Abstract
Purpose
Immunodeficiency, centromeric instability, and facial anomalies (ICF) syndrome is an extremely rare autosomal recessive disease. The immune phenotype is characterized by hypogammaglobulinemia in the presence of B cells. T cell lymphopenia also develops in some patients. We sought to further investigate the immune defect in an ICF patient with a novel missense mutation in DNMT3B and a severe phenotype.
Methods
Patient lymphocytes were examined for subset counts, immunoglobulin levels, T and B cell de novo production (via excision circles) and receptor repertoire diversity. Mutated DNMT3B protein structure was modeled to assess the effect of a mutation located outside of the catalytic region on protein function.
Results
A novel homozygous missense mutation, Ala585Thr, was found in DNMT3B. The patient had decreased B cell counts with hypogammaglobulinemia, and normal T cell counts. CD4+ T cells decreased over time, leading to an inversion of the CD4+ to CD8+ ratio. Excision circle copy numbers were normal, signifying normal de novo lymphocyte production, but the ratio between naïve and total B cells was low, indicating decreased in vivo B cell replication. T and B cell receptor repertoires displayed normal diversity. Computerized modeling of the mutated Ala585 residue suggested reduced thermostability, possibly affecting the enzyme kinetics.
Conclusions
Our results highlight the existence of a T cell defect that develops over time in ICF patient, in addition to the known B cell dysfunction. With intravenous immunoglobulin (IVIG) treatment ameliorating the B cell defect, the extent of CD4+ lymphopenia may determine the severity of ICF immunodeficiency.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
The immunodeficiency, centromeric instability, and facial anomalies (ICF) syndrome is an extremely rare autosomal recessive disease [1], with only several dozen patients diagnosed worldwide to date [2]. The majority of ICF patients display mutations in one of two genes, the DNA methyltransferase 3B gene (DNMT3B), termed ICF-1, and the Zinc finger and BTB (bric-a-bric, tramtrack, broad complex)-domain-containing 24 (ZBTB24) gene [3, 4], termed ICF-2. Additional complicit genes were recently discovered [5], and the genetic etiology in some patients remains unknown (termed ICF-X). Mutations in DNMT3B, an enzyme responsible for de novo methylation, result in reduced methylation across the genome, primarily in transcriptionally inactive regions [6], such as the centromere-adjacent heterochromatic regions of chromosomes 1, 16, and 9. Hypomethylation leads to increased DNA rearrangements, which in turn lead to breaks, whole-arm deletions, and branching of this region, the cytogenetic hallmark of ICF [7, 8].
The facial anomalies seen in ICF patients are typically mild and often consist of hypertelorism, flat nasal bridge, epicanthal folds, and low set ears [9]. Congenital defects such as cleft palate and syndactyly are present in a minority of patients, however, with increased prevalence compared with the general population. Growth and developmental delay and mild to moderate mental retardation have also been reported.
Immunodeficiency, characterized primarily by low or absent immunoglobulins in the presence of B cells, is almost universal in ICF patients, and recurrent infection is often the presenting symptom [2, 9]. T cell counts and function, normal at birth, decline in some patients over time [10]. Despite being the leading cause of death in ICF patients, the extent and the underlying mechanisms of this immunodeficiency remain poorly understood.
Here, we present a female patient with ICF syndrome caused by a novel mutation in DNMT3B. Using advanced molecular and genomic assays, we attempt to better characterize the immunologic phenotype of ICF-1 within the context of the current literature. In addition, we highlight new insights into DNMT3B structure-function relationship. As it has in other extremely rare diseases, it is our hope that incremental advances in the understanding of ICF pathogenesis will add up to a quicker diagnosis and better, more informed management.
Methods
Clinical Data
Patient record was obtained from the e-record registry of our hospital. The patient was examined by the authors. Informed consent was obtained, and all procedures performed were in accordance with the ethical standards of the institutional and/or national research committee and with the 1964 Helsinki declaration and its later amendments or comparable ethical standards.
