Abstract
The ability of a small molecule to interact with multiple target proteins provides the molecular basis of polypharmacology. So-defined compound promiscuity is intensely investigated in drug discovery. For example, for kinase inhibitors, the interplay between target selectivity and promiscuity plays a decisive role for different therapeutic applications. The “promiscuity cliff” (PC) concept was introduced previously to aid in promiscuity analysis. A PC is defined as a pair of structurally similar compounds with a large difference in promiscuity. Accordingly, PCs can reveal small structural modifications that might be responsible for selectivity or multi-target activity. In network representations, PCs form clusters of varying size and complexity that are difficult to analyze interactively. Herein, we introduce a computational method to systematically identify PC pathways, which are particularly rich in structure-promiscuity information, and extract them from PC clusters. PC pathways provide informative templates for experimental design. In a proof-of-concept investigation, we have applied the new computational approach to systematically identify pathways in more than 600 PC clusters formed by inhibitors of the human kinome, demonstrating the utility of the method and revealing many interesting promiscuity patterns.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Possible origins of compound promiscuity continue to be debated in the drug discovery community. Promiscuity is often due to non-specific binding resulting from aggregation effects and other assay artifacts and thus highly undesirable [1,2,3,4,5]. On the other hand, compound promiscuity may originate from true binding events when a small molecule interacts with multiple targets in a defined way. Such multi-target activities form the basis of polypharmacology with its associated functional effects [6,7,8]. The polypharmacology concept has gradually revised and further extended the long-standing single-target specificity paradigm in drug discovery [9, 10]. However, achieving target specificity of small molecules will continue to be a guiding principle for many therapeutic applications including, among others, the treatment of chronic diseases or development of anti-infective agents. Target specificity is also of critical relevance in other areas such as chemical biology where the development of high-quality chemical probes to interrogate target-dependent functional effects is a major focal point [11, 12]. By contrast, compounds with multi-kinase activity have been successfully applied in oncology [13, 14]. Other multi-target compounds show promise in therapeutic areas such as neurological disorders [15].
There are several computational and experimental avenues to explore molecular promiscuity. Compounds with multi-target activity can be identified through computational analysis of curated activity data [6, 7, 16] from medicinal chemistry that is available in major repositories such as ChEMBL [17] or biological screening data available in PubChem [18]. In addition, compound profiling and array experiments are a major source of multi-target activity information [18,19,20,21,22,23].
The “promiscuity cliff” (PC) concept [24,25,26] was originally introduced to bridge between computational and experimental approaches and aid in the analysis of compound array data [24]. A PC is defined as a pair of structurally analogous compounds, i.e., compounds that are only distinguished by a single substitution (R-group replacement), having a significant difference in the number of targets they are active against [24,25,26]. Accordingly, PCs reveal small chemical modifications that are implicated in causing promiscuity [25, 26]. Furthermore, differences in apparent promiscuity between PC compounds might be influenced by varying test (assay) frequencies. Thus, PCs also suggest additional target hypotheses for structural analogs of highly promiscuous compounds [26]. For meaningful applications of the PC concept, it must be ensured that compounds with assay liabilities and resulting frequent hitter characteristics are excluded from consideration [26, 27]. Going beyond the analysis of compound array experiments, PCs were identified on a large scale in publicly available active compounds from different sources [27, 28].
Kinase inhibitors are a prime target for promiscuity analysis because the vast majority of currently available inhibitors target the adenosine triphosphate (ATP) (cofactor)-binding site that is largely conserved across the human kinome [29, 30]. Hence, these inhibitors are expected to be promiscuous [31]. However, general promiscuity and lack of selectivity of kinase inhibitors is neither supported by profiling experiments [22, 23], nor compound activity data analysis [32,33,34].
To quantify differences in kinase activities of ATP site-directed compounds, a systematic search for PCs was carried out in a large collection of more than 112,000 inhibitors of 426 human kinases (82% of the human kinome) that were assembled from several public compound databases [28]. Nearly 16,000 PCs were identified. In a global network representation, these PCs formed more than 600 clusters of varying composition [28].
