Abstract
Data validation plays an important role in ensuring the reliability and reproducibility of studies. NMR investigations of the functional properties, dynamics, chemical kinetics, and structures of proteins depend critically on the correctness of chemical shift assignments. We present a novel probabilistic method named ARECA for validating chemical shift assignments that relies on the nuclear Overhauser effect data . ARECA has been evaluated through its application to 26 case studies and has been shown to be complementary to, and usually more reliable than, approaches based on chemical shift databases. ARECA is available online at http://areca.nmrfam.wisc.edu/.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Nuclear magnetic resonance (NMR) spectroscopy is used routinely for studying molecular interactions (Goldflam et al. 2011; Nishida and Shimada 2011), structural dynamics (Baldwin and Kay 2009; Kim et al. 2012; Kleckner and Foster 2011), and three-dimensional structures (Wüthrich 1986) of proteins. A necessary key step in such studies is associating spectral frequencies with atoms, i.e., finding a solution to the chemical shift assignment problem. Currently, most manual and automated approaches to NMR studies derive assignments from through-bond coupling in proteins labeled uniformly with 13C and 15N. The valid interpretation of NMR studies is dependent on the correctness of the assignments. For example, protein structure determination relies on assignments for the correct interpretation of through-space contact information contained in nuclear Overhauser effect (NOE) spectra (Wüthrich 1986). Algorithms have been developed to address the concern of the validity of chemical shift assignments on the basis of chemical shift statistics with or without information on the 3D structure of the protein (Moseley et al. 2004; Rieping and Vranken 2010; Shen and Bax 2010; Wang et al. 2010; Zhang et al. 2003). Because these approaches rely on chemical shift statistics from Biological Magnetic Resonance data Bank (BMRB, Ulrich et al. 2008), the errors they report are only indicators of deviations averaged chemical shifts. In a given protein, the effects of ring-currents (Vernet and Boekelheide 1974; Wannere and Schleyer 2003), hydrogen-bonds (Yao et al. 2010), and other local conformational effects often lead to substantial deviations from chemical shift predictions. As a result, correct assignments can be scored as invalid or incorrect assignments as valid. Validation methods that use 3D structures are only applicable when a highly accurate structure is available, and this is not the case for the many NMR studies concerned with molecular interactions, conformational changes, dynamics, or intrinsically disordered proteins. Current chemical shift validation methods suffer from their reliance on chemical shift statistics and 3D structures of uncertain structural quality (Buchner and Güntert 2015).
Every entry in BMRB contains a validation report generated by the AVS method (Moseley et al. 2004), which lists chemical shift assignment outliers. On average, every entry contains more than 19 outlier assignments with a standard deviation about 94 flagged atoms (Fig. 1). These large numbers of outliers clearly indicate the need for an independent approach for chemical shift validation. NOE experiments can provide an alternative source of information for validating chemical shift assignments through the detection of short-range through-space interactions between protons. NOE information has been used for automated assignment schemes (Schmidt and Güntert 2012; Xu et al. 2006) and for validating protein structures (Huang et al. 2005); however, the independent validation of assignments has been limited to a manual approach for backbone protons (Serrano et al. 2012). We describe here a probabilistic method called ARECA for ‘Assessment of the Reliability of Chemical shift Assignments’ that is applicable to diverse NMR studies (Fig. 2) and provides tools for examining and correcting suspicious assignments.
