Abstract
Small molecules with multi-target activity, also termed promiscuous compounds, are increasingly considered for pharmaceutical applications. The use of promiscuous chemical entities represents a departure from the compound specificity paradigm, one of the pillars of modern drug discovery. The popularity of promiscuous compounds is due to the concept of polypharmacology; another more recent drug discovery paradigm. It refers to insights that the efficacy of drugs often depends on interactions with multiple targets. Views concerning the extent to which small molecules might form well-defined interactions with multiple targets often differ, but comprehensive experimental investigations of promiscuity are currently rare. On the other hand, large volumes of active compounds and experimental measurements are becoming available and enable data-driven analyses of compound selectivity versus promiscuity. In this perspective, we discuss computational methods and data structures designed for promiscuity analysis. In addition, findings from large-scale exploration of activity profiles of inhibitors covering the human kinome are summarized. Although many kinase inhibitors are expected to be promiscuous, they are frequently found to be selective, which provides opportunities for target-directed drug discovery (rather than polypharmacology). We also discuss that machine learning yields evidence for the existence of structure–promiscuity relationships.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Facets of promiscuity and approaches for its assessment
The ability of small molecules to specifically interact with multiple targets is referred to as promiscuity [1, 2]. Unlike non-specific binding events that originate from compound aggregation or assay interference [3,4,5,6,7], genuine multi-target activity is often desirable and forms the basis of polypharmacology [8,9,10]. The polypharmacology paradigm states that bioactive compounds frequently interact with multiple targets in vivo and thereby elicit their therapeutic effects. Accordingly, polypharmacology has become a major discovery strategy in a number of therapeutic areas such as cardiovascular, metabolic, or oncological diseases where the typically multi-factorial nature of disorders and development of drug resistance affect therapeutic success [9, 11].
Experimental and complementary computational approaches have been introduced for compound promiscuity analysis. For example, microarray and target profiling experiments are a major source of multi-target activity data, as exemplified by kinase inhibitor profiling studies [12,13,14]. However, comprehensive cell-based or in vivo profiling analyses in model organisms are currently rare [15]. On the other hand, systematic computational analysis of rapidly growing amounts of compound activity data from medicinal chemistry and biological screening sources makes it possible to explore promiscuity in a data-driven manner on a large scale [16,17,18]. Given currently available activity data volumes, such analyses are expected to yield statistically sound trends, despite data incompleteness [2, 16, 17]. Furthermore, other computational approaches complementing compound data analysis have been developed to assess or predict compound promiscuity. For example, various statistical models based on ligand similarity were derived to predict new targets for known active compounds [19,20,21,22]. In addition, machine learning models were developed to distinguish between highly, weakly, or non-promiscuous molecules [23, 24]. Furthermore, known promiscuous compounds were used to establish previously unknown chemical links between distantly related or unrelated target proteins [25]. However, confirming new compound-based target relationships on the basis of experimental activity data is often hindered by uncertainty of assay readouts and potential artifacts. Accordingly, various computational filters have been developed to detect potential false-positive assay results [26, 27]. Such rule-based computational filters are often viewed controversially in the field. However, they provide helpful alerts raising awareness of potential artifacts that need to be considered carefully.
Compound promiscuity has also been investigated at the protein structure level where binding site similarity was determined and used to rationalize multi-target engagement of ligands [28]. Furthermore, the choice of appropriate protein conformations for the design of polypharmacological ligands is considered pivotal to success. Therefore, potential advantages and limitations of different structure selection methods were evaluated to foster multi-target drug development [29]. Moreover, systematic analysis of X-ray data identified ligands bound to multiple target proteins from different families, hence providing templates for polypharmacology-oriented ligand design [30]. Another analysis revealed that promiscuous compounds contained in multiple X-ray structures often formed different interaction hotspots in binding sites of unrelated proteins, but displayed overall similar binding modes [31]. A recent perspective details current structure-based approaches for compound promiscuity analysis [32].
