Abstract
In a majority of environments, microbes live as interacting communities. Microbial communities are composed of a mix of microbes with often unknown functions. Polymicrobial diseases represent the clinical and pathological manifestations induced by the presence of multiple infectious agents. These diseases are difficult to diagnose and treat and usually are more severe than monomicrobial infections. The interaction relationship between Enterococcus faecalis and Escherichia coli was researched using a Calvet calorimeter. Three mixtures of both bacteria were prepared in the following proportions: 20 + 80 % (0.2 mL E. faecalis + 0.8 mL E. coli), 50 + 50 % (0.5 mL E. faecalis + 0.5 mL E. coli) and 80 + 20 % (0.8 mL E. faecalis + 0.2 mL E. coli). Experiments were carried out at concentration of 106 CFU mL−1 and a constant temperature of 309.65 K. The differences in shape of graph of E. faecalis, E. coli and their mixtures were compared. Also, the thermokinetic parameters such as detection time (t d), growth constant (k), generation time (G) and the amount of heat released (Q) were calculated.
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
Introduction
Microorganisms are rarely found isolated in nature but rather exist as a part of a community [1]. Microbial communities can live almost anywhere and contain different types of organisms (bacteria, fungi, viruses and parasites). Abundance and diversity of microbes are controlled by the resources (food) and conditions (temperature, pH, oxygen concentration, etc.) that exist in the environment [2]. Microbial interactions often determine the state of the habitat in which they live. When these habitats are located on the human body, these interactions can affect health and cause disease [3].
Polymicrobial diseases represent the clinical and pathologic manifestations induced by the presence of multiple microorganisms. These are serious diseases whose aetiologic agents are sometimes difficult to diagnose and treat. They are often called complex infections, complicated infections, dual infections, mixed infections, secondary infections, co-infections, synergistic infections, concurrent infections or polymicrobial infections. The multiple aetiologies often induce a characteristic set of clinical signs and lesions referred to as “complexes” or syndromes [4]. The most common types of these infections include oral cavity (odontogenic infections), abdominal abscess or secondary peritonitis, solid organ abscesses (brain, liver, lungs), chronic infections (sinusitis, mastoiditis, otitis media), skin and soft tissue infections (diabetic foot wound, cellulitis, necrotic fasciitis) and bacteraemia [5–7].
Polymicrobial infections are being recognised with increasing frequency. In these infections, the presence of one microorganism generates a niche for other pathogenic microorganisms to colonise, one microorganism predisposes the host to colonisation by other microorganisms, or two or more non-pathogenic microorganisms together cause disease. Microbial interference can occur between potential pathogens or between probiotic organisms and pathogens [6]. These infections are often worse than monomicrobial infections, because it can modify the clinical course of the disease and increased its severity, affecting the selection of antimicrobial therapy, especially when it comes to pathogens that commonly exhibit antimicrobial resistance [3].
Polymicrobial diseases are difficult to reproduce and to study. In vivo and in vitro models are often used to study specific microbial interactions or parameters associated with infection and disease [4]. In the last few years, there have been several studies that try to understand the mechanism of these interactions through the development of in vivo models and biochemical or molecular techniques to study microbial communities [8–13].
Growth is an essential response to the physico-chemical environment by microorganisms. They can grow under a variety of physical, chemical and nutritional conditions [14].
Metabolism is a complex process involving numerous anabolic (synthesis of cell constituents and metabolites) and catabolic (breakdown of cell constituents and metabolites) reactions [1]. These reactions involve an increase in the size of organisms and an increase in their number. However, whatever the balance between these two process, the net effect is an increase in the total mass (biomass) on the culture medium [15].
Microcalorimetry is a technique that is based on the measurement of heat flow as a result of bacterial metabolism [16], which can be applied to detect bacterial growth in a few hours, even from samples with a low concentration, e.g. 10 CFU mL−1 [17, 18].
However, most of the published reports focused on the study of a single microorganism [17–23]; few studies tried to research the relationship between two microorganisms using the microcalorimetry [24–28].
In this study, we have used the microcalorimetry as a tool to investigate how two different bacteria interact. We believe that this technique could complement other methods.
Enterococcus faecalis is a Gram-positive cocci, typically arranged in pairs and shorts chains. This organism is found in the large intestine in high concentrations and in the genitourinary tract. It is an important pathogen in hospitalised patients; indeed, it is one of the common causes of infections acquired in the hospital (nosocomial infection). E. faecalis produce diseases as urinary tract infections, peritonitis, wound infections and bacteremia with or without endocarditis [15, 29].