PC clusters represent a rich source of information for promiscuity analysis. For example, from clusters, PC pathways (PCPs) can be isolated that represent sequences of compounds with alternating low promiscuity -or selectivity- and high promiscuity. Hence, inspection of PCPs makes it possible to follow stepwise structural modifications that strongly influence apparent promiscuity levels [28]. However, the large number of increasingly complex PC clusters quickly limits manual analysis of PCPs and makes it essentially impossible to comprehensively study pathways in an interactive manner. Hence, there is a need to automate this process and enable systematic analysis of PC clusters and PCPs.
Herein, we present a computational approach to systematically identify PCPs in clusters, prioritize most informative PCPs, and extract them. In addition, an entropy-based measure is applied to assess the distribution of pathway-associated kinase activities across the kinome.
Materials and methods
Data set
The previously reported set of kinase inhibitor PC clusters [28] was taken for method development and subjected to systematic analysis. Transformation size-restricted matched molecular pairs (MMPs) [35, 36] were calculated to generate pairs of structurally analogous kinase inhibitors. An MMP is defined as a pair of compounds that are only distinguished by a chemical modification (transformation) at a single site [36, 37]. For inhibitors forming MMPs, the promiscuity degree (PD) was determined as the number of kinase annotations on the basis of curated activity data, applying a potency threshold of 10 µM to IC50, Ki, or Kd values. An MMP was considered a PC if the absolute difference of inhibitor PD values (ΔPD) was at least 5, i.e., if one inhibitor was active against five more kinases than the other. In addition, the PD value of the less promiscuous inhibitor was required to be between 1 and 4 such that PCs could not be formed by pairs of highly promiscuous inhibitors. Accordingly, the smallest possible PC involved an inhibitor with PD = 1 and a structural analog with PD = 6. Applying these criteria, a total of 15,939 PCs were obtained that involved 10,741 kinase inhibitors, including 1653 inhibitors with PD values between 6 and 295. These inhibitors were capable of participating in PCs as highly promiscuous cliff partners. The global network representation of the 15,939 PCs (nodes: compounds, edges: pairwise PC relationships) contained 622 disjoint PC clusters [28].
Computational extraction of PC pathways
For computational analysis, PCP was defined as the shortest path between two nodes from a PC cluster. When multiple shortest paths existed between two nodes, ΔPD of the edges was considered and the path yielding the largest cumulative ΔPD value was chosen. In addition, to eliminate path redundancy, only a single path was retained if multiple shortest paths contained the same set of promiscuous compounds (PD ≥ 6). So-defined PCPs were systematically generated for all pairs of promiscuous non-terminal nodes (i.e., inhibitors forming at least two PC relationships with others). For each qualifying path, three parameters were calculated:
-
1.
Length (number of nodes)
-
2.
Total number of PCs involving promiscuous inhibitors with PD ≥ 6
-
3.
Cumulative ΔPD of edges of the path.
We note that the application of criterion 2 makes it possible to prioritize PCPs that contain “promiscuity hubs”, i.e., pathway compounds that form large numbers of PCs with others outside the PCP. Pathway hubs are further discussed below.
In addition to applying criteria for PCP prioritization, a frequency model for n kinase groups [29] associated with a path is obtained by counting the frequency of occurrence of kinases belonging to each represented group. From frequency counts, the Shannon entropy (SE) [38] was calculated:
Here, the \({p}_{i}\) is the relative frequency of occurrence of each kinase group. Low SE values indicate that kinases associated with a path belong to a single group while increasing values indicate that associated kinases belong to multiple (and increasing numbers of) groups.
PCPs were ranked separately in decreasing order according to criteria 1–3 specified above. Then, rank fusion was applied. Therefore, the three ranks of each path were sorted in ascending order yielding a tuple \(({r}_{a},{r}_{b},{r}_{c})\) with \({r}_{a}\le {r}_{b}\le {r}_{c}\). The PCPs were ranked according to the lexicographic order of the tuples. Initially, only the highest rank \({r}_{a}\) was considered and only in case of a tie, the second best rank \({r}_{b}\)was used; if there was a tie for both ranks, \({r}_{c}\) was taken into consideration. Lexicographic ranking ensured that the highest ranked pathways according to each criterion appeared near the top of the final ranking.