Lymphocyte Subset Determination
Cell surface markers of peripheral blood mononuclear cells (PBMCs) were determined by immunofluorescent staining and flow cytometry (Epics V; Coulter Electronics, Hialeah, FL) using anti-CD3, anti-CD19, anti-CD16, and anti-CD56 from BD Biosciences and anti-CD4 and anti-CD8 from Beckman Coulter Immunotech.
Quantification of sjTRECs, sjKRECs, and iRSS-Kde Rearrangements
Signal joint excision circles are by-products of V(D)J recombination at the TCR α/β locus (sjTREC) or IGH locus (sjKREC) and are present only in recent emigrants from the thymus and bone marrow, respectively. Their enumeration allows the quantification of naïve T and B cell production. An intronic rearrangement termed iRSS-Kde is present in all mature B cells. Thus, B cell replication history can be deduced by dividing iRSS-Kde (all B cells) by sjKREC (newly formed B cells). Signal Joint T cell Receptor Excision Circles (sjTREC) copy numbers were determined using quantitative real-time PCR (RQ-PCR). PCR reactions were performed as previously described [11] using as template 0.5 μg genomic DNA (gDNA) extracted from the patients’ PBMCs. RQ-PCR was carried out using an ABI PRISM 7900 Sequence Detector System (Applied Biosystems). A standard curve was constructed by using serial dilutions containing 103 to 106 copies of a known sjTREC plasmid. Patient and control samples were tested in triplicate, and the number of sjTRECs in a given sample was calculated by comparing the obtained cycle threshold (Ct) value of the sample to the standard curve using an absolute quantification algorithm. The amount of sjKRECs and intron Recombination Signal Sequence Kappa deleting element (iRSS-Kde) rearrangements was determined by RQ-PCR as previously described [12]. The threshold for Ct determination was positioned at the same level each time. Amplification of RNAseP (TaqMan assay, Applied Biosystems) served as a quality control of DNA amplification for both the sjTREC and the sjKREC/iRSS-Kde assays.
Cell Surface Analysis of T Cell Receptor Vβ Expression
Surface expression of individual TCR-Vβ families was analyzed using flow cytometry and a set of 24 Vβ-specific fluorochrome-labeled monoclonal antibodies (Beckman Coulter) as described [11]. Normal control values were obtained from the IOTest Beta Mark-Quick Reference Card (Beckman Coulter).
B Cell Receptor Repertoire
Complementarity determining region 3 (CDR3) length distribution for a cluster of B cell receptor heavy chain (IGH) V and J gene segment pairings was assessed by means of PCR amplification according to the standardized BIOMED-2 protocol [13]. Fluorescence-labeled PCR products (1 μL) were added to a mixture of 8.5 μL of deionized formamide and 0.5 μL of GeneScan 500 Rox internal lane standard (Applied Biosystems) and separated with the 3500 Genetic Analyzer (Applied Biosystems). Results were analyzed with Gene Mapper software (Applied Biosystems).
Exome Sequencing Analysis
The BWA mem algorithm (version 0.7.12) [14] was used for alignment of the sequence reads to the human reference genome (hg19). The HaplotypeCaller algorithm of GATK version 3.4 was applied for variant calling, as recommended in the best practice pipeline [15]. KGG-seq v.08 [16] was used for annotation of detected variants, and in house scripts were applied for further filtering.
Structural Bioinformatics
The DNMT3B protein model was obtained using ModWeb [17] server which applies the modeler homology modeling program. The DNMT3B protein structure served as a template in this procedure.
SCCOMP [18] was used for side chain modeling over a fixed backbone. This program uses contact surface areas as the core of its scoring function. Structural alignment was done using the TriangleMatch structural alignment program [19]. Jmol (www.jmol.org) was used for 3D visualization and graphics.
Thermostability predictions, based on the structural data, were performed using Duet [20], MCSM [21], SDM [22], Maestro [23], and i-mutant2 [24].