Escherichia coli is a facultative anaerobic, motile, Gram-negative rod that exists singly or in pairs. This bacterium is the most inhabitant common in the gastrointestinal tract. E. coli possesses a broad range of virulence factors. It is associated with a variety of diseases, including gastroenteritis and extraintestinal diseases, such as bacteremia, urinary tract infections, neonatal meningitis and intraabdominal infections [15, 29].
Materials and methods
Bacterial strains and sample preparation
The bacteria were supplied by the American Type Culture Collection (ATCC): E. faecalis (ATCC 29212) and E. coli (ATCC 25922). These bacterial strains were streaked onto blood agar plates and incubated at 309.65 K for 24 h. The blood agar plates with multiple bacterial colonies were then used to prepare a bacterial suspension with 0.9 % saline solution, whose concentration was adjusted to the corresponding 0.5 on the McFarland scale, using optical densitometer. This solution was then used to prepare further dilutions with the sterile saline to obtain final concentrations of 106 CFU mL−1. The inoculum concentration was confirmed using the plate count method. Later, mixtures of both bacteria were prepared in the following proportions: 20 + 80 % (0.2 mL E. faecalis + 0.8 mL E. coli), 50 + 50 % (0.5 mL E. faecalis + 0.5 mL E. coli) and 80 + 20 % (0.8 mL E. faecalis + 0.2 mL E. coli).
Experimental equipment and data collection
The microcalorimetric bacterial growth curves were obtained using Calvet microcalorimeter equipment. This device allows operation in the absence of vapour phase and is equipped with two screw-capped stainless steel cells of approximately 10 cm3. One of these cells contains the reference solution, named the reference cell, and the other one the sample, named the experimental cell. A Philips PM2535 multimeter and a data acquisition system were linked to the microcalorimeter. Calibration was performed electrically using a Setaram EJP30 stabilized current source. The precision in calorimetric signal was ±1 µV [30].
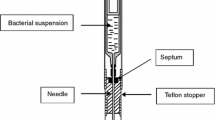
The external media of the calorimeter were maintained at a constant temperature of 309.65 K. The reference cell was filled with 7 mL of culture medium + 1 mL of sterile saline, while the experimental cell was injected with 7 mL of culture medium. The culture medium used was a liquid enriched with soybean–casein [28]. Both cells were introduced, from the upper part of the calorimeter in the internal thermopile chamber through two cylindrical holes aligned in parallel. The large distance that separates the cells from the entrance ensures the minimisation of heat flow to the exterior. The system is then allowed to stabilise during 2 h, after which 1 mL of the sample is introduced into the experimental cell.
The experiment was also carried out with a sample not containing any bacteria (control). All the experiments were realised in triplicate.
The reference cell and the experimental cell were cleaned and sterilised by autoclaving before using (20 min at 394.15 K).
Data were collected using the data collection and processing system, at intervals of 20 s, throughout the duration of the experiment.
Results
The difference in voltage generated between the experimental and reference cells over time allowed us to obtain the exothermic growth curves of E. faecalis, E. coli and their mixtures at 106 CFU mL−1.
The graph of E. faecalis (Fig. 1) presented a unique energetic phase in which stand out four peaks of electromotive force, and the signal is recorded for about 6 h, returning to baseline levels after this time. The curve of E. coli (Fig. 2) had two metabolic phases, the first with greater energy, but shorter duration, followed by a period of latency that preceded a second period of lower energy that is prolonged over time.
When we compare the curves of E. faecalis, E. coli and their mixtures (Fig. 3a, b, c), two metabolic phases are observed: a first peak with higher energy and shorter duration and a second peak of lower energy and longer duration that becomes less intense as decreasing the amount of E. coli in the sample. In graphs 80 % E. faecalis + 20 % E. coli (Fig. 3a) and 50 % E. coli + 50 % E. faecalis (Fig. 3b), a small peak adjacent to the peak of greater intensity than the recording time can be attributed to E. faecalis. However, in curve of 20 % E. faecalis + 80 % E. coli (Fig. 3c) only the growth profile of E. coli is observed.