All calculations were carried out using the Python-implemented NetworkX package [39]. Shortest path calculations of the unweighted network were performed using a breadth-first search strategy similar to Dijkstra’s algorithm [40]. The method organizes nodes of a network in layers of increasing distance around a source node. Each node in a layer represents a target node that contains pointers to all nodes of the previous layer, which extend possible shortest paths to the target node. Thus, all shortest paths from a source node to an arbitrary target node can be determined and prioritized according to the criteria outlined above.
Pathway visualization
Highly-ranked PCPs in PC clusters were visualized. Clusters were drawn using NetworkX [39] applying the Kamada–Kawai force-directed layout algorithm [41]. Cluster nodes were color-coded according to PD value ranges. In clusters, selected PCPs were traced using a thick black line. In addition, PCP compounds forming hubs with other nodes were identified. For kinases associated with PCP nodes, the frequency of occurrence was counted. For each selected PCP, a phylogenetic tree was drawn using KinMap [42], in which each dot represented a kinase associated with a PCP compound. Dots were scaled in size according to the frequency of kinase annotations.
Results and discussion
The new methodology for PCP extraction from PC clusters was tested on kinase inhibitor PCs identified on the basis of medicinal chemistry data. For these active compounds, no test frequencies were available. We note that PCs have also been identified on the basis of publicly available screening compounds for which test frequencies were available [43]. These PCs also extensively formed clusters [43], similar to the kinase inhibitor PCs used herein. For the development of our method, the source of PCs (medicinal chemistry data or biological screening) made no difference.
Promiscuity cliff clusters
The 15,939 PCs formed by 10,741 kinase inhibitors were organized in a PC network in which nodes represented inhibitors and edges pairwise PC relationships. This network contained 622 isolated clusters. Figure 1 reports the distribution of inhibitors, PCs, and mean ΔPD values for the clusters. About half of the clusters contained small numbers of compounds and PCs, with median values of 6.5 and 6.0, respectively. However, about 25% of the clusters contained 20 or more compounds and PCs, representing increasingly large and complex clusters. The median ΔPD value for PC clusters was close to 10 and the third quartile value was close to 25. Thus, PC clusters captured large differences in compound promiscuity.
Distribution of inhibitors, PCs, and mean ΔPD values for PC clusters. Boxplots report distribution of compounds, PCs, and mean ΔPD values for 622 kinase inhibitor PC clusters. Median values are reported and red diamond markers indicate the mean values of the distributions. Boxplots report the smallest value (bottom line), first quartile (lower boundary of the box), median value (thick line), third quartile (upper boundary of the box), largest value (top line), and outliers (points below the smallest or above the largest value)
Promiscuity cliff pathways
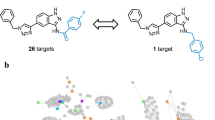
An exemplary PCP is shown in Fig. 2a. The PCP data structure is particularly attractive for the analysis of promiscuity patterns because PCPs consist of sequences of PC compounds with alternating large and small PD values. Hence, along a path iterative structural modifications can be examined that lead to large differences in promiscuity between structurally analogous compounds. In addition, as also shown in Fig. 2a, promiscuous PCP compounds frequently represent promiscuity hubs forming multiple PCs with other structural analogs outside the path that are only weakly promiscuous or non-promiscuous, which provides additional information. Thus, for the exploration of structure-promiscuity relationships, PCPs represent an informative data structure.
Promiscuity cliff pathways from cluster A. In a, the top ranked PCP is traced. Nodes are color-coded according to PD value ranges and nodes of PCP compounds are numbered. Below the cluster, structures of PCP compounds are shown and their PD values are reported in corresponding nodes. Structural modifications distinguishing pairs of inhibitors along the path are colored red. In b, a promiscuity hub from the PCP is depicted that forms multiple PCs to other inhibitors with one or two kinase annotations. Structures of exemplary analogs are shown. In c, mapping of kinase annotations from the top ranked PCP onto a phylogenetic tree of the human kinome is shown. Each kinase associated with the PCP is represented as a red dot. The dots are scaled in size according to the number of kinase annotations along the path. In d, a lower ranked PCP from cluster A is traced
Computational identification of pathways
Manually tracing PCPs is cumbersome and becomes essentially impossible when PC clusters grow in size beyond a few compounds such as the exemplary cluster shown in Fig. 2a. PC clusters contain many possible PCPs that need to be systematically examined to identify most informative paths. To these ends, computational analysis is essential and we introduce a new computational method for systematically identifying PCPs and extracting them from clusters. The approach relies on shortest path calculations between nodes in networks using breadth-first search akin to the Dijkstra’s algorithm [40]. Application of this approach makes it possible to exhaustively mine PC clusters for PCPs and automate their extraction, guided by criteria to prioritize PCPs according to their structure-promiscuity relationship information content. PCPs were extracted from all kinase inhibitor PC clusters containing at least two promiscuous compounds with PD ≥ 6. In the following, exemplary cases are presented.