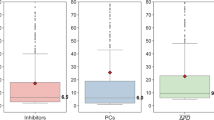
Despite the role of polypharmacology for the efficacy of many drugs, it currently remains unclear to which extent bioactive compounds are promiscuous. Analyses of currently available compound activity data do not support assumptions that drugs and other bioactive compounds might generally be promiscuous [17, 18]. Clearly, target selectivity of active compounds as a drug discovery goal cannot be disregarded. For promiscuity of drugs, expectation values have been put forward. On the basis of drug-target network analysis using different data sets and drug classes, it was estimated early on that drugs might interact on average with three to 13 targets, depending on the data sources that were used [33, 34]. Data incompleteness inevitably affects promiscuity assessment as long as drugs have not been tested against all possible protein targets [33], which will most likely remain an elusive goal. However, the consideration of test frequencies of compounds provides valuable insights. Screening compounds that were extensively tested in hundreds of assays were found to interact on average with two to three targets and also contained many consistently inactive molecules [17]. In addition, inhibitors of the human kinome, which are often expected to be promiscuous, as further discussed below, also displayed only limited global promiscuity [18]. Hence, more work will be required to systematically quantify promiscuity among bioactive compounds and further explore relationships between multi-target activity and target selectivity or specificity. Exploring such relationships continues to be critically important for many therapeutic applications.
For activity data-driven assessment of compound promiscuity, different chemoinformatic data structures have been introduced. In the following, key concepts leading to the derivation of these data structures are highlighted, their importance in analyzing structure–promiscuity relationships is discussed, and exemplary applications to inhibitors of the human kinome are presented.
Data structures for computational promiscuity analysis
In analogy to activity cliffs, which were defined as pairs of structurally similar active compounds with large potency differences [35, 36], promiscuity cliffs (PCs) have been defined as pairs of structurally analogous compounds with a large difference in the number of targets they are active against [37]. Furthermore, the promiscuity degree (PD) is defined as the number of targets a compound is active against [37]. Figure 1a shows exemplary PCs. By definition, PCs reveal small structural modifications of compounds that are associated with large differences in promiscuity. Thus, PCs enable the exploration of structure–promiscuity relationships and the derivation of new target hypotheses for structural analogues. Defining PCs requires the consideration of a compound similarity and a promiscuity difference (ΔPD) criterion. As a similarity criterion, the formation of a matched molecular pair (MMP) [38] is preferred [37]. An MMP is a pair of compounds that are only distinguished by a chemical modification at a single site [38]. The ΔPD criterion can be variably set, depending on the desired magnitude of PCs and specific requirements of applications.
Data structures for promiscuity analysis. a Exemplary promiscuity cliffs (PCs) are shown that are formed by compounds from biological screening (top; tested in 358 and 339 assays, respectively) and medicinal chemistry (bottom). b A section of a PC network is shown in which nodes represent compounds and edges pairwise PC relationships. Nodes are color-coded by promiscuity degrees (PD values). A large and heterogeneous PC cluster is highlighted. c A sequence of compounds forming a PC pathway is displayed that is traced in the highlighted cluster shown below. For each pathway compound, the PD value is reported. d A prominent promiscuity hub of the pathway is shown together with off-pathway compounds with which it forms PCs. In the cluster below, the corresponding subgraph from which the PCs originate is highlighted
Occurrence of PCs has been confirmed on the basis of experimental data by analyzing extensively assayed screening compounds [39]. High-confidence PCs were determined by taking assay frequency and overlap information for compounds into account [39]. PCs were frequently formed by compounds tested in hundreds of shared assays. Moreover, through large-scale analysis of activity data, thousands of PCs were identified in active compounds from biological screening or medicinal chemistry [39, 40].
PCs can be systematically assessed and visualized in PC networks (PCNs). In a PCN, nodes represent compounds and edges pairwise PC relationships (i.e., the pairwise formation of PCs by compounds) [39,40,41] (Fig. 1b). In addition, PCNs reveal the formation of PC clusters (disjoint network components) of varying size and topology (Fig. 1b). From these clusters, PC pathways (PCPs) can be isolated. PCP is defined as a sequence of PCs that consists of alternating highly and weakly promiscuous (or non-promiscuous) compounds [41]. An exemplary PCP is shown in Fig. 1c. Given their composition, PCPs are rich in structure–promiscuity relationship information. A characteristic feature of many PCPs is the presence of promiscuity hubs (PHs). Following network terminology, a hub refers to a densely connected node in a network. Hence, a PH is defined as a highly promiscuous PCP compound that forms many PCs with weakly or non-promiscuous compounds outside the pathway [41]. Accordingly, PHs suggest many target hypotheses for weakly or non-promiscuous structural analogues whose low PD values might be due to data sparseness. Figure 1d shows an example of a highly promiscuous hub and its PCN environment.