We determine the value of the maximum voltage (V max) and the time of its registration (t max) in the curves of E. faecalis, E. coli and their mixtures in different proportions. Although we cannot establish a quantitative proportionality relationship of the maximum peaks and their appearance time between pure cultures and their mixtures, all experiments show a maximum peak before 5 h (Table 1).
The detection time of the signal (t d) of E. faecalis, E. coli and their mixtures is presented in Table 1. As can be seen, the sample 20 % E. faecalis + 80 % E. coli has a t d shorter than 80 % E. faecalis + 20 % E. coli, because the t d of E. coli is approximately half that of E. faecalis. In all curves, growth can be detected in less than 4 h.
In general, microorganisms grow and divide up to a maximum during the logarithmic stage. In this phase of the growth curve, we fitted the data to an exponential equation that can be expressed as [31, 32]:
where n 0 is the number of bacteria at time 0, n t is the number of bacteria at time t, and k is the growth constant.
If P w is the thermal power of each cell, then:
Taking into account that P 0 and P t are the energies released at onset and at time t, respectively:
Thus, the growth constant (k) can be obtained by selecting two points from the exponential phase of a bacterial growth curve.
The generation time (G) is defined as the time taken by a population to duplicate its number and is expressed as:
Table 2 summarises the values of k and G of single microorganisms and their mixtures. As can been observed, E. faecalis has a lower k and greater G than E. coli. This would explain that the samples 50 % E. faecalis + 50 % E. coli and 80 % E. faecalis + 20 % E. coli have a lower k and greater G than the sample 20 % E. faecalis + 80 % E. coli. In all cases, the fitting equation in the exponential phase presented correlation coefficients (R 2) higher than 0.9865, indicating a good linear relationship.
From the curve, we can also calculate the amount of heat released (Q) over the culture time:
where A (µV h) is the area, calculated by the trapezoidal method, and K represents a constant whose value is 23.8 J µV−1 h−1, which was calculated from the electric calibration performed by the Joule effect on the equipment.
Also, we have compared the amount of heat exchanged by E. faecalis, E. coli and their mixtures during the first 24 h (Q 24). In Table 3, the Q 24 by E. faecalis is approximately three times less than E. coli, because the metabolic activity of E. faecalis is shorter than E. coli. However, the mixture 50 % E. faecalis + 50 % E. coli had a higher Q 24 than expected, due to the interaction between both bacteria.
Discussion
In this study, we have investigated the interaction relationship between two pathogenic bacteria, E. faecalis and E. coli, which are important microorganisms responsible for infectious diseases. Microcalorimetry proved to be an excellent tool for the detection and study of polymicrobial bacterial infections. Also, we were allowed us obtain the voltage–time curves of single microorganisms and their mixtures in three different proportions. From growth curves, we could calculate the thermokinetic parameters to assess the interaction between E. faecalis and E. coli. When we mixed both bacteria, we could observe that in the curves 80 % E. faecalis + 20 % E. coli (Fig. 3a) and 50 % E. faecalis + 50 % E. coli (Fig. 3b), presented a first peak of high intensity characteristic of growth curve of E. coli (Fig. 2). Next to this, second one appears less intense than its recording time can be attributed to E. faecalis. Finally, the graph 20 % E. faecalis + 80 % E. coli (Fig. 3c) is very similar to the E. coli (Fig. 2), because of its higher percentage in the sample; therefore, the growth of E. faecalis was not observable.
In relation to this work, our group had previously published different studies about the mechanism of interaction between E. faecalis and Klebsiella pneumoniae [25] and E. faecalis and Pseudomonas aeruginosa [28].
In case of E. faecalis and K. pneumoniae, the results revealed that when E. faecalis and K. pneumoniae were mixed, in the graphs of 20 % E. faecalis + 80 % K. pneumoniae and 50 % E. faecalis + 50 % K. pneumoniae, the characteristic growth of E. faecalis was not present, due to K. pneumoniae being completely inhibited its growth. In the sample 80 % E. faecalis + 20 % K. pneumoniae, we could observe in the first part of the curve the characteristic growth of E. faecalis, while the second phase was similar to the curve of K. pneumoniae. This is because 80 % of the sample was Enterococci, predominantly its growth profile [25].