Pathway analysis
Table 1 reports the composition of two representative clusters A and B from the global PC network and their pathway statistics resulting from computational analysis. Cluster A contained 132 kinase inhibitors and 42 computationally identified PCPs meeting the criteria specified above and cluster B contained 117 inhibitors and 21 PCPs. The comparison illustrates that the number of PCPs does not necessarily scale with the number of compounds. Rather, the topology of clusters and content of hubs are major factors determining the number of PCPs. For cluster A and B, PCPs with up to seven and five inhibitors were identified, respectively. Figure 2a depicts cluster A and the top ranked PCP identified by computational analysis. It consists of seven structural analogs with substitutions at three sites. The PCP compounds include two densely connected hubs (compounds 1 and 5) and have striking difference in promiscuity including four in part highly promiscuous inhibitors, especially compound 1 (PD = 62), and three others with single kinase annotations. Large differences in promiscuity along the path are accompanied by confined structural modifications. In Fig. 2b, a part of the hub configuration around highly promiscuous compound 1 is displayed, which forms PCs with numerous inhibitors having mostly single kinase annotations. These analogs are distinguished from the highly promiscuous inhibitor by only minor chemical modifications leading to very large differences in apparent promiscuity. These observations are puzzling and this PCP alone would provide a basis for extensive follow-up experiments to better understand possible origins of large-magnitude differences in promiscuity. For example, inhibitors with apparent specificity (PD = 1) might be tested against other PCP-associated kinases and/or additional analogs might be generated to probe the influence of selected and combined chemical modifications on promiscuity. Without the identification and analysis of PCPs, many of these puzzling structure-promiscuity relationships would most likely remain unnoticed, illustrating the utility of the PCP data structure.
Figure 2c shows that kinase annotations of inhibitors forming the top ranked PCP are widely distributed across the human kinome. The distribution of large dots indicates that a variety of distantly related kinases have multiple annotations originating from inhibitors of the PCP, suggesting additional target hypotheses for PCP compounds and hub analogs.
Figure 2d depicts a lower ranked PCP from cluster A that overlaps with the top ranked path. This PCP consists of five inhibitors including two densely connected hubs (compound 1 and 5) and one highly promiscuous inhibitor (compound 1, PD = 47). The lower rank of this PCP compared to the top ranked path is mainly due to its smaller size and lower cumulative ΔPD value. The kinome coverage of kinase annotations from both PCPs is comparable. Despite its lower rank, this PCP also reveals a variety of structure-promiscuity patterns and represents another informative template for experimental design.
Figure 3a depicts cluster B and its top ranked PCP. It consists of five inhibitors including three promiscuity hubs and two inhibitors with dual kinase activity. With 140 kinase annotations, PCP compound 1 is one of the most promiscuous kinase inhibitors we have identified. The PCP contains a close structural analog of this inhibitor with dual kinase activity (compound 2) that only differs by a hydroxyl to fluoro substitution. In addition, as shown in Fig. 3b, the hub environment of compound 1 also contains a variety of close analogs with only two or three kinase annotations. Thus, at a first glance, one might hypothesize that many analogs of compound 1 would also be more promiscuous but might have not been sufficiently tested. However, this immediate and plausible assumption of data sparseness as a cause of apparent differences in promiscuity is called into question when analyzing the kinome distribution of PCP kinase annotations, shown in Fig. 3c. In this case, kinome-wide activities only result from the pan-kinase inhibitor (compound 1), whereas activities of the other PCP compounds and PCP-associated inhibitors are strongly focused on the Src family within the tyrosine kinase (TK) group. This is a characteristic of inhibitors comprising cluster B, as also illustrated by considering another lower ranked PCP from this cluster, depicted in Fig. 3d. This PCP comprises five inhibitors and includes three promiscuity hubs (with a maximum of 14 kinase annotations). As revealed in Fig. 3e, these inhibitors are exclusively active against members of the TK group. Taken together, these observations suggest that it is unlikely that data sparseness alone would account for the apparent difference in promiscuity between compound 1 in Fig. 3a and other inhibitors in cluster B. Accordingly, exploring possible structural origins of pan-kinase versus TK promiscuity should, in this case, also be an attractive opportunity for follow-up investigation. Cluster A and B are representative of many PC clusters formed by kinase inhibitors that can be studied in detail on the basis of computational PCP analysis.