Results
Clinical Description
The proband is the first born child of consanguineous parents (first cousins, Fig. 1a) of Palestinian descent. She was born small for gestational age (2 kg) at term following an uncomplicated, but minimally monitored, pregnancy. She was noted to have mild facial dysmorphism (Fig. 1b), with no limb malformations. She encountered feeding difficulties immediately after birth, and over the course of the first year of her life, she developed severe gastroesophageal reflux disease and intractable diarrhea. Repeated stool exams were negative for pathogenic bacteria and parasites, as well as Clostridium Difficile toxin. Proteinuria, present on several urine analyses, prompted sonographic examination of the kidneys which demonstrated a bilaterally echogenic cortex with mild left pelvis dilatation (data not shown), suggestive of parenchymatic renal disease. The patient suffered from severe, intractable failure to thrive (4 kg at 2 years old), and after multiple dietary adjustments yielded minimal weight gain, was placed on total parenteral nutrition (TPN). She continued to show minimal weight gain, and at times even weight loss, on TPN.
Genetics and clinical phenotype of DNMT3B mutated patient. a Family pedigree of the affected family. Slashed solid symbol indicates the deceased affected patient P1, and open symbol unaffected sister S1. Square indicates male subject (Father), and circles female subjects. b Patient mild facial dysmorphism: Hypertelorism, epicanthal folds, low and posteriorly rotated ears, and micrognathia
The patient suffered from recurrent respiratory infections and was hospitalized several times for pneumonia, treated with antibiotics. Hypogammaglobulinemia of all gamma globulin isotypes was detected, and she began treatment with monthly IVIG infusions. An extensive immunological workup was performed, the results of which are detailed below.
At the age of one and a half years old, the patient developed sepsis with septic shock. Blood cultures revealed polymicrobia, including Klebsiella pneumoniae, Escherichia coli, and Enterobacter. She was treated with antibiotics with resolution of the infection and normalization of blood pressures. Additional, possibly iatrogenic infections, with Staphylococcus epidermidis and Candida parapsilosis, were successfully treated with antibiotics and antifungals. However, following the septic episode, she began exhibiting a neurologic decline and hand tremor at rest. Brain MRI showed expansion of cisterns and ventricles with signs of atrophy (data not shown), as well as multiple hypointensity foci in the left cerebellum and left frontal lobe. EEG demonstrated encephalopathy without epileptic activity (data not shown). Several months following her release from our hospital, she had passed away at home.
Immunological Assessment
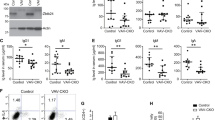
Phenotypic analysis of the patient’s lymphocytes revealed decreased B cell counts with normal to mildly elevated T cell subsets and normal counts of NK cells (Table 1). Serum immunoglobulins were consistently absent or, when taken for trough values during IVIG treatment, extremely decreased (Table 2). Serum albumin levels were consistently normal, indicating the hypogammaglobulinemia was unrelated to gastrointestinal loss from diarrhea or renal loss from the patient’s kidney disease. CD4 to CD8 ratio, normally between 0.9 and 1.9 and known to be reversed on occasion in ICF patients, was normal at 1.7 years of age (1.18), but reversed at 2.4 years of age (0.46) (Table 1). The patient’s de novo lymphopoiesis capability, as measured by sjTREC and sjKREC copy numbers, was normal (Table 3). iRSS-Kde/sjKREC in the patient reported here suggested a limited B cell replication history, despite a very significant infectious history (Table 3). In order to assess diversity of the T cell receptor repertoire, surface membrane expression of 24 different TCR Vβ (TRBV) families was analyzed by flow cytometry compared to healthy controls. All 24 Vβ families were expressed on patient’s T cells, with four Vβ families mildly under-expressed compared with healthy controls (Fig. 2a). Overall, the patient exhibited a normal T cell receptor repertoire. B cell receptor diversity was characterized by measuring the CDR3 length distribution for a cluster of IGH V and J gene segment pairings. The patient’s CDR3 length distribution exhibited a normal, Gaussian-like distribution, similar to that of a healthy control (Fig. 2b).
