metabolite analysis of urine samples. J Chromatogr A 1705:464167. https://doi.org/10.1016/j.chroma.2023.464167
Cassani S, Kovarich S, Papa E, Roy PP, van der Wal L, Gramatica P (2013) Daphnia and fish toxicity of (benzo) triazoles: validated QSAR models, and interspecies quantitative activity–activity modelling. J Hazard Mater 258:50–60. https://doi.org/10.1016/j.jhazmat.2013.04.025
Adeniji SE, Shallangwa GA, Arthur DE, Abdullahi M, Mahmoud A, Haruna A (2020) Quantum modelling and molecular docking evaluation of some selected quinoline derivatives as anti-tubercular agents. Heliyon. https://doi.org/10.1016/j.heliyon.2020.e03639
Pandey SK, Roy K (2023) Development of a read-across-derived classification model for the predictions of mutagenicity data and its comparison with traditional QSAR models and expert systems. Toxicology. https://doi.org/10.1016/j.tox.2023.153676
Thangapandian S, John S, Sakkiah S, Lee K (2010) Docking-enabled pharmacophore model for histone deacetylase 8 inhibitors and its application in anti-cancer drug discovery. J Mol Graph Model 29:382–395. https://doi.org/10.1016/j.jmgm.2010.07.007
Thangapandian S, John S, Sakkiah S, Lee KW (2010) Ligand and structure-based pharmacophore modeling to facilitate novel histone deacetylase 8 inhibitor design. Eur J Med Chem 45:4409–4417. https://doi.org/10.1016/j.ejmech.2010.06.024
Thangapandian S, John S, Lee Y, Kim S, Lee KW (2011) Dynamic structure-based pharmacophore model development: a new and effective addition in the histone deacetylase 8 (HDAC8) inhibitor discovery. Int J Mol Sci 12:9440–9462. https://doi.org/10.3390/ijms12129440
Halder AK, Mallick S, Shikha D, Saha A, Saha KD, Jha T (2015) Design of dual MMP-2/HDAC-8 inhibitors by pharmacophore mapping, molecular docking, synthesis and biological activity. RSC Adv 5:72373–72386. https://doi.org/10.1039/C5RA12606A
Hou X, Du J, Liu R, Zhou Y, Li M, Xu W, Fang H (2015) Enhancing the sensitivity of pharmacophore-based virtual screening by incorporating customized ZBG features: a case study using histone deacetylase 8. J Chem Inf Model 55:861–871. https://doi.org/10.1021/ci500762z
Debnath S, Debnath T, Majumdar S, Arunasree M, Aparna V (2016) A combined pharmacophore modeling, 3D QSAR, virtual screening, molecular docking, and ADME studies to identify potential HDAC8 inhibitors. Med Chem Res 25:2434–2450. https://doi.org/10.1007/s00044-016-1652-5
Kim S, Lee Y, Kim S, Lee SJ, Heo PK, Kim S, Kwon YJ, Lee KW (2018) Identification of novel human HDAC8 inhibitors by pharmacophore-based virtual screening and density functional theory approaches. Bull Korean Chem Soc 39:197–206. https://doi.org/10.1002/bkcs.11366
Kashyap K, Kakkar R (2019) An insight into selective and potent inhibition of histone deacetylase 8 through induced-fit docking, pharmacophore modeling and QSAR studies. J Biomol Struct Dyn. https://doi.org/10.1080/07391102.2019.1567388
Debnath S, Debnath T, Bhaumik S, Majumdar S, Kalle AM, Aparna V (2019) Discovery of novel potential selective HDAC8 inhibitors by combine ligand-based, structure-based virtual screening and in-vitro biological evaluation. Sci Rep 9:17174. https://doi.org/10.1038/s41598-019-53376-y
Vema A, Debnath S, Kalle AM (2022) Identification of novel HDAC8 selective inhibitors through ligand and structure-based studies: exploiting the acetate release channel differences among class I isoforms. Arab J Chem 15:103863. https://doi.org/10.1016/j.arabjc.2022.103863
Karplus M, McCammon JA (2002) Molecular dynamics simulations of biomolecules. Nat Struct Biol 9:646–652. https://doi.org/10.1038/nsb0902-646
Acknowledgements
SK gratefully acknowledges Council of Scientific & Industrial Research, Human Resource Development Group (CSIR-HRDG), New Delhi, India for awarding the Senior Research Fellowship (Appl. No.-111-2149-5423/2K23/1). SG is thankful to SERB, India for funding under MATRICS Scheme [Ref. No.: MTR/2022/000286]. The authors thank National Institute of Pharmaceutical Education and Research (NIPER), Kolkata, Department of Pharmaceutical Technology, Jadavpur University, Kolkata and Department of Pharmaceutical Technology, JIS University, Agarpara, Kolkata, India for providing research facilities.
Author information
Authors and Affiliations
Contributions
Conceptualization: [Shovanlal Gayen], [Sk. Abdul Amin]; Methodology: [Samima Khatun], [Indrasis Dasgupta], [Rakibul Islam]; Formal analysis and investigation: [Samima Khatun], [Indrasis Dasgupta], [Rakibul Islam], [Sk. Abdul Amin]; Writing—original draft preparation: [Samima Khatun], [Indrasis Dasgupta], [Rakibul Islam]; Writing—review and editing: [Shovanlal Gayen], [Devendra Kumar Dhaked], [Sk. Abdul Amin]; Funding acquisition: [Shovanlal Gayen]; Resources: [Tarun Jha], [Devendra Kumar Dhaked], [Shovanlal Gayen]; and Supervision: [Devendra Kumar Dhaked], [Tarun Jha], [Shovanlal Gayen].
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no known competing financial interests or personal relationships that could have appeared to influence the work reported in this paper.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Khatun, S., Dasgupta, I., Islam, R. et al. Unveiling critical structural features for effective HDAC8 inhibition: a comprehensive study using quantitative read-across structure–activity relationship (q-RASAR) and pharmacophore modeling. Mol Divers (2024). https://doi.org/10.1007/s11030-024-10903-y
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s11030-024-10903-y