The PC, PCN, PCP, and PH data structures provide a basis for detailed computational promiscuity analysis. Increasing size and complexity of PC clusters quickly limits interactive analysis of PCPs. Therefore, a computational approach to systematically identify, extract, and prioritize informative PCPs from PC clusters has been recently reported [42]. The methodology relied on the detection of the shortest path between any two nodes from a PC cluster. For the identification of shortest paths, a breadth-first search strategy akin to Dijkstra’s algorithm was applied [43]. PCPs were systematically identified for all pairs of promiscuous non-terminal nodes (i.e., nodes forming at least two PC relationships). For detected PCPs, three parameters were calculated including the pathway length (number of nodes), total number of PCs, and cumulative ΔPD value of all pathway edges. Redundant pathways were eliminated after identifying multiple pathways consisting of the same set of promiscuous nodes. Then, PCPs were prioritized based upon fusion of individual pathway rankings for the three parameters. This search method enabled fully automated analysis of PC clusters, PCPs, and PHs on the basis of PC network representations and was applied to systematically analyze promiscuity patterns among human kinase inhibitors [42].
Promiscuity analysis of kinase inhibitors
Inhibitors of the human kinome were subjected to systematic promiscuity analysis. Exploring these compounds on a large scale was of particular interest since clinical kinase inhibitors used in oncology typically have high promiscuity. Accordingly, these promiscuous kinase inhibitors have become a paradigm for polypharmacological compounds [44]. By extrapolating from these compounds, it is often assumed that ATP site directed kinase inhibitors might generally be promiscuous, as further discussed below.
For promiscuity analysis, kinase inhibitors and their activity data were systematically collected from several public compound repositories, curated, and combined. These efforts yielded more than 112,000 inhibitors with well-defined activity measurements [41]. For all curated inhibitors, kinase-based PD values were determined. Taken together, these inhibitors were found to be active against a total of 426 human kinases, hence providing 82% coverage of the kinome. The analysis of this unprecedentedly large data set revealed that nearly 40% of human kinase inhibitors had multi-kinase activity, but that only 4% were known to be active against five or more kinases. More than 60% of the inhibitors were only annotated with a single kinase activity. Therefore, global promiscuity among kinase inhibitors was not higher than observed for other compound classes, with mean and median PD values of 2.1 and 1.0, respectively [2, 41]. Overall, kinase inhibitor promiscuity was thus much lower than determined for the subset of clinical kinase inhibitors used in cancer treatment [41].
However, structurally analogous kinase inhibitors frequently displayed significant PD differences, leading to the formation of nearly 16,000 PCs (ΔPD ≥ 5) [41]. Representative examples of large-magnitude PCs formed by human kinase inhibitors are provided in Fig. 2a, b. In a global PCN representation for the human kinome, more than 600 distinct PC clusters of greatly varying composition emerged. Computational analysis of PC clusters yielded 8900 unique PCPs, ranging in length from three to 17 inhibitors [42]. Moreover, 520 kinase inhibitors qualified as PHs (with at least 10 PCs per hub). These PHs formed a total of 12,131 PCs (76% of all PCs) that involved nearly 7300 weakly or non-promiscuous analogues (with PD values of 1–4) [45]. Overall, large numbers of PCs, PCPs, and PHs were isolated from the comprehensive kinase inhibitor collection. Greatly varying PD values were observed and many inhibitors with single-kinase activity were detected using PC-based data structures. These findings also raised the question how kinase inhibitor promiscuity and selectivity might compare, as discussed in the following.
Promiscuity cliffs formed by human kinase inhibitors. In a, b, exemplary PCs formed by inhibitors of the human kinome are shown. Structural modifications are colored blue. For each inhibitor, its PD value is reported and a phylogenetic tree representation of the kinome is shown where its targets are represented as purple circles
Promiscuity versus selectivity of kinase inhibitors
Inhibitors of the human kinome are currently among the most intensely studied compounds in drug discovery [46,47,48]. The majority of current kinase inhibitors binds to the largely conserved adenosine triphosphate (ATP) cofactor binding site or, alternatively, less conserved regions proximal to this site [49,50,51,52]. Accordingly, the inhibitors are anticipated to display different degrees of promiscuity depending on their binding sites, which has also been analyzed on the basis of activity data [53, 54]. Promiscuity or selectivity of these inhibitors determines their potential for different therapeutic applications [44, 47, 55], which continues to be an intensely debated topic [55]. The active site-directed type I, I½, and II inhibitors display different binding modes that are characterized by different “in” and “out” combinations of the tripeptide DFG motif in the activation loop and the αC-helix in the active site region [52]. On the other hand, type III and IV inhibitors bind to different regions, which are often distant from the active site, and are allosteric in nature. Therefore, these inhibitors are typically more selective than other types [56]. Allosteric inhibitors are mostly discovered serendipitously and only a limited number of such inhibitors has been reported thus far [56]. The majority of current kinase inhibitors are type I inhibitors [51].