T and B cell receptor repertoire analyses. a Flow cytometry analysis of surface membrane expression of 24 T cell receptor β variable segments in our ICF-1 patient compared to average expression on healthy controls. Error bars indicate standard deviation (SD) across controls. b Polyclonal (gaussian) distribution of CDR3 length at the immunoglobulin heavy chain (IGH) locus across patient and control B cell populations
Genetic Evaluation
In order to identify the genetic cause of the clinical manifestations of our patient, whole exome sequencing (WES) was performed. Due to parental consanguinity, autosomal recessive analysis was performed and yielded 4293 homozygous variants which affect protein sequences. This list of variants was subsequently reduced to 105 (Table S1, Supplementary Appendix) rare variants, by filtering out variants present in ≥0.01 of our in-house exomes (n = 250) and variants present with a minor allele frequency (MAF) ≥ 0.01 in either the 1000 Genomes Project (1KG; http://browser.1000genomes.org/index.html) or dbSNP 135 database or the NHLBI Exome Sequencing Project (ESP) (http://evs.gs.washington.edu/EVS/).
This list of variants included a novel missense mutation in exon 16 of the DNA methyltransferase 3B (DNMT3B) gene (c.1753G > A; p. Ala585Thr, GCG to ACG) (NM_006892) (Table S1 marked in red). Mutations in DNMT3B are known to cause ICF type 1, a syndrome whose clinical manifestations, in particular the immunological phenotype, are similar to those found in our patient. The mutation was validated in the patient’s family by Dideoxy Sanger sequencing (Fig. 3a), segregated with the disease and occurred in an evolutionary highly conserved position in vertebrate species (Fig. 3b).
Mutation validation and conservation. a Chromatograms of the different DNMT3B genotypes detected in the studied pedigree. Arrows indicate the position of the mutation (c.1753G > A). b Alignment analysis of vertebrate DNMT3B proteins using the NCBI HomoloGene tool was performed. The mutated residue (A535 in human) is boxed. The aligned vertebrate orthologues from top to bottom are human, chimpanzee, gorilla, rhesus macaque, wolf, bovine, mouse, rat, chicken, and zebrafish
Mutated Protein Structure
Pathogenic mutations in the DNMT3B catalytic domain have been previously reported [25, 26]. Most of these mutations are located in the methyltransferase domain in the c-terminal part of the protein (Fig 4a). The mutations can be divided into mutations which affect interaction with the methyl donor, affect interaction with DNMT3L, a required partner for the enzymatic activity, affect interaction with the DNA, or affect protein stability [26]. Currently, there is no resolved structure of DNMT3B protein product (DNMT3B_HUMAN). The most reliable structural information is based on structures of DNMT3A_HUMAN which were recently published [27]. Human DNMT3B and DNMT3A share about 70 % identity in the methyltransferase domain. We modeled the DNMT3B catalytic domain based on the DNMT3A domain using ModWeb [17]. A general view of the model is shown in Fig. 4b. Alanine 585, which is the mutated residue in our patient, is not located in a very close proximity to the DNA or to the area directly involved in catalysis. It is located close to the cofactor (methyl donor) binding site but does not create direct contact with it. The side chain is facing the opposite direction of the cofactor. The catalytic domain in general and the Ala585 region are structurally highly conserved throughout evolution. Figure 4c shows structural alignment between the human DNMT3B methyltransferase domain and its equivalent from Haemophilus haemolyticus, a gram negative bacteria [28]. Despite low sequence identity of about 18 %, the structures are well aligned with a root mean square deviation of 1.5 Å. The cofactors, present in both structures, are perfectly superimposed. The residue in equivalent position to DNMT3B 585 in the bacteria protein is Glycine, which is even smaller than the Alanine found in the human protein. A mutation in Ala585 has been previously reported (Ala585Val) in an ICF patient [29] and even though Val and Thr have different physicochemical properties, both have similar size, significantly larger than the wild type Alanine. We modeled our mutation while rearranging the side chain conformations in the vicinity, using the SCCOMP side chain modeling program [18]. Even in the most optimized combination of side chain conformations, there is still clashing between the side chain of the mutated Thr585 and spatially adjacent residues—Gly614 and Ile622 (Fig. 4d). SCCOMP, like other side chain modeling programs, cannot predict backbone changes, but the clashing suggests that backbone rearrangements must take place. Such rearrangements are very likely to affect the conformation of the nearby cofactor binding site. We also examined the changes in thermostability due to the Ala585Thr mutation (Table S2, Supplementary Appendix). Most of the tools indeed suggest modest reduced thermostability for this variant.