However, in the study of E. faecalis and P. aeruginosa, when both pathogens were put together, in the curves 20 % E. faecalis + 80 % P. aeruginosa and 50 % E. faecalis + 50 % P. aeruginosa, we could identify the characteristic growth profile of P. aeruginosa during the duration of the experiment. However, in both cases, even at low rates of E. faecalis in the sample, the growth profile of P. aeruginosa is modified. Perhaps this is because E. faecalis can use nutrients more effectively or its growth is not completely inhibited by Pseudomonas. In the case of sample of 80 % E. faecalis + 20 % P. aeruginosa, the growth profiles of both bacteria can be identified clearly. In the first stretch, we could observe the growth curve of E. faecalis, with three peaks of voltage, which signal was recorded during the first 10 h of the experiment, and the second stretch where an increase in potential is maintained for the duration of the experiment is characteristic of Pseudomonas [28].
Conclusions
Microcalorimetry has allowed us to study bacterial growth of mixtures of E. faecalis and other bacteria (E. coli, K. pneumoniae and P. aeruginosa) from the heat generated by its metabolism. Also, this technique allows us to obtain in a few hours the voltage–time curves a real time and thermokinetic parameters such as the growth constant (k), generation time (G), detection time (t d) and the amount of heat released (Q).
Microcalorimetric investigations of interaction relationships between two bacteria are interesting, because it could complement other methods to research how bacteria behave in microbial communities and establish antibiotic treatment strategies in a more rapid and effective way.
References
Pepper IL, Gerba CP, Gentry TJ. Environmental microbiology. 3rd ed. Amsterdam: Academic Press; 2015.
Ausina RV, Moreno GS. Tratado SEIMC de Enfermedades Infecciosas y Microbiología Clínica. 1st ed. Madrid: Médica Panamericana; 2006.
Murray JL, Connell JL, Stacy A, Turner KH, Whiteley M. Mechanisms of synergy in polymicrobial infections. J Microbiol. 2014;52(3):188–99.
Brogden KA, Guthmiller JM. Polymicrobial diseases. Washington (DC): ASM Press; 2002.
Pavlović M, Nikolić J. Antimicrobial treatment of polymicrobial infections. Med Pregl. 2010;63(Suppl 1):37–46.
Brogden KA, Guthmiller JM, Taylor CE. Human polymicrobial infections. Lancet. 2005;365(9455):253–5.
Peters BM, Jabra-Rizk MA, O’May GA, Costerton JW, Shirtliff ME. Polymicrobial interactions: impact on pathogenesis and human disease. Clin Microbiol. 2012;25(1):193–213.
Dalton T, Dowd SE, Wolcott RD, Sun Y, Watters C, Griswold JA, Rumbaugh KP. An in vivo polymicrobial biofilm wound infection model to study interspecies interactions. PLoS One. 2011;6(11):e27317.
Mastropaolo MD, Evans NP, Byrnes MK, Stevens AM, Robertson JL, Melville SB. Synergy in polymicrobial infections in a mouse model of type 2 diabetes. Infect Immun. 2005;73(9):6055–63.
Bakaletz LO. Developing animal models for polymicrobial diseases. Nat Rev Microbiol. 2004;2(7):552–68.
Watrous J, Roach P, Alexandrov T, Heath BS, Yang JY, Kersten RD, et al. Mass spectral molecular networking of living microbial colonies. Proc Natl Acad Sci USA. 2012;109(26):E1743–52.
Yang JY, Phelan VV, Simkovsky R, Watrous JD, Trial RM, Fleming TC, et al. Primer on agar-based microbial imaging mass spectrometry. J Bacteriol. 2012;194(22):6023–8.
Wessel AK, Hmelo L, Parsek MR, Whiteley M. Going local: technologies for exploring bacterial microenvironments. Nat Rev Microbiol. 2013;11(5):337–48.
Prescott LM, Harley JP, Klein DA. Microbiology. 5th ed. New York: Mc Graw Hill; 2004.
Greenwood D, Slack R, Peutherer J, Barer M. Medical microbiology: a guide to microbial infections: pathogenesis, immunity, laboratory diagnosis, and control. 17th ed. Edinburgh: Churchill Livingstone/Elsevier; 2007.
Braissant O, Wirz D, Göpfert B, Daniels AU. Use of isothermal microcalorimetry to monitor microbial activities. FEMS Microbiol Lett. 2010;303:1–8.
Lago N, Legido JL, Paz-Andrade MI, Arias I, Casás LM. Microcalorimetric study of the growth and metabolism of Pseudomonas aeruginosa. J Therm Anal Calorim. 2011;105:651–5.