Experimental studies of active site-directed inhibitors have revealed different degrees of selectivity or promiscuity. Type I inhibitors directly bind to the conserved ATP site and are ATP-competitive. Thus, they are expected to be more promiscuous than type II inhibitors that target a less conserved hydrophobic pocket adjacent to the ATP binding site. However, both promiscuous and selective type I and II inhibitors were identified in profiling assays [12,13,14,15]. Furthermore, systematic computational analysis of activity data available for type I and II inhibitors including clinical candidates yielded similar results [53, 54]. Hence, there was no detectable selectivity advantage of type II over type I inhibitors, contrary to expectations. Two clinical kinase inhibitors with different promiscuity are shown in Fig. 3a [54]. Extensively assayed kinase inhibitors from biological screens were found to include specific inhibitors and others displaying different degrees of promiscuity at varying data confidence levels [18]. Corresponding observations were made for designated kinase probes from chemical biology. Chemical probes should ideally be target-specific, but kinase probes exhibited a wide range of activities and included both highly selective and highly promiscuous inhibitors [57]. Figure 3b shows exemplary kinase inhibitors designated as chemical probes with extremely different promiscuity [57].
Promiscuity of clinical kinase inhibitors and chemical probes. a Examples of clinical kinase inhibitors are shown that are annotated with a single kinase (capmatinib) or multiple kinases (lapatinib). b Two kinase inhibitors designated as chemical probes are shown, NVS-PAK1-1 and ruxolitinib, which display a very large difference in promiscuity. Kinase annotations are derived from publicly available high-confidence activity data. For each inhibitor, the PD value is reported in a circle
Taken together, these findings indicate that binding site conservation alone is not a major promiscuity determinant and that other effects such as binding kinetics and compound residence times are likely to contribute to promiscuity or selectivity of kinase inhibitors. Clearly, kinase inhibitors are not categorically promiscuous, but display a wide spectrum of activity profiles, which provide many opportunities for drug discovery as well as for future research.
Evidence for structure–promiscuity relationships through machine learning
Recently, machine learning has been applied to predict activity profiles of kinase inhibitors and their potential for polypharmacology [58, 59]. Furthermore, an online platform has been introduced for kinome-wide virtual compound screening using multi-task deep neural networks to guide multi-kinase drug design [60]. In addition to such applications, machine learning has been employed to investigate promiscuity from a more principal point of view, as discussed in the following.
Observations such as the low global promiscuity of kinase inhibitors or the frequent occurrence of PCs and PHs reinforce the question to which extent data incompleteness might affect promiscuity assessment. Naturally, data incompleteness also influences the analysis of kinase inhibitors as long as not all available inhibitors have been tested against all 518 kinases comprising the human kinome.
For bioactive compounds, it often remains difficult to rationalize why structural analogues often display large differences in promiscuity, as exemplified by the many PCs, PCPs, and PHs we have identified among kinase inhibitors. Importantly, if PD differences are a consequence of structural features or patterns, i.e., if true structure–promiscuity relationships exist, such structural patterns should be detectable using machine learning, even if they are difficult to uncover on the basis of expert analysis. Hence, if observed differences in compound promiscuity result from structural characteristics, it should be possible to build machine learning models to distinguish between promiscuous and non-promiscuous compounds. By contrast, if observed promiscuity differences would be strongly influenced by data incompleteness or experimental inconsistencies, no structure–promiscuity relationships would exist that could be detected via machine learning on the basis of molecular structure. In this case, machine learning models would inevitably fail.
To evaluate this conjecture, it was investigated whether or not predictive models could be derived to systematically distinguish between highly promiscuous and weakly or non-promiscuous screening compounds and, in addition, between promiscuous and non-promiscuous kinase inhibitors [24]. To assemble training and test sets for machine learning, structural analogues with different promiscuity were selected from PCs or randomly selected following alternative strategies. Using PCs as a source of training and test compounds further challenged the predictions because, in this case, promiscuous and non-promiscuous compounds included close structural analogs.
Different machine learning approaches were applied to build classification models on the basis of structural fingerprints. These methods included random forest (RF) [61], support vector machine (SVM) [62], deep neural network (DNN) [63], and graph convolutional network (GCN) [64] algorithms. As a control, nearest neighbor (1-NN) relationships between training and test compounds were analyzed on the basis of fingerprint Tanimoto similarity. In this case, the class label of the most similar training compound was assigned to each test compound.