Histogram of numbers of flagged atoms by the AVS method on the BMRB entries. The data were retrieved from BMRB in September 2015. The inset region shows that most of the entries have fewer than 40 flagged atoms. BMRB entry bmr16632 with 2489 flagged atoms has the highest number of flagged atoms (likely as the result of incorrect chemical shift referencing)
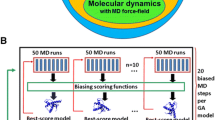
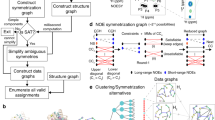
Application of ARECA to different types of NMR studies. ARECA is capable of validating protein chemical shift assignments in NMR studies with a variety of goals (e.g. molecular dynamics, chemical kinetics, structure calculations) and with samples containing different labeling schemes. The first column shows the protein labeling patterns of the NMR sample. The second column indicates the atom types whose assignments can be validated with a given labeling pattern. The third column indicates the types of input NOE data required. The fourth column indicates the expected NOE contacts that ARECA validates for each type of study
Methods
ARECA’s software package consists of five modules (Fig. 3): inputs, statistical analyses, probability calculations, outputs, and external resources.
Overall structure of ARECA. Users interact with the web-interface of ARECA to upload the inputs. These inputs are submitted to the probability calculations module and also to the external resource (LACS package). The statistical analyses module is a preprocessed unit that provides a truth model and a CDF to the probabilistic validation part of the probability calculations module. The probability calculations module uses the truth model and the CDF for assigning the short-range NOESY cross-peaks and also validating the inputs. The output module provides the validation results and the LACS results on chemical shift referencing
Input module
The input to ARECA consists of chemical shift assignments in either BMRB or XEASY format and 13C-edited and/or 15N-edited NOESY peak lists in either SPARKY or XEASY format. Alternatively, ARECA accepts input files generated by PONDEROSA-C/S (Lee et al. 2014), which performs peak picking on the NOESY spectra and generates a single compact file containing assignments and peak lists.
Statistical analyses
This module provides the necessary statistical information used in ARECA’s calculations. The truth model was constructed using the experimental database (PACSY, Lee et al. 2012) and was verified against the theoretical database generated using the Tinker molecular modeling software package (http://dasher.wustl.edu/tinker/) (Kundrot et al. 1991; Pappu et al. 1998; Ponder and Richards 1987; Ponder et al. 2010; Ren and Ponder 2003; Ren et al. 2011; Shen and Bax 2013). Supplementary Information S1 explains the process of constructing and evaluating the truth model. The cumulative distribution function (CDF) was generated from imperfect input and was used to calculate a p value for rejecting the null hypothesis that ARECA’s output is a result of validating imperfect inputs (chemical shift assignments and peak lists). The CDF is explained in Supplementary Information S2.
Probability calculation module
This module calculates the assignment probabilities and confidences using the truth model. The assignment probability can be thought of as a supporting factor that represents how well an assignment is validated by experimental NOESY peaks matching those expected from the truth model. A low probability (low supporting factor) means either the assignment is incorrect or that the experimental NOESY peak lists are missing expected peaks. Supplementary Information S3 describes this module in greater details.
External resource module
The correct chemical shift referencing of the heavy atoms of the backbone (plus CB) is crucial for protein NMR studies. The LACS (Linear Analysis of Chemical Shifts) package (Wang et al. 2005) is a commonly used tool for identifying and correcting chemical shift referencing errors. The LACS package applies a regression function to the chemical shifts of CA, CB, HA, and C, and suggests referencing adjustments. This report is available as an output of ARECA and can be used to identify and correct chemical shift referencing errors.
Output module
When the calculations are completed, ARECA sends an email to the user containing (a) the percentages of the flagged residues and atoms, (b) the calculated p value, which provides a quick assessment of the correctness of the inputs, (c) a hyperlink to an xml-formatted report of the probabilities and confidences of chemical shift assignments for the residues and atoms within, and (d) a hyperlink to a compressed file that contains a comprehensive report (in pdf format) with information (as described in Supplementary Information S3) on the reasoning behind each calculated probability, along with assigned NOESY peak lists and the LACS report. ARECA’s extension in NMRFAM-SPARKY (Lee et al. 2015) enables the user to view the peaks in the NOESY spectrum colored according to their assignment probability (Fig. 4).