Promiscuity cliff pathways from cluster B. In a, the top ranked PCP is traced. In b, a promiscuity hub is shown in detail. In c, the phylogenetic tree representation of kinase annotations associated with the top ranked PCP is depicted. In d, a lower ranked PCP from cluster B is shown. In e, the phylogenetic tree representation of the lower ranked PCP is displayed. The representation is according to Fig. 2
For the promiscuity hub examples reported in Figs. 2b and 3b, no comparative X-ray data are available to further investigate promiscuity differences. However, other examples of promiscuous compounds have recently been discussed on the basis of structural data [44], which are well worth considering in the context of PC analysis.
Conclusions
PC clusters from network representations represent a rich source of structure-promiscuity relationship information. The PCP data structure is particularly informative for promiscuity analysis and suitable to aid in experimental design. However, interactive graphical analysis of PC clusters and manual delineation of PCPs is difficult and limits PC analysis. Therefore, we have introduced a new computational approach to systematically extract and organize PCPs from PC clusters. The methodology makes it possible to exhaustively identify PCPs in data sets, as exemplified by our analysis of PC clusters formed by inhibitors of the human kinome. Systematically identified PCPs reveal many structure-promiscuity relationships that would be difficult, if not impossible to detect on the basis of interactive case-by-case analysis. PCPs provide a basis for exploring structural modifications that are implicated in triggering promiscuity versus selectivity and identify compound subsets in which apparent differences in promiscuity are likely due to data sparseness. Accordingly, the computational approach introduced herein enables a thorough investigation of promiscuity patterns on the basis of PCPs and associated promiscuity hubs. PCPs covering the human kinome we have identified as a part of our study will be made freely available for follow-up investigations as an open access deposition on the ZENODO platform [45].
References
McGovern SL, Caselli E, Grigorieff N, Shoichet BK (2002) A common mechanism underlying promiscuous inhibitors from virtual and high-throughput screening. J Med Chem 45:1712–1722
Feng BY, Shelat A, Doman TN, Guy RK, Shoichet BK (2005) High-throughput assays for promiscuous inhibitors. Nat Chem Biol 1:146–148
Shoichet BK (2006) Screening in a spirit haunted world. Drug Discov Today 11:607–615
Baell JB, Holloway GA (2010) New substructure filters for removal of pan assay interference compounds (PAINS) from screening libraries and for their exclusion in bioassays. J Med Chem 53:2719–2740
Baell J, Walters MA (2014) Chemistry: chemical con artists foil drug discovery. Nature 513:481–483
Hu Y, Bajorath J (2013) Compound promiscuity: what can we learn from current data? Drug Discov Today 18:644–650
Hu Y, Bajorath J (2013) High-resolution view of compound promiscuity. F1000Research 2:e144
Anighoro A, Bajorath J, Rastelli G (2014) Polypharmacology: challenges and opportunities in drug discovery. J Med Chem 57:7874–7887
Nurse P (1997) The ends of understanding. Nature 387:657–657
Roukos DH (2011) Networks medicine: from reductionism to evidence of complex dynamic biomolecular interactions. Pharmacogenomics 12:695–698