For both screening compounds and kinase inhibitors selected from PCs, models obtained with all machine learning methods were found to be predictive, with an overall accuracy approaching or exceeding 70%. For randomly selected compounds, prediction accuracy was higher than 70%, approaching 80% in a number of instances. Hence, there was a clear and consistent tendency to distinguish between promiscuous and non-promiscuous compounds on the basis of machine learning. Differences between alternative methods were only small and there was no detectable advantage of deep learning compared to RF and SVM. Surprisingly, the simple 1-NN classifier consistently approached the performance level of machine learning. These findings indicated that machine learning calculations were dominated by nearest neighbor effects and provided further evidence for the presence of structural patterns that distinguished promiscuous from non-promiscuous compounds [24].
As a first step to elucidate relevant structural patterns, the influence of individual fingerprint features on the predictions of promiscuous versus non-promiscuous compounds was analyzed using an SVM-based feature weighting and ranking method [65]. For SVM models, features were weighted according to their contributions to correct predictions of promiscuous or non-promiscuous kinase inhibitors and ranked on the basis of cumulative feature weights. Fingerprint features were clearly differentiated by weighting and top-ranked features were further analyzed. Four features were identified that consistently contributed to the correct prediction of promiscuous kinase inhibitors and four different features that consistently contributed to the prediction of non-promiscuous inhibitors. These consensus features were mapped onto exemplary promiscuous and non-promiscuous kinase inhibitors, respectively, and found to form distinct coherent substructures [24]. These findings further rationalized successful predictions at the structural level and revealed the first structural patterns that were characteristic of promiscuous compounds.
Concluding remarks
Exploring multi-target activities of small molecules is an attractive area of research. At the molecular level, it is equally challenging and interesting to understand how a compound can form well-defined interactions in different binding sites and how interaction patterns of promiscuous and target-specific compounds compare. Moreover, given the link between promiscuity and polypharmacology, the question arises how promiscuous drugs and bioactive compounds really are. The jury is still out but we are gaining insights into the distribution of promiscuous compounds across therapeutic targets, also taking experimental test frequencies into consideration. In the study of promiscuity, experimental profiling and computational approaches complement each other, providing opportunities for data-driven computational analysis and predictive modeling. Herein, we have discussed data structures designed to uncover structure–promiscuity relationships. In this context, the concept of promiscuity cliffs plays a central role, based upon which other data structures have evolved. Given the popularity of polypharmacology, opinions are often voiced that pharmaceutically relevant small molecules might generally have multi-target activity. However, such assumptions are currently unsubstantiated on the basis of available experimental data. As long as promiscuity is not systematically explored in profiling campaigns at the cellular level or in vivo using model organisms, we are required to rely on currently available data and knowledge extracted from them. Given the increasingly large volumes of compounds and activity data that are becoming available, computational analysis represents an attractive approach to detect promiscuity trends. Large-scale exploration of compound activity data has shown that promiscuity cannot generally be assumed for small molecules, despite data incompleteness. However, data-driven analysis has also detected many puzzling structure–promiscuity relationships that merit further investigation. On the basis of currently available profiling experiments and other activity data, the picture is emerging that active compounds cover a wide spectrum of activities, ranging from target-specific or selective to highly promiscuous chemical entities. Inhibitors of the human kinome provide a representative example, as discussed herein. Although the efficacy of small sets of clinical kinase inhibitors used in oncology is known to rely on extensive promiscuity, providing a paradigm for polypharmacology, promiscuity of kinase inhibitors cannot generally be assumed, not even for those targeting the conserved ATP site. Rather, a wealth of different activity profiles is observed for kinase inhibitors, consistent with observations made for other compound classes. This provides opportunities for drug discovery, for example, the development of highly selective kinase inhibitors for long-term treatment of chronic diseases. Moreover, these findings also provide opportunities for future research to further explore and better understand molecular determinants of multi-target activity on the one hand and of selectivity or specificity on the other. We have also discussed that machine learning has successfully been used to generate indirect evidence for the existence of valid structure–promiscuity relationships and the presence of structural patterns that differentiate promiscuous and non-promiscuous compounds. Therefore, machine learning provides a basis for systematic exploration and mapping of distinguishing structural features, which is a current topic of research in our laboratory. Furthermore, exploring structural signatures of promiscuity should also aid in predicting compounds with desired multi-target activities, which would further advance polypharmacology-based drug discovery.