Screen shots of the ARECA extension to NMRFAM-SPARKY. NMRFAM-SPARKY, which is available from the NMRFAM website, reads ARECA’s assigned peaks lists and overlays the peaks onto experimental NOE spectra color-coded according to their probabilities. Peaks with a probability higher than or equal to the ‘good’ ARECA cutoff are shown in green; peaks with a probability between the ‘good’ and ‘intermediate’ ARECA cutoffs are shown in blue; peaks with probabilities less than the intermediate cutoff are shown in yellow; and peaks that were expected but were not in the given NOE peak lists are shown in red. a ARECA window showing a partial 15N-NOESY peak list. The table shows the peak assignments, their spectral frequencies (ω1, ω2, ω3), and their calculated probabilities (in this example all 1.000). b NOESY region showing peaks colored according to their probabilities. This example shows the strip plot for residue Q182 N–H of VDRLBD (see Supplementary Information S4). Peaks corresponding to the inter-residue contacts of Q182H with V181H and D183H were present in the peak lists with the probability = 1 and are colored green. However, the intra-residue contact is missing its representative peak (in this case because of spectral folding known as ‘aliasing’), and its assigned probability is zero. ARECA’s extension placed the expected peak and colored it red to signal the inconsistency between assignments and NOE peak lists
In ARECA, each proposed chemical shift assignment is validated against NOESY peak lists on the basis of a truth model (Supplementary Information S1) generated from a curated database of peptide or protein structures. A list of expected NOESY contacts is generated from the truth model, and these are used to calculate for each residue (Supplementary Information S3) (1) the number of expected NOESY contacts that agree with the experimental NOESY peak lists (a measure of experimental support for the assignment or its probability) and (2) the number of assigned atoms out of the theoretical maximum (a measure of assignment completeness or its confidence). ARECA flags suspicious assignments: cases when fewer than half of the expected contacts could be verified (probability <0.5) or cases when more than half of the atoms expected to contact the atom in question were unassigned (confidence <0.5). ARECA also calculates the percentage of flagged atoms (the number of flagged atoms over the total number of assigned atoms) and reports the p value of rejecting the null hypothesis that the percentage is a result of validating imperfect inputs (chemical shift assignments and peak lists). The p value is calculated using a CDF of the percentages of 10,000 incorrect inputs. In Supplementary Information S2, we discuss (1) the construction of the CDF, (2) the advantages of ARECA over other validation methods, and (3) the evaluation of ARECA’s performance with incomplete or incorrect inputs that resulted in 98 % precision and 96 % recall.
Results and discussion
Over a range of 26 case studies, we compared ARECA’s performance against two validation packages that rely on the chemical shift statistics: AVS (Moseley et al. 2004) and PANAV (Wang et al. 2010). In addition, for proteins with structures deposited into the Protein Data Bank (PDB) (Berman et al. 2007), we compared ARECA’s performance against three validation packages that rely on 3D structures: VASCO (Rieping and Vranken 2010; Vranken and Rieping 2009), SHIFTCOR (Zhang et al. 2003), and SPARTA+ (Shen and Bax 2007, 2010). These comparisons were conducted on a set of 16 proteins that included 10 target proteins from the second round of CASD-NMR (Rosato et al. 2015; Wassenaar et al. 2012) and 6 other proteins from the BMRB. The other validation methods rely on chemical shift statistics as the basis for flagging outlier atoms that could have valid or incorrect assignments. By contrast, ARECA flags atoms with inconsistencies between chemical shift assignments and experimental NOE peak lists, as can result from incorrect assignments or incomplete or erroneous NOE peak lists.
The p value in ARECA addresses the question of whether the assignments and NOE peak lists are reliable or require further investigation. From our analysis, we concluded that a p value less than 0.05 is indicative of a reliable data set. In situations where the p value is higher than this threshold, additional investigations can be performed using the ARECA’s extension in NMRFAM-SPARKY (Supplementary Information S4). Table 1 shows the p values for the data sets examined here.