Structure view of DNMT3B Ala585Thr variant. a Schematic view of the protein domain and distribution of known ICF pathogenic variants. Position 585 is shown in red. Most of the mutations occur in the methyltransferase domain and in the C-terminal part of the protein. b Structural view of the DNMT3B model. The DNA is shown in gold. Note the methylated base which flipped toward the enzyme. The S-Adenosyl Homocysteine (SAH) methyl donor is shown in blue and Cys 651 which is known to be the catalytic nucleophile of the reaction is shown in purple. Ala585 is shown in red. c Structural alignment of the catalytic domains of human DNMT3A and Haemophilus haemolyticus methyltransferase demonstrates high structural conservation throughout evolution, despite the low sequence similarity. The binding configurations of the methyl donor are highly similar. The human methyltransferase appears in pink and the bacteria protein in light blue. The SAH appear in red and blue in the two structures respectively in stick representation. Ala585 and its equivalent Gly22 in the bacteria protein appear in spacefill representation. d After modeling, the side chain conformation of the mutated Threonine, steric clashes exist between the Thr side chain and two other residues: Gly614 and Ile622 in the most energetically favorable conformation, suggesting that backbone changes, which are not accounted for by our modeling procedure, must take place in the area
Discussion
Here, we present a girl with immunodeficiency, centromeric instability, and facial anomalies syndrome type 1 (ICF-1), caused by a novel mutation in the DNMT3B gene. As this is an extremely rare syndrome, with only several dozen diagnosed patients on record to date, we sought to review the literature and draw new insights into the clinical presentation, immunological phenotype, and protein structure-function relationship in ICF.
The clinical manifestations of the syndrome in our patient were prototypical of ICF. Her mild facial dysmorphism, which consisted of hypertelorism, micrognathia, epicanthal folds, and low and posteriorly rotated ears, is consistent with previous reports [30]. Recurrent infections of both viral and bacterial etiologies were accentuated in our patient by the drastic failure to thrive (FTT) to which they contributed. FTT strongly correlates with poor prognosis in ICF patients [2, 9], as it did in oursas it did in ours, and was likely the result of the increased metabolic demands of recurrent infection, chronic diarrhea, and feeding difficulties. Chronic diarrhea, of both infectious and non-infectious etiologies, is present in many ICF patients [9]. No viral or parasitic etiology was found in our patient. The patient had feeding difficulties as a neonate, possibly mechanical due to her micrognathia. Though difficult to assess, neurological development in our patient appeared to be age appropriate prior to developing cortical atrophy following a septic episode, which manifested clinically in acute neurologic decline and hand tremor at rest. Cortical atrophy had been reported in several ICF patients [9, 10], albeit without a specified infectious trigger, and may warrant a lower threshold for brain imaging in the event of suspected neurologic decline in these patients. Proteinuria and sonographic findings consistent with parenchymatic renal disease found in our patient are not commonly found in ICF patients.
The immunological hallmark of ICF is hypo- or agammaglobulinemia in the presence of B cells which display an immature phenotype with paucity of class switched, memory and plasma cells, accounting for the lack of antibody production [31–33]. Accordingly, our patient had consistently low to absent immunoglobulin levels, with decreased B cell counts. The immunologic defect, however, appears to go beyond hypogammaglobulinemia, as IVIG treatment, while certainly beneficial [33], fails to completely ameliorate the disease, and many ICF patients, ours amongst them, still succumb to infection [10]. This would suggest that an additional, unknown, element of immunodeficiency contributes to ICF patients’ susceptibility to infections. In support of this claim, successful hematopoietic stem cell transplant, unlike IVIG treatment, results in full immunocompetence [10, 33].
A review of the current literature suggests that a subtle T cell defect develops over time in ICF patients who survive past their early infancy [10]. T cell counts, normal at birth, decline in some patients during the second decade of life, along with neutrophil and thrombocyte counts [10]. It has been hypothesized that the cause of T cell decline is decreased proliferation capacity due to hypomethylation of subtelomeric regions [34], a hypothesis supported both by a decreased in vitro response to mitogen-induced proliferation with phytohemagglutinin (PHA) or IL2 [10], and an increased rate of apoptosis following activation, particularly in CD4+ cells [35]. Mice with homozygous missense mutations in the catalytic domain of DNMT3B likewise display normal T cells counts (and thymocyte counts) at birth, but an increased rate of apoptosis leads to rapidly developing T cell lymphopenia [36]. An inversion of the ratio between CD4+ and CD8+ T cells has also been reported in older ICF patients and was present even in our patient, who died at the early age of 3 years old. It is possible that this inversion, or simply a decline in CD4+, could be prognostic of deterioration in immunological competence, as it renders patients more susceptible to opportunistic infections.