References
Hu Y, Bajorath J (2013) Compound promiscuity: what can we learn from current data? Drug Discov Today 18:644–650
Hu Y, Bajorath J (2017) Entering the ‘big data’ era in medicinal chemistry: molecular promiscuity analysis revisited. Future Sci OA 3:FSO179
McGovern SL, Caselli E, Grigorieff N, Shoichet BK (2002) A common mechanism underlying promiscuous inhibitors from virtual and high-throughput screening. J Med Chem 45:1712–1722
Feng BY, Shelat A, Doman TN, Guy RK, Shoichet BK (2005) High-throughput assays for promiscuous inhibitors. Nat Chem Biol 1:146–148
Shoichet BK (2006) Screening in a spirit haunted world. Drug Discov Today 11:607–615
Baell JB, Holloway GA (2010) New substructure filters for removal of pan assay interference compounds (PAINS) from screening libraries and for their exclusion in bioassays. J Med Chem 53:2719–2740
Baell J, Walters MA (2014) Chemistry: chemical con artists foil drug discovery. Nature 513:481–483
Anighoro A, Bajorath J, Rastelli G (2014) Polypharmacology: challenges and opportunities in drug discovery. J Med Chem 57:7874–7887
Proschak E, Stark H, Merk D (2019) Polypharmacology by design: a medicinal chemist’s perspective on multitargeting compounds. J Med Chem 62:420–444
Bolognesi ML (2019) Harnessing polypharmacology with medicinal chemistry. ACS Med Chem Lett 10:273–275
Mei Y, Yang B (2018) Rational application of drug promiscuity in medicinal chemistry. Future Med Chem 10:1835–1851
Karaman MW, Herrgard S, Treiber DK, Gallant P, Atteridge CE, Campbell BT, Chan KW, Ciceri P, Davis MI, Edeen PT, Faraoni R, Floyd M, Hunt JP, Lockhart DJ, Milanov ZV, Morrison MJ, Pallares G, Patel HK, Pritchard S, Wodicka LM, Zarrinkar PP (2008) A quantitative analysis of kinase inhibitor selectivity. Nat Biotechnol 26:127–132
Anastassiadis T, Deacon SW, Devarajan K, Ma H, Peterson JR (2011) Comprehensive assay of kinase catalytic activity reveals features of kinase inhibitor selectivity. Nat Biotechnol 29:1039–1045
Elkins JM, Fedele V, Szklarz M, Abdul Azeez KR, Salah E, Mikolajczyk J, Romanov S, Sepetov N, Huang XP, Roth BL, Al Haj Zen A, Fourches D, Muratov E, Tropsha A, Morris J, Teicher BA, Kunkel M, Polley E, Lackey KE, Atkinson FL, Overington JP, Bamborough P, Müller S, Price DJ, Willson TM, Drewry DH, Knapp S, Zuercher WJ (2016) Comprehensive characterization of the published kinase inhibitor set. Nat Biotechnol 34:95–103
Klaeger S, Heinzlmeir S, Wilhelm M, Polzer H, Vick B, Koenig PA, Reinecke M, Ruprecht B, Petzoldt S, Meng C, Zecha J, Reiter K, Qiao H, Helm D, Koch H, Schoof M, Canevari G, Casale E, Depaolini SR, Feuchtinger A, Wu Z, Schmidt T, Rueckert L, Becker W, Huenges J, Garz AK, Gohlke BO, Zolg DP, Kayser G, Vooder T, Preissner R, Hahne H, Tõnisson N, Kramer K, Götze K, Bassermann F, Schlegl J, Ehrlich HC, Aiche S, Walch A, Greif PA, Schneider S, Felder ER, Ruland J, Médard G, Jeremias I, Spiekermann K, Kuster B (2017) The target landscape of clinical kinase drugs. Science 358:eaan4368
Hu Y, Bajorath J (2013) High-resolution view of compound promiscuity. F1000Research 2:e144
Jasial S, Hu Y, Bajorath J (2016) Determining the degree of promiscuity of extensively assayed compounds. PLoS ONE 11:e0153873
Stumpfe D, Tinivella A, Rastelli G, Bajorath J (2017) Promiscuity of inhibitors of human protein kinases at varying data confidence levels and test frequencies. RSC Adv 7:41265–41271
Keiser MJ, Setola V, Irwin JJ, Laggner C, Abbas AI, Hufeisen SJ, Jensen NH, Kuijer MB, Matos RC, Tran TB, Whaley R, Glennon RA, Hert J, Thomas KLH, Edwards DD, Shoichet BK, Roth BL (2009) Predicting new molecular targets for known drugs. Nature 462:175–181
Keiser MJ, Roth BL, Armbruster BN, Ernsberger P, Irwin JJ, Shoichet BK (2007) Relating protein pharmacology by ligand chemistry. Nat Biotechnol 25:197–206