The refined peak lists of all the CASD-NMR proteins had acceptable p values (<1.0E−4), which indicated that the inputs were accurate and required no further investigation. ARECA flagged significantly more assignments with the raw NOE peak lists as input than with the refined NOE peak lists (except for HR8254A, whose values remained the same with p values of 1.0E−4). Three of the non-CASD-NMR proteins examined (HR2869A, BvR155, RhR5) had unacceptable p values (>0.05); our investigation of the flagged atoms indicated that the low probabilities most likely were the result of errors in the peak lists deposited at BMRB. The remaining three proteins had acceptable p values (1.0E−4), and no further investigation was required. These results are discussed in greater details in Supplementary Information S4.
The number of atoms flagged by ARECA does not necessarily indicate the correctness or completeness of the data; nevertheless, we compared this number with those flagged by the other validation methods (Fig. 5). In parentheses are the ratio of the number of flagged atoms in common between ARECA and that method divided by the total number of atoms flagged by that method. These ratios indicate that the level of agreement between ARECA and the other validation approaches is low. The atoms flagged by the other validation methods include carbonyl carbons, which are not evaluated by ARECA. A similar analysis was carried out with raw peak lists for 10 CASD-NMR proteins (Fig. 6); in all but one case, the agreement was worse than with the refined peak lists as input.
The bars indicate the number of atoms flagged by the different validation methods for 10 CASD-NMR proteins (ARECA used the final peak lists) and 6 non-CASD-NMR proteins. If the method was not applicable because of the lack of a 3D structure, its name is followed by ‘(-)’. The numbers on top of the bars for the other methods provide the ratio between the atoms flagged in common by ARECA and that method divided by the total number of atoms flagged by that method
The bars indicate the number of atoms flagged by the different validation methods for 10 CASD-NMR proteins (ARECA used the raw peak lists). If the method was not applicable because of the lack of a 3D structure, its name is followed by ‘(-)’. The numbers on top of the bars for the other methods provide the ratio between the atoms flagged in common by ARECA and that method divided by the total number of atoms flagged by that method
Conclusions
In contrast to methods that report outliers that may represent valid or invalid assignments, ARECA flags assignments with low probability or confidence and offers tools for determining whether they result from missing data or an incorrect assignment; ARECA facilitates the correction of assignments on the basis of reinterpretation of experimental spectral data. In addition, the p values calculated by ARECA provide a criterion for the validity of the input data. Unique strengths of ARECA are (1) its provision of a metric for overall assignment validity and (2) its associated visualization tools that enable users to verify or correct the assignments of flagged atoms. The ARECA extension to the NMRFAM-SPARKY software suite (Lee et al. 2015) can be used to investigate atoms with low assignment probabilities in the context of experimental protein NMR data. Such investigations can diagnose whether a low assignment probability arose from missing NOE peaks (as can result from chemical exchange, peak overlap, or spectral artifacts) or from errors in data interpretation (such as uncorrected spectral aliasing or inaccurate peak picking). Verification of the consistency between NOESY data and chemical shift assignments can help to simplify and accelerate the tedious process of structure calculation and refinement from NMR data. Even for protein NMR studies that do not involve 3D structure determination, NOESY data can be collected and used to validate chemical shift assignments. ARECA shows promise for future incorporation into strategies for automated protein NMR assignment and structure determination.