In an attempt to locate additional insults to the adaptive immune system in ICF patients, we examined immunoglobulin and T cell receptor assembly and repertoire diversity. Normal receptor assembly via V(D)J recombination was inferred by Ehrlich et al., who detected IgM and/or IgD on a high percentage of B cells [32] and deduced that recombination, a process upstream to side chain expression, must have occurred. We sought to demonstrate V(D)J recombination more directly and quantify it. T and B cell excision circles, episomal byproducts of successful V(D)J recombination, were found in standard levels, indicating normal de novo production of naïve, rearranged B and T cell in our patient. When a lymphocyte replicates following activation, its episomal excision circle does not replicate, and the proportion of excision circle containing lymphocytes is diluted. The ratio between B cells carrying an excision circle and total B cells can thus provide an estimate for in vivo replication history. The patient’s total to new B cell ratio was low compared with that of healthy controls, in particular given the patient’s extensive infectious history. This is consistent with recently reported reduced in vitro proliferative capacity in B cells in ICF patients [33]. Similar to the reduced proliferation capacity described in T cells, this phenomenon in B cells may likewise result from subtelomeric hypomethylation. Receptor diversity, as estimated by differential usage of variable (V) gene segments for T cell receptor and CDR3 region length distribution for immunoglobulins, was normal in our patient.
Taken together, these data rule out defects in B cell production and receptor repertoire diversification as causes of B cell dysfunction in ICF, suggesting a combination of increased B cell apoptosis and a maturation block prior to class switching [31] are at fault. Regarding the presumed T cell defect, these data help narrow the focus to T cell response, in terms of proliferation and killing capacity, rather than recognition of antigens. This is implied by our finding that the TCR repertoire was intact in our patient.
Based on the bioinformatics analysis of the mutated DNMT3B protein structure (Fig. 4) in our patient, we suggest that structural constraints preclude mutations to large amino acids in the mutated Ala585 position, located on the boundary of helix and loop, which actively participate in the cofactor binding. It is likely that only Alanine (in humans and other vertebrates) and Glycine (as in the Haemophilus haemolyticus orthologue) are allowed in this position due to their small size. Several other pathogenic mutations have already been reported in the vicinity of Ala585 residue, around the methyl donor binding site, most of them indeed change side chain size. These include positions 603 [37], 606, and 613 [38]. The accumulation of mutations in this conserved region, and its high degree of structural similarity throughout evolution, both suggest that the region is highly structurally optimized for cofactor binding and catalytic efficiency.
In conclusion, our results highlight the fact that ICF immunodeficiency extends past hypogammaglobulinemia and may be better characterized as a combined immunodeficiency. With IVIG treatment ameliorating the B cell defect, the extent of CD4+ T cell decline may determine the severity of ICF immunodeficiency and should factor into patient management. ICF patients like ours, with early CD4+ decline, may benefit from HSCT over prophylactic antibiotics regimens.
References
Maraschio P, Zuffardi O, Dalla Fior T, Tiepolo L. Immunodeficiency, centromeric heterochromatin instability of chromosomes 1, 9, and 16, and facial anomalies: the ICF syndrome. J Med Genet. 1988;25:173–80.
Ehrlich M, Jackson K, Weemaes C. Immunodeficiency, centromeric region instability, facial anomalies syndrome (ICF). Orphanet J Rare Dis. 2006;1:2.
Hansen RS, Wijmenga C, Luo P, et al. The DNMT3B DNA methyltransferase gene is mutated in the ICF immunodeficiency syndrome. Proc Natl Acad Sci U S A. 1999;96:14412–7.