Wang L, Ma C, Wipf P, Liu H, Su W, Xie X-Q (2013) TargetHunter: an in silico target identification tool for predicting therapeutic potential of small organic molecules based on chemogenomic database. AAPS J 15:395–406
Awale M, Reymond J-L (2017) The polypharmacology browser: a web-based multi-fingerprint target prediction tool using ChEMBL bioactivity data. J Cheminform 9:e11
Jasial S, Gilberg E, Blaschke T, Bajorath J (2018) Machine learning distinguishes with high accuracy between pan-assay interference compounds that are promiscuous or represent dark chemical matter. J Med Chem 61:10255–10264
Blaschke T, Miljković F, Bajorath J (2019) Prediction of different classes of promiscuous and nonpromiscuous compounds using machine learning and nearest neighbor analysis. ACS Omega 4:6883–6890
Miljković F, Kunimoto R, Bajorath J (2017) Identifying relationships between unrelated pharmaceutical target proteins on the basis of shared active compounds. Future Sci OA 3:FSO212
Reker D, Bernardes GJL, Rodrigues T (2019) Computational advances in combating colloidal aggregation in drug discovery. Nat Chem 11:402–418
Dantas RF, Evangelista TCS, Neves BJ, Senger MR, Andrade CH, Ferreira SB, Silva-Junior FP (2019) Dealing with frequent hitters in drug discovery: a multidisciplinary view on the issue of filtering compounds on biological screenings. Expert Opin Drug Discov. https://doi.org/10.1080/17460441.2019.1654453
Haupt JV, Daminelli S, Schroeder M (2013) Drug promiscuity in PDB: protein binding site similarity is key. PLoS ONE 8:e65894
Pinzi L, Caporuscio F, Rastelli G (2018) Selection of protein conformations for structure-based polypharmacology studies. Drug Discov Today 23:1889–1896
Gilberg E, Stumpfe D, Bajorath J (2018) X-ray structure-based identification of compounds with activity against targets from different families and generation of templates for multitarget ligand design. ACS Omega 3:106–111
Gilberg E, Gütschow M, Bajorath J (2019) Promiscuous ligands from experimentally determined structures, binding conformations, and protein family-dependent interaction hotspots. ACS Omega 4:1729–1737
Gilberg E, Bajorath J (2019) Recent progress in structure-based evaluation of compound promiscuity. ACS Omega 4:2758–2765
Mestres J, Gregori-Puigjané E, Valverde S, Solé RV (2008) Data completeness—the Achilles heel of drug-target networks. Nat Biotechnol 26:983–984
Mestres J, Gregori-Puigjané E, Valverde S, Solé RV (2009) The topology of drug-target interaction networks: implicit dependence on drug properties and target families. Mol BioSyst 5:1051–1057
Maggiora GM (2006) On outliers and activity cliffs—why QSAR often disappoints. J Chem Inf Model 46:1535
Stumpfe D, Bajorath J (2012) Exploring activity cliffs in medicinal chemistry. J Med Chem 55:2932–2942
Dimova D, Bajorath J (2018) Rationalizing promiscuity cliffs. ChemMedChem 13:490–494
Kenny PW, Sadowski J (2005) Structure modification in chemical databases. In: Oprea TI (ed) Chemoinformatics in drug discovery. Wiley-VCH, Weinheim, pp 271–285
Hu Y, Jasial S, Gilberg E, Bajorath J (2017) Structure-promiscuity relationship puzzles—extensively assayed analogs with large differences in target annotations. AAPS J 19:856–864
Dimova D, Gilberg E, Bajorath J (2017) Identification and analysis of promiscuity cliffs formed by bioactive compounds and experimental implications. RSC Adv 7:58–66
Miljković F, Bajorath J (2018) Computational analysis of kinase inhibitors identifies promiscuity cliffs across the human kinome. ACS Omega 3:17295–17308
Miljković F, Vogt M, Bajorath J (2019) Systematic computational identification of promiscuity cliff pathways formed by inhibitors of the human kinome. J Comput Aided Mol Des 33:559–572
Dijkstra EW (1959) A note on two problems in connexion with graphs. Numer Math 1:269–271