Tan MR, Ren YS, Yan D, Meng XH, Cheng LH, Ll Qiu, et al. Detection of microorganisms in different states based on microcalorimetry. J Therm Anal Calorim. 2012;109(2):1069–75.
Zaharia DC, Iancu C, Steriade AT, Muntean AA, Balint O, Popa VT, et al. MicroDSC study of Staphylococcus epidermidis growth. BMC Microbiol. 2010;10:322.
Rivero NL, Soto JLL, Casás LM, Santos IA. Microcalorimetric study of the growth of Enterococcus faecalis in an enriched culture medium. J Therm Anal Calorim. 2012;108(2):665–70.
Rivero NL, Legido JL, Santos IA, Casás LM. Comparative study of microcalorimetric behavior of Escherichia coli, Proteus mirabilis and Klebsiella pneumoniae. Pol J Microbiol. 2012;61(3):199–204.
Bonkat G, Bachman A, Solokhina A, Widmer AF, Frei R, Gasser T, et al. Growth of mycobacteria in urine determined by isothermal microcalorimetry: implications for urogenital tuberculosis and other mycobacterial infections. Urology. 2012;80(5):1163e9–12.
Bonkat G, Braissant O, Malte R, Solokhina A, Widmer AF, Frei R, et al. Standardization of isothermal microcalorimetry in urinary tract detection by using artificial urine. World J Urol. 2013;31(3):553–7.
Kong W-J, Xing X-Y, Xiao XH, Zhao Y-L, Wei J-H, Wang JB, et al. Effect of berberine on Escherichia coli, Bacillus subtilis, and their mixtures as determined by isothermal microcalorimetry. Appl Microbiol Biotechnol. 2012;96:503–10.
Vázquez C, Lago N, Legido JL, Arias I, Casás LM, Mato MM. Microcalorimetric study of the growth of Enterococcus faecalis, Klebsiella pneumoniae and their mixtures in an enriched culture medium. J Therm Anal Calorim. 2013;113(3):1415–20.
Vázquez C, Lago N, Mato MM, Casás LM, Esarte L, Legido JL, et al. Microcalorimetric performance of the growth in culture of Escherichia coli, Proteus mirabilis and their mixtures in different proportions. J Therm Anal Calorim. 2014;116(1):107–12.
Wang F, Yao J, Yu C, Chen H, Yi Z. Investigating Pseudomonas putida-Candida humicola interactions as affected by chelate fe(III) in soil. Bull Environ Contam Toxicol. 2014;92(3):358–63.
Vázquez C, Lago N, Mato MM, Legido JL, Esarte L. Microcalorimetric study of the growth of Enterococcus faecalis, Pseudomonas aeruginosa and their mixtures in an enriched culture medium. J Therm Anal Calorim. 2015;121(1):463–8.
Murray PR, Rosenthal KS, Pfaller MA. Medical microbiology. 7th ed. Philadelphia: Elsevier Saunders; 2013.
Verdes PV, Mato MM, Paz Andrade MI, Legido JL. Contribution to study of the thermodynamics properties of mixtures containing 2-methoxy-2-methylpropane, alkanol, alkane. J Chem Therm. 2014;73:224–31.
Ma J, Qi WT, Yang LN, Yu WT, Xie YB, Wang W, et al. Microcalorimetric study on the growth and metabolism of microencapsulated microbial cell culture. J Microbiol Methods. 2007;68:172–7.
Braissant O, Bonkat G, Wirz D, Bachmann A. Microbial growth and isothermal microcalorimetry: growth models and their application to microcalorimetric data. Thermochim Acta. 2013;555:64–71.
Acknowledgements
We thank María Perfecta Salgado Gonzalez and Sofia Baz Rodríguez for their collaboration with the technical measures. We are also thankful for the financial support provided by the project EM 2012/141 by “Xunta de Galicia” and the project FIS 2011-23322 funded by Ministry of Science and Innovation of Spain. All these projects are co-financed with FEDER funds.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Vazquez, C., Lago, N., Mato, M.M. et al. Study of the growth of Enterococcus faecalis, Escherichia coli and their mixtures by microcalorimetry. J Therm Anal Calorim 125, 739–744 (2016). https://doi.org/10.1007/s10973-015-5203-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10973-015-5203-y