De Greef JC, Wang J, Balog J, et al. Mutations in ZBTB24 are associated with immunodeficiency, centromeric instability, and facial anomalies syndrome type 2. Am J Hum Genet. 2011;88:796–804.
Thijssen PE, Ito Y, Grillo G, et al. Mutations in CDCA7 and HELLS cause immunodeficiency-centromeric instability-facial anomalies syndrome. Nat Commun. 2015;6:7870.
Heyn H, Vida E, Sayols S, et al. Whole-genome bisulfite DNA sequencing of a DNMT3B mutant patient. Epigenetics. 2012;7:542–50.
Simo-Riudalbas L, Diaz-Lagares A, Gatto S, et al. Genome-wide DNA methylation analysis identifies novel hypomethylated non-pericentromeric genes with potential clinical implications in ICF syndrome. PLoS One. 2015;10, e0132517.
Walton EL, Francastel C, Velasco G. Dnmt3b prefers germ line genes and centromeric regions: lessons from the ICF syndrome and cancer and implications for diseases. Biology (Basel). 2014;3:578–605.
Hagleitner MM, Lankester A, Maraschio P, et al. Clinical spectrum of immunodeficiency, centromeric instability and facial dysmorphism (ICF syndrome). J Med Genet. 2008;45:93–9.
Weemaes CMR, van Tol MJD, Wang J, et al. Heterogeneous clinical presentation in ICF syndrome: correlation with underlying gene defects. Eur J Hum Genet. 2013;21:1219–25.
Lev A, Simon AJ, Bareket M, et al. The kinetics of early T and B cell immune recovery after bone marrow transplantation in RAG-2-deficient SCID patients. PLoS One. 2012;7, e30494.
Lev A, Simon AJ, Broides A, et al. Thymic function in MHC class II-deficient patients. J Allergy Clin Immunol. 2013;131:831–9.
Kraus M, Lev A, Simon A. Disturbed B and T cell homeostasis and neogenesis in patients with ataxia telangiectasia. J Clin Immunol. 2014;34:561–72.
Li H, Durbin R. Fast and accurate short read alignment with Burrows-Wheeler transform. Bioinformatics. 2009;25:1754–60.
McKenna A, Hanna M, Banks E, et al. The Genome Analysis Toolkit: a map reduce framework for analyzing next-generation DNA sequencing data. Genome Res. 2010;20:393–303.
Li MX, Gui HS, Kwan JSH, Bao SY, Sham PC. A comprehensive framework for prioritizing variants in exome sequencing studies of Mendelian diseases. Nucleic Acids Res. 2012;40(7):e53.
Webb B, Sali A. Protein structure modeling with MODELLER. Methods Mol Biol. 2014;1137:1–15.
Eyal E, Najmanovich R, McConkey BJ, Edelman M, Sobolev V. Importance of solvent accessibility and contact surfaces in modeling side-chain conformations in proteins. J Comput Chem. 2004;25:712–24.
Nussinov R, Wolfson HJ. Efficient detection of three-dimensional structural motifs in biological macaromolecules by computer vision techniques. Proc Natl Acad Sci. 1991;88:10495–9.
DE Pires V, Ascher DB, Blundell TL. DUET: a server for predicting effects of mutations on protein stability using an integrated computational approach. Nucleic Acids Res. 2014;42:314–9.
DE Pires V, Ascher DB, Blundell TL. MCSM: predicting the effects of mutations in proteins using graph-based signatures. Bioinformatics. 2014;30:335–42.
Worth CL, Preissner R, Blundell TL. SDM—a server for predicting effects of mutations on protein stability and malfunction. Nucleic Acids Res. 2011;39:215–22.
Laimer J, Hofer H, Fritz M, Wegenkittl S, Lackner P. MAESTRO-multi agent stability prediction upon point mutations. BMC Bioinformatics. 2015;16:116.
Capriotti E, Fariselli P, Casadio R. I-Mutant2.0: predicting stability changes upon mutation from the protein sequence or structure. Nucleic Acids Res. 2005;33:306–10.
Hamidi T, Singh AK, Chen T. Genetic alterations of DNA methylation machinery in human diseases. Epigenomics. 2015;7:247–65.