References
Baldwin AJ, Kay LE (2009) NMR spectroscopy brings invisible protein states into focus. Nat Chem Biol 5:808–814
Berman H, Henrick K, Nakamura H, Markley JL (2007) The worldwide Protein Data Bank (wwPDB): ensuring a single, uniform archive of PDB data. Nucleic Acids Res 35:D301–D303
Buchner L, Güntert P (2015) Increased reliability of nuclear magnetic resonance protein structures by consensus structure bundles. Structure 23:425–434
Goldflam M, Tarrago T, Gairi M, Giralt E (2011) NMR studies of protein-ligand interactions. Methods Mol Biol 831:233–259
Huang YJ, Powers R, Montelione GT (2005) Protein NMR recall, precision, and F-measure scores (RPF Scores): structure quality assessment measures based on information retrieval statistics. JACS 127:1665–1674
Kim JH, Tonelli M, Markley JL (2012) Disordered form of the scaffold protein IscU is the substrate for iron-sulfur cluster assembly on cysteine desulfurase. PNAS 109:454–459
Kleckner IR, Foster MP (2011) An introduction to NMR-based approaches for measuring protein dynamics. Biochim Biophys Acta 1814:942–968
Kundrot CE, Ponder JW, Richards FM (1991) Algorithms for calculating excluded volume and its derivatives as a function of molecular conformation and their use in energy minimization. J Comput Chem 12:402–409
Lee W, Yu W, Kim S, et al (2012) PACSY, a relational database management system for protein structure and chemical shift analysis. J Biomol NMR 54:169–179. doi:10.1007/s10858-012-9660-3
Lee W, Stark JL, Markley JL (2014) PONDEROSA-C/S: client-server based software package for automated protein 3D structure determination. J Biomol NMR 60:73–75
Lee W, Tonelli M, Markley JL (2015) NMRFAM-SPARKY: enhanced software for biomolecular NMR spectroscopy. Bioinformatics 31:1325–1327
Moseley HN, Sahota G, Montelione GT (2004) Assignment validation software suite for the evaluation and presentation of protein resonance assignment data. J Biomol NMR 28:341–355
Nishida N, Shimada I (2011) An NMR method to study protein-protein interactions. Methods Mol Biol 757:129–137
Pappu RV, Hart RK, Ponder JW (1998) Analysis and application of potential energy smoothing and search methods for global optimization. J Phys Chem B 102:9725–9742
Ponder JW, Richards FM (1987) An efficient newton-like method for molecular mechanics energy minimization of large molecules. J Comput Chem 8:1016–1024
Ponder JW, Wu C, Ren P, Pande VS, Chodera JD, Schnieders MJ, Haque I, Mobley DL, Lambrecht DS, DiStasio RA Jr, Head-Gordon M, Clark GN, Johnson ME, Head-Gordon T (2010) Current status of the AMOEBA polarizable force field. J Phys Chem B 114:2549–2564
Ren P, Ponder JW (2003) Polarizable atomic multipole water model for molecular mechanics simulation. J Phys Chem B 107:5933–5947
Ren P, Wu C, Ponder JW (2011) Polarizable atomic multipole-based molecular mechanics for organic molecules. J Chem Theory Comput 7:3143–3161
Rieping W, Vranken WF (2010) Validation of archived chemical shifts through atomic coordinates. Proteins 78:2482–2489
Rosato A, Vranken W, Fogh R, Ragan T, Tejero R, Pederson K, Lee H-W, Prestegard J, Yee A, Wu B, Lemak A, Houliston S, Arrowsmith C, Kennedy M, Acton T, Xiao R, Liu G, Montelione G, Vuister G (2015) The second round of critical assessment of automated structure determination of proteins by NMR: CASD-NMR-2013. J Biomol NMR 62:413–424
Schmidt E, Güntert P (2012) A new algorithm for reliable and general NMR resonance assignment. J Am Chem Soc 134:12817–12829
Serrano P, Pedrini B, Mohanty B, Geralt M, Herrmann T, Wuthrich K (2012) The J-UNIO protocol for automated protein structure determination by NMR in solution. J Biomol NMR 53:341–354