Arrowsmith CH, Audia JE, Austin C, Baell J, Bennett J, Blagg J, Bountra C, Brennan PE, Brown PJ, Bunnage ME, Doepner-Buser C, Campbell RM, Carter AJ, Cohen P, Copeland RA, Cravatt B, Dahlin JL, Dhanak D, Edwards AM, Frederiksen M, Frye SV, Gray N, Grimshaw CE, Hepworth D, Howe T, Huber KVM, Jin J, Knapp S, Kotz JD, Kruger RG, Lowe D, Mader MM, Marsden B, Mueller-Fahrnow A, Müller S, O’Hagan RC, Overington JP, Owen DR, Rosenberg SH, Ross R, Roth B, Schapira M, Schreiber SL, Shoichet B, Sundström M, Superti-Furga G, Taunton J, Toledo-Sherman L, Walpole C, Walters MA, Willson TM, Workman P, Young RN, Zuercher WJ (2015) The promise and peril of chemical probes. Nat Chem Biol 11:536–541
Miljković F, Bajorath J (2018) Data-driven exploration of selectivity and off-target activities of designated chemical probes. Molecules 23:e2434
Knight ZA, Lin H, Shokat KM (2010) Targeting the cancer kinome through polypharmacology. Nat Rev Cancer 10:130–137
Gross S, Rahal R, Stransky N, Lengauer C, Hoeflich KP (2015) Targeting cancer with kinase inhibitors. J Clin Invest 125:1780–1789
Bolognesi ML, Cavalli A (2016) Multitarget drug discovery and polypharmacology. ChemMedChem 11:1190–1192
Stumpfe D, Tinivella A, Rastelli G, Bajorath J (2017) Promiscuity of inhibitors of human protein kinases at varying data confidence levels and test frequencies. RSC Adv 7:41265–41271
Gaulton A, Hersey A, Nowotka M, Bento AP, Chambers J, Mendez D, Mutowo P, Atkinson F, Bellis LJ, Cibrián-Uhalte E, Davies M, Dedman N, Karlsson A, Magariños MP, Overington JP, Papadatos G, Smit I, Leach AR (2017) The ChEMBL database in 2017. Nucleic Acids Res 45(Database issue):D945–D954
Kim S, Thiessen PA, Bolton EE, Chen J, Fu G, Gindulyte A, Han L, He J, He S, Shoemaker BA, Wang J, Yu B, Zhang J, Bryant SH (2016) PubChem substance and compound databases. Nucleic Acids Res 44(Database issue):D1202–D1213
Karaman MW, Herrgard S, Treiber DK, Gallant P, Atteridge CE, Campbell BT, Chan KW, Ciceri P, Davis MI, Edeen PT, Faraoni R, Floyd M, Hunt JP, Lockhart DJ, Milanov ZV, Morrison MJ, Pallares G, Patel HK, Pritchard S, Wodicka LM, Zarrinkar PP (2008) A quantitative analysis of kinase inhibitor selectivity. Nat Biotechnol 26:127–132
Anastassiadis T, Deacon SW, Devarajan K, Ma H, Peterson JR (2011) Comprehensive assay of kinase catalytic activity reveals features of kinase inhibitor selectivity. Nat Biotechnol 29:1039–1045
Clemons PA, Bodycombe NE, Carrinski HA, Wilson JA, Shamji AF, Wagner BK, Koehler AN, Schreiber SL (2010) Small molecules of different origins have distinct distributions of structural complexity that correlate with protein-binding profiles. Proc Natl Acad Sci USA 107:18787–18792
Elkins JM, Fedele V, Szklarz M, Abdul Azeez KR, Salah E, Mikolajczyk J, Romanov S, Sepetov N, Huang XP, Roth BL, Al Haj Zen A, Fourches D, Muratov E, Tropsha A, Morris J, Teicher BA, Kunkel M, Polley E, Lackey KE, Atkinson FL, Overington JP, Bamborough P, Müller S, Price DJ, Willson TM, Drewry DH, Knapp S, Zuercher WJ (2016) Comprehensive characterization of the published kinase inhibitor set. Nat Biotechnol 34:95–103
Klaeger S, Heinzlmeir S, Wilhelm M, Polzer H, Vick B, Koenig PA, Reinecke M, Ruprecht B, Petzoldt S, Meng C, Zecha J, Reiter K, Qiao H, Helm D, Koch H, Schoof M, Canevari G, Casale E, Depaolini SR, Feuchtinger A, Wu Z, Schmidt T, Rueckert L, Becker W, Huenges J, Garz AK, Gohlke BO, Zolg DP, Kayser G, Vooder T, Preissner R, Hahne H, Tõnisson N, Kramer K, Götze K, Bassermann F, Schlegl J, Ehrlich HC, Aiche S, Walch A, Greif PA, Schneider S, Felder ER, Ruland J, Médard G, Jeremias I, Spiekermann K, Kuster B (2017) The target landscape of clinical kinase drugs. Science 358:eaan4368