Knight ZA, Lin H, Shokat KM (2010) Targeting the cancer kinome through polypharmacology. Nat Rev Cancer 10:130–137
Miljković F, Bajorath J (2019) Data structures for compound promiscuity analysis: promiscuity cliffs, pathways and promiscuity hubs formed by inhibitors of the human kinome. Fut Sci OA 5:FSO404
Cohen P (2002) Protein kinases—the major drug targets of the twenty-first century? Nat Rev Drug Discovery 1:309–315
Simmons DL (2013) Targeting kinases: a new approach to treating inflammatory rheumatic diseases. Curr Opin Pharmacol 13:426–434
Laufer S, Bajorath J (2014) New frontiers in kinases: second generation inhibitors. J Med Chem 57:2167–2168
Gavrin LK, Saiah E (2013) Approaches to discover non-ATP site kinase inhibitors. Med Chem Commun 4:41–51
Zhao Z, Wu H, Wang L, Liu Y, Knapp S, Liu Q, Gray NS (2014) Exploration of type II binding mode: a privileged approach for kinase inhibitor focused drug discovery? ACS Chem Biol 9:1230–1241
Hu Y, Furtmann N, Bajorath J (2015) Current compound coverage of the kinome. J Med Chem 58:30–40
Miljković F, Rodríguez-Pérez R, Bajorath J (2019) Machine learning models for accurate prediction of kinase inhibitors with different binding modes. J Med Chem. https://doi.org/10.1021/acs.jmedchem.9b00867
Miljković F, Bajorath J (2018) Exploring selectivity of multikinase inhibitors across the human kinome. ACS Omega 3:1147–1153
Miljković F, Bajorath J (2018) Reconciling selectivity trends from a comprehensive kinase inhibitor profiling campaign with known activity data. ACS Omega 3:3113–3119
Levitzki A (2013) Tyrosine kinase inhibitors: views of selectivity, sensitivity, and clinical performance. Annu Rev Pharmacol Toxicol 53:161–185
Wu P, Dinér P, Bunch L (2018) The screening and design of allosteric kinase inhibitors. In: Ward RA, Goldberg FW (eds) Kinase drug discovery: modern approaches. RSC, Cambridge, pp 34–60
Miljković F, Bajorath J (2018) Data-driven exploration of selectivity and off-target activities of designated chemical probes. Molecules 23:e2434
Rodríguez-Pérez R, Bajorath J (2019) Multitask machine learning for classifying highly and weakly potent kinase inhibitors. ACS Omega 4:4367–4375
Li X, Li Z, Wu X, Xiong Z, Yang T, Fu Z, Liu X, Tan X, Zhong F, Wan X, Wang D, Ding X, Yang R, Hou H, Li C, Liu H, Chen K, Jiang H, Zheng M (2019) Deep learning enhancing kinome-wide polypharmacology profiling: model construction and experiment validation. J Med Chem. https://doi.org/10.1021/acs.jmedchem.9b00855
Li Z, Li X, Liu X, Fu Z, Xiong Z, Wu X, Tan X, Zhao J, Zhong F, Wan X, Luo X, Chen K, Jiang H, Zheng M (2019) KinomeX: a web application for predicting kinome-wide polypharmacology effect on small molecules. Bioinformatics. https://doi.org/10.1093/bioinformatics/btz519
Breiman L (2001) Random forests. Mach Learn 45:5–32
Joachims T (1999) Making large-scale SVM learning practical. In: Schölkopf B, Burges CJC, Smola AJ (eds) Advances in kernel methods: support vector learning. MIT Press, Cambridge, pp 169–184
Goodfellow I, Bengio Y, Courville A (2016) Deep learning. MIT Press, Cambridge
Duvenaud D, Maclaurin D, Aguilera-Iparraguirre J, Gomez-Bombarelli R, Hirzel T, Aspuru-Guzik A, Adams RP (2015) Convolutional networks on graph for learning molecular fingerprints. Neural Inf Proc Sys 28:2224–2232
Balfer J, Bajorath J (2015) Visualization and interpretation of support vector machine activity predictions. J Chem Inf Model 55:1136–1147
Author information
Authors and Affiliations
Corresponding author
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Miljković, F., Bajorath, J. Data structures for computational compound promiscuity analysis and exemplary applications to inhibitors of the human kinome. J Comput Aided Mol Des 34, 1–10 (2020). https://doi.org/10.1007/s10822-019-00266-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10822-019-00266-0