Shen Y, Bax A (2007) Protein backbone chemical shifts predicted from searching a database for torsion angle and sequence homology. J Biomol NMR 38:289–302
Shen Y, Bax A (2010) SPARTA+: a modest improvement in empirical NMR chemical shift prediction by means of an artificial neural network. J Biomol NMR 48:13–22
Shen Y, Bax A (2013) Protein backbone and sidechain torsion angles predicted from NMR chemical shifts using artificial neural networks. J Biomol NMR 56:227–241
Ulrich EL, Akutsu H, Doreleijers JF, Harano Y, Ioannidis YE, Lin J, Livny M, Mading S, Maziuk D, Miller Z, Nakatani E, Schulte CF, Tolmie DE, Kent Wenger R, Yao H, Markley JL (2008) BioMagResBank. Nucleic Acids Res 36:402–408
Vernet RD, Boekelheide V (1974) Nuclear magnetic resonance spectroscopy. ring-current effects on carbon-13 chemical shifts. PNAS 71:2961–2964
Vranken WF, Rieping W (2009) Relationship between chemical shift value and accessible surface area for all amino acid atoms. BMC Struct Biol 9:20
Wang L, Eghbalnia HR, Bahrami A, Markley JL (2005) Linear analysis of carbon-13 chemical shift differences and its application to the detection and correction of errors in referencing and spin system identifications. J Biomol NMR 32:13–22
Wang B, Wang Y, Wishart DS (2010) A probabilistic approach for validating protein NMR chemical shift assignments. J Biomol NMR 47:85–99
Wannere CS, Schleyer PV (2003) How do ring currents affect (1)h NMR chemical shifts? Org Lett 5:605–608
Wassenaar T, van Dijk M, Loureiro-Ferreira N, van der Schot G, de Vries S, Schmitz C, van der Zwan J, Boelens R, Giachetti A, Ferella L, Rosato A, Bertini I, Herrmann T, Jonker HA, Bagaria A, Jaravine V, Güntert P, Schwalbe H, Vranken W, Doreleijers J, Vriend G, Vuister G, Franke D, Kikhney A, Svergun D, Fogh R, Ionides J, Laue E, Spronk C, Jurkša S, Verlato M, Badoer S, Dal Pra S, Mazzucato M, Frizziero E, Bonvin AJJ (2012) WeNMR: structural biology on the grid. J Grid Comput 10:743–767
Wüthrich K (1986) NMR of proteins and nucleic acids. Wiley, Hoboken
Xu Y, Zheng Y, Yang D (2006) NOESY-based strategy for assignments of backbone and side chain resonances of large proteins without deuteration. Nat Protoc Exch. doi:10.1038/nprot.2006.410
Yao L, Grishaev A, Cornilescu G, Bax A (2010) The impact of hydrogen bonding on amide 1H NMR spectroscopy. JACS 132:10866–10875
Zhang H, Neal S, Wishart DS (2003) RefDB: a database of uniformly referenced protein chemical shifts. J Biomol NMR 25:173–195
Acknowledgments
HD thanks Alireza Siahpirani (Wisconsin Institute of Discovery) for invaluable discussions about the computational aspects of the introduced method and Dr. Vincent Chen (National Magnetic Resonance Facility at Madison) for helpful discussions concerning applications of the MolProbity program. For the use of web portals, computing and storage facilities, we thank the WeNMR project (European FP7 e-Infrastructure grant, contract no. 261572, www.wenmr.eu), supported by the European Grid Initiative (EGI) through the national GRID Initiatives of Belgium, France, Italy, Germany, the Netherlands, Poland, Portugal, Spain, UK, South Africa, Malaysia, Taiwan, the Latin America GRID infrastructure via the Gisela project, the International Desktop Grid Federation (IDGF) with its volunteers, and the US Open Science Grid (OSG). This study was carried out at the National Magnetic Resonance Facility at Madison, which is supported by US National Institutes of Health (NIH) grant P41GM103399.
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Dashti, H., Tonelli, M., Lee, W. et al. Probabilistic validation of protein NMR chemical shift assignments. J Biomol NMR 64, 17–25 (2016). https://doi.org/10.1007/s10858-015-0007-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10858-015-0007-8