Cheng X, Blumenthal RM. Mammalian DNA methyltransferases: a structural perspective. Structure. 2008;16:341–50.
Guo X, Wang L, Li J, et al. Structural insight into autoinhibition and histone H3-induced activation of DNMT3A. Nature. 2015;517:640–4.
Klimasauskas S, Kumar S, Roberts RJ, Cheng X. Hhal methyltransferase flips its target base out of the DNA helix. Cell. 1994;76:357–69.
Wijmenga C, Hansen RS, Gimelli G, et al. Genetic variation in ICF syndrome: evidence for genetic heterogeneity. Hum Mutat. 2000;16:509–17.
Ehrlich M, Sanchez C, Shao C, et al. ICF, an immunodeficiency syndrome: DNA methyltransferase 3B involvement, chromosome anomalies, and gene dysregulation. Autoimmunity. 2008;41:253–71.
Blanco-Betancourt CE, Moncla A, Milili M, et al. Defective B-cell-negative selection and terminal differentiation in the ICF syndrome. Blood. 2004;103:2683–90.
Ehrlich M, Buchanan KL, Tsien F, et al. DNA methyltransferase 3B mutations linked to the ICF syndrome cause dysregulation of lymphogenesis genes. Hum Mol Genet. 2001;10:2917–31.
Sterlin D, Velasco G, Moshous D, et al. Genetic, cellular and clinical features of ICF syndrome: a French national survey. J Clin Immunol. 2016;36:149–59.
Yehezkel S, Segev Y, Viegas-Péquignot E, Skorecki K, Selig S. Hypomethylation of subtelomeric regions in ICF syndrome is associated with abnormally short telomeres and enhanced transcription from telomeric regions. Hum Mol Genet. 2008;17:2776–89.
Pezzolo A, Prigione I, Facchetti P, et al. T-cell apoptosis in ICF syndrome. J Allergy Clin Immunol. 2001;108:310–2.
Ueda Y, Okano M, Williams C, Chen T, Georgopoulos K, Li E. Roles for Dnmt3b in mammalian development: a mouse model for the ICF syndrome. Development. 2006;133:1183–92.
Landrum MJ, Lee JM, Benson M, et al. ClinVar: public archive of interpretations of clinically relevant variants. Nucleic Acids Res. 2015;44:D862–8.
Stenson PD, Ball EV, Mort M, et al. Human gene mutation database (HGMD): 2003 update. Hum Mutat. 2003;21:577–81.
Shearer WT, Rosenblatt HM, Gelman RS, et al. Lymphocyte subsets in healthy children from birth through 18 years of age: the pediatric AIDS clinical trials group P1009 study. J Allergy Clin Immunol. 2003;112:973–80.
Acknowledgments
We thank the patient’s family for their cooperation. Raz Somech is supported by the Jeffrey Modell Foundation (JMF).
Authorship Contributions
B.P.S, Y.A. and R.S. diagnosed, followed, and treated the patient.
E.R., A.L., R.S, and A.J.S. conceived and designed the experiments.
E.R., A.L. A.J.S., and S.F.B. performed the experiments.
E.E., O.B., and N.K., performed the bioinformatic analysis
E.R., A.L., E.E., R.S., and A.J.S. analyzed and interpreted results and wrote the paper.
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
All procedures performed in studies involving human participants were in accordance with the ethical standards of the institutional and/or national research committee and with the 1964 Helsinki Declaration and its later amendments or comparable ethical standards.
Informed consent was obtained from all individual participants included in the study.
Conflict of Interest
The authors declare that they have no conflict of interests.
Additional information
Erez Rechavi and Atar Lev have equally contributed to the study.
Electronic Supplementary Material
Below is the link to the electronic supplementary material.
ESM 1
(DOCX 50.7 KB)
Rights and permissions
About this article
Cite this article
Rechavi, E., Lev, A., Eyal, E. et al. A Novel Mutation in a Critical Region for the Methyl Donor Binding in DNMT3B Causes Immunodeficiency, Centromeric Instability, and Facial Anomalies Syndrome (ICF). J Clin Immunol 36, 801–809 (2016). https://doi.org/10.1007/s10875-016-0340-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10875-016-0340-z