Dimova D, Hu Y, Bajorath J (2012) Matched molecular pair analysis of small molecule microarray data identifies promiscuity cliffs and reveals molecular origins of extreme compound promiscuity. J Med Chem 55:10220–10228
Bajorath J (2017) From activity cliffs to promiscuity cliffs. Fut Sci OA 3:FSO227
Dimova D, Bajorath J (2018) Rationalizing promiscuity cliffs. ChemMedChem 13:490–494
Dimova D, Gilberg E, Bajorath J (2017) Identification and analysis of promiscuity cliffs formed by bioactive compounds and experimental implications. RSC Adv 7:58–66
Miljković F, Bajorath J (2018) Computational analysis of kinase inhibitors identifies promiscuity cliffs across the human kinome. ACS Omega 3:17295–17308, 2018
Manning G, Whyte DB, Martinez R, Hunter T, Sudarsanam S (2002) The protein kinase complement of the human genome. Science 298:1912–1934
Noble MEM, Endicott JA, Johnson LN (2004) Protein kinase inhibitors: insights into drug design from structure. Science 303:1800–1805
Levitzki A (2013) Tyrosine kinase inhibitors: views of selectivity, sensitivity, and clinical Performance. Annu Rev Pharmacol Toxicol 53:161–185
Miljković F, Bajorath J (2018) Exploring selectivity of multikinase inhibitors across the human kinome. ACS Omega 3:1147–1153
Miljković F, Bajorath J (2018) Reconciling selectivity trends from a comprehensive kinase inhibitor profiling campaign with known activity data. ACS Omega 3:3113–3119
Miljković F, Bajorath J (2018) Evaluation of kinase inhibitor selectivity using cell-based profiling data. Mol Inform 37:e1800024
Hu X, Hu Y, Vogt M, Stumpfe D, Bajorath J (2012) MMP-cliffs: systematic identification of activity cliffs on the basis of matched molecular pairs. J Chem Inf Model 52:1138–1145
Kenny PW, Sadowski J (2004) Structure modification in chemical databases. In: Oprea TI (ed) Chemoinformatics in drug discovery. Wiley-VCH, Weinheim, pp 271–285
Hussain J, Rea C (2010) Computationally efficient algorithm to identify matched molecular pairs (MMPs) in large data sets. J Chem Inf Model 50:339–348
Shannon CE (1948) A mathematical theory of communication. Bell Syst Techn J 27:379–423
Hagberg AA, Schult DA, Swart PJ (2008) Exploring network structure, dynamics, and function using NetworkX. In: Varoquaux G, Vaught T, Millman J (eds) Proceedings of the 7th python in science conference (SciPy 2008), Pasadena, CA, Aug 19–24, pp 11–15
Dijkstra EW (1959) A note on two problems in connexion with graphs. Numer Math 1:269–271
Kamada T, Kawai S (1989) An algorithm for drawing general undirected graphs. Inf Process Lett 31:7–15
Eid S, Turk S, Volkamer A, Rippmann F, Fulle S (2017) KinMap: a web-based tool for interactive navigation through human kinome data. BMC Bioinform 18:e16
Hu Y, Jasial S, Gilberg E, Bajorath J (2017) Structure-promiscuity relationship puzzles—extensively assayed analogs with large differences in target annotations. AAPS J 19:856–864
Gilberg E, Bajorath J (2019) Recent progress in structure-based evaluation of compound promiscuity. ACS Omega 4:2758–2765
Author information
Authors and Affiliations
Corresponding author
Additional information
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Miljković, F., Vogt, M. & Bajorath, J. Systematic computational identification of promiscuity cliff pathways formed by inhibitors of the human kinome. J Comput Aided Mol Des 33, 559–572 (2019). https://doi.org/10.1007/s10822-019-00198-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10822-019-00198-9